Abstract
Non-coding RNAs (ncRNAs) account for the majority of the widespread transcripts of mammalian genomes. They rarely encode proteins and peptides, but their regulatory role is crucial in numerous physiological and pathological processes. The m6A (N6-methyladenosine) modification is one of the most common internal RNA modifications in eukaryotes and is associated with all aspects of RNA metabolism. Accumulating researches have indicated a close association between m6A modification and ncRNAs, and suggested m6A-modified ncRNAs played a crucial role in tumor progression. The correlation between m6A modification and ncRNAs offers a novel perspective for investigating the potential mechanisms of cancer pathological processes, which suggests that both m6A modification and ncRNAs are critical prognostic markers and therapeutic targets in numerous malignancies. In the present report, we summarized the interaction between m6A modification and ncRNA, emphasizing how their interaction regulates pathological processes in cancer.
Similar content being viewed by others
Facts
-
1.
Non-coding RNAs rarely encode proteins and peptides but play an important role in the pathological process of human cancer.
-
2.
N6-methyladenosine modification of ncRNA is a dynamic and reversible process, which is regulated by writers, erasers, and readers.
-
3.
N6-methyladenosine modification can influence non-coding RNA metabolism.
-
4.
N6-methyladenosine plays an important role in the cancer development.
-
5.
Small molecule inhibitors of m6A-related proteins have great therapeutic potential in human cancer.
Open questions
-
1.
Are there any writers, erasers, and readers that specifically target ncRNA?
-
2.
Can m6A-modified ncRNAs be effective diagnostic biomarkers in human cancer?
-
3.
What other functions are affected by m6A-modified ncRNAs in cancer pathology?
-
4.
Can m6A-modified ncRNAs be effective therapeutic targets in human cancer?
Introduction
In the human genome, two percent of genome is used to transcribe mRNAs, and the rest are used to transcribe ncRNAs [1]. NcRNAs can control gene expression during the growth and development of organisms. An increasing number of studies have suggested that dysregulated ncRNAs are associated with various diseases, especially cancer. Therefore, ncRNAs are expected to be targets for cancer diagnosis and therapy.
The N6-methyladenosine (m6A) modification is one of the most abundant RNA modification types [2], and it has been proven to be a crucial factor in the pathological process of cancer. M6A occurs most frequently in the stop codons and 3ʹ-untranslated region (3ʹ-UTR) of mRNA (Fig. 1A), which has a consistent classical motif RRACH (R = G or A and H = A, C, or U) (Fig. 1B) [3].
A Schematic representation of the distribution of m6A, m5C, m7G, m1A, and m6Am in the mammalian transcriptome. The m6A modification is generally enriched in long internal exons, stop codons, and 3ʹ-UTR. B The motifs of m6A, m5C, and m1A are conserved. C Molecular structures of RNA methylation: m6A, m5C, m6Am, m7G, and m1A.
The new literature also demonstrates abundant m6A modifications in ncRNAs, similar to mRNA. Accumulating researches have shown that m6A-modified ncRNAs play an important role in the pathological process of cancer, such as metastasis, tumor microenvironment, and therapy resistance. The present review summarized the most recent developments in m6A modification of ncRNAs, especially their molecular mechanisms of action and their functional roles in cancer, to provide some clues for the development of new strategies for cancer intervention.
Non-coding RNAs (ncRNAs)
NcRNA accounts for the majority of total RNAs in humans, and rarely encodes proteins or peptides [4]. Here, we mainly introduce miRNAs, lncRNAs, and circRNAs.
MiRNAs have only 22 to 25 nucleotides, a type of small ncRNAs [5], which can participate in the formation of RNA-induced silencing complex and interact with targeted mRNAs for further post-transcriptional inhibition. Recent mechanistic studies also suggest that miRNAs regulate the pathological process of tumor by forming competing endogenous RNA (ceRNA) signaling pathways with other lncRNA/circRNA.
LncRNAs have been identified as ncRNAs with a minimum length of 200 nucleotides a few years ago [6]. They have multiple biological functions, including acting as molecular scaffolds, interfering with the transcription of nearby pre-mRNA, and regulating gene expression in transcriptional, post-transcriptional or post-translational processes. In addition, a few lncRNAs can translate proteins or peptides with biological functions.
Additionally, circRNAs are produced by pre-mRNA back-splicing of introns/exons in eukaryotes [7]. Accumulating evidence has indicated that circRNAs could function as gene regulators and even encode functional proteins/peptides. Furthermore, numerous studies have reported that circRNAs might be potential prognostic markers or therapeutic targets in cancer.
m6A modification and detection methods
The regulatory molecular mechanisms of m6A modification
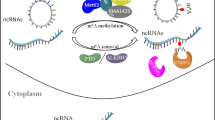
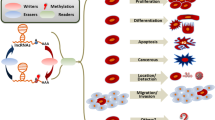
Over 160 different types of RNA modifications are currently known [8, 9], including m6A, m5C, m6Am, m7G, and m1A (Fig. 1C) [10, 11]. The priority regions and motifs of m5C, m6Am, m7G, and m1A are conserved (Fig. 1A, B). Since m6A is the most common RNA modification type in eukaryotic cells, we will focus on summarizing the role of m6A-modified ncRNAs in this study [12]. Recently, it has been discovered that m6A is dynamic and reversible [13, 14], and this fundamental characteristic of m6A is given by “writers”, “erasers”, and “readers” [15] (Fig. 2, Table 1). The m6A writers, erasers, and readers are m6A methylases, demethylases, and recognition proteins, respectively, which install m6A, remove m6A, and recognize m6A [16, 17].
The m6A is catalyzed by methyltransferase complexes (MTCs), including methyltransferase-like 3 (METTL3) [18], methyltransferase-like 14 (METTL14) [19, 20], KIAA1429 [21], Wilms tumor 1 associated protein (WTAP) [22], RNA binding motif protein 15 (RBM15) [23], and zinc finger CCCH domain-containing protein 13 (ZC3H13) [24]. METTL3 is the sole catalyst subunit in MTC. However, it becomes inactive in the absence of METTL14 during the formation of m6A [25, 26]. METTL14 primarily functions to stabilize the MTC and to recognize a specific RNA sequence (“RRACH”) as a catalytic substrate [25, 26]. METTL3 and METTL14 form a heterodimer in nuclear speckles with the assistance of WTAP. RBM15 mediates the binding of METTL3/14-WTAP complex to RNA for m6A modification [27]. In addition, KIAA1429 and ZC3H13 are components of MTC [28, 29]. However, current open questions include the exact physical relationship of the MTCs and the detailed mechanism of these methyltransferases. It is also noteworthy whether these writers are the potential diagnostic biomarkers and novel therapeutic targets for cancer.
RNA m6A modification could be reversibly removed, which required demethylases, also named erasers. Currently, the most widely studied m6A erasers are fat mass and obesity-associated enzyme (FTO) and AlkB homolog 5 (ALKBH5). Both FTO and ALKBH5 belong to the family of Fe (II)- and 2-oxoglutarate (2OG)-dependent AlkB dioxygenases [30, 31], which can remove m6A modification in the presence of Fe (II) and 2OG. Mechanically, m6A is oxidized to N6-hydroxymethyladenosine (hm6A), which is then transformed to N6-formyladenosine (f6A). Finally, the demethylation process is completed after the conversion of f6A to adenosine (A) [32]. Interestingly, recent research showed that FTO also possesses demethylase activity to N6,2’-O-dimethyladenosine (m6Am), indicating that FTO can catalyze the demethylation process of different substrates [33]. Undoubtedly, these erasers exert a critical role in m6A modifications, so additional efforts are needed to gain a more in-depth understanding.
The m6A modification plays different biological roles by being recognized by m6A readers. At present, there are three widely studied types of readers: YT521-B homology (YTH) domain family, insulin-like growth factor 2 mRNA-binding proteins (IGF2BPs), and heterogeneous nuclear ribonucleoproteins (HNRNPs) [34,35,36,37]. Members of the YTH domain family comprise YTH domain family protein 1–3 (YTHDF1-3) and YTH domain containing 1–2 (YTHDC1-2), with a conserved m6A binding domain that recognizes m6A modification [38]. The first discovered m6A reader is YTHDF2, which recognizes a specific m6A motif through its C-terminal region to regulate the stability of m6A-modified RNA. Moreover, its N-terminal recruits the CCR4-NOT deadenylase complex by binding to the SH domain of CCR4-NOT transcriptional complex subunit 1 (CNOT1), finally promoting the instability of m6A-modified RNA. YTHDF1 recognizes the m6A motif and binds to the translation initiation complex, which facilitates the translation of the m6A-modified RNA in a cap-independent manner [39, 40]. YTHDF3 assists YTHDF2 to accelerate m6A-modified RNA degradation or works together with YTHDF1 to promote m6A-modified RNA translation [41, 42]. YTHDC1 is mainly located in the nucleus, which not only facilitates exon inclusion but also accelerates the export of m6A-modified RNA from the nucleus to the cytoplasm [43,44,45]. YTHDC2 can facilitate the translation of m6A-modified RNA after recognizing the m6A motif [46, 47]. HNRNPA2/B1 is the most known m6A reader protein in the HNRNP family, which promotes the processing of primary microRNA (pri-miRNA) by recognizing m6A modification of some pri-miRNA and interacting with drosha ribonuclease III (DROSHA) and DiGeorge syndrome critical region 8 (DGCR8) [48]. In addition, IGF2BPs (including IGF2BP1-3) also recognize m6A modification to enhance the stability of mRNA and translation efficiency [49]. Since the biological function of m6A is required to be recognized by readers, inhibiting readers or blocking the recognition of readers to m6A may be a new strategy for tumor therapy.
