Abstract
Single-cell transcriptomics provide a systematic map of gene expression in different human cell types. The next challenge is to systematically understand cell-type-specific gene function. The integration of CRISPR-based functional genomics and stem cell technology enables the scalable interrogation of gene function in differentiated human cells. Here we present the first genome-wide CRISPR interference and CRISPR activation screens in human neurons. We uncover pathways controlling neuronal response to chronic oxidative stress, which is implicated in neurodegenerative diseases. Unexpectedly, knockdown of the lysosomal protein prosaposin strongly sensitizes neurons, but not other cell types, to oxidative stress by triggering the formation of lipofuscin, a hallmark of aging, which traps iron, generating reactive oxygen species and triggering ferroptosis. We also determine transcriptomic changes in neurons after perturbation of genes linked to neurodegenerative diseases. To enable the systematic comparison of gene function across different human cell types, we establish a data commons named CRISPRbrain.








Similar content being viewed by others
Data availability
All screen datasets are publicly available on the CRISPRbrain website (https://crisprbrain.org/) (associated with Figs. 1 and 2 and Extended Data Figs. 1 and 2). The accession number for the RNA-seq datasets reported in this paper is GSE152988, and mapping of sgRNAs to single cells is available at https://kampmannlab.ucsf.edu/crop-seq (associated with Figs. 4, 6 and 7 and Extended Data Fig. 4). The DisGeNET database is available at https://www.disgenet.org/ (associated with Fig. 6). There are no restrictions on data availability. Source data are provided with this paper.
Code availability
All data analyses were performed using published computational pipelines and standard Python/R packages as described in the Methods. Our pipeline for analysis of the CROP-seq data is available at https://kampmannlab.ucsf.edu/crop-seq.
References
Tian, R. et al. CRISPR interference-based platform for multimodal genetic screens in human iPSC-derived neurons. Neuron 104, 239–255.e12 (2019).
Gilbert, L. A. et al. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159, 647–661 (2014).
Kampmann, M. CRISPR-based functional genomics for neurological disease. Nat. Rev. Neurol. 16, 465–480 (2020).
Weltner, J. et al. Human pluripotent reprogramming with CRISPR activators. Nat. Commun. 9, 2643 (2018).
Wang, C. et al. Scalable production of iPSC-derived human neurons to identify tau-lowering compounds by high-content screening. Stem Cell Rep. 9, 1221–1233 (2017).
Horlbeck, M. A. et al. Compact and highly active next-generation libraries for CRISPR-mediated gene repression and activation. eLife 5, e19760 (2016).
Yilmaz, A., Peretz, M., Aharony, A., Sagi, I. & Benvenisty, N. Defining essential genes for human pluripotent stem cells by CRISPR–Cas9 screening in haploid cells. Nat. Cell Biol. 20, 610–619 (2018).
Hart, T. et al. Evaluation and design of genome-wide CRISPR/SpCas9 knockout screens. G3 (Bethesda) 7, 2719–2727 (2017).
Colacurcio, D. J. & Nixon, R. A. Disorders of lysosomal acidification—the emerging role of v-ATPase in aging and neurodegenerative disease. Ageing Res. Rev. 32, 75–88 (2016).
Dandage, R. & Landry, C. R. Paralog dependency indirectly affects the robustness of human cells. Mol. Syst. Biol. 15, e8871 (2019).
Kügler, S. et al. The X-linked inhibitor of apoptosis (XIAP) prevents cell death in axotomized CNS neurons in vivo. Cell Death Differ. 7, 815–824 (2000).
Schulz, A. et al. The stress-responsive gene GDPGP1/mcp-1 regulates neuronal glycogen metabolism and survival. J. Cell Biol. 219, e201807127 (2020).
Chen, X. et al. Antiapoptotic and trophic effects of dominant-negative forms of dual leucine zipper kinase in dopamine neurons of the substantia nigra in vivo. J. Neurosci. 28, 672–680 (2008).
Smith, C. D. et al. Excess brain protein oxidation and enzyme dysfunction in normal aging and in Alzheimer disease. Proc. Natl Acad. Sci. USA 88, 10540–10543 (1991).
Hensley, K. et al. Brain regional correspondence between Alzheimer’s disease histopathology and biomarkers of protein oxidation. J. Neurochem. 65, 2146–2156 (1995).
Yao, Y. et al. Enhanced brain levels of 8,12-iso-iPF2α-VI differentiate AD from frontotemporal dementia. Neurology 61, 475–478 (2003).
