Abstract
The common myna (Acridotheres tristis) is one of the most invasive bird species in the world, yet its colonisation history is only partly understood. We identified the introduction history and population structure, and quantified the genetic diversity of myna populations from the native range in India and introduced populations in New Zealand, Australia, Fiji, Hawaii, and South Africa, based on thousands of single nucleotide polymorphism markers in 814 individuals. We were able to identify the source population of mynas in several invasive locations: mynas from Fiji and Melbourne, Australia, were likely founded by individuals from a subpopulation in Maharashtra, India, while mynas in Hawaii and South Africa were likely independently founded by individuals from other localities in India. Our findings suggest that New Zealand mynas were founded by individuals from Melbourne, which, in turn, were founded by individuals from Maharashtra. We identified two genetic clusters among New Zealand mynas, divided by New Zealand’s North Island’s axial mountain ranges, confirming previous observations that mountains and thick forests may form barriers to myna dispersal. Our study provides a foundation for other population and invasion genomic studies and provides useful information for the management of this invasive species.
Similar content being viewed by others
Introduction
Invasive species are organisms which are not native to a particular area that may reproduce and expand demographically and spatially (Pyšek and Richardson 2010), and potentially cause economic or environmental harm or harm to human health (Matheson and McGaughran 2022). In the last 50 years, the financial cost of invasive species has been estimated to exceed $USD 1.288 trillion globally (1970–2020; Zenni et al. 2021). Invasive species are one of the leading causes of biodiversity loss and ecosystem degradation, with considerable impacts on native species via predation and competition (Clavero and García-Berthou 2005; Pyšek and Richardson 2010; Simberloff et al. 2013; Luque et al. 2014). The impacts of invasive species are a growing concern as increases in global connectivity, habitat modification and climate change have led to an increase in the number and distribution of invasive species in recent decades (Hulme 2009; Turbelin et al. 2017).
Information on invasive species, such as their origin, dispersal characteristics, and population dynamics informs their management and can help predict risk of invasion into novel areas (Rollins et al. 2009; Cassey et al. 2014; Evans et al. 2018; Fournier et al. 2019). Population genetic methods can be used to provide information on invasion history, population connectivity and demographics (Rollins et al. 2006; Marrs et al. 2008; Kekkonen et al. 2011), and can provide insight into the existing or novel adaptations that have enabled invasion success (Prentis et al. 2008).
Common mynas (Acridotheres tristis, hereafter mynas) have established from their native range in Central to Southeast Asia to become a globally invasive species and are one of only three bird species on the IUCN ‘100 of the World’s Worst Invasive Alien Species’ list (Global Invasive Species Database 2021). In the 18th century, the French deliberately introduced Indian mynas to Mauritius and Reunion Island to control insect pests (Cheke and Hume 2008), making this one of the world’s first attempts at biological pest control (Safford and Hawkins 2013). Mynas were subsequently intentionally introduced to at least 13 locations across the world, but have also successfully self-introduced globally (Long 1981). Mynas are now invasive in Africa, North America, Europe, the Middle East, Australia, and numerous island nations including Madagascar, Fiji, Maldives, and New Zealand (Feare and Craig 1999; Peacock et al. 2007; Hart et al. 2020; CAB International 2021).
Mynas have a generalist, omnivorous diet and can themselves be agricultural pests (Dawson and Bull 1970; Hone 1978; Peacock et al. 2007) and can have negative impacts on native ecosystems, especially island ecosystems with high level of endemism (Hart et al. 2020). These impacts include direct competition with native birds and mammals for food and nests, including taking over native birds’ nest cavities, predation on eggs and chicks (Hart et al. 2020), and assisting with the dispersal of invasive and exotic plants (Pimentel et al. 2000; Parkes 2006; Saavedra 2009). In addition, mynas may also impact human health and livelihood through the spread of disease, noise pollution, and property damage (Yap et al. 2002; Parkes 2006; Saavedra 2009; Clark et al. 2015). Eradication of mynas on islands has been found to positively impact the native fauna, for example, the increase in numbers of the native Seychelles magpie robins and Seychelles paradise flycatchers accelerated after removal of 90% of mynas from Denis Island, Seychelles (Feare et al. 2021).
Mynas are thought to have been introduced to New Zealand from Australia in the 1870s (Thomson 1922; Long 1981; CAB International 2021), where they had been introduced from the native range, likely India, in the 1860s (McCoy et al. 1885; Ewart et al. 2019). However, a review of historical data and literature by Beesley et al. (2023) on myna introduction history and distribution in New Zealand suggests some uncertainty in the timing, the number of individuals, the number of introductions, and their source. For instance, there is some evidence that the first introduction of mynas in New Zealand may be earlier than the 1870s (e.g. Huddleston 1868), and it is unclear which of the initial introductions to New Zealand survived and formed the current populations. Since their initial introduction into New Zealand, mynas have been translocated/introduced around the country (e.g. to Whanganui, New Plymouth, Hawke’s Bay), but their exact colonisation history remains unclear. In addition, mynas are no longer seen at many of their initial introduction points into New Zealand (e.g. Nelson, Christchurch, Dunedin, and Wellington).
Population genetic data can help elucidate the colonisation history, and by extension invasion pathways, and identify gene flow between populations. Restriction-site associated DNA sequencing (RADseq) and similar genotyping by sequencing (GBS) methods subsample the genome by using restriction enzymes to fragment genomic DNA and then sequence adjacent to the cut sites with next-generation sequencing (Baird et al. 2008; Wright et al. 2019). This generates genome-wide high-throughput sequencing data, making it a suitable genotyping method for obtaining individual-level genotype information that can be compared across individuals and populations. Consequently, RADseq has been used in numerous ecological, evolutionary, and conservation genomic studies to address varied questions (Andrews et al. 2016), such as describing population structure and connectivity, detecting hybrids, resolving phylogenies, and identifying genome regions under selection (Near et al. 2018; Ewart et al. 2019; Hofmeister et al. 2021; Stuart et al. 2021; Forsdick et al. 2021). Although RADseq does not require a reference genome (Baird et al. 2008; Peterson et al. 2012), the use of a reference genome typically improves the reliability of genotype calls and downstream inferences (Shafer et al. 2017).
Here, we utilise single nucleotide polymorphism (SNP) markers – identified from RADseq reads mapped against a draft myna genome assembly - from 814 myna individuals from six countries to 1) identify the likely sources of invasive population(s), and more specifically of the New Zealand myna population, 2) clarify our current understanding of the myna’s colonisation history in New Zealand, and 3) identify population structure and genetic diversity in New Zealand mynas. This study provides a foundation for further invasion genomic studies and useful insights for species management.
Materials and methods
Data filtering and analyses performed in R were conducted in R version 4.1.2 (R Core Team 2021), with figures plotted using the ‘ggplot2’ R package version 3.3.6 (Wickham 2016) unless noted. See Table S2.1 for a summary of the software and packages used.