In addition to the m6A-related enzymes already mentioned, several new m6A-related enzymes have recently been reported. METTL4 [50], METTL5 [51], METTL16 [52], and ZCCHC4 [53] are recently recognized as the m6A writers. Besides, FMRP Translational Regulator 1 (FMR1) and Proline rich coiled-coil 2 A (PRRC2A) are two novel m6A readers [54, 55]. Furthermore, a novel m6A eraser, AlkB homolog 3 (ALKBH3), has been reported [56]. The majority of the m6A writers, erasers, and readers target both mRNA and ncRNA, but few studies have reported m6A writers, erasers, and readers that specifically target ncRNA. Therefore, it is worth investigating whether there are m6A-related enzymes that specifically target ncRNA.
Detecting methods of m6A modification
In recent years, with the continuous exploration and research on m6A, a number of methods to detect m6A modification have been developed, which further promoted m6A research [57, 58].
Antibody-dependent methods
The m6A-specific antibody is most often necessary in high-throughput sequencing approaches for m6A [59]. For example, methylated RNA immunoprecipitation sequencing (MeRIP) is the first generation of m6A detection methods, which has facilitated the research progress of m6A [60, 61].
Cross-linking immunoprecipitation (CLIP) and high-throughput sequencing (genome-wide CLIP) have recently enabled the investigation of genome-wide RNA binding protein (RBP)-RNA binding at single base-pair resolution [62]. These approaches have evolved through the development of three distinct versions: high-throughput sequencing crosslinking immunoprecipitation (HITS-CLIP), photoactivatable ribonucleoside enhanced crosslinking and immunoprecipitation (PARCLIP), and individual-nucleotide crosslinking and immunoprecipitation (iCLIP) [63]. To more accurately detect the m6A modification site, two similar m6A-seq methods were developed: photo-cross-linking-assisted m6A sequencing strategy (PA-m6A-seq) [64, 65] and m6A individual-nucleotide resolution UV crosslinking and immunoprecipitation (miCLIP) [64, 66]. In addition, Molinie et al. exploited m6A-level and isoform-characterization sequencing (m6A-LAIC-seq) [60, 67, 68].
Although the m6A antibody-dependent sequencing methods mentioned above have been widely used, they still have several unavoidable disadvantages. The reproducibility (30%–60%) of detection results is poor because of the uneven quality of commercial antibodies, and the low specificity of some antibodies may lead to a high false-positive rate of test results [69]. Therefore, these findings indicate that it is necessary to find more effective and accurate methods for the detection of m6A modification (Table 2).
Antibody-independent methods
Many endoribonuclease-based approaches exist for detecting m6A, including MAZTER-Seq and m6A-sensitive RNA-endoribonuclease-facilitated sequencing (m6A-REF-seq), which is an example of an antibody-independent m6A sequencing strategy [70, 71]. MazF, an m6A-sensitive endoribonuclease, can cleave the ACA sequence but cannot cleave the m6ACA sequence [72,73,74]. MAZTER-seq and m6A-REF-seq were invented because of this characteristic.
Deamination adjacent to RNA modification targets (DART-Seq), an additional antibody-independent method for m6A sequencing, uses APOBEC1-YTH protein to induce C-to-U editing at sites next to m6A, thus identifying m6A sites [75].
Lately, two chemical labeling approaches (m6A-label-seq and m6A-SEAL) have been developed [76, 77]. By metabolically labeling target substrate adenosines in the m6A generation process, m6A-label-seq detects m6A modification at the base resolution, which is applicable to all m6A motif sequences [76]. The FTO-assisted m6A selective chemical labeling method (m6A-SEAL) specifically detects transcriptome-wide m6A [77].
In conclusion, although these approaches are no longer dependent on m6A antibodies, they can only identify specific m6A motifs, and their recognition efficiency is greatly affected by the efficiency of chemical reactions. Therefore, the development of new m6A recognition methods is still needed (Table 2).
Predicting m6A sites by databases
The bioinformatics field can significantly enhance research efficiency through the prediction of m6A modification sites. In this study, we summarized these prediction tools (Table 3).
While most studies have focused on the roles of m6A in mRNA, it is clear that we are just scratching the surface about the role of m6A-modified ncRNAs. The poor signal-to-noise ratio associated with commonly used m6A-mapping techniques is a major barrier to progress on that front, particularly since some ncRNAs tend to be less abundant. With the development of technology, other new technologies are also expected to promote a new understanding about m6A modification of ncRNAs, such as third-generation sequencing technology, newly designed mass spectrometry protocols technology and improved chromatography. Moreover, single-cell technologies are being applied to epitranscriptomics, which can reveal the heterogeneity and the spatial differences in m6A-modified ncRNAs. In a word, the development of a method for specifically detecting m6A modification of ncRNA is also a direction in our future exploration.
The biological role of m6A-modified ncRNAs
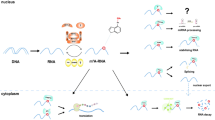
The m6A-modified ncRNAs can play different biological roles in intracellular and extracellular environments. We provide a brief introduction to these roles below (Fig. 3).
Top: In the nucleus, m6A modification can regulate splicing, processing, stability, interacting with RNA-binding proteins (RBP), and nucleus export of ncRNAs. The m6A-modified lncRNA mediates gene remodeling. In the cytoplasm, m6A modification mediates ncRNA translation, stability, ceRNA function, and interacting with RBP. Extracellular m6A modification can regulate circRNA immunity. Bottom: The m6A modification could regulate the processing, maturation, and ceRNA function of miRNAs. The m6A modification could regulate the stability, RBP interaction, and ceRNA function of lncRNAs, and m6A-modified lncRNA could regulate gene remodeling. The m6A modification could regulate the splicing, stability, translation, immunity, RBP interaction, and ceRNA function of circRNAs.
The role of m6A modification of ncRNA in the extracellular compartment
Mammalian cells possess innate immunity against foreign circRNAs, but it remains unclear what determines self-versus-foreign identity in circRNA immunity. However, recent research has proved that m6A modification of circRNAs could inhibit innate immunity. Specifically, foreign circRNAs (without m6A) induce a wide range of immune responses as potent adjuvants in vivo, including antigen-specific T cell activation, antibody production, and antitumor immunity, but m6A modification eliminates the adjuvant activity of foreign circRNAs. Mechanically, foreign circRNAs (without m6A) directly activate the RNA pattern recognition receptor RIG-I to activate the downstream transcription factor IRF3 (interferon regulatory factor 3). Activated IRF3 then forms a dimer for transport to the nucleus where it interacts with other transcription factors to activate the immune system. In contrast, m6A modification of foreign circRNAs can abrogate the activation of RIG-I. Thus, m6A-mediated disruption of RIG activation suppresses the activation of the immune system. For example, it was reported that m6A modification marked exogenous circFOREIGN as “self”, thereby inhibiting the activation of IRF3. Disruption of IFR3 further blocks activation of the antitumor immune system [78]. Taken together, these results suggest that m6A modification both sequesters and blocks exogenous circRNAs by activating the RIG-I pathway.
The role of m6A modification of ncRNA between the nucleus and cytoplasm
The m6A modification regulates the stability of ncRNAs
Recently, accumulating studies have verified that m6A modification play an important role in the regulation of the stability of ncRNAs.
It has been reported that human heat-responsive protein 12 (HRSP12) linked m6A “reader” YTHDF2 to RNase P/MRP (endoribonucleases) forming a YTHDF2-HRSP12-RNase P/MRP complex to influence the stability of m6A-modified circRNAs. Mechanically, the m6A-modified circRNAs that are preferentially targeted for endoribonucleolytic cleavage possess an HRSP12-binding site and an RNase P/MRP-directed cleavage site upstream and downstream of the YTHDF2-binding site, respectively. Therefore, HRSP12 can function as a connector to connect YTHDF2 and RNase P/MRP, eliciting endoribonucleolytic cleavage of YTHDF2-bound circRNAs [79, 80]. Although m6A modification regulating RNA stability has been widely reported, its molecular mechanisms still need to be clarified.
