Abstract
Background
Biliary tract cancers (BTC) are rare but highly aggressive tumours with poor prognosis, usually detected at advanced stages. Herein, we aimed at identifying BTC-specific DNA methylation alterations.
Methods
Study design included statistical power and sample size estimation. A genome-wide methylation study of an explorative cohort (50 BTC and ten matched non-tumoral tissue samples) has been performed. BTC-specific altered CpG islands were validated in over 180 samples (174 BTCs and 13 non-tumoral controls). The final biomarkers, selected by a machine-learning approach, were validated in independent tissue (18 BTCs, 14 matched non-tumoral samples) and bile (24 BTCs, five non-tumoral samples) replication series, using droplet digital PCR.
Results
We identified and successfully validated BTC-specific DNA methylation alterations in over 200 BTC samples. The two-biomarker panel, selected by an in-house algorithm, showed an AUC > 0.97. The best-performing biomarker (chr2:176993479-176995557), associated with HOXD8, a pivotal gene in cancer-related pathways, achieved 100% sensitivity and specificity in a new series of tissue and bile samples.
Conclusions
We identified a novel fully efficient BTC biomarker, associated with HOXD8 gene, detectable both in tissue and bile by a standardised assay ready-to-use in clinical trials also including samples from non-invasive matrices.
Similar content being viewed by others
Background
Biliary tract cancer (BTC) comprises a group of highly aggressive malignancies clinically classified as intrahepatic and extrahepatic cholangiocarcinomas (CCAs) and gallbladder cancers (GBCs).
BTC incidence and mortality vary according to geographic regions and are related to the distribution of risk factors associated with this cancer [1]. In Western countries, the main risk factors for CCA include biliary tract diseases such as benign stenosis, primary sclerosing cholangitis (PSC), hepatolithiasis and choledochal cysts, and these tumours display low but gradually increasing incidence rates [2]. Early detection of CCA in PSC patients is difficult since the associated-inflammatory process leads to biliary strictures mimicking early neoplastic changes [3].
Due to its silent evolution and clinical manifestations only at advanced stages, BTCs are usually diagnosed when the tumour is locally advanced or metastatic, thus unresectable.
The current diagnostic strategy for BTC includes a combination of clinical, radiological, biochemical and histological approaches [4]. Endoscopic retrograde cholangiopancreatography (ERCP) combined with biliary brush cytology and cyto-histological analysis of tumour tissue could be performed to confirm a suspected case of BTC [5].
Regrettably, current diagnostic methods have shown limited specificity and sensitivity [6, 7]. The use of biomarkers is a promising alternative for the detection of BTC and some of them have already been implemented in the clinic, i.e., the carcinoembryonic antigen and the carbohydrate antigen 19-9 (CA 19-9). However, elevated levels of these markers have also been found in benign conditions challenging their specificity [4]. As a consequence, accurate diagnosis may prove challenging, highlighting the need for a detection method for BTC.
DNA methylation alterations are early events during tumorigenesis and may be detected as early as in preneoplastic lesions in many types of tumours [8,9,10,11,12], including CCA [13,14,15] and even several years prior to tumour diagnosis [16].
Several DNA methylation-based biomarkers, with the specificity of 100% and sensitivity values ranging between 75 and 89%, have been proposed to detect BTC in tissue samples [17,18,19]. However, many of these studies focused on biomarkers that are frequently hypermethylated also in other types of cancers, often with a higher incidence than BTC.
Genome-wide methylation profiling represents a promising strategy for the discovery of new biomarkers specific to BTC. To our knowledge, very few studies have conducted a global methylation analysis on BTC samples [20, 21].
An important advantage for clinical implementation is that methylation alterations can also be detected in cell-free DNA (cfDNA) from different matrices such as blood, urine and stool [8, 22,23,24,25], greatly facilitating their implementation in the clinical setting.
Several studies demonstrated that DNA methylation alterations can be detected in bile [13, 26], biliary brush cytology specimens [27, 28], plasma [18] and serum samples [29] from patients with BTC, although not always representing BTC-specific biomarkers.
In order to select BTC methylation alterations also detectable in non-invasive matrices such as blood and faeces, it becomes of crucial importance to identify extremely specific biomarkers for BTC and not for other cancers, especially those of gastrointestinal origin.
The primary aim of this work was to identify BTC-specific DNA methylation biomarkers, with high sensitivity and specificity regardless of the tumour localisation. We performed a whole-genome methylation profiling of 50 BTC tissue samples from different localisation (intrahepatic, extrahepatic and gallbladder) and ten matched-normal tissue samples using Illumina EPIC® arrays. Secondly, we aimed at assessing the performance of the best biomarker, associated with HOXD8 gene, in bile samples from BTC patients, using droplet digital PCR (ddPCR), currently the most sensitive technology. Finally, we tested whether this biomarker was specific for BTC or could also be detected in subjects at high risk of developing BTC, such as patients with benign stenosis.
Methods
Samples and data collection
Tissue samples
Discovery cohort for whole-genome methylation assay
Fifty formalin-fixed paraffin-embedded (FFPE) tumour tissue samples and ten matched-normal controls from BTC patients (25 males and 25 females, mean age at diagnosis: 70.4 ± 10.9, 22 intrahepatic, 20 gallbladders and 8 extrahepatic) were obtained from the Oncology Service, Department of Medical Sciences and Public Health of the University of Cagliari (Italy) and IRCCS-Istituto Romagnolo per lo Studio dei Tumori (IRST) “Dino Amadori”, Meldola (Italy). Multiple haematoxylins and eosin slides were reviewed by an expert pathologist and sections including at least 50% of neoplastic tissue were selected as tumour samples, while sections devoid of malignant cells were selected as normal samples.