Sampling, DNA extraction, sequencing and processing
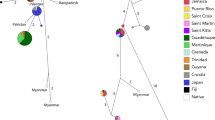
A total of 183 myna tissue samples in ethanol from India, New Zealand, Australia, South Africa, Hawaii and Fiji between 1975–1989 (Baker and Moeed 19801987; Fleischer et al. 1991) were received from the Royal Ontario Museum (ROM). A further 193 euthanized mynas were obtained from myna control programmes from contributors in New Zealand between 2017–2020, and muscle tissue was subsampled from each individual. DNA was extracted from the ROM tissue samples using the DNeasy Blood & Tissue Kit (Qiagen) following the manufacturer’s protocols. DNA was extracted from the New Zealand tissue samples using the Monarch Genomic DNA Purification Kit (NEB) following the manufacturer’s protocols. DNA concentration was measured using a Qubit 2.0 Fluorometer (Thermo Fisher Scientific). DNA was diluted to standardised concentrations of 50–100 ng/μL, and sent to Diversity Arrays Technology Pty Ltd company (DArT P/L) for further processing (Kilian et al. 2012; Cruz et al. 2013). Samples from 363 individuals were successfully sequenced using the proprietary Diversity Arrays Technology platform and protocol (DArTseq™). We included 13 duplicate samples. DArTseq also includes internal replicates of samples as part of its protocol. DArTseq is a restriction enzyme‐based genome complexity reduction method that has been used to generate SNP data for a range of studies (Melville et al. 2017; Ketema et al. 2020) including a previous population genomic study on Australian mynas (Ewart et al. 2019). We utilised the PstI-SphI restriction enzyme double digest following Ewart et al. (2019), enabling our samples to be processed and co-analysed with the DArTseq data from 451 mynas from Australia from the aforementioned study (mynas sampled in 2014–2015). In total, our study dataset comprised 814 mynas sampled from six countries (Fig. 1, Tables S1.1 and S1.2).
Blue represents the native range and orange represents the invasive range. Note that this figure does not cover the entire invasive range of the myna. All samples from India, Fiji, South Africa, and Hawaii were provided by the Royal Ontario Museum (ROM). Some samples from Australia and New Zealand were also provided by the ROM. Map of sample locations include: A entire dataset, B India, and C New Zealand. Within New Zealand’s North island (C), early introduction points pre-1980 are marked in bold brown numbers, namely, (1) Hawke’s Bay (Napier), (2) Wellington, (3), Whanganui, and (4) New Plymouth.
Variant calling
Barcodes in DArTseq raw read sequences were removed using the process_radtags program in STACKS version 2.58 (Catchen et al. 2013) and adaptors trimmed using fastp version 0.20.0 (Chen et al. 2018). Quality control checks were performed using fastqc version 0.11.9 (Andrews 2010) and multiqc version 1.9 (Ewels et al. 2016). Reads were then aligned to a draft common myna reference genome generated from an Australian individual (see step 3 in Appendix S2 for more details) using BWA version 0.7.17 (Li and Durbin 2009). Reads were subsequently sorted and indexed using SAMTOOLS version 1.12 (Li et al. 2009). As detailed in Appendix S2.1, SNPs were called through the mpileup and call commands in BCFtools version 1.13 (Li 2011) based on three subsets of individuals – the entire dataset (ALL dataset: 814 individuals; 1024 samples including replicates), a New Zealand-only dataset (NZ dataset: 226 individuals, 282 samples) and an India-only dataset (IND dataset: 78 individuals, 80 samples). All analyses described below were performed on the genotypes called by BCFtools, but to ensure results were consistent, population structure analyses were also performed on SNP datasets called from other pipelines (STACKS and DArT P/L DArTsoft14 SNP calling algorithm). Population structure was congruent across the three SNP datasets (see Fig. S2.1 and Appendix S10 for more details).
Data filtering
As described below, the BCFtools SNP datasets were filtered based on locus data quality (e.g. SNP quality score, read depths, genotype quality scores), sample data missingness, locus data missingness, and linkage disequilibrium, and finally filtered to only retain SNPs that are under Hardy-Weinberg equilibrium (HWE). Depending on the analyses performed, the datasets were filtered using different stringencies (see Fig. S2.2 for more details). Variants were output in binary variant call format (BCF) and were converted to variant call format (VCF) and compressed VCF (vcf.gz) using BCFtools for uses in different software.
SNP quality filters
The SNP datasets were filtered to only retain biallelic SNPs using the view and filter commands in BCFtools. VCFtools version 0.1.15 (Danecek et al. 2011) was used to filter the dataset to only retain SNPs with SNP quality scores (QUAL) ≥30 and recode genotype calls to NA where genotype quality scores (GQ) were <30 or where the coverage fell outside of minimum (<15) and maximum read depth thresholds. The maximum read depth filters varied according to the population dataset due to variation in the distribution of read depths across populations (see Appendix S3.2 for more details). A maximum depth of 100 was applied to the New Zealand and India datasets and 125 to the entire dataset.
Removal of replicates, admixed Australian samples, and samples with heterozygote excess
Ewart et al. (2019) suggests three introduction points in Australia (Melbourne, Sydney, and Gold Coast) with some levels of admixture in some populations (Northern NSW and southern Queensland). The inclusion of admixed individuals is known to affect some downstream analysis and assignment of admixed individuals to a single population can be arbitrary (Pritchard et al. 2000; Luu et al. 2017). Further, the introduction of mynas to New Zealand likely occurred in the 1870s and 1880s before the establishment of some of the other populations in Australia. To simplify global population structure analyses and the identification of introduction source(s) of New Zealand mynas, we used BCFtools to retain only Australian samples from Melbourne, Sydney, and Gold Coast in the ALL dataset prior to further filtering and subsampling (see Fig. S1.1 for locations of Australian samples retained).
One sample from each set of the replicates (including DArTseq internal replicates) was retained at random prior to further SNP filtering. A further three samples (two from Sydney, and one from Odisha) were removed from the ALL and IND dataset due to the samples exhibiting heterozygote excess (highly negative FIS) (see Appendix S3 for more details). A total of 226, 77, and 467 individuals were retained in the NZ, IND, and ALL datasets respectively.
SNP data missingness, and singletons/doubletons
NZ, IND, and ALL datasets were filtered to remove SNPs with >20% missingness using BCFtools. For data used in the population structure analyses, singletons and doubletons (SNPs only occurring in one sample) were removed from the dataset using VCFtools to remove potential artefactual alleles (see section 3.5 and 3.6 in Appendix S3 for more details). For data used for calculating the site frequency spectrum, filters for singletons and doubletons were not performed to prevent introducing SNP ascertainment bias and altering the site frequency spectrum.
Linkage disequilibrium
We calculated pairwise linkage disequilibrium (LD) and plotted this against pairwise distance using PopLDdecay (Zhang et al. 2019) to visualise LD in our datasets (Figs. S4.1–4.3). To remove SNPs in LD, we retained 1 SNP per every 100,000 base pairs when found on the same reference contig using the --thin option in VCFtools (see Appendix S4 for more details). After thinning, we retained 5474, 5978, and 5904 SNPs in the NZ, IND, and ALL dataset, respectively.
Population delineations
Further downstream analyses, such as population pairwise-FST calculations and tests for Hardy Weinberg Equilibrium (HWE), require prior knowledge of the population structure. However, incorrect delineation of populations may result in inaccurately clumping genetically distinct populations, or separating genetically homogenous populations.
Consequently, initial principal component analysis (PCA) and sparse non-negative matrix factorisation (sNMF) population genetic structure analysis were performed on the NZ, IND, and ALL dataset prior to HWE filtering to avoid the inadvertent introduction of a Wahlund effect (Pearman et al. 2022). PCA was performed and visualised using the ‘dartR’ R package version 1.9.9.1 (Gruber et al. 2018). sNMF was performed using the ‘LEA’ R package version 3.2.0 (Frichot and François 2015) with default settings, regularisation parameter (α)= 100, and 10 replicates per K value (number of genetic groups) and plotted using the ‘pophelper’ R package version 2.3.1 (Francis 2017).