The m6A modification affects the interactions of ncRNAs and RNA binding proteins (RBPs)
Some studies showed that m6A modification on ncRNAs could affect the interaction between ncRNAs and RBPs. Mechanically, m6A modification prevents the formation of RNA local secondary structures and make RNA more easily recognized by RBPs through the “m6A switch” mechanism. For example, the mutation or upregulation of MALAT1, a conserved lncRNA, has been consistently associated with tumorigenesis and metastasis. The m6A modification of MALAT1 could increase the accessibility of RBPs (such as HNRNPC) by preventing the formation of its local secondary structures to expose its purine-rich sequences [36, 81]. In addition, recent accumulating researches reported that ncRNAs can regulate downstream target genes through interacting with m6A modulators (writers, erasers, readers). For example, PACERR, an upregulated lncRNA in pancreatic ductal adenocarcinoma (PDAC), contributes to increasing the number of M2-polarized cells and facilizing the malignant phenotype in PDAC by binding to m6A reader IGF2BP2 to increase the stability of downstream target genes KLF12 and c-myc [82].
The roles of m6A modification of ncRNA in the cytoplasm
The m6A modification promotes the translation of ncRNAs
NcRNAs are generally reported to be incapable of encoding proteins and peptides. However, with the study of m6A modification and ncRNAs, it was found that m6A modification can promote the translation of ncRNAs in a cap-independent manner, and the proteins/peptides produced by ncRNAs may be involved in the pathological process of cancer. Mechanically, m6A “reader” YTHDF3 could identify m6A modification. It promoted translation initiation factors (such as eIF3A and eIF4G2) and ribosomes to bind to the internal ribosome entry site (IRES), hence initiating the ncRNA translation process in a cap-independent manner. In addition, m6A methyltransferase and demethylase can enhance and inhibit m6A-dependent translation of circRNAs, respectively [83, 84]. For example, m6A modification makes circ-ZNF609 capable of translation by recruiting the translation initiation factor eIF4G2 [85]. M6A modification is one of the mechanisms underlying the translational potential of circRNA, which has been confirmed. However, how to identify circRNAs that encode proteins and how to judge whether these proteins have biological functions have not been illuminated. Therefore, m6A-modified circRNA might become a new hot spot for oncology research.
The m6A modification regulates the competing endogenous RNA (ceRNA) mechanism of ncRNAs
Many studies have reported that lncRNAs and circRNAs can function as molecular sponges of miRNAs to regulate the downstream target mRNAs of miRNAs, which is called the ceRNA mechanism [86]. Conversely, m6A modification could influence the ceRNA mechanism by regulating the stability of lncRNAs or circRNAs [87]. For instance, elevated m6A levels promoted circRNA-SORE stability, thereby upregulating circRNA-SORE. Subsequently, upregulated circRNA-SORE facilitated the hepatocellular carcinoma (HCC) progression by sponging miR-103a-2-5p and miR-660-3p, further competitively activating the Wnt/β-catenin signaling pathway [88]. In addition, m6A modification could control the ceRNA mechanism by influencing the maturation of miRNAs [89]. For example, m6A modification accelerated the splicing of immature miR-221/222 by recruiting Drosha and DiGeorge Critical Region 8 (DGCR8). The target gene PTEN expression of miR-221/222 was downregulated, which contributed to the proliferation of bladder cancer cells [90].
The role of m6A modification of ncRNA in the nucleus
The m6A modification promotes the nuclear export of ncRNAs
The m6A “reader” YTHDC1 has been shown to interact with nuclear export adaptor protein SRSF3, suggesting that m6A modification was responsible for the export of m6A-modified ncRNAs from the nucleus to the cytoplasm [44, 91]. Chen et al. found that silencing YTHDC1 increased the content of circNSUN2 in the nucleus. What’s more, the upregulation of wild-type YTHDC1 rescued the nuclear export deficiency of circNSUN2. Thus, the nuclear export of circNSUN2 is dependent on m6A modification [92].
The m6A modification accelerates the biogenesis of ncRNAs
The m6A modification can also regulate the biogenesis of ncRNAs by modulating their splicing. For circRNAs, reverse complementary sequences in transposable elements (TEs) promote cyclization, but the detailed mechanism remains elusive [93, 94]. Studies showed that m6A “writers” METTL3/14 bound to TEs, and the TEs in flanking introns of pre-mRNAs can form a stem-loop in back splicing to promote the cyclization of pre-mRNAs. As a result, this might be the potential mechanism by which m6A promotes circRNA biogenesis [80, 95]. A recent study demonstrated that METTL3 installed m6A in the reverse complementary sequences of flanking introns of circ1662, and facilitated the back splicing of circ1662 based on the intron pairing-driven circularization pattern [96].
The m6A modification promotes the maturation of miRNAs
The microprocessor complex comprising the endonuclease Drosha and DiGeorge Critical Region 8 (DGCR8) protein can cleave primary miRNA (pri-miRNA) into precursor miRNA (pre-miRNA). Pre-miRNAs are subsequently transported to the cytoplasm via exportin. In the cytoplasm, pre-miRNAs are further cleaved into mature miRNAs by Dicer [97]. Intriguingly, several reports demonstrated that m6A is a regulatory factor for promoting the maturation of pri-miRNAs. Mechanically, m6A “writer” METTL3 accounts for methylating pri-miRNA to accelerate its maturation via recruiting DGCR8 and m6A “reader” HNRNPA2/B1. Furthermore, HNRNPA2/B1 interacts with DGCR8 to facilitate the binding of DGCR8 to pri-miRNA, which increases the continuous generation of pre-miRNA [48, 98]. There are a number of examples of this regulatory pattern. For example, METTL3 promotes cell proliferation by facilitating the maturation of pri-miR221/222 in bladder cancer, which targets PTEN [90]. In colorectal cancer, METTL3 leads to an abnormal m6A level and promotes the production of mature miR-1246, which mediates cancer progression by inhibiting the SPRED/MAPK signaling pathway [99].
The m6A modification facilitates chromatin remodeling
Chromatin remodeling is a switch of chromatin structure. Specifically, the packaging state of chromatin, histones in nucleosomes, and corresponding DNA molecules change during gene expression. Currently, numerous studies have indicated the relationship between m6A modification and chromatin remodeling. For example, lncRNA X-inactive specific transcript (XIST) mediates X chromosome remodeling/silencing [23] and loss of m6A “writers” RBM15/RBM15B disrupt XIST-mediated X chromosome gene silencing in an m6A-dependent manner, demonstrating that m6A-modified ncRNAs promote chromatin remodeling.
Here we mainly reviewed the function of m6A modification of lncRNA, miRNA and circRNA in cancer. In addition to the common miRNAs, circRNAs, and lncRNAs, ncRNAs also include ribosomal RNA (rRNA), small nuclear RNA (snRNA), small nucleolar RNA (snoRNA), etc. However, the effect of the interaction between m6A and these ncRNAs is rarely studied in cancer pathology. On the other hand, the tumor immune microenvironment (TME) plays a critical role in cancer pathology and can affect responsiveness to immunotherapy. Despite many studies found that some relationships between m6A-modified ncRNAs and immunogenicity in cancer cells, there are few studies about the m6A-modified ncRNAs in immune cells in the TME. Therefore, more efforts are needed to uncover the roles and mechanisms of m6A-modified ncRNAs in cancer pathology.
The role of m6A-modified ncRNAs in common tumors
Since m6A and ncRNAs are both closely related to cancers, it is natural to speculate that m6A modification regulates the function of ncRNAs in various cancers. Herein, we summarize the latest findings of m6A modified ncRNAs in common cancers (Table 4).
Lung cancer
M6A modification affects lung cancer progression by regulating the stability of ncRNAs. For example, Qian et al. revealed that LCAT3, a novel lncRNA, was stabilized by m6A modification and subsequently activated c-MYC to promote lung cancer progression [100]. In addition, m6A modified circRNAs participated in the tumor immunity of lung cancer. The m6A-modified circNDUFB2 activates the RIG-I-MAVS signaling cascade and recruits immune cells into the TME to trigger anti-tumor immunity in NSCLC [101]. However, m6A-modified circIGF2BP3 restrains CD8 + T-cell responses and leads to tumor immune evasion by promoting the deubiquitylation of PD-L1 in NSCLC [102]. Moreover, m6A modification could accelerate lung cancer metastasis by facilitating the maturation of pri-miRNAs. Wang et al. showed that m6A-modified miR-143-3p promotes brain metastasis of lung cancer by increasing the splicing of precursor miR-143-3p to facilitate its biogenesis [103]. Additionally, recent studies have reported that m6A-related lncRNAs could serve as potential biomarkers for predicting prognosis and immune response in patients with lung adenocarcinoma (LUAD) [104]. Xu et al. constructed an m6A-related lncRNA risk model comprising 12 m6A-related lncRNAs in LUAD. They concluded that the risk model was identified as an independent predictor of prognosis, which might be promising for the clinical prediction of prognosis and immunotherapeutic response in LUAD patients [104].