Samples were tested for CA 19-9 in the respective clinical centres.
Clinical characteristics of BTC patients are shown in Supplementary Table S1.
Validation cohort for droplet digital PCR assay
Explorative tissue test series: For an explorative analysis, 32 samples (14 paired BTC/normal samples, four BTC samples) were included. Nine out of 18 BTC were GBC and nine were CCA (6 extrahepatic, 3 intrahepatic). Five were overlapping with those analysed by Illumina EPIC arrays. FFPE samples were collected at Meldola centre mentioned above (Italy) (N = 28) and at the Department of Gastroenterology and Hepatology, Navarra University Hospital Complex, Pamplona (Spain) (N = 4).
Samples were tested for CA 19-9 in the respective clinical centres.
Clinical data are presented in Supplementary Table S2.
Publicly available datasets
Methylation data from The Cancer Genome Atlas (TCGA) and GEO portal dataset were retrieved for the validation of the identified methylation alterations. Additional details are provided in Supplementary Document S1.
Bile samples
Twenty-nine bile samples, comprising 24 samples from a new series of CCA patients (21 extrahepatic and 3 intrahepatic; four of which were paired with tissue samples), and five from patients with benign stenosis, were included. Bile samples were collected during ERCP at Pamplona (Spain) centre mentioned above as previously described [30]. After collection bile samples were maintained at 4 °C, centrifuged for 10 min (4 °C) at 3500 × g and stored in aliquots at −80 °C in a biobank facility. The whole process was performed in less than 2 h.
Samples were tested for CA 19-9.
Clinical data are reported in Supplementary Table S3.
Experimental assays
DNA extraction and quantification
DNA was extracted from FFPE tissues using QIAamp DNA FFPE Tissue kit (Qiagen) or the QIAamp DNA kit (Qiagen). DNA was extracted by microdissection of five FFPE tissue slides of 10 µm and 20 µm. DNA concentration was quantified by UV spectrophotometry (NanoDrop Products, Thermo Scientific) and by fluorometric reading (Quant-iT™ PicoGreen® dsDNA Assay Kit).
cfDNA was extracted from 1 ml of bile. Prior to cfDNA isolation, bile was thawed at 4 °C and centrifugated at 14,000 × g for 10 min at 4 °C to ensure the removal of impurities in the supernatant. Bile cfDNA was extracted using the Maxwell RSC ccfDNA Plasma Kit (Promega) according to the manufacturer’s instructions. Bile cfDNA concentrations were determined using a QuantiFluor dsDNA System (Promega), and cfDNA size distributions were analysed by Agilent 2100 Bioanalyzer (Agilent Technologies).
Genome-wide methylation assay
The quality of DNA extracted from FFPE samples was evaluated prior to bisulfite conversion using Infinium HD FFPE QC Assay (Illumina). DNA samples that passed this quality control step were treated with bisulfite using EZ DNA Methylation Gold Kit (Zymo Research). Bisulfite-converted DNA samples were subjected to a DNA restoration process using Infinium FFPE DNA Restore Kit (Illumina).
DNA samples were analysed by Illumina Infinium Human Methylation EPIC BeadChips (EPIC) interrogating over 850,000 CpG sites, according to the Illumina Infinium® HD Methylation protocol. Illumina iScan was used to scan and record high-resolution images of the emitted fluorescence.
Droplet digital PCR assay
Bisulfite treatment
DNA samples were treated with bisulfite using EZ DNA Methylation Gold Kit (Zymo Research).
Droplet digital PCR DNA methylation assays
Primer and probes were designed for the two assays targeting CpG islands (CGIs) at chr2:176993479-176995557 and chr5:145713641-145713913.
The assays were designed on the genomic regions selected based on the methylation information of the CpG sites interrogated by the methylation array probes in the CGIs of interest. In particular, only regions including CpG sites displaying low methylation values in normal samples and high methylation values in tumour samples from the Discovery, TCGA-CHOL and GSE89803 datasets, were selected. DNA methylation status was analysed by ddPCR using the QX200TM Droplet DigitalTM PCR System (BioRad) as previously described [31]. The ddPCR reaction comprised primers (900 nM each), probes (250 nM each), 30 ng bisulfite-converted DNA template from tissues and 70 ng bisulfite-converted DNA template from bile and 1× ddPCR Supermix for Probes (BioRad) in a final volume of 22 µL. The 4Plex control was included in all wells (for sequences for the 4Plex control see ref. [31]). Droplets were generated in the QX200 Droplet Generator (BioRad), with 70 µL of droplet generation oil (BioRad) and 20 µL ddPCR mixture and the PCR was carried out in a T100 Thermal Cycler (BioRad) using the cycling programme recommended by the manufacturer. Finally, the QX200 Droplet Reader (BioRad) was used to read the fluorescence signals.
Data analyses
The detailed analysis of genome-wide and ddPCR methylation data is described in Supplementary Document S1.
For the selection of the most informative biomarkers, an in-house algorithm named TASTOPAL (The Accurate System TO Predict A Lump) was developed. Additional details and the selection pipeline are available in Supplementary Document S1. Supplementary Fig. S4 shows an example of methods applied by the algorithm to select the best biomarkers.
Statistical analyses and power calculation
Chi-square and Fisher’s exact tests (two-sided) were used to determine statistically significant associations between DNA methylation patterns and vs clinical parameters. Statistical analyses of methylation data and power calculation are described in the relative data analysis paragraphs in Supplementary Document S1.