Dalfó, E. et al. Evidence of oxidative stress in the neocortex in incidental Lewy body disease. J. Neuropathol. Exp. Neurol. 64, 816–830 (2005).
Subbarao, K. V., Richardson, J. S. & Ang, L. C. Autopsy samples of Alzheimer’s cortex show increased peroxidation in vitro. J. Neurochem. 55, 342–345 (1990).
Yang, W. S. et al. Regulation of ferroptotic cancer cell death by GPX4. Cell 156, 317–331 (2014).
Wang, C. et al. Elevated p62/SQSTM1 determines the fate of autophagy-deficient neural stem cells by increasing superoxide. J. Cell Biol. 212, 545–560 (2016).
Taguchi, K. et al. Keap1 degradation by autophagy for the maintenance of redox homeostasis. Proc. Natl Acad. Sci. USA 109, 13561–13566 (2012).
Reho, J. J., Guo, D.-F. & Rahmouni, K. Mechanistic target of rapamycin complex 1 signaling modulates vascular endothelial function through reactive oxygen species. J. Am. Heart Assoc. 8, e010662 (2019).
Delgado-Camprubi, M., Esteras, N., Soutar, M. P., Plun-Favreau, H. & Abramov, A. Y. Deficiency of Parkinson’s disease-related gene Fbxo7 is associated with impaired mitochondrial metabolism by PARP activation. Cell Death Differ. 24, 120–131 (2017).
Doll, S. et al. ACSL4 dictates ferroptosis sensitivity by shaping cellular lipid composition. Nat. Chem. Biol. 13, 91–98 (2017).
Yuan, H., Li, X., Zhang, X., Kang, R. & Tang, D. Identification of ACSL4 as a biomarker and contributor of ferroptosis. Biochem. Biophys. Res. Commun. 478, 1338–1343 (2016).
Ishimoto, T. et al. CD44 variant regulates redox status in cancer cells by stabilizing the xCT subunit of system xc− and thereby promotes tumor growth. Cancer Cell 19, 387–400 (2011).
Weber, R. A. et al. Maintaining iron homeostasis is the key role of lysosomal acidity for cell proliferation. Mol. Cell 77, 645–655 (2020).
Haack, T. B. et al. Exome sequencing reveals de novo WDR45 mutations causing a phenotypically distinct, X-linked dominant form of NBIA. Am. J. Hum. Genet. 91, 1144–1149 (2012).
Diaz-Font, A. et al. A mutation within the saposin D domain in a Gaucher disease patient with normal glucocerebrosidase activity. Hum. Genet. 117, 275–277 (2005).
Radha Rama Devi, A. et al. Acute Gaucher disease-like condition in an Indian infant with a novel biallelic mutation in the prosaposin gene. J. Pediatr. Genet. 8, 81–85 (2019).
Spiegel, R. et al. A mutation in the saposin A coding region of the prosaposin gene in an infant presenting as Krabbe disease: first report of saposin A deficiency in humans. Mol. Genet. Metab. 84, 160–166 (2005).
Oji, Y. et al. Variants in saposin D domain of prosaposin gene linked to Parkinson’s disease. Brain 143, 1190–1205 (2020).
Jain, I. H. et al. Genetic screen for cell fitness in high or low oxygen highlights mitochondrial and lipid metabolism. Cell 181, 716–727(2020).
Bartz, F. et al. Identification of cholesterol-regulating genes by targeted RNAi screening. Cell Metab. 10, 63–75 (2009).
Puri, V. et al. Sphingolipid storage induces accumulation of intracellular cholesterol by stimulating SREBP-1 cleavage. J. Biol. Chem. 278, 20961–20970 (2003).
Glaros, E. N. et al. Glycosphingolipid accumulation inhibits cholesterol efflux via the ABCA1/apolipoprotein A-I pathway: 1-phenyl-2-decanoylamino-3-morpholino-1-propanol is a novel cholesterol efflux accelerator. J. Biol. Chem. 280, 24515–24523 (2005).
Brunk, U. T., Jones, C. B. & Sohal, R. S. A novel hypothesis of lipofuscinogenesis and cellular aging based on interactions between oxidative stress and autophagocytosis. Mutat. Res. 275, 395–403 (1992).
Ross, D. & Siegel, D. Functions of NQO1 in cellular protection and Coq10 metabolism and its potential role as a redox sensitive molecular switch. Front. Physiol. 8, 595 (2017).
Asher, G., Lotem, J., Kama, R., Sachs, L. & Shaul, Y. NQO1 stabilizes p53 through a distinct pathway. Proc. Natl Acad. Sci. USA 99, 3099–3104 (2002).