These findings revealed the effect of m6a-modified ncRNAs on the progression of lung cancer and indicated that m6A-modified ncRNAs are potential predictive markers and therapeutic targets for lung cancer.
Hepatocellular carcinoma
Recent studies have demonstrated that a considerable number of m6A-modified ncRNAs are involved in the pathological process of HCC. The majority of researches have shown that m6A affects HCC progression or chemotherapy resistance by regulating the expression of ncRNAs. For instance, Wu et al. revealed that the downregulation of lncRNA MEG3 in m6A induced degradation manner accelerated the proliferation, migration, and invasion of HCC cells through the miR-544b/BTG2 (B-cell translocation gene) signal pathway. Zuo et al. showed that METTL3-mediated m6A modification led to LINC00958 upregulation by stabilizing its RNA transcript. The upregulated LINC00958 sponged miR-3619-5p to increase hepatoma-derived growth factor (HDGF) expression, thus facilitating HCC lipogenesis and progression. Furthermore, several studies have shown that many m6A-modified lncRNAs are partially overexpressed in tumor tissues and could be used to predict HCC prognosis in prognostic models, independent of other clinical features [105]. Yu et al. utilized LASSO regression to construct a prognostic model for m6A-modified lncRNAs in HCC. They discovered that several m6A-modified lncRNAs were partially upregulated in tumor tissues and could be used as independent prognostic markers in HCC [105].
Collectively, these reports demonstrate an essential role for m6A modified ncRNAs in HCC malignant behaviors and some m6A-modified ncRNAs as new therapeutic targets and predictors in HCC patients.
Glioma
M6A-modified ncRNAs have been shown to affect glioma progression by regulating either these ncRNAs stability or their target mRNAs. Chang et al. demonstrated that the stability of MALAT1, a classic oncogenic lncRNA, is highly dependent on m6A in glioma and the m6A-modified MALAT1 became more stable and promoted glioma progression [106].
Furthermore, several studies have reported that m6A-related lncRNAs are potential prognostic biomarkers to predict the progression of glioma [107, 108]. Tu et al. reported the use of m6A-related lncRNA in low-grade glioma (LGG) samples from the Cancer Genome Atlas (TCGA) and the Chinese Glioma Genome Atlas (CGGA) datasets to construct a prognostic model. As a result, 24 m6A-related lncRNAs were confirmed as independent prognostic marks for LGG in this prognostic model [108].
Taken together, these results show that m6A-related ncRNAs participate in the pathological process and serve as potential biomarkers in glioma.
Gastric cancer
Liu et al. reported that m6A modification post-transcriptionally stabilized lncRNA ThAP7-AS1, thereby promoting cancer progression in GC [109]. Moreover, m6A-mediated upregulation of lncRNA LINC01320 induces the proliferation, migration, and invasion of GC [110].
Furthermore, m6A modified ncRNAs have been shown to improve chemotherapy sensitivity by promoting the maturation of pri-miRNAs. As reported by Sun et al., m6A-dependent pri-miR-17-92 maturation increased the sensitivity to chemotherapy (everolimus) in GC [111]. Additionally, several studies have demonstrated that m6A-related lncRNAs have a robust signature for prognostic characterization and immunotherapy response in GC [112,113,114].
Collectively, these reports showed that m6A-modified ncRNAs were strongly associated with the pathological process of GC.
Brest cancer
In breast cancer (BC), it has been reported that m6A modification of ncRNAs facilitates translation initiation and also affects their stability. Rong et al. revealed that the N6 methylation at adenosine 1832 (m6A1832) of mammalian 18 S rRNA, which is pivotal in the decoding center, was modified by a conserved methyltransferase, METTL5. This modification triggers translation initiation by inducing conformational changes in the ribosomal decoding center, which facilitates mRNAs binding to the ribosomal decoding center [115].
Rong et al. showed that m6A modification increased the stability of LINC00958, which acted as a ceRNA of miR-378a-3p to facilitate BC occurrence via upregulation of YY1 transcription factor [116]. Likewise, m6A modification regulates the expression of circMETTL3 and circMETTL3 promotes BC progression by acting as a ceRNA of miR-31-5p to upregulate its target gene cyclin-dependent kinases (CDK1) [87].
In summary, these studies have laid a foundation for a better understanding of BC pathogenesis and development from the perspective of m6A-modified ncRNAs.
Colorectal cancer
Many reports have shown an important role of m6A-modified ncRNAs in colorectal cancer (CRC) development and progression through several potential mechanisms. One of which is m6A methylation modification of pri-miRNAs promoting their maturation. For example, Peng et al. demonstrated that m6A methyltransferase METTL3 installed m6A on pri-miR-1246 and facilitated the maturation of pri-miR-1246. Mature miR-1246 plays a pivotal role in tumor metastasis by downregulating its target gene SPRED2 in CRC. In contrast, Chen et al. revealed that the m6A writer METTL14 inhibited CRC progression by accelerating m6A-dependent pri-miR-375 maturation [117].
Another mechanism is m6A modification enhancing the stability of ncRNAs. Chen et al. found that m6A modification increased the stability of circ1662, and the accumulated circ1662 accelerated CRC cell invasion and migration by promoting YAP1 nuclear transport [96]. Moreover, m6A modification regulates the CRC progression by regulating the ceRNA mechanism of ncRNAs. Ma et al. found that LncRNA LBX2-AS1 promotes CRC progression and chemotherapy resistance (5-fluorouracil) by acting as a ceRNA to sponge miR-422a, which was enhanced by m6A methylation of LBX2-AS1 in a METTL3-dependent manner [118].
Accumulating studies have also shown that m6A-related lncRNAs could serve as prognostic markers in CRC [119,120,121]. Zeng et al. classified 473 CRC patients from TCGA into two subgroups through consensus clustering based on significant differences in survival. As result, a prognostic model was constructed by choosing 16 m6A-related lncRNAs. They concluded that CRC patients who had downregulated m6A-related lncRNAs expression had a higher risk score, indicating a poor prognosis [120].
The above studies have shown that despite m6A-modified ncRNAs being involved in the occurrence and progression of some cancers, there are few studies about m6A-modified ncRNAs in other cancers, such as sarcoma, uterine corpus endometrial carcinoma, and cholangiocarcinoma, etc. Additionally, the levels of m6A modification and ncRNAs have been regarded as potential diagnostic biomarkers. In addition, whether the expression profiles of m6A-modified ncRNAs can be used into the classification and stage or grade of cancers is a new direction. The expression profiles of m6A-modified ncRNAs combined with radiomics to analyze clinical phenotypes, therapeutic efficacy and clinical outcomes of patients is also a completely new field. Based on the discovered biological functions of m6A-modified ncRNAs, m6A modification showed its “double-edged sword” function. Thus, we suggest that future research on m6A-modified ncRNAs could help to elucidate their molecular mechanisms in cancer.
Clinical significance of m6A-modified ncRNAs
m6A-modified ncRNAs and cancer metastasis
One of the horrors of malignancy is its ability to metastasize. In recent years, researchers have found that m6A-modified ncRNAs are involved in cancer metastasis. For example, m6A-modified circMDK was significantly upregulated in HCC and could promote cancer metastasis in vivo and in vitro [122]. Therefore, we believe that m6A-modified ncRNAs play a role in the process of cancer metastasis, which suggests that m6A-modified ncRNAs may be used in clinical diagnosis to predict cancer metastasis.
m6A-modified ncRNAs and cancer chemoradiotherapy resistance
Many malignancies are resistant to chemoradiotherapy, thus resulting in poor prognosis of patients. Therefore, it is urgent to elucidate the underlying mechanisms of cancer chemoradiotherapy resistance. It has been reported that increased m6A levels in adipocyte exosomal LncRNAs can mediate myeloma drug resistance [123]. In contrast, increased m6A levels of lncRNA DBH-AS1 can inhibit gemcitabine resistance in pancreatic cancer [124]. In a word, these findings suggest that m6A-modified ncRNAs may be a potential direction for addressing cancer chemoradiotherapy resistance in the future.
m6A-modified ncRNAs and cancer immunotherapy
Immunotherapy is a new strategy of cancer therapy that improves the ability of immune system to kill cancer cells. There are also several studies about m6A-modified ncRNAs in cancer immunotherapy. For example, m6A-modified circRHBDD1 can restrict immunotherapy efficacy in HCC [125]. In addition, Lei et al. constructed a risk model, a 11-m6A-related lncRNAs, which can monitor immunotherapy for gastric cancer [126]. These studies suggest that m6A-modified ncRNAs play a critical role in cancer immunotherapy.