Based on these preliminary population clustering results, a total of 32 populations were delineated for further analyses. These included separating the six individuals from Maharashtra (India) as a separate population (‘IND: Maharashtra subpopulation A’ or ‘Maharashtra subpopulation A,’ hereafter) from other Maharashtra individuals due to separate clustering. Other populations were also separated based on the location and timing of the sample collection (samples from the ROM were collected in 1970s–1980s, while the rest were collected in 2014–2020). This delineation of populations will be referred to as ‘popdef1’ hereafter (see Tables S1.2 and S1.3 for details).
Hardy Weinberg Equilibrium
Using populations as delineated above (popdef1), loci were filtered to remove non-neutral loci by testing for HWE using the HWExactMat function from the ‘HardyWeinberg’ R package version 1.7.2 (Graffelman and Camarena 2008; Graffelman 2015), which tests for departure from HWE using the Fisher’s exact test. All loci with p values < 0.01 in any population were removed from the dataset. A p value threshold of < 0.01 was chosen instead of a correction for multiple testing because the distributions of the p values did not follow a uniform distribution (i.e., the p value distribution had a peak near 1 and 0; see Appendix S6). All subsequent analyses except for the site frequency spectrum calculations were conducted with the HWE filtered SNP dataset (5037, 5900, 5584 SNPs for the NZ, IND, ALL dataset).
Subsampling of the global (ALL) dataset for further PCA analyses
Uneven sample sizes across genetic populations have been shown to potentially affect analyses such as principal component analysis (PCA) (Privé et al. 2020) and population structure analyses (Puechmaille 2016). Therefore, we explored the effects of subsampling and the delineation of populations in the global (ALL) dataset. PCA on full and subsampled datasets suggests that there is an effect of uneven sampling in the ALL dataset, distorting the principal components and inflating the differences between the more well-sampled and inbred populations (i.e., Australian and New Zealand samples) and the other populations (Appendix S5).
Consequently, samples in the ALL dataset from the same geographic region that were genetically very similar based on the preliminary clustering analyses (section 2.3.5, above) were grouped together and populations were delineated as follows for subsampling: IND: Maharashtra subpopulation A, IND: Other, AUS: Melbourne, AUS: Sydney, AUS: Gold Coast, NZ: Napier, NZ: Other, South Africa, Fiji and Hawaii. This delineation of populations will be referred to as ‘popdef2’ hereafter (see Tables S1.2 and S1.3 for more details). We explored different subsampling strategies (Appendix S5), and decided to subsample the ALL dataset so that we retained 20 individuals per introduced population when we performed the PCA. The uneven sampling is not expected to affect the results of other analyses so the full ALL dataset was used for these (see below). Further, given relatively even sample numbers across populations within India and New Zealand, we did not subsample the IND and NZ datasets for PCA.
Population structure
PCA analyses were repeated on the HWE-filtered SNP datasets using the popdef2 delineated populations. Unlike ADMIXTURE and STRUCTURE, sNMF does not assume HWE. However, the final sNMF analysis were performed on the HWE-filtered dataset to ensure that we were detecting neutral population structure. We also performed both PCA and sNMF on different subsets of the dataset (MAF > 0, MAF > 0.05, MAF > 0.1) to ensure consistency of results (Appendix S8.6 and S8.7).
Inter-population genetic differences with the HWE-filtered dataset were examined across the dataset using pairwise-FST values calculated using the gl.fst.pop function in the ‘dartR’ R package (100 bootstraps performed) for all population combinations.
Isolation by distance analysis based on the Mantel test (IBD) was performed using the gl.ibd function in the ‘dartR’ R package to determine whether genetic differentiation between populations within India and within New Zealand can be explained by geographical distance. To do so, we correlated the transformed population pairwise-FST (\({F}_{{ST}}/(1-{F}_{{ST}})\)) and the natural log-transformed geographical distance (km) (Rousset 1997). Mantel tests were performed on the populations from India and the populations from within the NZ: Other cluster with more than five individuals. Given high genetic differentiation between Maharashtra and Maharashtra subpopulation A (from the same sampling location), we excluded Maharashtra subpopulation A from the Mantel test.
Genetic diversity
Genetic diversity indices were calculated for each population using the HWE-filtered dataset. In the NZ and IND dataset, the popdef1 population delineation was used. For the ALL dataset, both the popdef1 and popdef2 delineations were used in order to investigate the fine-scale and broader-scale diversity patterns across India and New Zealand. Mean observed and expected heterozygosity were calculated using the gl.report.heterozygosity function in the ‘dartR’ R package. The proportion of polymorphic loci was calculated after randomly subsampling the dataset to n = 6 per population, 100 times. This subsampling accounts for the uneven sampling per population, as populations with more samples will likely have more polymorphic loci. Mean allelic richness was calculated using a rarefaction method to account for uneven sampling using the allel.rich function in the ‘PopGenReport’ R package version 3.0.4 (Adamack and Gruber 2014; Gruber and Adamack 2015). Mean allelic richness, and private allelic richness were also calculated using another rarefaction method using the software HP-rare version 1.1 (Kalinowski 2005). Finally, the shape of the folded site frequency spectrum was used alongside other genetic diversity metrics to visualise the genetic bottlenecks experienced in the different introduced populations. The folded site frequency spectrum (SFS) was plotted based on the minor allele frequency calculated by the gl.percent.freq function in the ‘dartR’ R package for the different populations within the ALL dataset, subsampling to n = 6 individuals per population and repeating the subsampling 100 times.
Results
Population structure
New Zealand dataset
The PCA of the New Zealand-only (NZ) dataset shows two clusters on PC1 (Fig. 2A): individuals from Napier, and individuals from the rest of New Zealand. Within the ‘rest of New Zealand’ cluster, three individuals from Leigh separate on PC2. Based on pairwise relatedness values, these represent a trio of likely first-degree relatives. Note that close relatives were not removed from the dataset to avoid misrepresenting the overall diversity within populations. The sNMF population structure analysis was best explained with K = 2 (Fig. 2C) and supports the two populations as shown by the PCA. In agreement with the population structure analysis, the Napier populations have moderate levels of genetic differentiation from all other New Zealand populations (pairwise-FST = 0.05–0.07), with low but significant differentiation between the ROM (collected in 1984) and modern (collected between 2017–2020) samples (pairwise-FST = 0.015; Fig. S8.5). The Great Barrier Island (GBI) population shows low but significant differentiation (pairwise-FST = 0.010–0.014) from the other New Zealand populations. Pairwise-FST values among the other New Zealand population are generally very low (pairwise-FST < 0.005) and/or insignificant (p values > 0.05).
A PC1 vs PC2 for the NZ dataset, based on 5037 SNPs. Colour and shape of points represent sample location and time of collections. ROM = Royal Ontario Museum samples which were collected between 1975–1989. Other samples were collected between 2017–2020. Axes are labelled with the variance explained. B Mantel test for samples from NZ: Other, excluding populations with fewer than 5 individuals, and sample M0367 from Auckland which had different sample locations from other Auckland samples. C Population structure estimated by sNMF analysis, using K = 2. Three-letter codes labelled beneath the figure (e.g. KKH, HEL, NGU, etc.) represent location and time of collection as labelled in (A).
The Mantel test on the populations within NZ: Other (excluding populations with n ≤ 5) found no significant isolation by distance (Fig. 2B, R2 = 0.00635, p = 0.333). The cluster of points with \({F}_{{ST}}/(1-{F}_{{ST}})\) > 0.01 is a result of the inclusion of GBI; removal of the GBI population did not result in significant isolation by distance (Fig. S8.4).