Ramsey, C. P. et al. Expression of Nrf2 in neurodegenerative diseases. J. Neuropathol. Exp. Neurol. 66, 75–85 (2007).
Imaizumi, Y. et al. Mitochondrial dysfunction associated with increased oxidative stress and α-synuclein accumulation in PARK2 iPSC-derived neurons and postmortem brain tissue. Mol. Brain 5, 35 (2012).
Tsherniak, A. et al. Defining a cancer dependency map. Cell 170, 564–576 (2017).
Oughtred, R. et al. The BioGRID database: a comprehensive biomedical resource of curated protein, genetic, and chemical interactions. Protein Sci. 30, 187–200 (2021).
Cui, Y. et al. CRISP-view: a database of functional genetic screens spanning multiple phenotypes. Nucleic Acids Res. 49, D848–D854 (2021).
Guo, X. et al. Mitochondrial stress is relayed to the cytosol by an OMA1-DELE1-HRI pathway. Nature 579, 427–432 (2020).
Topol, A., Tran, N. N. & Brennand, K. J. A guide to generating and using hiPSC derived NPCs for the study of neurological diseases. J. Vis. Exp. e52495 (2015).
Cheng, C., Fass, D. M., Folz-Donahue, K., MacDonald, M. E. & Haggarty, S. J. Highly expandable human iPS cell-derived neural progenitor cells (NPC) and neurons for central nervous system disease modeling and high-throughput screening. Curr. Protoc. Hum. Genet 92, 21.8.1–21.8.21 (2017).
Tcw, J. et al. An efficient platform for astrocyte differentiation from human induced pluripotent stem cells. Stem Cell Rep. 9, 600–614 (2017).
Brownjohn, P. W. et al. Functional studies of missense TREM2 mutations in human stem cell-derived microglia. Stem Cell Rep. 10, 1294–1307 (2018).
Piñero, J. et al. DisGeNET: a discovery platform for the dynamical exploration of human diseases and their genes. Database (Oxf.) 2015, bav028 (2015).
Bersuker, K. et al. The CoQ oxidoreductase FSP1 acts parallel to GPX4 to inhibit ferroptosis. Nature 575, 688–692 (2019).
Komarov, P. G. et al. A chemical inhibitor of p53 that protects mice from the side effects of cancer therapy. Science 285, 1733–1737 (1999).
Zhao, C. et al. Dual regulatory switch through interactions of Tcf7l2/Tcf4 with stage-specific partners propels oligodendroglial maturation. Nat. Commun. 7, 10883 (2016).
Hauser, M. et al. The spectrin-actin-based periodic cytoskeleton as a conserved nanoscale scaffold and ruler of the neural stem cell lineage. Cell Rep. 24, 1512–1522 (2018).
Wojcik, M., Hauser, M., Li, W., Moon, S. & Xu, K. Graphene-enabled electron microscopy and correlated super-resolution microscopy of wet cells. Nat. Commun. 6, 7384 (2015).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Hill, A. J. et al. On the design of CRISPR-based single-cell molecular screens. Nat. Methods 15, 271–274 (2018).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Adamson, B. et al. A multiplexed single-cell CRISPR screening platform enables systematic dissection of the unfolded protein response. Cell 167, 1867–1882 (2016).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Soneson, C., Love, M. I. & Robinson, M. D. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000Res. 4, 1521 (2015).
Pang, X. P., Hershman, J. M., Chung, M. & Pekary, A. E. Characterization of tumor necrosis factor-α receptors in human and rat thyroid cells and regulation of the receptors by thyrotropin. Endocrinology 125, 1783–1788 (1989).
Liao, Y., Wang, J., Jaehnig, E. J., Shi, Z. & Zhang, B. WebGestalt 2019: gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Res. 47, W199–W205 (2019).