Small-molecule inhibitors of m6A-related proteins
Two METTL3 small-molecule inhibitors, STM2457 [127] and UZH1a [128], have recently been reported. Several potential inhibitors of FTO have also been identified including Rhein [129, 130], meclofenamic acid (MA) [131], bifunctional fluorescein derivatives [132], N-CDPCB [133], CHTB [134], FB23 [135], and CS1/2 [136]. Furthermore, two small molecule compounds, MV1305 [137] and ALK-04 [138], have been shown to inhibit ALKBH5. In addition, the BTYNB [139] compound selectively targets IGF2BP1. The information about these small-molecule inhibitors were summarized in Table 5.
Although the biological roles of m6A-modified ncRNAs in cancer metastasis, chemoradiotherapy resistance and immunotherapy have been extensively investigated in recent years, cancer therapy strategies based on m6A-modified ncRNAs are still virgin territory in the field of clinical application. Thus, we need to pay more attention to the potential of m6A-modified ncRNAs in clinical applications.
Conclusion and perspectives
In the present review, we briefly discussed the various key players and detection methods of m6A modification. We also outlined the biological roles of m6A-modified ncRNAs in some cancers. Some small-molecule inhibitors of m6A-related proteins are also summarized in detail. Altogether, we recapitulated the most recent advances in understanding the critical roles of m6A-modified ncRNAs in cancer. However, some specific issues should be clarified in future studies.
Firstly, the majority of the m6A writers, erasers, and readers target both mRNA and ncRNA, but few studies have reported m6A writers, erasers, and readers that specifically target ncRNA. Therefore, we hypothesized whether there might be some writers, erasers, and readers specifically targeting ncRNA, which needs to be focused on in future studies. Future studies addressing the cooperation of writers, erasers, and readers with the specifically targeting ncRNA will reveal the details of its association with cancer and the possibility of a therapeutic target.
Secondly, the potential diagnostic biomarkers will be not only the deregulated ncRNAs in cancer patients but also the chemically modified ncRNAs, such as m6A-modified ncRNAs. In addition, m6A can promote many ncRNAs to encode the specific proteins/peptides, which may also be the promising diagnostic markers in cancer. Moreover, the m6A-modified ncRNAs have recently been found in peripheral blood, which makes m6A-modified ncRNAs as diagnostic biomarkers in clinic application more promising and convenient. In a word, m6A-modified ncRNAs could be the effective diagnostic biomarkers in human cancer for the above reasons.
Thirdly, accumulating studies have shown that tumor cell stemness and metabolic reprogramming play an important role in tumor occurrence and progression. However, the interaction between m6A modified ncRNAs and tumor cell stemness or metabolism reprogramming is also rarely studied. Future studies are also expected to elucidate the heterogeneity and complexity of m6A-modified ncRNA in cancer tissues. Because the existing studies on m6A-modified ncRNAs are just the tip of the iceberg, we should pay more attention to the relationship between m6A-modified ncRNAs and other biological functions in cancer in the future.
Lastly, as mentioned above, the majority of drugs currently developed based on m6A are small molecule inhibitors of m6A-related enzymes. However, epitranscriptome editing, similar to genome editing, can also be used for editing m6A motifs in cancer, and such technology will likely be applied to the clinical treatment of cancer in the future. Recently, accumulating research has displayed that CRISPR technology can introduce or remove m6A at precise ncRNA sites. Thus, we hypothesized that CRISPR technology could make m6A-modified ncRNAs as the therapeutic targets for cancers by editing m6A motifs.
Overall, an increasing number of studies have validated that m6A-modified ncRNAs play a crucial role in human cancer occurrence and progression and m6A-modified ncRNA is a hot topic in oncology research, with potential prognostic and therapeutic prospects for a broad range of cancers.
Data availability
All data are available upon request.
References
Bikle DD. Vitamin D regulation of and by long non coding RNAs. Mol Cell Endocrinol. 2021;532:111317.
Wei J, Yu X, Yang L, Liu X, Gao B, Huang B, et al. FTO mediates LINE1 m(6)A demethylation and chromatin regulation in mESCs and mouse development. Science. 2022;376:968–73.
Oerum S, Meynier V, Catala M, Tisné C. A comprehensive review of m6A/m6Am RNA methyltransferase structures. Nucleic Acids Res. 2021;49:7239–55.
Wan G, Liu Y, Han C, Zhang X, Lu X. Noncoding RNAs in DNA repair and genome integrity. Antioxid Redox Signal. 2014;20:655–77.
Grosshans H, Slack FJ. Micro-RNAs: small is plentiful. J Cell Biol. 2002;156:17–21.
Zhang Y, Tao Y, Liao Q. Long noncoding RNA: a crosslink in biological regulatory network. Brief Bioinform. 2018;19:930–45.
Xue C, Li G, Zheng Q, Gu X, Bao Z, Lu J, et al. The functional roles of the circRNA/Wnt axis in cancer. Mol Cancer. 2022;21:108.
Begik O, Lucas MC, Liu H, Ramirez JM, Mattick JS, Novoa EM. Integrative analyses of the RNA modification machinery reveal tissue- and cancer-specific signatures. Genome Biol. 2020;21:97.
Barbieri I, Kouzarides T. Role of RNA modifications in cancer. Nat Rev Cancer. 2020;20:303–22.
Sun T, Wu R, Ming L. The role of m6A RNA methylation in cancer. Biomed Pharmacother. 2019;112:108613.
Zhang L, Hou C, Chen C, Guo Y, Yuan W, Yin D, et al. The role of N(6)-methyladenosine (m(6)A) modification in the regulation of circRNAs. Mol Cancer. 2020;19:105.
Gu Y, Wu X, Zhang J, Fang Y, Pan Y, Shu Y, et al. The evolving landscape of N(6)-methyladenosine modification in the tumor microenvironment. Mol Ther. 2021;29:1703–15.
Zhang S, Zhao BS, Zhou A, Lin K, Zheng S, Lu Z, et al. m(6)A Demethylase ALKBH5 Maintains Tumorigenicity of Glioblastoma Stem-like Cells by Sustaining FOXM1 Expression and Cell Proliferation Program. Cancer Cell. 2017;31:591–606. e596
Wang J, Li Y, Wang P, Han G, Zhang T, Chang J, et al. Leukemogenic chromatin alterations promote AML leukemia stem cells via a KDM4C-ALKBH5-AXL signaling axis. Cell Stem Cell. 2020;27:81–97. e88
Chen M, Wong CM. The emerging roles of N6-methyladenosine (m6A) deregulation in liver carcinogenesis. Mol Cancer. 2020;19:44.
Wang T, Kong S, Tao M, Ju S. The potential role of RNA N6-methyladenosine in Cancer progression. Mol Cancer. 2020;19:88.
Li HB, Tong J, Zhu S, Batista PJ, Duffy EE, Zhao J, et al. m(6)A mRNA methylation controls T cell homeostasis by targeting the IL-7/STAT5/SOCS pathways. Nature. 2017;548:338–42.
Chai RC, Chang YZ, Chang X, Pang B, An SY, Zhang KN, et al. YTHDF2 facilitates UBXN1 mRNA decay by recognizing METTL3-mediated m(6)A modification to activate NF-κB and promote the malignant progression of glioma. J Hematol Oncol. 2021;14:109.
Wang Y, Li Y, Toth JI, Petroski MD, Zhang Z, Zhao JC. N6-methyladenosine modification destabilizes developmental regulators in embryonic stem cells. Nat Cell Biol. 2014;16:191–8.
Ping XL, Sun BF, Wang L, Xiao W, Yang X, Wang WJ, et al. Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res. 2014;24:177–89.
Jiang X, Liu B, Nie Z, Duan L, Xiong Q, Jin Z, et al. The role of m6A modification in the biological functions and diseases. Signal Transduct Target Ther. 2021;6:74.
Jia GX, Lin Z, Yan RG, Wang GW, Zhang XN, Li C, et al. WTAP Function in Sertoli Cells Is Essential for Sustaining the Spermatogonial Stem Cell Niche. Stem Cell Rep. 2020;15:968–82.
Patil DP, Chen CK, Pickering BF, Chow A, Jackson C, Guttman M, et al. m(6)A RNA methylation promotes XIST-mediated transcriptional repression. Nature. 2016;537:369–73.
Wen J, Lv R, Ma H, Shen H, He C, Wang J, et al. Zc3h13 regulates nuclear RNA m(6)A methylation and mouse embryonic stem cell self-renewal. Mol Cell. 2018;69:1028–38. e1026
Wang X, Feng J, Xue Y, Guan Z, Zhang D, Liu Z, et al. Structural basis of N(6)-adenosine methylation by the METTL3-METTL14 complex. Nature. 2016;534:575–8.