Results
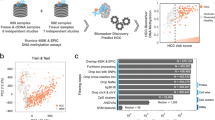
The experimental strategy to identify highly specific and sensitive BTC methylation-based biomarkers consisted of: (1) global DNA methylation analysis of the Discovery cohort samples; (2) validation of the results in TCGA-CHOL cohort and GSE89803 cohort samples; (3) selection of methylation alterations not shared with other gastrointestinal tumours, particularly those with high incidence, i.e. colon cancer (TCGA-COAD), rectal cancer (TCGA-READ) and gastric cancer (TCGA-STAD); (4) application of a machine-learning approach to select the minimum number of the most specific and sensitive alterations; (5) identification of the best-performing biomarker, tested in bile samples.
Figure 1 summarises the analysis workflow.
Identification of DNA methylation alterations in the Discovery dataset
A genome-wide methylation analysis of 50 tumours and ten matched-normal tissue samples from 50 BTC patients was performed using Illumina EPIC arrays. According to sample size calculation (see Supplementary Document S1), the analysed number of samples would guarantee to identify biologically relevant differences in methylation (|Δβ | = 0.2) with a statistical power of 100%. After filtering samples based on the β-values distribution (Supplementary Fig. S1), 26 good-quality samples, comprising 17 tumours and nine matched-normal samples, were selected for the downstream analyses.
The differential methylation analysis between tumour and matched-normal samples identified 648 differentially methylated (|Δβ| > 0.20, combined P value < 0.05) CpG islands (CGIs), comprising 631 hypermethylated (Δβ > 0.20) and 17 hypomethylated (Δβ < −0.20) CGIs (Fig. 2a, b). No statistically significant differentially methylated CGI was detected after P value correction for multi-hypothesis testing (using FDR). This was not entirely unexpected given the reduced sample size in the analysis [8, 16, 32, 33].
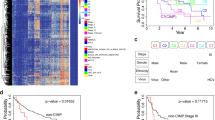
a Scatter plot displaying average CGI β-values distribution in normal and tumour samples of the Discovery dataset. Red dots indicate statistically significant differentially methylated (|Δβ| > 0.20, P value < 0.05) CGIs. b Volcano plot of CGIs Δβ of the Discovery dataset. In the x axis, the difference of methylation between tumours and normal samples. In the y axis, the -logarithm of the P value. Red dots indicate hypermethylated CGIs (Δβ > 0.2) and green dots represent hypomethylated CGIs (Δβ < −0.2). c, d Discovery dataset unsupervised hierarchical clustering analysis based on the average CGI β values (c) and on the average CGI somatic changes, defined as the difference between the β-value of every tumour and the average of the normal samples for each of the aberrantly methylated CGIs (d). e Scatter plot displaying average CGI β-values distribution in normal and tumour samples of the TCGA-CHOL dataset. Red dots indicate statistically significant differentially methylated CGIs. f Volcano plot distribution of CGIs Δβ of the TCGA-CHOL dataset. Red dots indicate hypermethylated CGIs (Δβ > 0.2) and green dots represent hypomethylated CGIs (Δβ < −0.2). g, h TCGA-CHOL dataset unsupervised hierarchical clustering analysis based on the average CGI β values (g) and somatic changes (h) for each of the aberrantly methylated CGIs.
Two heatmaps were generated using CGI methylation values (Fig. 2c, d). Unsupervised hierarchical clustering analysis (UHC) yielded two main clusters: a small cluster comprising the most hypermethylated tumour samples (cluster a) and a second cluster further divided into a cluster of tumours showing intermediate methylation values (cluster b) and a cluster including the normal samples along with three tumour samples showing low methylation values (cluster c) (Fig. 2c, d). No statistically significant association was observed between clusters and tumour location. Stage I and II tumours predominantly clustered with normal controls (cluster c) compared with tumours of higher Stages (III and IV) that were in the cluster including only tumours (clusters a and b) (Fisher’s test P value = 0.015). Grade 2 tumours were predominantly in cluster a, while grade 3 tumours were in cluster b (Fisher’s test P value = 0.029).
We focused on CGIs that become hypermethylated in tumours, under the rationale that they would be more easily detected in liquid biopsies than CGIs that become hypomethylated. Moreover, CGIs showing β-values higher than 0.20 in normal samples (likely reflecting methylation heterogeneity among non-tumour cells) were filtered out, obtaining a final set of 171 somatically hypermethylated CGIs.
Methylation analysis using TCGA-CHOL dataset
To increase the robustness of the identified methylation alterations, methylation data from TCGA-CHOL dataset (including 36 tumour samples and nine normal tissue samples) were analysed. Differential methylation analysis revealed 2203 differentially methylated CGIs (|Δβ| > 0.20, adjusted combined P value < 0.05) (Fig. 2e, f). This dataset yielded a higher number of both hypermethylated (2078) and hypomethylated (125) CGIs compared to the Discovery dataset.
UHC of the TCGA-CHOL samples was similar to that observed in the Discovery dataset, with only one tumour sample clustering in the normal sample subgroup (Fig. 2g, h). No significant associations with tumour stage or grade were observed.
Again, we focused on hypermethylated CGIs in tumours. CGIs showing β-values higher than 0.20 in normal samples were filtered out resulting in 998 hypermethylated CGIs. Notably, 125 out of 171 hypermethylated CGIs (P value = 1.5 × 10−9) identified in our Discovery dataset were validated in the TCGA-CHOL dataset (Fig. 1).
