Abstract
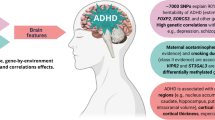
Attention deficit hyperactivity disorder (ADHD) is a complex disorder that manifests variability in long-term outcomes and clinical presentations. The genetic contributions to such heterogeneity are not well understood. Here we show several genetic links to clinical heterogeneity in ADHD in a case-only study of 14,084 diagnosed individuals. First, we identify one genome-wide significant locus by comparing cases with ADHD and autism spectrum disorder (ASD) to cases with ADHD but not ASD. Second, we show that cases with ASD and ADHD, substance use disorder and ADHD, or first diagnosed with ADHD in adulthood have unique polygenic score (PGS) profiles that distinguish them from complementary case subgroups and controls. Finally, a PGS for an ASD diagnosis in ADHD cases predicted cognitive performance in an independent developmental cohort. Our approach uncovered evidence of genetic heterogeneity in ADHD, helping us to understand its etiology and providing a model for studies of other disorders.





Similar content being viewed by others
Data availability
The consent structure of iPSYCH and Danish law prevent individual genotype and phenotype data from being shared publicly. Reasonable requests to access individual-level data to verify the findings in this article can be accommodated with permission from the Danish Scientific Ethics Committee, the Danish Health Data Authority, the Danish Data Protection Agency and the Danish Neonatal Screening Biobank Steering Committee; interested parties can contract T.M.W. and expect a response within 1 week. Approvals for access can take several months and are governed by strict data use agreements. The ABCD study data can be accessed, by request, from the NIMH Data Archive (https://nda.nih.gov/abcd). The GWAS summary statistics used in this work were downloaded from and are available in public repositories as described in Supplementary Table 28. Leave-one-study-out meta-analysis summary statistics for psychiatric disorders are available upon request from the Psychiatric Genomics Consortium Disorder Working Group chairs (https://pgc.unc.edu/for-researchers/data-access-committee/data-access-information/). The eQTL and sQTL visualizations and the data used for the analyses described in this article were obtained from the GTEx portal (https://gtexportal.org/) on 10 March 2021 and 23 October 2023.
Code availability
The code for the multinomial regression tests and supplementary simulations is available at https://github.com/AndrewSchork/. Other software used for the analyses are publicly available as described in the Methods and Reporting Summary. The wrappers and pipelines used to link tools with individual-level data are available on request; interested parties should contact A.J.S.
References
American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders 5th edn (CBS Publishers & Distributors, 2017).
World Health Organization. The ICD-10 Classification of Mental and Behavioural Disorders: Clinical Descriptions and Diagnostic Guidelines (World Health Organization, 1992).
World Health Organization. The ICD-10 Classification of Mental and Behavioural Disorders: Diagnostic Criteria for Research (World Health Organization, 1993).
Franke, B. et al. Live fast, die young? A review on the developmental trajectories of ADHD across the lifespan. Eur. Neuropsychopharmacol. 28, 1059–1088 (2018).
Luo, Y., Weibman, D., Halperin, J. M. & Li, X. A review of heterogeneity in attention deficit/hyperactivity disorder (ADHD). Front. Hum. Neurosci. 13, 42 (2019).
Thapar, A., Cooper, M. & Rutter, M. Neurodevelopmental disorders. Lancet Psychiatry 4, 339–346 (2017).
Dalsgaard, S., Mortensen, P. B., Frydenberg, M. & Thomsen, P. H. Long-term criminal outcome of children with attention deficit hyperactivity disorder. Crim. Behav. Ment. Health 23, 86–98 (2013).
Dalsgaard, S., Østergaard, S. D., Leckman, J. F., Mortensen, P. B. & Pedersen, M. G. Mortality in children, adolescents, and adults with attention deficit hyperactivity disorder: a nationwide cohort study. Lancet 385, 2190–2196 (2015).
Dalsgaard, S. et al. Association of mental disorder in childhood and adolescence with subsequent educational achievement. JAMA Psychiatry 77, 797–805 (2020).