Wang P, Doxtader KA, Nam Y. Structural basis for cooperative function of Mettl3 and Mettl14 methyltransferases. Mol Cell. 2016;63:306–17.
Knuckles P, Lence T, Haussmann IU, Jacob D, Kreim N, Carl SH, et al. Zc3h13/Flacc is required for adenosine methylation by bridging the mRNA-binding factor Rbm15/Spenito to the m(6)A machinery component Wtap/Fl(2)d. Genes Dev. 2018;32:415–29.
Yue Y, Liu J, Cui X, Cao J, Luo G, Zhang Z, et al. VIRMA mediates preferential m(6)A mRNA methylation in 3’UTR and near stop codon and associates with alternative polyadenylation. Cell Disco. 2018;4:10.
Rong ZX, Li Z, He JJ, Liu LY, Ren XX, Gao J, et al. Downregulation of fat mass and obesity associated (FTO) promotes the progression of intrahepatic cholangiocarcinoma. Front Oncol. 2019;9:369.
Gerken T, Girard CA, Tung YC, Webby CJ, Saudek V, Hewitson KS, et al. The obesity-associated FTO gene encodes a 2-oxoglutarate-dependent nucleic acid demethylase. Science. 2007;318:1469–72.
Kurowski MA, Bhagwat AS, Papaj G, Bujnicki JM. Phylogenomic identification of five new human homologs of the DNA repair enzyme AlkB. BMC Genomics. 2003;4:48.
Fu Y, Jia G, Pang X, Wang RN, Wang X, Li CJ, et al. FTO-mediated formation of N6-hydroxymethyladenosine and N6-formyladenosine in mammalian RNA. Nat Commun. 2013;4:1798.
Mauer J, Luo X, Blanjoie A, Jiao X, Grozhik AV, Patil DP, et al. Reversible methylation of m(6)A(m) in the 5’ cap controls mRNA stability. Nature. 2017;541:371–5.
Yang Y, Hsu PJ, Chen YS, Yang YG. Dynamic transcriptomic m(6)A decoration: writers, erasers, readers and functions in RNA metabolism. Cell Res. 2018;28:616–24.
Zaccara S, Ries RJ, Jaffrey SR. Reading, writing and erasing mRNA methylation. Nat Rev Mol Cell Biol. 2019;20:608–24.
Huang H, Weng H, Chen J. m(6)A modification in coding and non-coding RNAs: roles and therapeutic implications in cancer. Cancer Cell. 2020;37:270–88.
Zhou KI, Pan T. An additional class of m(6)A readers. Nat Cell Biol. 2018;20:230–2.
Wang X, Lu Z, Gomez A, Hon GC, Yue Y, Han D, et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature. 2014;505:117–20.
Wang X, Zhao BS, Roundtree IA, Lu Z, Han D, Ma H, et al. N(6)-methyladenosine modulates messenger RNA translation efficiency. Cell. 2015;161:1388–99.
Meyer KD, Patil DP, Zhou J, Zinoviev A, Skabkin MA, Elemento O, et al. 5’ UTR m(6)A promotes cap-independent translation. Cell. 2015;163:999–1010.
Li A, Chen YS, Ping XL, Yang X, Xiao W, Yang Y, et al. Cytoplasmic m(6)A reader YTHDF3 promotes mRNA translation. Cell Res. 2017;27:444–7.
Shi H, Wang X, Lu Z, Zhao BS, Ma H, Hsu PJ, et al. YTHDF3 facilitates translation and decay of N(6)-methyladenosine-modified RNA. Cell Res. 2017;27:315–28.
Xiao W, Adhikari S, Dahal U, Chen YS, Hao YJ, Sun BF, et al. Nuclear m(6)A Reader YTHDC1 Regulates mRNA Splicing. Mol Cell. 2016;61:507–19.
Roundtree IA, Luo GZ, Zhang Z, Wang X, Zhou T, Cui Y, et al. YTHDC1 mediates nuclear export of N(6)-methyladenosine methylated mRNAs. Elife. 2017;6:e31311.
Lesbirel S, Viphakone N, Parker M, Parker J, Heath C, Sudbery I, et al. The m(6)A-methylase complex recruits TREX and regulates mRNA export. Sci Rep. 2018;8:13827.
Wojtas MN, Pandey RR, Mendel M, Homolka D, Sachidanandam R, Pillai RS. Regulation of m(6)A transcripts by the 3’→5’ RNA helicase YTHDC2 is essential for a successful meiotic program in the Mammalian germline. Mol Cell. 2017;68:374–87. e312
Hsu PJ, Zhu Y, Ma H, Guo Y, Shi X, Liu Y, et al. Ythdc2 is an N(6)-methyladenosine binding protein that regulates mammalian spermatogenesis. Cell Res. 2017;27:1115–27.
Alarcón CR, Goodarzi H, Lee H, Liu X, Tavazoie S, Tavazoie SF. HNRNPA2B1 is a mediator of m(6)A-dependent nuclear RNA processing events. Cell. 2015;162:1299–308.
Huang H, Weng H, Sun W, Qin X, Shi H, Wu H, et al. Recognition of RNA N(6)-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Biol. 2018;20:285–95.
Gu L, Wang L, Chen H, Hong J, Shen Z, Dhall A, et al. CG14906 (mettl4) mediates m(6)A methylation of U2 snRNA in Drosophila. Cell Disco. 2020;6:44.
Ignatova VV, Stolz P, Kaiser S, Gustafsson TH, Lastres PR, Sanz-Moreno A, et al. The rRNA m(6)A methyltransferase METTL5 is involved in pluripotency and developmental programs. Genes Dev. 2020;34:715–29.
Aoyama T, Yamashita S, Tomita K. Mechanistic insights into m6A modification of U6 snRNA by human METTL16. Nucleic Acids Res. 2020;48:5157–68.
Ma H, Wang X, Cai J, Dai Q, Natchiar SK, Lv R, et al. N(6-)Methyladenosine methyltransferase ZCCHC4 mediates ribosomal RNA methylation. Nat Chem Biol. 2019;15:88–94.
Zhang G, Xu Y, Wang X, Zhu Y, Wang L, Zhang W, et al. Dynamic FMR1 granule phase switch instructed by m6A modification contributes to maternal RNA decay. Nat Commun. 2022;13:859.
Wu R, Li A, Sun B, Sun JG, Zhang J, Zhang T, et al. A novel m(6)A reader Prrc2a controls oligodendroglial specification and myelination. Cell Res. 2019;29:23–41.
Liu C, Yang S, Zhang Y, Wang C, Du D, Wang X, et al. Emerging roles of N6-methyladenosine demethylases and its interaction with environmental toxicants in digestive system cancers. Cancer Manag Res. 2021;13:7101–14.
Jora M, Lobue PA, Ross RL, Williams B, Addepalli B. Detection of ribonucleoside modifications by liquid chromatography coupled with mass spectrometry. Biochim Biophys Acta Gene Regul Mech. 2019;1862:280–90.
Reuter JA, Spacek DV, Snyder MP. High-throughput sequencing technologies. Mol Cell. 2015;58:586–97.
Zhao LY, Song J, Liu Y, Song CX, Yi C. Mapping the epigenetic modifications of DNA and RNA. Protein Cell. 2020;11:792–808.
Dominissini D, Moshitch-Moshkovitz S, Schwartz S, Salmon-Divon M, Ungar L, Osenberg S, et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature. 2012;485:201–6.
Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR. Comprehensive analysis of mRNA methylation reveals enrichment in 3’ UTRs and near stop codons. Cell. 2012;149:1635–46.
Zhang Z, Xing Y. CLIP-seq analysis of multi-mapped reads discovers novel functional RNA regulatory sites in the human transcriptome. Nucleic Acids Res. 2017;45:9260–71.
Wang T, Xiao G, Chu Y, Zhang MQ, Corey DR, Xie Y. Design and bioinformatics analysis of genome-wide CLIP experiments. Nucleic Acids Res. 2015;43:5263–74.
Linder B, Grozhik AV, Olarerin-George AO, Meydan C, Mason CE, Jaffrey SR. Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat Methods. 2015;12:767–72.
Körtel N, Rücklé C, Zhou Y, Busch A, Hoch-Kraft P, Sutandy FXR, et al. Deep and accurate detection of m6A RNA modifications using miCLIP2 and m6Aboost machine learning. Nucleic Acids Res. 2021;49:e92.
Chen K, Lu Z, Wang X, Fu Y, Luo GZ, Liu N, et al. High-resolution N(6) -methyladenosine (m(6) A) map using photo-crosslinking-assisted m(6) A sequencing. Angew Chem Int Ed Engl. 2015;54:1587–90.
Meyer KD, Jaffrey SR. The dynamic epitranscriptome: N6-methyladenosine and gene expression control. Nat Rev Mol Cell Biol. 2014;15:313–26.