Selection of BTC-specific methylation alterations
To select only BTC-specific alterations, we analysed the methylation changes of the putative biomarkers in other gastrointestinal cancer types using data from TCGA. Of the 125 previously validated CGIs, we excluded those that also exhibited differential methylation (|Δβ| > 0.20) in any of the colon (COAD), rectal (READ) or gastric (STAD) cancer datasets of the TCGA, obtaining a set of 30 BTC-specific somatically hypermethylated CGIs (Supplementary Table S4). Heatmaps generated with methylation values of these CGIs in the Discovery (Fig. 3a) and TCGA-CHOL (Fig. 3b) datasets showed very similar clustering (Fig. 2c, g).
a, b Discovery dataset (a) and TCGA-CHOL dataset (b) unsupervised hierarchical clustering analysis based on the average CGI β-values for the 30 BTC-specific altered CGIs. c ROC curves for the 11 CGIs showing an AUC ≥ 0.90 in the Discovery dataset. Red arrows indicate CGIs showing an AUC ≥ 0.90 also in TCGA-CHOL dataset.
Specificity and sensitivity of the 30 BTC-specific methylation-based biomarkers were evaluated by ROC analysis. In the Discovery dataset, 11 CGIs showed an area under curve (AUC) equal or higher than 0.90 (Fig. 3c), while 21 CGIs had an AUC ≥ 0.90 in TCGA-CHOL dataset. Seven CGIs had an AUC equal or higher than 0.90 in both datasets (Supplementary Table S5).
BTC-specific altered CGIs in the excluded samples
To explore the behaviour of the selected methylation alterations in the 33 tumour samples that were initially excluded due to their abnormal β-value distributions (Supplementary Fig. S1), UHC was carried out using CGI methylation values of the 30 BTC-specific altered CGIs. The rationale of this analysis was that despite the abnormal genome-wide β-value distribution of these samples, the β-values exclusive of the selected CGIs could still provide useful information. The results revealed that the clustering of these samples (Fig. 4b) was similar to that of the good-quality selected tumour samples (Fig. 4a), reinforcing the potential value of these alterations for BTC detection.
a, b Discovery dataset good-quality selected samples (a) and excluded samples (b) unsupervised hierarchical clustering analysis based on the average CGI β-values for the 30 BTC-specific altered CGIs. c GSE89803 dataset unsupervised hierarchical clustering analysis based on the average CGI β-values for the 27 validated CGIs.
Validation in a large BTC dataset
In order to validate alterations of the 30 selected CGIs, methylation data of a large dataset (GSE89803) including 138 tumours and four normal tissue controls from different ethnic groups were analysed. Methylation alterations (|Δβ| > 0.20) were confirmed for 27 out of 30 CGIs. Of note, two of the three CGIs that were not validated displayed high methylation values in both tumour and normal samples (Supplementary Table S6).
UHC of GSE89803 samples yielded similar results to those observed for the Discovery and TCGA-CHOL datasets, with only five tumour samples clustering along with the normal ones (Fig. 4c).
Selection of best-performing biomarkers
The application of a Machine-Learning approach, using the Discovery dataset as training set to prioritise methylation biomarkers and build a diagnostic model on them, and TCGA-CHOL and GSE89803 as validation datasets, resulted in an extremely compact model which generated a ranking list of the best combinations of biomarkers (based on AUC, sensitivity and specificity). We selected the best combination in terms of very high performance and technical assay feasibility. The two biomarkers combination (CGIs mapping on chr2:176993479-176995557 and chr5:145713641-145713913), achieved a promising AUC = 0.972, sensitivity = 0.944 and specificity=1.00 on the TCGA dataset, and AUC = 0.982, sensitivity = 0.964 and specificity = 1.00, on the GSE89803 dataset. Figure 5 shows methylation values of the CpG sites interrogated by EPIC probes in the selected CGIs across the three different datasets.
a, b Methylation values obtained from the Discovery (EPIC array), TCGA-CHOL (450 K array) and GSE89803 (450 K array) datasets. Mean β-values, resulting from the average of the samples (normal indicated with blue dots and tumours indicated with red dots), of each probe, belonging to CGI chr2:176993479-176995557 (a) and CGI chr5:145713641-145713913 (b). The red arrows indicate the CpG sites included in ddPCR experimental assay design. c, d Box plots of the CGI mean β-values for CGI chr2:176993479-176995557 (c) and CGI chr5:145713641-145713913 (d) in tumour and normal tissues obtained from the Discovery, TCGA-CHOL and GSE89803 datasets.
Validation of best-performing biomarkers
As a further step towards the future implementation of these biomarkers in clinical settings, we explored the application of digital PCR DNA methylation assays on ten (five matched tumour and normal) samples previously profiled by Illumina EPIC arrays and 69 additional tissue and bile samples from new case series.
According to sample size calculation (see Supplementary Document S1), the estimated statistical power to identify biologically relevant differences in methylation (|Δβ | = 0.2) was 100% for the tissue cohort and 99.97% for the bile cohort. Removing the 10 tissue samples previously analysed by EPIC arrays, thus reducing the sample size to 13 BTC and 9 normal tissues, the estimated statistical power remained very high (99.996%).
Exploratory analyses—tissue samples
The two candidate biomarkers were tested in a series of BTC (N = 18) and paired normal (N = 14) tissue samples. Assay chr2:176993479-176995557 showed a sensitivity of 100% (N = 17/17), a specificity of 100% (N = 14/14) and AUC of 1.00 (Fig. 6a), while the sensitivity of assay chr5:145713641-145713913 was 76% (N = 13/17), the specificity was 93% (N = 13/14) and AUC was 0.870 (Fig. 6a). One tumour tissue sample was excluded since it resulted negative for both biomarkers. Of note, this sample was already excluded in the whole-genome methylation analysis because of its abnormal β-value distribution (Supplementary Fig. S2). The combined two-biomarker panel resulted in 100% (N = 17/17) sensitivity and 93% (N = 13/14) specificity.