Daley, D., Jacobsen, R. H., Lange, A.-M., Sørensen, A. & Walldorf, J. Costing Adult Attention Deficit Hyperactivity Disorder: Impact on the Individual and Society (OUP, 2015).
Plana-Ripoll, O. et al. Exploring comorbidity within mental disorders among a Danish national population. JAMA Psychiatry 76, 259–270 (2019).
McClellan, J. & King, M.-C. Genetic heterogeneity in human disease. Cell 141, 210–217 (2010).
Dahl, A. & Zaitlen, N. Genetic influences on disease subtypes. Annu. Rev. Genomics Hum. Genet. 21, 413–435 (2020).
Charney, A. W. et al. Evidence for genetic heterogeneity between clinical subtypes of bipolar disorder. Transl. Psychiatry 7, e993 (2017).
Ruderfer, D. M. et al. Genomic dissection of bipolar disorder and schizophrenia, including 28 subphenotypes. Cell 173, 1705–1715 (2018).
Faraone, S. V. & Larsson, H. Genetics of attention deficit hyperactivity disorder. Mol. Psychiatry 24, 562–575 (2019).
Williams, N. M. et al. Rare chromosomal deletions and duplications in attention-deficit hyperactivity disorder: a genome-wide analysis. Lancet 376, 1401–1408 (2010).
Olsen, L. et al. Prevalence of rearrangements in the 22q11.2 region and population-based risk of neuropsychiatric and developmental disorders in a Danish population: a case-cohort study. Lancet Psychiatry 5, 573–580 (2018).
Satterstrom, F. K. et al. Autism spectrum disorder and attention deficit hyperactivity disorder have a similar burden of rare protein-truncating variants. Nat. Neurosci. 22, 1961–1965 (2019).
Demontis, D. et al. Discovery of the first genome-wide significant risk loci for attention deficit/hyperactivity disorder. Nat. Genet. 51, 63–75 (2019).
Sullivan, P. F. et al. Psychiatric genomics: an update and an agenda. Am. J. Psychiatry 175, 15–27 (2018).
Sullivan, P. F. & Geschwind, D. H. Defining the genetic, genomic, cellular, and diagnostic architectures of psychiatric disorders. Cell 177, 162–183 (2019).
Cai, N. et al. Minimal phenotyping yields genome-wide association signals of low specificity for major depression. Nat. Genet. 52, 437–447 (2020).
Wray, N. R., Lee, S. H. & Kendler, K. S. Impact of diagnostic misclassification on estimation of genetic correlations using genome-wide genotypes. Eur. J. Hum. Genet. 20, 668–674 (2012).
Wimberley, T. et al. Genetic liability to ADHD and substance use disorders in individuals with ADHD. Addiction 115, 1368–1377 (2020).
Jansen, A. G. et al. Psychiatric polygenic risk scores as predictor for attention deficit/hyperactivity disorder and autism spectrum disorder in a clinical child and adolescent sample. Behav. Genet. 50, 203–212 (2020).
Martin, J. et al. A genetic investigation of sex bias in the prevalence of attention-deficit/hyperactivity disorder. Biol. Psychiatry 83, 1044–1053 (2018).
Rovira, P. et al. Shared genetic background between children and adults with attention deficit/hyperactivity disorder. Neuropsychopharmacology 45, 1617–1626 (2020).
Liley, J., Todd, J. A. & Wallace, C. A method for identifying genetic heterogeneity within phenotypically defined disease subgroups. Nat. Genet. 49, 310–316 (2017).
Nadeau, J. H. Modifier genes in mice and humans. Nat. Rev. Genet. 2, 165–174 (2001).
Fanous, A. H. & Kendler, K. S. Genetic heterogeneity, modifier genes, and quantitative phenotypes in psychiatric illness: searching for a framework. Mol. Psychiatry 10, 6–13 (2005).
Pedersen, C. B. et al. The iPSYCH2012 case-cohort sample: new directions for unravelling genetic and environmental architectures of severe mental disorders. Mol. Psychiatry 23, 6–14 (2018).
