Abstract
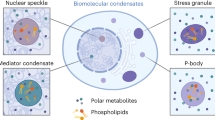
Diverse mechanisms have been described for selective enrichment of biomolecules in membrane-bound organelles, but less is known about mechanisms by which molecules are selectively incorporated into biomolecular assemblies such as condensates that lack surrounding membranes. The chemical environments within condensates may differ from those outside these bodies, and if these differed among various types of condensate, then the different solvation environments would provide a mechanism for selective distribution among these intracellular bodies. Here we use small molecule probes to show that different condensates have distinct chemical solvating properties and that selective partitioning of probes in condensates can be predicted with deep learning approaches. Our results demonstrate that different condensates harbor distinct chemical environments that influence the distribution of molecules, show that clues to condensate chemical grammar can be ascertained by machine learning and suggest approaches to facilitate development of small molecule therapeutics with optimal subcellular distribution and therapeutic benefit.






Similar content being viewed by others
Data availability
Source and probe partitioning screen data are available at https://doi.org/10.6084/m9.figshare.23736693, https://doi.org/10.6084/m9.figshare.23535258.
Code availability
Code is available on GitHub: machine learning tools, https://github.com/pgmikhael/ChemicalGrammar; in vitro droplet assays, https://github.com/jehenninger/in_vitro_droplet_assay; RDKit Calculations, https://github.com/hrkilgore/rdkit_scripts and PLSR Model, https://github.com/uberholzer/partitioning_PLSR.
Change history
25 October 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41589-023-01491-3
References
Banani, S. F., Lee, H. O., Hyman, A. A. & Rosen, M. K. Biomolecular condensates: organizers of cellular biochemistry. Nat. Rev. Mol. Cell Biol. 18, 285–285 (2017).
Alberts, B. et al. Molecular Biology of the Cell 7th edn (W. W. Norton & Company, 2022).
Klein, I. A. et al. Partitioning of cancer therapeutics in nuclear condensates. Science 368, 1386–1392 (2020).
Kilgore, H. R. & Young, R. A. Learning the chemical grammar of biomolecular condensates. Nat. Chem. Biol. 18, 1298–1306 (2022).
Shin, Y. & Brangwynne, C. P. Liquid phase condensation in cell physiology and disease. Science 357, eaaf4382 (2017).
Langdon, E. M. & Gladfelter, A. S. A new lens for RNA localization: liquid-liquid phase separation. Annu. Rev. Microbiol. 72, 255–271 (2018).
Lyon, A. S., Peeples, W. B. & Rosen, M. K. A framework for understanding the functions of biomolecular condensates across scales. Nat. Rev. Mol. Cell Biol. 22, 215–235 (2021).
Alberti, S. & Hyman, A. A. Biomolecular condensates at the nexus of cellular stress, protein aggregation disease and ageing. Nat. Rev. Mol. Cell Biol. 22, 196–213 (2021).
Feric, M. & Misteli, T. Phase separation in genome organization across evolution. Trends Cell Biol. 31, 671–685 (2021).
Forman-Kay, J. D., Ditlev, J. A., Nosella, M. L. & Lee, H. O. What are the distinguishing features and size requirements of biomolecular condensates and their implications for RNA-containing condensates? RNA 28, 36–47 (2022).
Du, M. & Zhijian, Chen J. DNA-induced liquid phase condensation of cGAS activates innate immune signaling. Science 361, 704–709 (2018).
Howard, T. P. & Roberts, C. W. M. Partitioning of chemotherapeutics into nuclear condensates-opening the door to new approaches for drug development. Mol. Cell 79, 544–545 (2020).
Lyons, H. et al. Functional partitioning of transcriptional regulators by patterned charge blocks. Cell 186, 327–345.e328 (2023).
Sabari, B. R. et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 361, eaar3958 (2018).
Larson, A. G. et al. Liquid droplet formation by HP1α suggests a role for phase separation in heterochromatin. Nature 547, 236–240 (2017).
Patel, A. et al. A liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell 162, 1066–1077 (2015).
Rogers, D. & Hahn, M. Extended-connectivity fingerprints. J. Chem. Inf. Model. 50, 742–754 (2010).
Mater, A. C. & Coote, M. L. Deep learning in chemistry. J. Chem. Inf. Model. 59, 2545–2559 (2019).
Walters, W. P. & Barzilay, R. Applications of deep learning in molecule generation and molecular property prediction. Acc. Chem. Res. 54, 263–270 (2021).
Hunter, C. A. & Sanders, J. K. M. The nature of π-π interactions. J. Am. Chem. Soc. 112, 5525–5534 (1990).
Thakuria, R., Nath, N. K. & Saha, B. K. The nature and applications of π–π interactions: a perspective. Cryst. Growth Des. 19, 523–528 (2019).
Dougherty, D. A. The cation−π interaction. Acc. Chem. Res. 46, 885–893 (2013).
Stewart, J. A., McCormack, J. J. & Krakoff, I. H. Clinical and clinical pharmacologic studies of mitoxantrone. Cancer Treat. Rep. 66, 1327–1331 (1982).