Exploratory analyses—bile samples
Since assay chr2:176993479-176995557 exhibited the highest sensitivity and specificity in tissue samples, bile samples were tested using only this best biomarker. Bile series comprised 24 samples from BTC patients and five samples from patients with benign biliary stenosis. Despite the reduced number of non-tumoral controls, both the sensitivity and the specificity were 100% and the AUC was 1.00 (Fig. 6b).
Discussion
BTCs are extremely aggressive tumours that metastasise in most of patients. It is therefore urgent to identify stable and easily traceable biomarkers ideally available by non-invasive or minimally invasive monitoring approaches. DNA methylation alterations respond to this need, representing specific tumour signatures and much more stable than RNA and proteins.
Moreover, DNA methylation alterations are very early events in carcinogenesis, thus representing extremely valuable biomarkers not only as sentinels of relapse, minimal residual disease or metastasis but also for primary diagnosis. BTCs are indeed diagnosed at an advanced stage due to the lack of accurate diagnostic methods.
While the identification of tumour location-specific methylation profiles may be important [34,35,36], in this study we identified DNA methylation alterations that could enable detection and tracking of biliary tract tumours regardless of their anatomical location and natural history. This suggests that the methylation profile of these tumours might present a lowest common denominator among different pathological subtypes.
The whole-genome methylation approach allowed us to identify and validate 27 BTC-specific methylation alterations on three large case series (Discovery, TCGA-CHOL and GSE89803 datasets).
Several DNA methylation biomarkers have been proposed to detect BTC. These biomarkers, however, are also frequently hypermethylated in other more prevalent gastrointestinal cancers [17, 18, 28]. To increase the specificity of our putative biomarkers, we selected CGIs not altered in other gastrointestinal tumours (colon, rectal and gastric cancer). The identification of BTC-specific alterations is crucial when investigated in non-invasive matrices, such as blood and stool samples, but also in minimally invasive matrices such as bile, to avoid misclassification with other malignancies.
A proprietary machine-learning algorithm (TASTOPAL) was developed to select the most informative CGIs among the altered ones. This approach proved successful because it rendered a panel of only two markers showing very high sensitivity and specificity in three independent datasets of BTC patients (from different ethnic groups, tumour locations, predisposing risk factors). Importantly, the same algorithm can be applied for the selection of markers of other tumour types and different diseases, with manifold advantages compared to the manual biomarker selection.
Illumina Methylation EPIC arrays have proven to be an excellent tool to discover novel potential biomarkers. We have assessed the translatability of the DNA methylation results obtained by EPIC arrays towards droplet digital PCR (ddPCR), an extremely sensitive, robust, fast and cost-effective technique, ideal for the absolute quantification of low-copies DNA molecules of interest.
The two-biomarker panel showed a very high performance in tissue samples (combined sensitivity of 100% and specificity of 93%). A perfect concordance between the results obtained for the same five paired tissue samples by the genome-wide and the targeted methylation assays, specifically designed for a selected region within the two altered CGIs, was observed (data not shown). Of note, BTC samples were equally represented by bile ducts (N = 9) and gallbladder (N = 9) tumours reinforcing the potential of this biomarker panel to detect tumours from different localisations.
The performance of the two individual ddPCR assays was not identical, possibly due to the assay design. The chr2:176993479-176995557 was more accurate (100% sensitivity and specificity) in tissue samples. For this reason, it was selected as the best biomarker to be tested in bile samples to verify its performance as a minimally invasive detection tool for BTC. Of note, the biomarker has shown excellent (>0.90) AUC values in all the datasets analysed. Interestingly, a positive correlation between the statistical power and the robustness of the biomarker was observed (Supplementary Fig. S3).
The biomarker showed 100% sensitivity and specificity in bile samples, proving to be an excellent candidate biomarker for non-invasive BTC investigation. Moreover, this biomarker yielded negative results in bile samples from patients with benign biliary disease, suggesting that this methylation alteration is specific of a state of malignancy. Interestingly, one sample from a patient with benign stenosis showed three positive droplets for the marker of interest and using a threshold based on the best accuracy (another parameter that can be used for setting the threshold), rather than the highest sum of sensitivity and specificity, it was classified as positive. This would suggest that benign biliary disease patients resulting positive for this alteration may be at higher risk of developing BTC compared with negative ones and thus deserve a closer clinical follow-up. Moreover, the screening of this biomarker in patients with benign biliary diseases would be recommended. In fact, the presence of DNA methylation alterations in these patients may potentially represent early triggers in the carcinogenesis process and/or that they harbour pre-cancerous-like features many years prior to tumour onset. The development of BTC after 10–20 years of a previous benign pathology is not rare [37]. This possible scenario is also in line with the value of DNA methylation biomarkers as predictive of neoplasia, managing to forecast the development of cancer even ten years before onset [16].
According to our results, HOXD8 hypermethylation is the best-performing biomarker to identify the presence of BTC regardless of tumour location. In fact, although previous works have demonstrated that DNA methylation alterations can be detected in bile and biliary brush samples, the proposed panels included a higher number of biomarkers and yet they showed lower sensitivity values. Moreover, the majority of these studies were focused on CCA or even on a particular CCA subtype, limiting the application of those panels for the detection of all BTC subtypes. For instance, Shin et al. proposed a five-biomarker panel, with a sensitivity of 76% and a specificity of 100%, to detect extrahepatic CCA in bile using MethyLight [26]. Another work found that p14ARF and p16INK4a methylation had respectively a sensitivity of 46% and 53% for the detection of CCA and GBC in bile [13]. However, similar or even higher percentages of PSC samples showed methylation of the two genes [13]. A six-gene panel allowed the distinction of malignant biliary strictures from benign samples with 77% sensitivity and 78% specificity [38]. Moreover, two studies analysing biliary brush samples, identified a three-biomarker panel, with specificity and sensitivity respectively of 86% and 80% [27], and a four-biomarker panel with 85% sensitivity and 98% specificity [28], distinguishing BTC patients from patients with other biliary diseases. Importantly, all the above-mentioned studies employed techniques with lower sensitivity compared to ddPCR [39].