Indian dataset
PC1 vs PC2 from the native-range India-only (IND) dataset shows two to four clusters (Fig. 3A). The most distinct cluster consists of samples from Maharashtra subpopulation A. The sNMF population structure analysis of the IND dataset was best explained with K = 1 (Fig. S8.11, see Fig. S8.12 for population structure plots at K = 2–5). With the exclusion of Gujarat (n = 1), and Maharashtra subpopulation A, a Mantel test on the populations within India found significant isolation by distance (Fig. 3B, R2 = 0.224, p = 0.008).
A PC1 vs PC2 for the IND dataset, based on 5900 SNPs. Axes are labelled with the variance explained. The lower left cluster of samples is a subpopulation, termed ‘subpopulation A’ of Maharashtra samples. B Mantel test for samples from India, excluding the samples from Gujarat (n = 1) and Maharashtra subpopulation A.
Population pairwise-FST values suggest low (FST = 0.001–0.03) but significant differentiation between all Indian populations except for Maharashtra subpopulation A which showed moderate levels of genetic differentiation from other Indian populations (FST = 0.042–0.073) (Fig. S8.13).
Global dataset
The global (ALL) dataset was subsampled due to the large variation in sample sizes from different countries. Across 10 replicates, PCA on this subsampled data showed consistent clustering of the populations (results not shown), with a representative run presented in Fig. 4A. PC1 vs PC2 from this subsampled dataset shows five clusters (Fig. 4A): 1) samples from the Australian Gold Coast, 2) samples from New Zealand (NZ: Napier and NZ: Other), AUS: Melbourne, AUS: Sydney, Fiji and IND: Maharashtra subpopulation A, 3) samples from the rest of India, 4) samples from Hawaii (clustering closely to cluster 3), and 5) samples from South Africa. PC1 vs PC2 on just samples from cluster 2 show three clusters (Fig. 4B): 2A) samples from Sydney, 2B) samples from NZ: Other, and 2C) samples from NZ: Napier, Melbourne, Fiji, and Maharashtra subpopulation A. The numbering of the clusters refers to the labels in Fig. 4C.
A PC1 vs PC2 for the ALL dataset, number of individuals subsampled to n = 20 per introduced populations, based on 5572 SNPs. Colour and shapes of points represent sample locations. B PC1 vs PC2 for the ALL dataset with only populations that clustered closely with New Zealand. This includes NZ: Other, NZ: Napier, Fiji, AUS: Melbourne, AUS: Sydney and IND: Maharashtra subpopulation A. In (A) and (B), axes are labelled with the variance explained. C Population structure estimated by sNMF analysis, using K = 8. Three-letter codes are abbreviations of population names, as shown in the legend of the PCA plots. The cluster number refers to the genetic clusters described in text as shown by the PCA and the sNMF population structure plot.
The sNMF population structure analysis of the ALL dataset was best explained with K = 8, and supports seven genetic clusters (Fig. 4C). The seven genetic clusters/groups are: 1) Gold Coast, 2A) Sydney, 2B) NZ: Other, 2C) Melbourne, Fiji, Napier, and Maharashtra subpopulation A, 3) IND: Other, 4) Hawaii, and 5) South Africa. These groupings are very similar to the five clusters identified in the first PCA (Fig. 4A), but split PCA grouping 2 into the three sNMF clusters (2A-C) which were also identified from the second PCA (Fig. 4B).
Population pairwise-FST between all populations showed high genetic differentiation (pairwise-FST > 0.247) among distant bottlenecked populations (Gold Coast, Hawaii, and South Africa) and low genetic differentiation (pairwise-FST = 0.005–0.037) between Maharashtra subpopulation A and Melbourne, Napier, and Fiji (Fig. 5) – lower than genetic differentiation between Maharashtra subpopulation A and IND: Other (pairwise-FST = 0.041).
Populations were delineated in line with PCA and sNMF (Fig. 4, popdef2). Maharashtra subpopulation A was delineated as a separate population from other Indian samples as it clusters closely with AUS: Melbourne, NZ: Napier, and Fiji. All FST values have p values < 0.01.
Maharashtra subpopulation A, Melbourne, Napier, and Fiji (Fig. 4, cluster 2C) show very low genetic differentiation (pairwise-FST < 0.05), comparable to populations from IND: Other (e.g. Odisha, Karnataka, etc.), and NZ: Other (e.g. Leigh, Waiheke, Thames, etc.) (see Fig. S8.18 for more details).
Population pairwise-FST between Hawaii and South Africa showed high genetic differentiation (pairwise-FST = 0.247). However, both populations showed lower population pairwise-FST with IND: Other (pairwise-FST = 0.105–0.131) than with other populations in this study.
Genetic diversity
All genetic diversity indices (mean observed heterozygosity (Ho), mean expected heterozygosity (He), mean allelic richness (AR), proportion of polymorphic loci and mean private allelic richness (PAR)) inferred that myna populations in India were more diverse than the introduced populations (Table 1), except for Maharashtra subpopulation A which is the least diverse among the native populations and shows similar diversity metrics to the Melbourne population. Within the introduced populations, the Melbourne population appeared to be most diverse, followed by Fiji/Napier, NZ: Other, Sydney, Hawaii, and South Africa and Gold Coast (Table 1).
Similar to the other genetic diversity indices, the shape of the SFS plot (Fig. 6) suggests that the IND: Other population is most diverse, followed by Maharashtra subpopulation A, Melbourne/Fiji, Napier, NZ: Other, Sydney, and Hawaii/South Africa/Gold Coast. Note that IND: Other and NZ: Other are a combination of samples from multiple locations and may alter some statistics (see Tables S9.1–9.3 and Fig. S9.1 for results with populations delineated based on popdef1 population delineations).
Discussion
The spread of invasive species may leave characteristic genetic signatures across the landscape. Given appropriate sampling, the source population(s) and population structure can be identified and compared to historical records and current observations. The differences in genetic diversity, differentiation, and composition between populations can elucidate the connectivity between populations and population introduction history. Using common myna samples from the native and the invasive range, we were able to utilise genomic analyses to identify the population structure, and evaluate and infer the myna introduction history from India to Australia and then to New Zealand.
Population structure, genetic diversity, and introduction history
Our extensive sampling from the native and invasive range has allowed, for the first time, inference of the origin of mynas introduced to New Zealand. Historical records suggests that mynas in New Zealand were founded by individuals from an established population in Australia which in turn was founded by individuals from the native range in India (Thomson 1922). The clustering of New Zealand mynas with Melbourne samples strongly supports importation of birds from Melbourne, and is in agreement with analysis of primary literature from the time of introduction (Beesley et al. 2023). The clustering of mynas from Melbourne and from Fiji with Maharashtra subpopulation A also suggests that the birds in Melbourne and Fiji were likely founded by individuals from Maharashtra.
Two genetic clusters in New Zealand were identified that appear to be consistent with the historical record of the species introduction and the distribution in New Zealand (Cunningham 1948; Robertson 2007; Heather and Robertson 2015). The two populations appear to be divided by North Island’s axial mountain ranges (Tararua, Ruahine, Kaweka, Kaimanawa and Raukūmara ranges), with the population to the east of the mountain range from Masterton to East Cape represented by the more genetically diverse samples from NZ: Napier, while the population to the west from Palmerston North to Northland is represented by samples from the rest of New Zealand (NZ: Other) (Figs. 1C and 2, and Tables S1.2, and S1.3).