Kildemoes, H. W., Sørensen, H. T. & Hallas, J. The Danish National Prescription Registry. Scand. J. Public Health 39, 38–41 (2011).
Lynge, E., Sandegaard, J. L. & Rebolj, M. The Danish National Patient Register. Scand. J. Public Health 39, 30–33 (2011).
Mors, O., Perto, G. P. & Mortensen, P. B. The Danish Psychiatric Central Research Register. Scand. J. Public Health 39, 54–57 (2011).
Pedersen, C. B. The Danish Civil Registration System. Scand. J. Public Health 39, 22–25 (2011).
Grove, J. et al. Identification of common genetic risk variants for autism spectrum disorder. Nat. Genet. 51, 431–444 (2019).
Lee, P. H. et al. Genomic relationships, novel loci, and pleiotropic mechanisms across eight psychiatric disorders. Cell 179, 1469–1482 (2019).
Wang, D. et al. Comprehensive functional genomic resource and integrative model for the human brain. Science 362, eaat8464 (2018).
Walker, R. L. et al. Genetic control of expression and splicing in developing human brain informs disease mechanisms. Cell 179, 750–771 (2019).
Lonsdale, J. et al. The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Mittelstaedt, T. & Schoch, S. Structure and evolution of RIM-BP genes: identification of a novel family member. Gene 403, 70–79 (2007).
Hibino, H. et al. RIM binding proteins (RBPs) couple Rab3-interacting molecules (RIMs) to voltage-gated Ca2+ channels. Neuron 34, 411–423 (2002).
Acuna, C., Liu, X., Gonzalez, A. & Südhof, T. C. RIM-BPs mediate tight coupling of action potentials to Ca2+-triggered neurotransmitter release. Neuron 87, 1234–1247 (2015).
Bucan, M. et al. Genome-wide analyses of exonic copy number variants in a family-based study point to novel autism susceptibility genes. PLoS Genet. 5, e1000536 (2009).
Barch, D. M. et al. Demographic, physical and mental health assessments in the adolescent brain and cognitive development study: rationale and description. Dev. Cogn. Neurosci. 32, 55–66 (2018).
Epstein, J. N. & Loren, R. E. A. Changes in the definition of ADHD in DSM-5: subtle but important. Neuropsychiatry 3, 455–458 (2013).
Xu, G., Strathearn, L., Liu, B., Yang, B. & Bao, W. Twenty-year trends in diagnosed attention-deficit/hyperactivity disorder among US children and adolescents, 1997–2016. JAMA Netw. Open 1, e181471 (2018).
Martin, J. et al. Biological overlap of attention-deficit/hyperactivity disorder and autism spectrum disorder: evidence from copy number variants. J. Am. Acad. Child Adolesc. Psychiatry 53, 761–770 (2014).
LaBianca, S. et al. Brief report: clusters and trajectories across the autism and/or ADHD spectrum. J. Autism Dev. Disord. 48, 3629–3636 (2018).
Ghirardi, L. et al. The familial co-aggregation of ASD and ADHD: a register-based cohort study. Mol. Psychiatry 23, 257–262 (2018).
Rommelse, N. N. J., Franke, B., Geurts, H. M., Hartman, C. A. & Buitelaar, J. K. Shared heritability of attention-deficit/hyperactivity disorder and autism spectrum disorder. Eur. Child Adolesc. Psychiatry 19, 281–295 (2010).
Mattheisen, M. et al. Identification of shared and differentiating genetic architecture for autism spectrum disorder, attention-deficit hyperactivity disorder and case subgroups. Nat. Genet. 54, 1470–1478 (2022).
Peyrot, W. J. & Price, A. L. Identifying loci with different allele frequencies among cases of eight psychiatric disorders using CC-GWAS. Nat. Genet. 53, 445–454 (2021).
Young, S. et al. Guidance for identification and treatment of individuals with attention deficit/hyperactivity disorder and autism spectrum disorder based upon expert consensus. BMC Med. 18, 146 (2020).
Pinto, R., Rijsdijk, F., Ronald, A., Asherson, P. & Kuntsi, J. The genetic overlap of attention-deficit/hyperactivity disorder and autistic-like traits: an investigation of individual symptom scales and cognitive markers. J. Abnorm. Child Psychol. 44, 335–345 (2016).