Interestingly, the selected altered CGI (chr2:176993479-176995557) is associated with HOXD8 gene, belonging to class I homeobox gene family, well known to be involved in carcinogenesis. This suggests that the selected alteration would fulfil the requirements of an optimal biomarker being the most informative but also very likely functionally relevant for the disease. Several studies have shown that HOX genes are either overexpressed or downregulated in a variety of cancers, acting respectively as proto-oncogenes or tumour suppressors depending on the tissue type. Epigenetic mechanisms, including DNA methylation and histone modifications, have been shown to be responsible for an altered expression of these genes in cancer. Specifically, HOXD8 is epigenetically downregulated in lung cancer [40] and used as a biomarker to detect prostate cancer in urine samples [41]. Interestingly, homeobox genes are among the most hypermethylated genes in BTC [42] and, as reported above, other studies have identified other HOX genes as DNA methylation-based biomarkers for BTC, although with lower performances [29, 43].
We acknowledge some limitations in our study. First of all, the Discovery cohort comprised FFPE samples. This kind of sample represents a precious source for research purposes in those cases where it is extremely difficult to obtain fresh tumour samples. In the case of a rare tumour such as BTC, the availability of FFPE samples is even of greater value since the collection of samples for a study of adequate statistical power requires extraordinary time and efforts. However, formaldehyde induces several types of DNA damage such as crosslinks, DNA fragmentation, abasic sites and deamination of cytosine bases [44]. Importantly, formaldehyde-induced crosslinks, inter-strand DNA crosslinks and protein-DNA crosslinks may affect the efficacy of bisulfite treatment, by hampering DNA denaturation, crucial for strand-specific bisulfite conversion [45]. To overcome this limitation, it is imperative to restore DNA integrity after bisulfite treatment as performed in this study. Although this strategy has been successful to obtain reliable data, we cannot exclude that formalin fixation negatively affected the quality of the results from samples. In fact, we observed abnormal β-value distributions in some samples that were consequently removed from the analysis. Due to this selection process, the size of the Discovery cohort was substantially reduced, preventing the identification of methylation alterations resisting multiple-testing correction. We overcame this limitation by selecting DNA methylation alterations shared between the Discovery dataset and TCGA-CHOL dataset. Therefore, the robustness of our results is confirmed by the validation of the selected alterations in multiple independent datasets. Moreover, we showed that even the tumour samples initially excluded clustered similarly to the good-quality selected samples, when the methylation information only from the selected CGIs was used, further reinforcing the validity of our findings.
Another limitation of this study is the unavailability of tissues and bile samples from healthy individuals, for obvious technical and ethical reasons. Thus, the normal tissue samples derived from a section of the paraffin block devoid of malignant cells. Normal tissue surrounding the tumour could already show cancer characteristic alterations [46]. However, the normal samples of the analysed datasets displayed methylation profiles distinct from tumour samples and very similar among them as found in the distribution of the standard deviation (SD) of CpG methylation probes from the TCGA-CHOL dataset (median SD = 0.02, 90th percentile = 0.09). Of note, statistical power calculation has shown that the discovery cohort including 50 tumours and only ten matched-normal controls would guarantee 100% power to detect methylation alterations (|Δβ | ≥ 0.2) conventionally considered as biologically relevant (Supplementary Document S1).
Finally, we acknowledge that ddPCR results on bile samples would benefit from further validation in a large case series from different tumour locations and controls. Nevertheless, the number of analysed bile samples would guarantee a statistical power of 99.97% for the validation of the identified alterations (Supplementary Document S1). In agreement with the pathway to bring a candidate DNA methylation biomarker from the laboratory into molecular diagnostics [47], we have successfully completed the preclinical phase, including the definition of the biomarker, its external validation (in publicly available datasets), the assay design and external validation in independent cohorts (including both tissue and bile samples). Therefore, the promising results obtained during all these steps indicate that the biomarker is ready to be tested in a clinical trial.
In summary, we present a novel DNA methylation-based BTC biomarker, CGI chr2:176993479-176995557, associated with HOXD8 gene, with excellent diagnostic capabilities, which can be applied both in tissue biopsies and bile samples, outperforming all previously reported biomarkers. The next challenge is to test this biomarker for the detection of BTC from completely non-invasive matrices, such as stools and blood. We also envision to begin a clinical trial to evaluate the impact of this biomarker in surveillance and early diagnostic tests of patients at risk for BTC development and to predict patient prognosis and response to treatments, improving patient stratification and personalised therapeutic strategies.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Code availability
The code to reproduce the results of this paper is available under reasonable request to the corresponding authors.
References
Bergquist A, Von Seth E. Epidemiology of cholangiocarcinoma. Best Pract Res Clin Gastroenterol. 2015;29:221–32.
Kongpetch S, Jusakul A, Ong CK, Lim WK, Rozen SG, Tan P, et al. Pathogenesis of cholangiocarcinoma: from genetics to signalling pathways. Best Pract Res Clin Gastroenterol. 2015;29:233–44.
Rizvi S, Eaton JE, Gores GJ. Primary sclerosing cholangitis as a premalignant biliary tract disease: surveillance and management. Clin Gastroenterol Hepatol. 2015;13:2152–65.