Historical records suggest that mynas were introduced to Napier in the 1870s/1880s and therefore represent an early population establishment (Acclimatisation 1877; ‘Hawke’s bay acclimatisation society.’ 1877; Cunningham 1948), while the populations in the rest of the North Island likely established later by northward colonisation from the Wellington, Whanganui or New Plymouth region (2, 3, and 4 in Fig. 1C) where they were first introduced in the 1870s/1880s (Thomson 1922; Long 1981; Beesley et al. 2023). Mynas only started to appear in the Waikato and Auckland region in the 1920s and only established themselves in these regions in the 1940s (Cunningham 1948). Cunningham (1948) did not report mynas north of Auckland, or from Waiheke and Great Barrier Island where some of our samples were collected. While Cunningham (1948) described four main areas of the myna distribution within the North Island, the 1947 distribution map in the study showed the two New Zealand populations west and east of the axial mountain range being clearly separated, and more so than in more recent distribution maps (Robertson 2007; Heather and Robertson 2015; Beesley et al. 2023).
Among the New Zealand populations, Great Barrier Island population (GBI) shows low but significant levels of genetic differentiation from other New Zealand populations (Fig. S8.5, pairwise-FST = 0.010–0.014). Mynas were only first recorded on GBI in 1960 (Bell and Brathwaite 1964), and were widespread by 1975 (Bell 1976; Ogle 1981). GBI clearly nests within NZ: Other in our population structure analyses but SFS and genetic diversity metrics show that GBI has the lowest diversity of any of our New Zealand sampling sites and is likely to have suffered some genetic bottlenecking and drifted from NZ: Other that may have founded it (Fig. S9.1, and Table S9.1). Continual gene flow with the mainland NZ: Other population may remove this exhibited bottleneck effect in the future. We investigated gene flow among the New Zealand populations (n ≥ 20) within NZ: Other, and between the Napier and Leigh population (representative of the NZ: Other population) using BA3-SNPs version 3.04 (Wilson and Rannala 2003; Mussmann et al. 2019), but were unable to detect any gene flow. However, it is likely the populations in our analysis are not differentiated enough for BA3-SNPs to deduce gene flow (see Appendix S7.2 for more details).
While NZ: Other is identified as a separate genetic cluster in the sNMF analysis (cluster 2B), it is most genetically similar to sNMF cluster 2C (pairwise-FST = 0.052, 0.055, 0.057 and 0.07 with Melbourne, Napier, Maharashtra subpopulation A, and Fiji, respectively). This clustering, and given that Napier has higher genetic diversity than NZ: Other, raises the hypothesis that Napier and NZ: Other were separate introductions from Melbourne, versus NZ: Other being established from Napier birds. Attempts were made to resolve the population history with phylogenetic trees and coalescent simulations using fastsimcoal2 version 2.7.09 (Excoffier et al. 2021), but key nodes were not well supported and the different scenarios in the coalescent simulations did not differ significantly (see Appendix S7.1 for more details). This may be due to the shared population history and very similar and recent coalescent times for each of the scenarios. Nonetheless, historical records, New Zealand’s proximity to Australia, Sydney population’s lower genetic diversity compared to NZ: Other, and the lower genetic differentiation between NZ: Other and AUS: Melbourne populations would suggest that mynas in NZ: Other may have also been founded by individuals from Melbourne (rather than Sydney) but have suffered a more severe genetic bottleneck compared to the Napier population.
Sydney is identified as a separate genetic cluster in the sNMF analysis (cluster 2A) and is most genetically similar to Melbourne, (pairwise-FST = 0.082 in Fig. 5, and pairwise-FST = 0.081–0.089 in Fig. S8.18). Interestingly, this suggests that, like mynas in New Zealand, mynas in Sydney may have also been founded by individuals from Melbourne. This result provides further context to the findings of Ewart et al. (2019), who found that Sydney and Melbourne were genetically distinct and likely represented two introduction points but, in the absence of data from the native range nor clear historic records, could not differentiate between Sydney being established from a separate introduction from India, or a translocation from Melbourne. Although potentially resolving the ultimate origin of Sydney and Melbourne populations, the present study does not resolve the origin of the Gold Coast population that perplexed Ewart et al. (2019). The high level of genetic distinctiveness in the Gold Coast population observed in this study implies either introduction from an as yet unidentified source, or extreme bottlenecking in a Melbourne-derived population.
Hawaii and South Africa showed high levels of genetic differentiation between each other, but both populations show lower genetic differentiation with IND: Other than with other populations in this study. This suggests that mynas in Hawaii and South Africa may have been independently founded by individuals from IND: Other.
Implications and future analyses
Mynas in New Zealand appear to have a clear population structure and are divided into two populations, east and west of North Island’s mountain range. While the two populations may share a common origin, the population in the east (NZ: Napier) is more diverse and has a smaller distribution. Further analyses can be built upon this study to provide more information for management of the species in New Zealand. Our gene flow analysis was unsuccessful, but further analyses on higher-density sequencing data (e.g. to enable inference of ancestral haplotypes) may help resolve the very recent establishment of the New Zealand populations. Additional samples from more locations will also help elucidate finer scale population structure, and reduce the chance that key populations are not sampled. For example, samples from the highways crossing the North Island axial mountain range can potentially help deduce if there is admixture between the NZ: Other and NZ: Napier populations, and if this requires any management interventions. Similarly, tracking individuals in key populations may provide an alternative source of information on the species’ dispersal ability and population connectivity, which may further help inform myna management programmes on the feasibility of myna eradication in some locations (e.g. island populations).
The identification of Maharashtra subpopulation A as the potential source of the mynas in Australia and Fiji also raised other questions regarding this potential source population. Maharashtra subpopulation A is genetically less diverse than the other Indian populations and has potentially experienced a population bottleneck. These samples were collected in 1975 from Maharashtra, although the exact location is unknown. Other samples from Maharashtra, and the rest of India were also collected from a similar time period (a West Bengal sample and two other Maharashtra samples were collected in 1975; other Indian samples, in 1983; see Table S1.2 for more details). Why is there a more bottlenecked population within the more-or-less panmictic Indian population? Where are these samples exactly from? To answer these questions, additional samples from the potential source populations of Maharashtra should be analysed.
We note that this dataset includes samples collected from the same locality from different time periods (Sydney, Melbourne, and Napier). In all cases, we found low but significant pairwise-FST between the samples collected in 1970s–1980s (ROM) and the modern samples collected in 2014–2020, suggesting that the genetic composition of mynas in these populations may have changed since the collection of the ROM samples. Although there may be some bias from sample size differences, in addition to variability introduced from stochastic sampling effects and differences between sampling locations, it is likely that much of this signal is due to drift and population size changes. Interestingly, unlike Melbourne and Napier where samples from different time periods show similar levels of genetic diversity, modern Sydney samples show higher genetic diversity compared to Sydney ROM samples, and modern Sydney samples are more genetically similar to modern Melbourne (and Melbourne ROM) than Sydney ROM (Fig. S8.18). These results may point to recent gene flow between Melbourne and Sydney, but equally could represent differences between sampling locations between the two timepoints. However, some changes between populations over time may have occurred in response to changing selection pressures (e.g. increased urbanisation, adaptation to the newly invaded environment, (Baker and Moeed 1979)). Analyses in the future may be able to identify candidate genes for adaptive traits that are under selection in the invasive populations, and help provide a biological explanation for their invasive success.