Schwartz S, Mumbach MR, Jovanovic M, Wang T, Maciag K, Bushkin GG, et al. Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5’ sites. Cell Rep. 2014;8:284–96.
Haussmann IU, Bodi Z, Sanchez-Moran E, Mongan NP, Archer N, Fray RG, et al. m(6)A potentiates Sxl alternative pre-mRNA splicing for robust Drosophila sex determination. Nature. 2016;540:301–4.
Garcia-Campos MA, Edelheit S, Toth U, Safra M, Shachar R, Viukov S, et al. Deciphering the “m(6)A code” via antibody-independent quantitative profiling. Cell. 2019;178:731–47. e716
Zhang Z, Chen LQ, Zhao YL, Yang CG, Roundtree IA, Zhang Z, et al. Single-base mapping of m(6)A by an antibody-independent method. Sci Adv. 2019;5:eaax0250.
Wang X, Tian L, Li Y, Wang J, Yan B, Yang L, et al. RBM15 facilitates laryngeal squamous cell carcinoma progression by regulating TMBIM6 stability through IGF2BP3 dependent. J Exp Clin Cancer Res. 2021;40:80.
Bezrukov F, Prados J, Renzoni A, Panasenko OO. MazF toxin causes alterations in Staphylococcus aureus transcriptome, translatome and proteome that underlie bacterial dormancy. Nucleic Acids Res. 2021;49:2085–101.
Pandey RR, Pillai RS. Counting the cuts: MAZTER-Seq quantifies m(6)A levels using a methylation-sensitive ribonuclease. Cell. 2019;178:515–7.
Meyer KD. DART-seq: an antibody-free method for global m(6)A detection. Nat Methods. 2019;16:1275–80.
Shu X, Cao J, Cheng M, Xiang S, Gao M, Li T, et al. A metabolic labeling method detects m(6)A transcriptome-wide at single base resolution. Nat Chem Biol. 2020;16:887–95.
Wang Y, Xiao Y, Dong S, Yu Q, Jia G. Antibody-free enzyme-assisted chemical approach for detection of N(6)-methyladenosine. Nat Chem Biol. 2020;16:896–903.
Chen YG, Chen R, Ahmad S, Verma R, Kasturi SP, Amaya L, et al. N6-methyladenosine modification controls circular RNA immunity. Mol Cell. 2019;76:96–109. e109
Park OH, Ha H, Lee Y, Boo SH, Kwon DH, Song HK, et al. Endoribonucleolytic cleavage of m(6)A-containing RNAs by RNase P/MRP complex. Mol Cell. 2019;74:494–507. e498
Zhou C, Molinie B, Daneshvar K, Pondick JV, Wang J, Van Wittenberghe N, et al. Genome-wide maps of m6A circRNAs identify widespread and cell-type-specific methylation patterns that are distinct from mRNAs. Cell Rep. 2017;20:2262–76.
Zheng G, Dahl JA, Niu Y, Fedorcsak P, Huang CM, Li CJ, et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol Cell. 2013;49:18–29.
Liu Y, Shi M, He X, Cao Y, Liu P, Li F, et al. LncRNA-PACERR induces pro-tumour macrophages via interacting with miR-671-3p and m6A-reader IGF2BP2 in pancreatic ductal adenocarcinoma. J Hematol Oncol. 2022;15:52.
Yang Y, Fan X, Mao M, Song X, Wu P, Zhang Y, et al. Extensive translation of circular RNAs driven by N(6)-methyladenosine. Cell Res. 2017;27:626–41.
He L, Man C, Xiang S, Yao L, Wang X, Fan Y. Circular RNAs’ cap-independent translation protein and its roles in carcinomas. Mol Cancer. 2021;20:119.
Di Timoteo G, Dattilo D, Centrón-Broco A, Colantoni A, Guarnacci M, Rossi F, et al. Modulation of circRNA metabolism by m(6)A modification. Cell Rep. 2020;31:107641.
Qi X, Lin Y, Chen J, Shen B. Decoding competing endogenous RNA networks for cancer biomarker discovery. Brief Bioinform. 2020;21:441–57.
Li Z, Yang HY, Dai XY, Zhang X, Huang YZ, Shi L, et al. CircMETTL3, upregulated in a m6A-dependent manner, promotes breast cancer progression. Int J Biol Sci. 2021;17:1178–90.
Xu J, Wan Z, Tang M, Lin Z, Jiang S, Ji L, et al. N(6)-methyladenosine-modified CircRNA-SORE sustains sorafenib resistance in hepatocellular carcinoma by regulating β-catenin signaling. Mol Cancer. 2020;19:163.
Wang H, Song X, Song C, Wang X, Cao H. m(6)A-seq analysis of microRNAs reveals that the N6-methyladenosine modification of miR-21-5p affects its target expression. Arch Biochem Biophys. 2021;711:109023.
Han J, Wang JZ, Yang X, Yu H, Zhou R, Lu HC, et al. METTL3 promote tumor proliferation of bladder cancer by accelerating pri-miR221/222 maturation in m6A-dependent manner. Mol Cancer. 2019;18:110.
Rong D, Sun G, Wu F, Cheng Y, Sun G, Jiang W, et al. Epigenetics: Roles and therapeutic implications of non-coding RNA modifications in human cancers. Mol Ther Nucleic Acids. 2021;25:67–82.
Tao M, Zheng M, Xu Y, Ma S, Zhang W, Ju S. CircRNAs and their regulatory roles in cancers. Mol Med. 2021;27:94.
Zhang XO, Wang HB, Zhang Y, Lu X, Chen LL, Yang L. Complementary sequence-mediated exon circularization. Cell. 2014;159:134–47.
Ashwal-Fluss R, Meyer M, Pamudurti NR, Ivanov A, Bartok O, Hanan M, et al. circRNA biogenesis competes with pre-mRNA splicing. Mol Cell. 2014;56:55–66.
Capel B, Swain A, Nicolis S, Hacker A, Walter M, Koopman P, et al. Circular transcripts of the testis-determining gene Sry in adult mouse testis. Cell. 1993;73:1019–30.
Chen C, Yuan W, Zhou Q, Shao B, Guo Y, Wang W, et al. N6-methyladenosine-induced circ1662 promotes metastasis of colorectal cancer by accelerating YAP1 nuclear localization. Theranostics. 2021;11:4298–315.
Gebert LFR, MacRae IJ. Regulation of microRNA function in animals. Nat Rev Mol Cell Biol. 2019;20:21–37.
Alarcón CR, Lee H, Goodarzi H, Halberg N, Tavazoie SF. N6-methyladenosine marks primary microRNAs for processing. Nature. 2015;519:482–5.
Peng W, Li J, Chen R, Gu Q, Yang P, Qian W, et al. Upregulated METTL3 promotes metastasis of colorectal Cancer via miR-1246/SPRED2/MAPK signaling pathway. J Exp Clin Cancer Res. 2019;38:393.
Qian X, Yang J, Qiu Q, Li X, Jiang C, Li J, et al. LCAT3, a novel m6A-regulated long non-coding RNA, plays an oncogenic role in lung cancer via binding with FUBP1 to activate c-MYC. J Hematol Oncol. 2021;14:112.
Li B, Zhu L, Lu C, Wang C, Wang H, Jin H, et al. circNDUFB2 inhibits non-small cell lung cancer progression via destabilizing IGF2BPs and activating anti-tumor immunity. Nat Commun. 2021;12:295.
Liu Z, Wang T, She Y, Wu K, Gu S, Li L, et al. N(6)-methyladenosine-modified circIGF2BP3 inhibits CD8(+) T-cell responses to facilitate tumor immune evasion by promoting the deubiquitination of PD-L1 in non-small cell lung cancer. Mol Cancer. 2021;20:105.
Wang H, Deng Q, Lv Z, Ling Y, Hou X, Chen Z, et al. N6-methyladenosine induced miR-143-3p promotes the brain metastasis of lung cancer via regulation of VASH1. Mol Cancer. 2019;18:181.
Xu F, Huang X, Li Y, Chen Y, Lin L. m(6)A-related lncRNAs are potential biomarkers for predicting prognoses and immune responses in patients with LUAD. Mol Ther Nucleic Acids. 2021;24:780–91.
Yu ZL, Zhu ZM. Comprehensive analysis of N6-methyladenosine -related long non-coding RNAs and immune cell infiltration in hepatocellular carcinoma. Bioengineered. 2021;12:1708–24.
Chang YZ, Chai RC, Pang B, Chang X, An SY, Zhang KN, et al. METTL3 enhances the stability of MALAT1 with the assistance of HuR via m6A modification and activates NF-κB to promote the malignant progression of IDH-wildtype glioma. Cancer Lett. 2021;511:36–46.