Macias RIR, Banales JM, Sangro B, Muntané J, Avila MA, Lozano E, et al. The search for novel diagnostic and prognostic biomarkers in cholangiocarcinoma. Biochim Biophys Acta - Mol Basis Dis. 2018;1864:1468–77.
Rizvi S, Khan SA, Hallemeier CL, Kelley RK, Gores GJ. Cholangiocarcinoma-evolving concepts and therapeutic strategies. Nat Rev Clin Oncol. 2018;15:95–111.
Charatcharoenwitthaya P, Enders FB, Halling KC, Lindor KD. Utility of serum tumor markers, imaging, and biliary cytology for detecting cholangiocarcinoma in primary sclerosing cholangitis. Hepatology. 2008;48:1106–17.
Kim K, Yoo D, Lee HS, Lee KJ, Park SB, Kim C, et al. Identification of potential biomarkers for diagnosis of pancreatic and biliary tract cancers by sequencing of serum microRNAs. BMC Med Genomics. 2019;12:1–11.
Fadda A, Gentilini D, Moi L, Barault L, Leoni VP, Sulas P, et al. Colorectal cancer early methylation alterations affect the crosstalk between cell and surrounding environment, tracing a biomarker signature specific for this tumor. Int J Cancer. 2018;143:907–20.
Luo Y, Wong CJ, Kaz AM, Dzieciatkowski S, Carter KT, Morris SM, et al. Differences in DNA methylation signatures reveal multiple pathways of progression from adenoma to colorectal cancer. Gastroenterology. 2014;147:418–29. e8.
Øster B, Thorsen K, Lamy P, Wojdacz TK, Hansen LL, Birkenkamp-Demtröder K, et al. Identification and validation of highly frequent CpG island hypermethylation in colorectal adenomas and carcinomas. Int J Cancer. 2011;129:2855–66.
Saavedra KP, Brebi PM, Roa JCS. Epigenetic alterations in preneoplastic and neoplastic lesions of the cervix. Clin Epigenetics. 2012;4:13.
Vega-Benedetti AF, Loi E, Moi L, Blois S, Fadda A, Antonelli M, et al. Clustered protocadherins methylation alterations in cancer. Clin Epigenet. 2019;11:1–20.
Klump B, Hsieh CJ, Dette S, Holzmann K, Kießlich R, Jung M, et al. Promoter methylation of INK4a/ARF as detected in bile-significance for the differential diagnosis in biliary disease. Clin Cancer Res. 2003;9:1773–8.
Ishikawa A, Sasaki M, Sato Y, Ohira S, Chen MF, Huang SF, et al. Frequent p16ink4a inactivation is an early and frequent event of intraductal papillary neoplasm of the liver arising in hepatolithiasis. Hum Pathol. 2004;35:1505–14.
Kim BH, Cho NY, Shin SH, Kwon HJ, Jang JJ, Kang GH. CpG island hypermethylation and repetitive DNA hypomethylation in premalignant lesion of extrahepatic cholangiocarcinoma. Virchows Arch. 2009;455:343–51.
Loi E, Moi L, Fadda A, Satta G, Zucca M, Sanna S, et al. Methylation alteration of SHANK1 as a predictive, diagnostic and prognostic biomarker for chronic lymphocytic leukemia. Oncotarget. 2019;10:4987–5002.
Amornpisutt R, Proungvitaya S, Jearanaikoon P, Limpaiboon T. DNA methylation level of OPCML and SFRP1: a potential diagnostic biomarker of cholangiocarcinoma. Tumor Biol. 2015;36:4973–8.
Branchi V, Schaefer P, Semaan A, Kania A, Lingohr P, Kalff JC, et al. Promoter hypermethylation of SHOX2 and SEPT9 is a potential biomarker for minimally invasive diagnosis in adenocarcinomas of the biliary tract. Clin Epigenet. 2016;8:1–11.
Andresen K, Boberg KM, Vedeld HM, Hektoen M, Wadsworth CA, Clausen OP, et al. Novel target genes and a valid biomarker panel identified for cholangiocarcinoma. Epigenetics. 2012;7:1249–57.
Farshidfar F, Zheng S, Gingras MC, Newton Y, Shih J, Robertson AG, et al. Integrative genomic analysis of cholangiocarcinoma identifies distinct IDH-mutant molecular profiles. Cell Rep. 2017;18:2780–94.
Jusakul A, Cutcutache I, Yong CH, Lim JQ, Huang MN, Padmanabhan N, et al. Whole-genome and epigenomic landscapes of etiologically distinct subtypes of cholangiocarcinoma. Cancer Discov. 2017;7:1116–35.
Church TR, Wandell M, Lofton-Day C, Mongin SJ, Burger M, Payne SR, et al. Prospective evaluation of methylated SEPT9 in plasma for detection of asymptomatic colorectal cancer. Gut. 2014;63:317–25.
Su SF, De Castro Abreu AL, Chihara Y, Tsai Y, Andreu-Vieyra C, Daneshmand S, et al. A panel of three markers hyper- And hypomethylated in urine sediments accurately predicts bladder cancer recurrence. Clin Cancer Res. 2014;20:1978–89.
Vega-Benedetti AF, Loi E, Moi L, Orrù S, Ziranu P, Pretta A, et al. Colorectal cancer early detection in stool samples tracing CPG islands methylation alterations affecting gene expression. Int J Mol Sci. 2020;21:1–16.