Kaur, R., Manjal, S. K., Rawal, R. K. & Kumar, K. Recent synthetic and medicinal perspectives of tryptanthrin. Bioorg. Med. Chem. 25, 4533–4552 (2017).
Wang, J. et al. A molecular grammar governing the driving forces for phase separation of prion-like RNA binding proteins. Cell 174, 688–699.e616 (2018).
Martin, E. W. et al. Valence and patterning of aromatic residues determine the phase behavior of prion-like domains. Science 367, 694–699 (2020).
Choi, J.-M., Holehouse, A. S. & Pappu, R. V. Physical principles underlying the complex biology of intracellular phase transitions. Annu. Rev. Biophys. 49, 107–133 (2020).
Landrum, G. et al. RDKit: Open-Source chemoinformatics. Zenodo https://doi.org/10.5281/zenodo.4750957 (2021).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Mevik, B.-H. & Wehrens, R. The pls package: principal component and partial least squares regression in R. J. Stat. Softw. 18, 1–23 (2007).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2022); https://www.R-project.org/
Mehmood, T., Liland, K. H., Snipen, L. & Sæbø, S. A review of variable selection methods in partial least squares regression. Chemom. Intell. Lab. Syst. 118, 62–69 (2012).
Yang, K. et al. Analyzing learned molecular representations for property prediction. J. Chem. Inf. Mod. 59, 3370–3388 (2019).
Jin, W., Barzilay, R. & Jaakkola, T. Multi-objective molecule generation using interpretable substructures. In Proc. 37th Int. Conf. Mach. Learn. (PMLR, 2020).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Acknowledgements
We thank J. Platt, J. H. Cheah, C. Soule, W. Salmon and C. Rogers for comments and discussion and A. Tubelli for graphic art. Support was provided by NIH grant nos. GM144283 (R.A.Y.) and CA155258 (R.A.Y.), NSF grant no. PHY2044895 (R.A.Y.), Damon Runyon Cancer Research Foundation Fellowship grant no. 2458-22 (H.R.K.), NSF Graduate Research Fellowship grant no. 1745302 (K.J.O.), the MIT Jameel Clinic for Machine Learning in Health (R.B., P.G.M.) and Eric and Wendy Schmidt Center, Broad Institute (P.G.M.), Basic Science Research Institute Fund grant no. 2021R1A6A1A10042944 (Y.-T.C.) and a National Research Foundation of Korea grant funded by the Korean government (MSIT) (no. 2023R1A2C300453411 to Y.-T.C.).
Author information
Authors and Affiliations
Contributions
H.R.K., R.B., A.B. and R.A.Y. conceived the idea. Y.-T.C. provided the fluorescent probes. H.R.K., K.J.O., A.B., N.M.H. and C.V.D. performed the experiments. H.R.K., P.G.M. and K.J.O. analyzed the data. H.R.K., P.G.M., K.J.O, T.I.L. and R.A.Y. prepared the manuscript.
Corresponding authors
Ethics declarations
Competing interests
R.A.Y. is a founder and shareholder of Syros Pharmaceuticals, Camp4 Therapeutics, Omega Therapeutics, Dewpoint Therapeutics and Paratus Sciences, and has consulting or advisory roles at Precede Biosciences and Novo Nordisk. R.B. has consulting or advisory roles at Dewpoint Therapeutics, J&J, Amgen, Outcomes4Me, Immunai and Firmenich. H.R.K. is a consultant of Dewpoint Therapeutics. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Live cell confocal and two-photon imaging of endogenously fluorescent drugs.
HCT-116 cells were incubated with a drug or natural product at 50 μM for 1 hour and then imaged with a confocal or two-photon microscope. Image for sunitinib is also shown in Fig. 1. Scale: 10 μm.
Extended Data Fig. 2 Subcellular distribution of small molecules in a breast cancer cell line.
Live MCF7 cells were incubated with a drug or natural product at 50 μM for 1 hour prior to confocal imaging. Scale: 10 μm R = CH2CH2OCH2CH2NH2, R2 = CH2CH2N(CH2CH3)2.
Extended Data Fig. 3 Subcellular distribution of small molecules in a prostate cancer cell line.
Live PC3 cells were incubated with a drug or natural product at 50 μM for 1 hour prior to confocal imaging. Scale: 10 μm. R = CH2CH2OCH2CH2NH2, R2 = CH2CH2N(CH2CH3)2.
Extended Data Fig. 4 Additional analysis of condensate selectivity in fluorescent probe partitioning.