Wang W, Li J, Lin F, Guo J, Zhao J. Identification of N(6)-methyladenosine-related lncRNAs for patients with primary glioblastoma. Neurosurg Rev. 2021;44:463–70.
Tu Z, Wu L, Wang P, Hu Q, Tao C, Li K, et al. N6-Methylandenosine-Related lncRNAs Are Potential Biomarkers for Predicting the Overall Survival of Lower-Grade Glioma Patients. Front Cell Dev Biol. 2020;8:642.
Liu HT, Zou YX, Zhu WJ, Sen L, Zhang GH, Ma RR, et al. lncRNA THAP7-AS1, transcriptionally activated by SP1 and post-transcriptionally stabilized by METTL3-mediated m6A modification, exerts oncogenic properties by improving CUL4B entry into the nucleus. Cell Death Differ. 2022;29:627–41.
Hu N, Ji H. N6-methyladenosine (m6A)-mediated up-regulation of long noncoding RNA LINC01320 promotes the proliferation, migration, and invasion of gastric cancer via miR495-5p/RAB19 axis. Bioengineered. 2021;12:4081–91.
Sun Y, Li S, Yu W, Zhao Z, Gao J, Chen C, et al. N(6)-methyladenosine-dependent pri-miR-17-92 maturation suppresses PTEN/TMEM127 and promotes sensitivity to everolimus in gastric cancer. Cell Death Dis. 2020;11:836.
Wang H, Meng Q, Ma B. Characterization of the prognostic m6A-related lncRNA signature in gastric cancer. Front Oncol. 2021;11:630260.
Han T, Xu D, Zhu J, Li J, Liu L, Deng Y. Identification of a robust signature for clinical outcomes and immunotherapy response in gastric cancer: based on N6-methyladenosine related long noncoding RNAs. Cancer Cell Int. 2021;21:432.
Yu ZL, Zhu ZM. N6-methyladenosine related long non-coding RNAs and immune cell infiltration in the tumor microenvironment of gastric cancer. Biol Proced Online. 2021;23:15.
Rong B, Zhang Q, Wan J, Xing S, Dai R, Li Y, et al. Ribosome 18S m(6)A methyltransferase METTL5 promotes translation initiation and breast cancer cell growth. Cell Rep. 2020;33:108544.
Rong D, Dong Q, Qu H, Deng X, Gao F, Li Q, et al. m(6)A-induced LINC00958 promotes breast cancer tumorigenesis via the miR-378a-3p/YY1 axis. Cell Death Disco. 2021;7:27.
Chen X, Xu M, Xu X, Zeng K, Liu X, Sun L, et al. METTL14 suppresses CRC progression via regulating n6-methyladenosine-dependent primary miR-375 processing. Mol Ther. 2020;28:599–612.
Ma YN, Hong YG, Yu GY, Jiang SY, Zhao BL, Guo A, et al. LncRNA LBX2-AS1 promotes colorectal cancer progression and 5-fluorouracil resistance. Cancer Cell Int. 2021;21:501.
Zuo L, Su H, Zhang Q, Wu WY, Zeng Y, Li XM, et al. Comprehensive analysis of lncRNAs N(6)-methyladenosine modification in colorectal cancer. Aging (Albany NY). 2021;13:4182–98.
Zeng H, Xu Y, Xu S, Jin L, Shen Y, Rajan KC, et al. Construction and analysis of a colorectal cancer prognostic model based on N6-methyladenosine-related lncRNAs. Front Cell Dev Biol. 2021;9:698388.
Song W, Ren J, Yuan W, Xiang R, Ge Y, Fu T N6-Methyladenosine-Related lncRNA Signature Predicts the Overall Survival of Colorectal Cancer Patients. Genes. 2021, 12.
Du A, Li S, Zhou Y, Disoma C, Liao Y, Zhang Y, et al. M6A-mediated upregulation of circMDK promotes tumorigenesis and acts as a nanotherapeutic target in hepatocellular carcinoma. Mol Cancer. 2022;21:109.
Wang Z, He J, Bach DH, Huang YH, Li Z, Liu H, et al. Induction of m(6)A methylation in adipocyte exosomal LncRNAs mediates myeloma drug resistance. J Exp Clin Cancer Res. 2022;41:4.
Ye X, Wang LP, Han C, Hu H, Ni CM, Qiao GL, et al. Increased m(6)A modification of lncRNA DBH-AS1 suppresses pancreatic cancer growth and gemcitabine resistance via the miR-3163/USP44 axis. Ann Transl Med. 2022;10:304.
Cai J, Chen Z, Zhang Y, Wang J, Zhang Z, Wu J, et al. CircRHBDD1 augments metabolic rewiring and restricts immunotherapy efficacy via m(6)A modification in hepatocellular carcinoma. Mol Ther Oncolytics. 2022;24:755–71.
Lei L, Li N, Yuan P, Liu D. A new risk model based on a 11-m(6)A-related lncRNA signature for predicting prognosis and monitoring immunotherapy for gastric cancer. BMC Cancer. 2022;22:365.
Yankova E, Blackaby W, Albertella M, Rak J, De Braekeleer E, Tsagkogeorga G, et al. Small-molecule inhibition of METTL3 as a strategy against myeloid leukaemia. Nature. 2021;593:597–601.
Moroz-Omori EV, Huang D, Kumar Bedi R, Cheriyamkunnel SJ, Bochenkova E, Dolbois A, et al. METTL3 inhibitors for epitranscriptomic modulation of cellular processes. ChemMedChem. 2021;16:3035–43.
Chen B, Ye F, Yu L, Jia G, Huang X, Zhang X, et al. Development of cell-active N6-methyladenosine RNA demethylase FTO inhibitor. J Am Chem Soc. 2012;134:17963–71.
Niu Y, Lin Z, Wan A, Chen H, Liang H, Sun L, et al. RNA N6-methyladenosine demethylase FTO promotes breast tumor progression through inhibiting BNIP3. Mol Cancer. 2019;18:46.
Huang Y, Yan J, Li Q, Li J, Gong S, Zhou H, et al. Meclofenamic acid selectively inhibits FTO demethylation of m6A over ALKBH5. Nucleic Acids Res. 2015;43:373–84.
Wang T, Hong T, Huang Y, Su H, Wu F, Chen Y, et al. Fluorescein derivatives as bifunctional molecules for the simultaneous inhibiting and labeling of FTO protein. J Am Chem Soc. 2015;137:13736–9.
He W, Zhou B, Liu W, Zhang M, Shen Z, Han Z, et al. Identification of a novel small-molecule binding site of the fat mass and obesity associated protein (FTO). J Med Chem. 2015;58:7341–8.
Qiao Y, Zhou B, Zhang M, Liu W, Han Z, Song C, et al. A novel inhibitor of the obesity-related protein FTO. Biochemistry. 2016;55:1516–22.
Huang Y, Su R, Sheng Y, Dong L, Dong Z, Xu H, et al. Small-molecule targeting of oncogenic FTO demethylase in acute myeloid leukemia. Cancer Cell. 2019;35:677–91. e610
Su R, Dong L, Li Y, Gao M, Han L, Wunderlich M, et al. Targeting FTO suppresses cancer stem cell maintenance and immune evasion. Cancer Cell. 2020;38:79–96. e11
Malacrida A, Rivara M, Di Domizio A, Cislaghi G, Miloso M, Zuliani V, et al. 3D proteome-wide scale screening and activity evaluation of a new ALKBH5 inhibitor in U87 glioblastoma cell line. Bioorg Med Chem. 2020;28:115300.
Li N, Kang Y, Wang L, Huff S, Tang R, Hui H, et al. ALKBH5 regulates anti-PD-1 therapy response by modulating lactate and suppressive immune cell accumulation in tumor microenvironment. Proc Natl Acad Sci USA. 2020;117:20159–70.
Müller S, Bley N, Busch B, Glaß M, Lederer M, Misiak C, et al. The oncofetal RNA-binding protein IGF2BP1 is a druggable, post-transcriptional super-enhancer of E2F-driven gene expression in cancer. Nucleic Acids Res. 2020;48:8576–90.
Acknowledgements
This research was supported by the National Natural Science Foundation of China (82173090, U1804172 to Xianzhi Liu, 81773187 to Lei Han and 81702465 to Zhenyu Zhang).
Author information
Authors and Affiliations
Contributions
LH, XL and ZZ conceived the review. LL and YZ drafted the manuscript and revised it before submission. DP, CW and XZ collected the references. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Luo, L., Zhen, Y., Peng, D. et al. The role of N6-methyladenosine-modified non-coding RNAs in the pathological process of human cancer. Cell Death Discov. 8, 325 (2022). https://doi.org/10.1038/s41420-022-01113-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-022-01113-2
- Springer Nature Limited
This article is cited by
-
Transcriptome-wide profiling identifies colon cancer-associated m6A transcripts and potential RNA methyl modifiers
Molecular Biology Reports (2024)