This study presents the most extensive population genomic studies of the common myna to date, which utilises samples from multiple locations across the world. We were able to identify the source population of mynas in a few invasive locations: populations in Melbourne and Fiji were likely founded by individuals from Maharashtra, and populations in New Zealand were likely founded by individuals from Melbourne. We were able to identify two distinct populations in New Zealand, east and west of North Island’s axial mountain ranges. This observation confirms previous observations that mountain ranges and thick forests may form barriers for myna dispersal and provide useful information for the species management. These findings support the use of thick forests and mountains as barriers of management units and a more localised management strategy of mynas in New Zealand (East and West of the North Island’s axial mountain range). This also highlights the potential additional benefits of reforestation programmes in New Zealand (e.g. One Billion Trees Programme (Te Uru Rākau 2020)). However, it must be noted that not all forest types may form barriers for myna dispersal, and further research would be needed to determine the forest features that act as barriers (e.g. old versus new growth forests, forest patch size). The population structure in New Zealand also allows management programmes to possibly identify the source(s) of newly established populations (e.g. populations south of 40°S), and vagrants, and limit their invasion pathways. With decreasing cost of sequencing, improving sequencing capabilities, and increasing availability of reference genomes, the identification of the source population(s), the introduction history, and the population structure from this study provides a strong basis for more detailed studies of the species in the future (e.g. adaptation genomics, genome-wide association scans, and gene-environment association analyses). This study has highlighted a common challenge faced in genetic studies of most invasive species – uneven sampling involving inbred populations and population delineations - and demonstrated how this issue may be handled. This study also demonstrates the value and utility of museum collections to address present-day challenges such as invasive species.
Data availability
All scripts used in data processing and analysis are available on GitHub (https://github.com/akamolphat/myna_popgen). Some code is also available in the Supplementary materials. Individual metadata can be found in Table S1.2 in the Supplementary Materials. Raw reads, barcodes for demultiplexing, and processed variant data (VCF format) can be found on Dryad Digital repository (https://doi.org/10.5061/dryad.xsj3tx9m7).
References
Acclimatisation (6 January 1877) Southland Times. Page 2. https://paperspast.natlib.govt.nz/newspapers/ST18770105.2.7. Accessed 21 January 2022
Adamack AT, Gruber B (2014) PopGenReport: simplifying basic population genetic analyses in R. Methods Ecol Evol 5:384–387. https://doi.org/10.1111/2041-210x.12158
Andrews S (2010) FastQC. A quality control tool for high throughput sequence data. https://www.bioinformatics.babraham.ac.uk/projects/fastqc/
Andrews KR, Good JM, Miller MR, Luikart G, Hohenlohe PA (2016) Harnessing the power of RADseq for ecological and evolutionary genomics. Nat Rev Genet 17:81–92. https://doi.org/10.1038/nrg.2015.28
Baird NA, Etter PD, Atwood TS, Currey MC, Shiver AL, Lewis ZA et al. (2008) Rapid SNP discovery and genetic mapping using sequenced RAD markers. PLOS ONE 3:e3376. https://doi.org/10.1371/journal.pone.0003376
Baker AJ, Moeed A (1979) Evolution in the introduced New Zealand populations of the common myna, Acridotheres tristis (Aves: Sturnidae). Can J Zool 57:570–584. https://doi.org/10.1139/z79-067
Baker AJ, Moeed A (1980) Morphometric variation in Indian samples of the Common myna, Acridotheres tristis (Aves: Sturnidae). Bijdragen tot de Dierkd 50:351–363. https://doi.org/10.1163/26660644-05002005
Baker AJ, Moeed A (1987) Rapid genetic differentiation and founder effect in colonizing populations of common mynas (Acridotheres tristis). Evolution 41:525–538. https://doi.org/10.1111/j.1558-5646.1987.tb05823.x
Beesley A, Whibley A, Santure AW, Battles HT (2023) The introduction and distribution history of the common myna (Acridotheres tristis) in New Zealand. N Z J Zool 1–13. https://doi.org/10.1080/03014223.2023.2182332
Bell BD (1976) Status of Great Barrier Island birds. Notornis 23:310–319
Bell BD, Brathwaite DH (1964) The birds of Great Barrier and Arid Islands. Notornis 10:363–383
BirdLife International and Handbook of the Birds of the World. (2016). Acridotheres tristis (spatial data). In The IUCN Red List of Threatened Species. Version 2021-3. https://www.iucnredlist.org
CAB International (2021) Acridotheres tristis (common myna). https://www.cabi.org/isc/datasheet/2994. Accessed 19 January 2022
Cassey P, Prowse TAA, Blackburn TM (2014) A population model for predicting the successful establishment of introduced bird species. Oecologia 175:417–428. https://doi.org/10.1007/s00442-014-2902-1
Catchen J, Hohenlohe PA, Bassham S, Amores A, Cresko WA (2013) Stacks: an analysis tool set for population genomics. Mol Ecol 22:3124–3140. https://doi.org/10.1111/mec.12354
Cheke AS, Hume JP (2008) Lost land of the dodo: an ecological history of Mauritius, Réunion & Rodrigues. Yale University Press, London
Chen S, Zhou Y, Chen Y, Gu J (2018) fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34:i884–i890. https://doi.org/10.1093/bioinformatics/bty560
Clark NJ, Olsson-Pons S, Ishtiaq F, Clegg SM (2015) Specialist enemies, generalist weapons and the potential spread of exotic pathogens: malaria parasites in a highly invasive bird. Int J Parasitol 45:891–899. https://doi.org/10.1016/j.ijpara.2015.08.008
Clavero M, García-Berthou E (2005) Invasive species are a leading cause of animal extinctions. Trends Ecol Evol 20:110–110. https://doi.org/10.1016/j.tree.2005.01.003
Cruz VMV, Kilian A, Dierig DA (2013) Development of DArT Marker Platforms and Genetic Diversity Assessment of the U.S. Collection of the New Oilseed Crop Lesquerella and Related Species. PLOS ONE 8:e64062. https://doi.org/10.1371/journal.pone.0064062
Cunningham JM (1948) Distribution of myna in N.Z. Notornis 3:57–64
Danecek P, Auton A, Abecasis G, Albers CA, Banks E, DePristo MA et al. (2011) The variant call format and VCFtools. Bioinformatics 27:2156–2158. https://doi.org/10.1093/bioinformatics/btr330
Dawson DG, Bull PC (1970) A questionnaire survey of bird damage to fruit. NZ J Agric Res 13:362–371. https://doi.org/10.1080/00288233.1970.10425409
Evans T, Kumschick S, Şekercioğlu ÇH, Blackburn TM (2018) Identifying the factors that determine the severity and type of alien bird impacts. Divers Distrib 24:800–810. https://doi.org/10.1111/ddi.12721
Ewart KM, Griffin AS, Johnson RN, Kark S, Magory Cohen T, Lo N et al. (2019) Two speed invasion: assisted and intrinsic dispersal of common mynas over 150 years of colonization. J Biogeogr 46:45–57. https://doi.org/10.1111/jbi.13473
Ewels P, Magnusson M, Lundin S, Käller M (2016) MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32:3047–3048. https://doi.org/10.1093/bioinformatics/btw354
Excoffier L, Marchi N, Marques DA, Matthey-Doret R, Gouy A, Sousa VC (2021) fastsimcoal2: demographic inference under complex evolutionary scenarios. Bioinformatics 37:4882–4885. https://doi.org/10.1093/bioinformatics/btab468
Feare CJ, Bristol RM, van de Crommenacker J (2021) Eradication of a highly invasive bird, the common myna Acridotheres tristis, facilitates the establishment of insurance populations of island endemic birds. Bird Conserv Int 32:439–459. https://doi.org/10.1017/s0959270921000435
Feare C, Craig A (1999) Starlings and mynas. Princeton University Press, Princeton, New Jersey.