(a-c) Dot plots comparing the partition ratios of probes above the 90th percentile in, a, MED1, b, NPM1, and c, HP1α against their partition ratios in other condensates. P values computed with a two-sided unpaired t-test, n = 50 and d.f.=98 for all groups, with t-statistics and effect sizes (η2) as follows: MED1-NPM1 (t = 22.87, η2 = 0.84), MED1- HP1α (t = 19.88, η2 = 0.80), NPM1-MED1 (t = 3.43, η2 = 0.11), NPM1- HP1α (t = 8.24, η2 = 0.41), HP1α -MED1 (t = 3.15, η2 = 0.09), HP1α -NPM1 (t = 4.09, η2 = 0.15). (d-f) Dot plots comparing the partition ratios of probes below the 10th percentile in d, MED1, e, NPM1, and f, HP1α against their partition ratios in other condensates. P values computed with a two-sided unpaired t-test, n = 50 and d.f.=98 for all groups, with test statistics as follows: MED1-NPM1 (t = 3.79, η2 = 0.13), MED1- HP1α (t = 4.65, η2 = 0.18), NPM1-MED1 (t = 3.12, η2 = 0.09), NPM1- HP1α (t = 2.60, η2 = 0.07), HP1α -MED1 (t = 3.76, η2 = 0.13), HP1α -NPM1 (t = 4.20, η2 = 0.15). (g-i) Dot plots comparing the percentile ranks of probes below the 10th percentile in g, MED1, h, NPM1, and i, HP1α against their percentiles in other condensates. P values computed with a two-sided Wilcoxon matched-pairs signed rank test, n = 50 for all groups, test statistic |W | (left to right): 915, 1003, 928, 1137, 1099, 681. For panels a-i, centerline and error bars represent mean ± s.d., P values were not adjusted for multiple comparisons, sample size n = 100 probes.
Extended Data Fig. 5 Probe features suggest a chemical grammar in condensates.
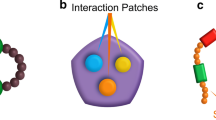
a, Cartoon depicting how similar molecules (here, sharing color) might interact with the same chemical environment. b, Schematic showing calculation of Tanimoto similarity matrices comparing fluorescent probes by their Morgan Fingerprints. c, Schematic and d, dot plots showing calculation of mean Tanimoto similarities from matrices of fluorescent probes compared against each other in high-to-high (H-H), high-to-low (H-L) and low-to-low (L-L) partitioning regions. e, Graphic and f, dot plots show the comparison of high partitioning probes between condensates through quantification of matrices, significance between groups was not assessed. Centerline and error bars represent mean ± s.d. Panel d, all comparisons were statistically significant with P value, P < 0.0001 (asterisks do not appear in figure), sample size MED1 n = 120. NPM1 n = 100. HP1α n = 100, without adjustment for multiple comparisons. Unpaired two-sided t-test statistic and degrees of freedom: MED1 H-H, t = 9.5, df = 238. MED1 H-L, t = 12.7, df = 238. MED1 L-L, t = 7.3, df = 238. NPM1 H-H, t = 12.17, df = 198. NPM1 H-L, t = 7.4, df = 198. NPM1 L-L, t = 9.4, df = 198. HP1α H-H, t = 4.8, df = 198. HP1α H-L, t = 10.7, df = 198. HP1α L-L, t = 8.3, df = 198.
Extended Data Fig. 6 Comparison of deep learning and Tanimoto similarity approaches toward predicting partitioning behaviors.
Receiver operating characteristic curves quantifying the success of deep learning and Tanimoto similarity approaches for determining partitioning behavior of small molecules into condensates. a, Deep learning-based approach to probe classification (compounds were considered to have partitioned if probes concentrated above 2.7 for MED1, 2.7 for NPM1, and 2.0 for HP1α). Tanimoto similarity approach to probe classification in b, MED1, c, NPM1, and d, HP1α condensates across different thresholds of Tanimoto similarity (compounds were considered to have concentrated into a condensate if probe partition ratio was K > 2.0 for MED1, NPM1, and HP1α).
Extended Data Fig. 8 Receiver operating characteristic curves comparing performance of Tanimoto similarity and deep learning classifiers on in vivo compounds.
a, Performance of NPM1 deep learning model (AUC-ROC = 0.62) and Tanimoto similarity (AUC-ROC = 0.52) at identifying drugs and natural products that concentrate in the nucleolus. b, Performance of HP1α deep learning model (AUC-ROC = 0.59) and Tanimoto similarity (AUC-ROC = 0.52) at identifying drugs and natural products that concentrate in chromocenters.
Extended Data Fig. 9 Live cell images showing the partitioning of small molecules into nuclear compartments.
a, Micrograph and line plot showing the signal intensity from mitoxantrone along that indicated gray arrow in HCT-116 cells. b, Mouse embryonic stem cells stained with the DNA dye Hoechst. (a) chromocenter, (b) perinuclear heterochromatin, (c) nucleolus. Zoom (2x). c, Micrograph and line plot showing the signal intensity from tryptanthrin along the indicated gray arrow in mouse embryonic stem cells. Scale: 10 μm. Images were recorded after 1 hour of incubation for tryptanthrin and mitoxantrone, and 10 minutes for Hoechst.
Supplementary information
Supplementary Information
Supplementary Figs. 1–12 and Tables 1–3.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kilgore, H.R., Mikhael, P.G., Overholt, K.J. et al. Distinct chemical environments in biomolecular condensates. Nat Chem Biol 20, 291–301 (2024). https://doi.org/10.1038/s41589-023-01432-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-023-01432-0
- Springer Nature America, Inc.
This article is cited by
-
MYC phase separation selectively modulates the transcriptome
Nature Structural & Molecular Biology (2024)
-
In search of chemical rationales
Nature Chemical Biology (2024)