Panagiotidi, M., Overton, P. G. & Stafford, T. Co-occurrence of ASD and ADHD traits in an adult population. J. Atten. Disord. 23, 1407–1415 (2019).
Aoki, Y. et al. Association of white matter structure with autism spectrum disorder and attention-deficit/hyperactivity disorder. JAMA Psychiatry 74, 1120–1128 (2017).
Asherson, P. & Agnew-Blais, J. Annual research review: does late-onset attention-deficit/hyperactivity disorder exist? J. Child Psychol. Psychiatry 60, 333–352 (2019).
Zaitlen, N. et al. Informed conditioning on clinical covariates increases power in case-control association studies. PLoS Genet. 8, e1003032 (2012).
Rajagopal, V. M. et al. Differences in the genetic architecture of common and rare variants in childhood, persistent and late-diagnosed attention-deficit hyperactivity disorder. Nat. Genet. 54, 1117–1124 (2022).
Yap, C. X. et al. Misestimation of heritability and prediction accuracy of male-pattern baldness. Nat. Commun. 9, 2537 (2018).
van Rheenen, W., Peyrot, W. J., Schork, A. J., Lee, S. H. & Wray, N. R. Genetic correlations of polygenic disease traits: from theory to practice. Nat. Rev. Genet. 20, 567–581 (2019).
Wray, N. R. et al. From basic science to clinical application of polygenic risk scores: a primer. JAMA Psychiatry 78, 101–109 (2021).
Lewis, C. M. & Vassos, E. Polygenic risk scores: from research tools to clinical instruments. Genome Med. 12, 44 (2020).
Laursen, T. M., Agerbo, E. & Pedersen, C. B. Bipolar disorder, schizoaffective disorder, and schizophrenia overlap: a new comorbidity index. J. Clin. Psychiatry 70, 1432–1438 (2009).
Nørgaard-Pedersen, B. & Hougaard, D. M. Storage policies and use of the Danish Newborn Screening Biobank. J. Inherit. Metab. Dis. 30, 530–536 (2007).
Schork, A. J. et al. A genome-wide association study of shared risk across psychiatric disorders implicates gene regulation during fetal neurodevelopment. Nat. Neurosci. 22, 353–361 (2019).
O’Connell, J. et al. Haplotype estimation for biobank-scale data sets. Nat. Genet. 48, 817–820 (2016).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Howie, B. N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006).
Manichaikul, A. et al. Robust relationship inference in genome-wide association studies. Bioinformatics 26, 2867–2873 (2010).
Fox, J. Polycor: Polychoric and polyserial correlations. R package version 0.7-8 https://cran.r-project.org/web/packages/polycor/index.html (2023).
Savalei, V. What to do about zero frequency cells when estimating polychoric correlations. Struct. Equ. Modeling 18, 253–273 (2011).
Galili, T., O’Callaghan, A., Sidi, J. & Sievert, C. heatmaply: an R package for creating interactive cluster heatmaps for online publishing. Bioinformatics 34, 1600–1602 (2018).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Morris, A. P. et al. A powerful approach to sub-phenotype analysis in population-based genetic association studies. Genet. Epidemiol. 34, 335–343 (2010).
Ripley, B. nnet: Feed-forward neural networks and multinomial log-linear models. R package version v. 7.3-16 https://cran.r-project.org/web/packages/nnet/index.html (2021).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Watanabe, K., Taskesen, E., van Bochoven, A. & Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 8, 1826 (2017).
Gusev, A. et al. Transcriptome-wide association study of schizophrenia and chromatin activity yields mechanistic disease insights. Nat. Genet. 50, 538–548 (2018).
Gusev, A. et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat. Genet. 48, 245–252 (2016).
Watanabe, K. et al. A global overview of pleiotropy and genetic architecture in complex traits. Nat. Genet. 51, 1339–1348 (2019).