Fleischer RC, Williams RN, Baker AJ (1991) Genetic variation within and among populations of the Common Myna (Acridotheres tristis) in Hawaii. J Hered 82:205–208. https://doi.org/10.1093/oxfordjournals.jhered.a111066
Forsdick NJ, Martini D, Brown L, Cross HB, Maloney RF, Steeves TE et al. (2021) Genomic sequencing confirms absence of introgression despite past hybridisation between a critically endangered bird and its common congener. Glob Ecol Conserv 28:e01681. https://doi.org/10.1016/j.gecco.2021.e01681
Fournier A, Penone C, Pennino MG, Courchamp F (2019) Predicting future invaders and future invasions. Proc Natl Acad Sci USA 116:7905–7910. https://doi.org/10.1073/pnas.1803456116
Francis RM (2017) pophelper: an R package and web app to analyse and visualize population structure. Mol Ecol Resour 17:27–32. https://doi.org/10.1111/1755-0998.12509
Frichot E, François O (2015) LEA: An R package for landscape and ecological association studies. Methods Ecol Evol 6:925–929. https://doi.org/10.1111/2041-210x.12382
Global Invasive Species Database (2021) Species profile: Acridotheres tristis. http://www.iucngisd.org/gisd/species.php?sc=108&thinsp. Accessed 14 November 2021
Graffelman J (2015) Exploring diallelic genetic markers: the HardyWeinberg package. J Stat Softw 64:1–23. https://doi.org/10.18637/jss.v064.i03
Graffelman J, Camarena JM (2008) Graphical tests for Hardy-Weinberg Equilibrium based on the ternary plot. Hum Hered 65:77–84. https://doi.org/10.1159/000108939
Gruber B, Adamack AT (2015) landgenreport: a new R function to simplify landscape genetic analysis using resistance surface layers. Mol Ecol Resour 15:1172–1178. https://doi.org/10.1111/1755-0998.12381
Gruber B, Unmack PJ, Berry OF, Georges A (2018) DARTR: An R package to facilitate analysis of SNP data generated from reduced representation genome sequencing. Mol Ecol Resour 18:691–699. https://doi.org/10.1111/1755-0998.12745
Hart LA, Rogers AM, van Rensburg BJ (2020) Common Myna (Acridotheres tristis Linnaeus, 1766). In: Downs CT, Hart LA (eds) Invasive birds: global trends and impacts, CAB International, Wallingford, pp 25–32.
Hawke’s bay acclimatisation society (20 March 1877) Hawke’s Bay Herald. Page 2. https://paperspast.natlib.govt.nz/newspapers/HBH18770320.2.11. Accessed 21 January 2022
Heather BD, Robertson HA (2015) The field guide to the birds of New Zealand, 4th edn. Penguin Random House New Zealand Limited, Auckland.
Hofmeister NR, Werner SJ, Lovette IJ (2021) Environmental correlates of genetic variation in the invasive european starling in North America. Mol Ecol 30:1251–1263. https://doi.org/10.1111/mec.15806
Hone J (1978) Introduction and spread of the common myna in New South Wales. Emu 78:227–230. https://doi.org/10.1071/mu9780227
Huddleston F Mr Huddleston and the acclimatisation society. Nelson Evening Mail. https://paperspast.natlib.govt.nz/newspapers/NEM18680901.2.10.1. Accessed 13 December 2021
Hulme PE (2009) Trade, transport and trouble: managing invasive species pathways in an era of globalization. J Appl Ecol 46:10–18. https://doi.org/10.1111/j.1365-2664.2008.01600.x
Kalinowski ST (2005) HP-RARE 1.0: a computer program for performing rarefaction on measures of allelic richness. Mol Ecol Notes 5:187–189. https://doi.org/10.1111/j.1471-8286.2004.00845.x
Kekkonen J, Seppä P, Hanski IK, Jensen H, Väisänen RA, Brommer JE (2011) Low genetic differentiation in a sedentary bird: house sparrow population genetics in a contiguous landscape. Heredity (Edinb) 106:183–190. https://doi.org/10.1038/hdy.2010.32
Ketema S, Tesfaye B, Keneni G, Amsalu Fenta B, Assefa E, Greliche N et al. (2020) DArTSeq SNP-based markers revealed high genetic diversity and structured population in Ethiopian cowpea [Vigna unguiculata (L.) Walp] germplasms. PLOS ONE 15:1–20. https://doi.org/10.1371/journal.pone.0239122
Kilian A, Wenzl P, Huttner E, Carling J, Xia L, Blois H et al. (2012) Diversity Arrays Technology: A Generic Genome Profiling Technology on Open Platforms. Methods Mol Biol 888:67–89. https://doi.org/10.1007/978-1-61779-870-2_5
Li H (2011) A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27:2987–2993. https://doi.org/10.1093/bioinformatics/btr509
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25:1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N et al. (2009) The Sequence Alignment/Map format and SAMtools. Bioinformatics 25:2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Long JL (1981) Introduced birds of the world: the worldwide history, distribution, and influence of birds introduced to new environments. David & Charles, Newton Abbot.
Luque GM, Bellard C, Bertelsmeier C, Bonnaud E, Genovesi P, Simberloff D et al. (2014) The 100th of the world’s worst invasive alien species. Biol Invasions 16:981–985. https://doi.org/10.1007/s10530-013-0561-5
Luu K, Bazin E, Blum MGB (2017) pcadapt: an R package to perform genome scans for selection based on principal component analysis. Mol Ecol Resour 17:67–77. https://doi.org/10.1111/1755-0998.12592
Marrs RA, Sforza R, Hufbauer RA (2008) When invasion increases population genetic structure: a study with Centaurea diffusa. Biol Invasions 10:561–572. https://doi.org/10.1007/s10530-007-9153-6
Matheson P, McGaughran A (2022) Genomic data is missing for many highly invasive species, restricting our preparedness for escalating incursion rates. Sci Rep. 12:1–8. https://doi.org/10.1038/s41598-022-17937-y
McCoy F, Ferguson JM, MacGillivray PH, Wild JJ (1885) Natural history of Victoria: Prodromus of the zoology of Victoria, or, figures and descriptions of the living species of all classes of the Victorian indigenous animals. By authority: J. Ferres, government printer, Melbourne.