Acknowledgements
We thank L. Gan and B. Conklin for support and advice; P. Kennedy and A. Nummy at Cayman Chemical for untargeted lipidomics; E. Chow (UCSF), D. Bogdanoff (UCSF), A. Detweiler (CZI Biohub), N. Neff (CZI Biohub) and M. Tan (CZI Biohub) for next-generation sequencing; A. Samelson and X. Guo for comments on this manuscript; and E. Li and J. Olzmann for discussions. We thank the staff at the University of California, Berkeley Electron Microscope Laboratory for advice and assistance in electron microscopy sample preparation and data collection. This research was supported by the Intramural Research Program of the NIH/NINDS, an NIH Director’s New Innovator Award (NIH/ NIGMS DP2 GM119139 to M.K.), NIH/NIA grants (R01 AG062359 and R56 AG057528 to M.K. and F30 AG066418 to K.L.), the NINDS Tau Center Without Walls (NIH/NINDS U54 NS100717 to M.K.), an Allen Distinguished Investigator Award (Paul G. Allen Family Foundation) to M.K., a Chan Zuckerberg Biohub Investigator Award (to M.K.) and a Tau Consortium Investigator Award (Rainwater Charitable Foundation) to M.K. K.X. is a Chan Zuckerberg Biohub investigator and acknowledges support from the National Institute of General Medical Sciences of the National Institutes of Health (DP2GM132681). The participation of A.B.S. was supported, in part, by the Intramural Research Program of the National Institute on Aging, National Institutes of Health, part of the Department of Health and Human Services (project number ZIA AG000957-16). CRISPRbrain development was supported, in part, by a collaboration among the Kampmann Lab, UCSF and Data Tecnica International, LLC.
Author information
Authors and Affiliations
Contributions
R.T. and M.K. conceived this study and wrote the manuscript with input from the other authors. R.T. designed and conducted experiments with help from A.A. and J.H. and guidance from M.K. R.T. performed data analyses. R.Y. performed STORM imaging with guidance from K.X. N.D. generated iPSC-derived microglia, and K.L. generated iPSC-derived astrocytes. S.H.H., M.A.N. and F.F. developed the CRISPRbrain data commons with critical input from R.T. and M.K. and feedback from A.B.S. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
M.K. has filed a patent application related to CRISPRi and CRISPRa screening (PCT/US15/40449) and serves on the Scientific Advisory Boards of Engine Biosciences, Casma Therapeutics and Cajal Neuroscience. The remaining authors declare no competing financial interests.
Additional information
Peer review information Nature Neuroscience thanks Kristen Brennand and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Karyotyping of the monoclonal CRISPRa-iPSC line, and analysis of CRISPRi and CRISPRa hits.
(a) A normal karyotype was confirmed for the monoclonal CRISPRa-iPSC line. (b,c) Comparison of CRISPRi (b) and CRISPRa (c) efficacy in iPSCs and iPSC-derived neurons. The relative mRNA level of each targeted gene was calculated as the ratio of its expression in cells expressing a targeting sgRNA as compared to a non-targeting control sgRNA measured by qPCR (mean +/s sd, n = 3 technical replicates). The housekeeping gene ACTB was used for normalization. (d,e) Top, heatmaps showing phenotype scores (Log2-fold change) of all 5 sgRNAs (x-axis) targeting each hit gene (y-axis) from the primary CRISPRi (left) and CRISPRa (right) survival screens. The five sgRNAs targeting a given gene are ranked by the significance of their P values and are shown from left to right. Bottom, bar graphs summarizing the percentage of hit genes that have a certain number of sgRNAs (x-axis) showing a significant phenotype (false discovery rate (FDR) < 0.1; P values were calculated by α-RRA in the MAGeCK pipeline and FDR values were calculated using the Benjamini-Hochberg method to adjust for multiple comparisons) in CRISPRi (left) and CRISPRa (right) survival screens. (f,g) Scatter plots showing the relationship between Gene Score and gene coding sequence (CDS) length (left) or gene length (right) for genome-wide CRISPRi (f) and CRISPRa (g) survival screens. (h, i) Top: Venn diagrams comparing CRISPRi (h) and CRISPRa (i) screen results for neuronal survival from this paper with other published survival screens for different human cell types. For CRISPRi, hit genes with toxic phenotypes for the survival of neurons were compared with those for cancer cells (‘gold-standard’ essential genes 14) and pluripotent stem cells 11–13 (genes that were identified as essential in more than one studies were retained for comparison). Protective hits for the survival of neurons were compared with those for human pluripotent stem cells 11,12 (genes that were identified as essential in both studies were retained for comparison). For CRISPRa, hits were compared with our published survival screen in K562 cells 10 reanalyzed using our MAGeCK-iNC pipeline. Bottom: Gene Ontology (GO) term enrichment analysis was conducted for hits resulting in increased survival (red) or decreased survival (blue); terms are shown up to an FDR of 0.05. (j) Neuronal expression levels of neuron-specific hit genes and other hit genes from CRISPRi (top) and CRISPRa (bottom) screens are shown, binned by order of magnitude.
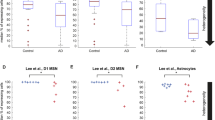
Extended Data Fig. 2 Comparing CRISPRa survival screens in +AO and -AO conditions.