Machiela, M. J. & Chanock, S. J. LDlink: a web-based application for exploring population-specific haplotype structure and linking correlated alleles of possible functional variants. Bioinformatics 31, 3555–3557 (2015).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
de la Torre-Ubieta, L. et al. The dynamic landscape of open chromatin during human cortical neurogenesis. Cell 172, 289–304 (2018).
Bryois, J. et al. Evaluation of chromatin accessibility in prefrontal cortex of individuals with schizophrenia. Nat. Commun. 9, 3121 (2018).
Finucane, H. K. et al. Heritability enrichment of specifically expressed genes identifies disease-relevant tissues and cell types. Nat. Genet. 50, 621–629 (2018).
Lloyd-Jones, L. R. et al. Improved polygenic prediction by Bayesian multiple regression on summary statistics. Nat. Commun. 10, 5086 (2019).
Smoller, J. W. et al. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet 381, 1371–1379 (2013).
Pilling, L. C. et al. Human longevity is influenced by many genetic variants: evidence from 75,000 UK Biobank participants. Aging 8, 547–560 (2016).
Savage, J. E. et al. Genome-wide association meta-analysis in 269,867 individuals identifies new genetic and functional links to intelligence. Nat. Genet. 50, 912–919 (2018).
Davies, G. et al. Genome-wide association study of cognitive functions and educational attainment in UK Biobank (N = 112 151). Mol. Psychiatry 21, 758–767 (2016).
Okbay, A. et al. Polygenic prediction of educational attainment within and between families from genome-wide association analyses in 3 million individuals. Nat. Genet. 54, 437–449 (2022).
Mills, M. C. et al. Identification of 371 genetic variants for age at first sex and birth linked to externalising behaviour. Nat. Hum. Behav. 5, 1717–1730 (2021).
Jansen, P. R. et al. Genome-wide analysis of insomnia in 1,331,010 individuals identifies new risk loci and functional pathways. Nat. Genet. 51, 394–403 (2019).
Erzurumluoglu, A. M. et al. Meta-analysis of up to 622,409 individuals identifies 40 novel smoking behaviour associated genetic loci. Mol. Psychiatry 25, 2392–2409 (2020).
Nagel, M. et al. Meta-analysis of genome-wide association studies for neuroticism in 449,484 individuals identifies novel genetic loci and pathways. Nat. Genet. 50, 920–927 (2018).
Trubetskoy, V. et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature 604, 502–508 (2022).
Walters, R. K. et al. Transancestral GWAS of alcohol dependence reveals common genetic underpinnings with psychiatric disorders. Nat. Neurosci. 21, 1656–1669 (2018).
Howard, D. M. et al. Genome-wide meta-analysis of depression identifies 102 independent variants and highlights the importance of the prefrontal brain regions. Nat. Neurosci. 22, 343–352 (2019).
Benyamin, B. et al. Childhood intelligence is heritable, highly polygenic and associated with FNBP1L. Mol. Psychiatry 19, 253–258 (2014).
Sanchez-Roige, S. et al. Genome-wide association study meta-analysis of the Alcohol Use Disorders Identification Test (AUDIT) in two population-based cohorts. Am. J. Psychiatry 176, 107–118 (2019).
Loughnan, R. J. et al. Intelligence polygenic score is more predictive of crystallized measures: evidence from the Adolescent Brain Cognitive Development (ABCD) study. Psychol. Sci. 34, 714–725 (2023).
Raj, A., Stephens, M. & Pritchard, J. K. fastSTRUCTURE: variational inference of population structure in large SNP data sets. Genetics 197, 573–589 (2014).