Melville J, Haines ML, Boysen K, Hodkinson L, Kilian A, Smith Date KL et al. (2017) Identifying hybridization and admixture using SNPs: application of the DArTseq platform in phylogeographic research on vertebrates. R Soc Open Sci 4:161061. https://doi.org/10.1098/rsos.161061
Mussmann SM, Douglas MR, Chafin TK, Douglas ME (2019) BA3-SNPs: Contemporary migration reconfigured in BayesAss for next-generation sequence data. Methods Ecol Evol 10:1808–1813. https://doi.org/10.1111/2041-210x.13252
Near TJ, MacGuigan DJ, Parker E, Struthers CD, Jones CD, Dornburg A (2018) Phylogenetic analysis of Antarctic notothenioids illuminates the utility of RADseq for resolving Cenozoic adaptive radiations. Mol Phylogenet Evol 129:268–279. https://doi.org/10.1016/j.ympev.2018.09.001
Ogle CC (1981) Great Barrier Island wildlife survey. TANE 27:177–200
Parkes J (2006) Feasibility plan to eradicate Common Mynas (Acridotheres tristis) from Mangaia Island, Cook Islands. Lincoln. http://www.issg.org/cii/Electronic.references/pii/project_docs/cooks_myna/mangaia_myna_feasibility_plan_2006.pdf. Accessed 19 January 2022
Peacock DS, van Rensburg BJ, Robertson MP (2007) The distribution and spread of the invasive alien common myna, Acridotheres tristis L. (Aves: Sturnidae), in southern Africa. S Afr J Sci 103:465–473. 10520/ejc96625
Pearman WS, Urban L, Alexander A (2022) Commonly used Hardy–Weinberg equilibrium filtering schemes impact population structure inferences using RADseq data. Mol Ecol Resour 22:2599–2613. https://doi.org/10.1111/1755-0998.13646
Peterson BK, Weber JN, Kay EH, Fisher HS, Hoekstra HE (2012) Double digest RADseq: an inexpensive method for de novo SNP discovery and genotyping in model and non-model species. PLOS ONE 7:e37135. https://doi.org/10.1371/journal.pone.0037135
Pimentel D, Lach L, Zuniga R, Morrison D (2000) Environmental and economic costs of nonindigenous species in the United States. Bioscience 50:53–65. https://doi.org/10.1641/0006-3568(2000)050[0053:eaecon]2.3.co;2
Prentis PJ, Wilson JRU, Dormontt EE, Richardson DM, Lowe AJ (2008) Adaptive evolution in invasive species. Trends Plant Sci 13:288–294. https://doi.org/10.1016/j.tplants.2008.03.004
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1093/genetics/155.2.945
Privé F, Luu K, Blum MGB, McGrath JJ, Vilhjálmsson BJ (2020) Efficient toolkit implementing best practices for principal component analysis of population genetic data. Bioinformatics 36:4449–4457. https://doi.org/10.1093/bioinformatics/btaa520
Puechmaille SJ (2016) The program structure does not reliably recover the correct population structure when sampling is uneven: subsampling and new estimators alleviate the problem. Mol Ecol Resour 16:608–627. https://doi.org/10.1111/1755-0998.12512
Pyšek P, Richardson DM (2010) Invasive Species, Environmental Change and Management, and Health. Annu Rev Environ Resour 35:25–55. https://doi.org/10.1146/annurev-environ-033009-095548
R Core Team (2021) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing. Vienna, Austria. https://www.R-project.org/
Robertson CJR, HP, FMJ, PCR (2007) Atlas of bird distribution in New Zealand 1999-2004. The Ornithological Society of New Zealand, Wellington.
Rollins LA, Woolnough AP, Sherwin WB (2006) Population genetic tools for pest management: a review. Wildl Res 33:251–261. https://doi.org/10.1071/wr05106
Rollins LA, Woolnough AP, Wilton AN, Sinclair R, Sherwin WB (2009) Invasive species can’t cover their tracks: using microsatellites to assist management of starling (Sturnus vulgaris) populations in Western Australia. Mol Ecol 18:1560–1573. https://doi.org/10.1111/j.1365-294x.2009.04132.x
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-Statistics under isolation by distance. Genetics 145:1219–1228. https://doi.org/10.1093/genetics/145.4.1219
Saavedra S (2009) First control for Common myna (Acridotheres tristis) on Ascension island 2009. Canary Island. http://indianmynaaction.org.au/wp-content/uploads/2021/08/Ascension-Island-2009-Final-report.pdf. Accessed 19 January 2022
Safford R, Hawkins F (2013) The birds of Africa: Volume VIII: The Malagasy Region: Madagascar, Seychelles, Comoros, Mascarenes. Bloomsbury Publishing, London.
Shafer ABA, Peart CR, Tusso S, Maayan I, Brelsford A, Wheat CW et al. (2017) Bioinformatic processing of RAD-seq data dramatically impacts downstream population genetic inference. Methods Ecol Evol 8:907–917. https://doi.org/10.1111/2041-210x.12700
Simberloff D, Martin J-L, Genovesi P, Maris V, Wardle DA, Aronson J et al. (2013) Impacts of biological invasions: what’s what and the way forward. Trends Ecol Evol 28:58–66. https://doi.org/10.1016/j.tree.2012.07.013
Stuart KC, Cardilini APA, Cassey P, Richardson MF, Sherwin WB, Rollins LA et al. (2021) Signatures of selection in a recent invasion reveal adaptive divergence in a highly vagile invasive species. Mol Ecol 30:1419–1434. https://doi.org/10.1111/mec.15601
Thomson GM (1922) The naturalisation of animals and plants in New Zealand. Cambridge University Press, New Zealand
Te Uru Rākau (2020) The one billion trees programme: our future, our billion trees. Te Uru Rākau, Wellington, New Zealand. https://www.mpi.govt.nz/dmsdocument/31860/direct. Accessed 5 April 2023
Turbelin AJ, Malamud BD, Francis RA (2017) Mapping the global state of invasive alien species: patterns of invasion and policy responses. Glob Ecol Biogeogr 26:78–92. https://doi.org/10.1111/geb.12517
Wickham H (2016) ggplot2: elegant graphics for data analysis. Springer-Verlag New York.
Wilson GA, Rannala B (2003) Bayesian inference of recent migration rates using multilocus genotypes. Genetics 163:1177–1191. https://doi.org/10.1093/genetics/163.3.1177
Wright B, Farquharson KA, McLennan EA, Belov K, Hogg CJ, Grueber CE (2019) From reference genomes to population genomics: comparing three reference-aligned reduced-representation sequencing pipelines in two wildlife species. BMC Genom 20:453. https://doi.org/10.1186/s12864-019-5806-y
Yap CA-M, Sodhi NS, Brook BW (2002) Roost characteristics of invasive mynas in Singapore. J Wildl Manag 66:1118. https://doi.org/10.2307/3802943
Zenni RD, Essl F, García-Berthou E, McDermott SM (2021) The economic costs of biological invasions around the world. NeoBiota 67:1–9. https://doi.org/10.3897/neobiota.67.69971
Zhang C, Dong SS, Xu JY, He WM, Yang TL (2019) PopLDdecay: a fast and effective tool for linkage disequilibrium decay analysis based on variant call format files. Bioinformatics 35:1786–1788. https://doi.org/10.1093/bioinformatics/bty875
Acknowledgements
We extend many thanks to thank Mark Peck at the Royal Ontario Museum for providing myna samples. Our thanks to the Australian Museum for providing us with myna samples and sequence data, and to the many collectors who contributed myna samples from Australia. We are very grateful to Adrian Turner, Richard Taylor, Mark Mitchell, Helen Smith, Jeremy Warden, Clinton Duffy, Chris Bindon, James Russell, and Margaret Stanley for providing samples as part of control efforts in New Zealand. Thank you to Katarina Stuart for helpful feedback and discussion on the analyses. We thank The Royal Society Te Apārangi Marsden Grant (grant number UOA1911), Australian Museum Foundation and City of Sydney Environmental Grants Programme for funding. Kamolphat Atsawawaranunt (K.A.) would also like to acknowledge the University of Auckland Doctoral Scholarship for funding. Finally, we are very grateful to the editor Gerald Heckel and three anonymous reviewers for their insightful comments and suggestions that significantly improved the manuscript.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions.
Author information
Authors and Affiliations
Contributions
AW and AS designed the research and provided the guidelines and the ideas on the analyses. AW performed DNA extractions and KA performed the analyses and wrote the first draft. All authors contributed to the interpretation of the results and the final draft.
Corresponding authors
Ethics declarations
Competing interests
The author declares no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Associate editor Gerald Heckel.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Atsawawaranunt, K., Ewart, K.M., Major, R.E. et al. Tracing the introduction of the invasive common myna using population genomics. Heredity 131, 56–67 (2023). https://doi.org/10.1038/s41437-023-00621-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-023-00621-w
- Springer Nature Switzerland AG
This article is cited by
-
Projecting the current and potential future distribution of New Zealand’s invasive sturnids
Biological Invasions (2024)