Each dot represents one gene, and its Gene Score in the +AO screen was plotted on the x-axis and Gene Score in the -AO screen on the y-axis. The Pearson correlation coefficient is shown.
Extended Data Fig. 3 Characterization of PSAP KO in other cell types.
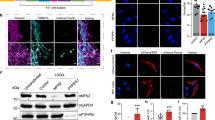
(a) qPCR validation of PSAP knockdown in neural progenitor cells (left), astrocytes (middle) and microglia (right) diffentiated from CRISPRi iPSCs expression a PSAP sgRNA as compared to a non-targeting control sgRNA (mean +/s sd, n = 3 technical replicates). The housekeeping gene ACTB was used for normalization. (b) Representative fluorescence microscopy images for neural progenitor cells (left), astrocytes (middle) and microglia (right) diffentiated from CRISPRi iPSCs expression a non-targeting sgRNA or a PSAP sgRNA, stained with LAMP2 and LC3B antibodies from 3 independent experiments. DRAQ5 was used for nuclear staining. Scale bar, 10 μm.
Extended Data Fig. 4 Examples of the CROP-seq classification method, and shared transcriptomic signatures of VPS54, PAXIP1, and PON2 knockdown in human iPSC-derived neurons.
(a,b) CROP-seq examples showing the application of the outlier detection-based classification method in cases where two sgRNAs targeting the same gene had heterogeneous efficacy (a, SOX5 in CRISPRa) or the expression level of the target gene was too low to quantify knockdown level (b, ZNF592 in CRISPRi). (c) Transcriptomic changes induced by knockdown of VPS54 (left), PAXIP1 (middle), and PON2 (right) in neurons. For each perturbation, the top 200 upregulated and downregulated genes compared to control (that is unperturbed cells) are shown in red and blue, respectively. Within this set, shared genes among all three perturbations are highlighted in green.
Supplementary information
Supplementary Information
Supplementary Fig. 1 and Supplementary Note.
Supplementary Table 1
Screen results for primary and pooled validation screens. Screens were analyzed using the MAGeCK-iNC pipeline (see Methods for details). Hit class values of 1, −1 or 0 were assigned to hit genes with positive phenotype scores, hit genes with negative phenotype scores or non-hits, respectively. P values were calculated using the Mann–Whitney U test in the MAGeCK-iNC pipeline. P values were not corrected for multiple hypothesis testing; instead, an empirical FDR was determined as described in the Methods. Each screen is provided in a separate tab.
Supplementary Table 3
Untargeted lipidomics data for WT and PSAP KO neurons. Untargeted lipidomics data for WT and PSAP KO neurons. P values were calculated using two-sided Student’s t-test and were corrected for multiple testing using the Benjamini–Hochberg method.
Supplementary Table 4
sgRNA sequences for pooled validation and CROP-seq libraries. These tables show the protospacer sequences for sgRNAs in the pooled validation and CROP-seq libraries. Each library is provided in a separate tab. sgRNA information for the genome-wide libraries was previously published10.
Supplementary Table 5
sgRNA cell counts for CROP-seq screens. These tables summarize the number of cells for sgRNAs in the CRISPRi and CRISPRa CROP-seq screens. First tab: CRISPRi; second tab: CRISPRa.
Source data
Source Data Fig. 3
Unprocessed western blots for Fig. 3c.
Source Data Fig. 5
Unprocessed western blots for Fig. 5l.
Source Data Fig. 7
Unprocessed western blots for Fig. 7e.
Rights and permissions
About this article
Cite this article
Tian, R., Abarientos, A., Hong, J. et al. Genome-wide CRISPRi/a screens in human neurons link lysosomal failure to ferroptosis. Nat Neurosci 24, 1020–1034 (2021). https://doi.org/10.1038/s41593-021-00862-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-021-00862-0
- Springer Nature America, Inc.
This article is cited by
-
Compact CRISPR genetic screens enabled by improved guide RNA library cloning
Genome Biology (2024)
-
Identification of genetic modifiers enhancing B7-H3-targeting CAR T cell therapy against glioblastoma through large-scale CRISPRi screening
Journal of Experimental & Clinical Cancer Research (2024)
-
An integrated toolkit for human microglia functional genomics
Stem Cell Research & Therapy (2024)
-
High-resolution genome-wide mapping of chromosome-arm-scale truncations induced by CRISPR–Cas9 editing
Nature Genetics (2024)
-
Bevacizumab induces ferroptosis and enhances CD8+ T cell immune activity in liver cancer via modulating HAT1 and increasing IL-9
Acta Pharmacologica Sinica (2024)