Acknowledgements
Data used in the preparation of this article were obtained from the ABCD study (https://abcdstudy.org), held by the National Institute of Mental Health (NIMH) Data Archive. This is a multisite longitudinal study designed to recruit more than 10,000 children aged 9–10 years and follow them over 10 years into early adulthood. The ABCD study is supported by the National Institutes of Health (NIH) and additional federal partners under award nos. U01DA041022, U01DA041028, U01DA041048, U01DA041089, U01DA041106, U01DA041117, U01DA041120, U01DA041134, U01DA041148, U01DA041156, U01DA041174, U24DA041123 and U24DA041147. A full list of supporters is available at https://abcdstudy.org/federal-partners.html. A listing of participating sites and a complete listing of the study investigators can be found at https://abcdstudy.org/principal-investigators.html. ABCD Consortium investigators designed and implemented the study or provided data but did not necessarily participate in the analysis or the writing of this article. This article reflects the views of the authors and may not reflect the opinions or views of the NIH or ABCD Consortium investigators. The ABCD data repository grows and changes over time. The iPSYCH initiative is funded by the Lundbeck Foundation (grant nos. R102-A9118 and R155-2014-1724), the Mental Health Services Capital Region of Denmark, the University of Copenhagen, Aarhus University and the University Hospital in Aarhus. Genotyping of iPSYCH samples was supported by grants from the Lundbeck Foundation, the Stanley Foundation, the Simons Foundation (SFARI 311789) and the NIMH (5U01MH094432-02). The IPSYCH initiative uses the Danish National Biobank resource, which is supported by the Novo Nordisk Foundation. IPSYCH data were stored and analyzed at the Computerome HPC facility (http://www.computerome.dtu.dk/); we are grateful for continuous support from the HPC team led by A. Syed of DTU Bioinformatics, Technical University of Denmark. We acknowledge funding from the Lundbeck Foundation under fellowship no. R335-2019-2318 (A.J.S.), the National Institute for Aging of the NIH under award nos. U19AG023122, U24AG051129S1, UH2AG064706 and UH2AG064706S1 (A.J.S.), the Research Fund of the Mental Health Services – Capital Region of Denmark R4A92 (S.L.), the Lundbeck Foundation R208-2015-3951 (S.L.), Fonden for Faglig Udvikling af Speciallægepraksis (The Foundation for the Professional Development of Specialist Medical Practice) 38850/16 (S.L.), a European Commission Horizon 2020 grant no. 667302 (S.D.), Helsefonden (Health Fund) grant no. 19-8-0260 (S.D.) and the European Union’s Horizon 2020 research and innovation program under grant no. 847879 (S.D.).
Author information
Authors and Affiliations
Contributions
S.L., I.B., S.D., T.M.W. and A.J.S. conceived the study. S.D., T.M.W. and A.J.S. supervised the study. S.L., I.B. and A.J.S. are responsible for overall study design, with several components of the manuscript carried out with input and guidance from collaborators. S.L., I.B. and D.H. extracted and defined the data from the registers, assisted and guided by E.A., M.G.P. and S.D. S.L. and A.J.S. conducted the SNP heritability, genetic correlations, GWAS and PGS generation, with assistance from J.R.G., V.A., M.V. and A.I. They were supervised by A.J.S. S.L. conducted the functional annotations, with assistance, design and supervision from R.W., D.H.G. and A.J.S. S.L., J.M. and A.J.S. conducted the single-locus and polygenic multinomial tests, using a statistical implementation from A.W.D. and N.Z., and supervised by A.W.D., N.Z. and A.J.S. R.L. and C.E.P. designed and conducted the analysis of the ABCD data and were supervised by T.L.J. A.J.S. conducted the simulations, with support from S.L., M.K. and K.S.K. A.D.B., D.M.H., O.M., M.N., P.B.M. and T.M.W. contributed the iPSYCH data. T.L.J. contributed the ABCD data. S.L. wrote the initial manuscript draft. S.L., I.B. and A.J.S. wrote subsequent versions of the manuscript. All authors discussed the results, commented on the drafts and provided critical feedback throughout.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Evangelos Vassos and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Notes and Figs. 1–43.
Supplementary Table
Supplementary Tables 1–29.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
LaBianca, S., Brikell, I., Helenius, D. et al. Polygenic profiles define aspects of clinical heterogeneity in attention deficit hyperactivity disorder. Nat Genet 56, 234–244 (2024). https://doi.org/10.1038/s41588-023-01593-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-023-01593-7
- Springer Nature America, Inc.
This article is cited by
-
Connecting clinical and genetic heterogeneity in ADHD
Nature Genetics (2024)