Barault L, Amatu A, Siravegna G, Ponzetti A, Moran S, Cassingena A, et al. Discovery of methylated circulating DNA biomarkers for comprehensive non-invasive monitoring of treatment response in metastatic colorectal cancer. Gut. 2018;67:1995–2005.
Shin SH, Lee K, Kim BH, Cho NY, Jang JY, Kim YT, et al. Bile-based detection of extrahepatic cholangiocarcinoma with quantitative DNA methylation markers and its high sensitivity. J Mol Diagnostics. 2012;14:256–63.
Parsi MA, Li A, Li CP, Goggins M. DNA methylation alterations in endoscopic retrograde cholangiopancreatography brush samples of patients with suspected pancreaticobiliary disease. Clin Gastroenterol Hepatol. 2008;6:1270–8.
Andresen K, Boberg KM, Vedeld HM, Honne H, Jebsen P, Hektoen M, et al. Four DNA methylation biomarkers in biliary brush samples accurately identify the presence of cholangiocarcinoma. Hepatology. 2015;61:1651–9.
Wasenang W, Chaiyarit P, Proungvitaya S, Limpaiboon T. Serum cell-free DNA methylation of OPCML and HOXD9 as a biomarker that may aid in differential diagnosis between cholangiocarcinoma and other biliary diseases. Clin Epigenet. 2019;11:39.
Urman JM, Herranz JM, Uriarte I, Rullán M, Oyón D, González B, et al. Pilot multi-omic analysis of human bile from benign and malignant biliary strictures: a machine-learning approach. Cancers. 2020;12:1–30.
Pharo HD, Andresen K, Berg KCG, Lothe RA, Jeanmougin M, Lind GE. A robust internal control for high-precision DNA methylation analyses by droplet digital PCR. Clin Epigenet. 2018;10:24.
Tong T, Zhao H. Practical guidelines for assessing power and false discovery rate for a fixed sample size in microarray experiments. Stat Med. 2008;27:1960–72.
Saffari A, Silver MJ, Zavattari P, Moi L, Columbano A, Meaburn EL, et al. Estimation of a significance threshold for epigenome-wide association studies. Genet Epidemiol. 2018;42:20–33.
Mukund K, Syulyukina N, Ramamoorthy S, Subramaniam S. Right and left-sided colon cancers-specificity of molecular mechanisms in tumorigenesis and progression. BMC Cancer. 2020;20:317.
Baran B, Mert Ozupek N, Yerli Tetik N, Acar E, Bekcioglu O, Baskin Y. Difference between left-sided and right-sided colorectal cancer: a focused review of literature. Gastroenterol Res. 2018;11:264–73.
Antonelli M, Fadda A, Loi E, Moi L, Zavattari C, Sulas P, et al. Integrated DNA methylation analysis identifies topographical and tumoral biomarkers in pilocytic astrocytomas. Oncotarget. 2018;9:13807–21.
Boonstra K, Weersma RK, van Erpecum KJ, Rauws EA, Spanier BWM, Poen AC, et al. Population-based epidemiology, malignancy risk, and outcome of primary sclerosing cholangitis. Hepatology. 2013;58:2045–55.
Zhang Y, Yang B, Du Z, Gao YT, Wang YJ, Jing X, et al. Identification and validation of specific methylation profile in bile for differential diagnosis of malignant biliary stricture. Clin Biochem. 2010;43:1340–4.
De Rubis G, Rajeev Krishnan S, Bebawy M. Liquid biopsies in cancer diagnosis, monitoring, and prognosis. Trends Pharmacol Sci. 2019;40:172–86.
Sato T, Arai E, Kohno T, Takahashi Y, Miyata S, Tsuta K, et al. Epigenetic clustering of lung adenocarcinomas based on DNA methylation profiles in adjacent lung tissue: Its correlation with smoking history and chronic obstructive pulmonary disease. Int J Cancer. 2014;135:319–34.
Zhao F, Olkhov-Mitsel E, van der Kwast T, Sykes J, Zdravic D, Venkateswaran V, et al. Urinary DNA methylation biomarkers for noninvasive prediction of aggressive disease in patients with prostate cancer on active surveillance. J Urol. 2017;197:335–41.
Sriraksa R, Zeller C, Dai W, Siddiq A, Walley AJ, Limpaiboon T, et al. Aberrant DNA methylation at genes associated with a stem cell-like phenotype in cholangiocarcinoma tumors. Cancer Prev Res. 2013;6:1348–55.
Shu Y, Wang B, Wang J, Wang JM, Zou SQ. Identification of methylation profile of HOX genes in extrahepatic cholangiocarcinoma. World J Gastroentero.l 2011;17:3407–19.
Do H, Dobrovic A. Sequence artifacts in DNA from formalin-fixed tissues: causes and strategies for minimization. Clin Chem. 2015;61:64–71.
Wen X, Jeong S, Kim Y, Bae JM, Cho NY, Kim JH, et al. Improved results of LINE-1 methylation analysis in formalin-fixed, paraffin-embedded tissues with the application of a heating step during the DNA extraction process. Clin Epigenetics. 2017;9:1.
Takeshima H, Ushijima T. Accumulation of genetic and epigenetic alterations in normal cells and cancer risk. npj Precis Oncol. 2019;3:1–8.
Mikeska T, Bock C, Do H, Dobrovic A. DNA methylation biomarkers in cancer: progress towards clinical implementation. Expert Rev Mol Diagn. 2012;12:473–87.
Acknowledgements
We wish to thank the pathologists Dr. Maurizio Puccetti and Dr. Francesco Limarzi for selecting and reviewing the hematoxylin and eosin slides of the tissues used for the study.
Funding
This work was supported by grants from Fondazione di Sardegna, Italy (Prot. 738/2012-0246 in 2012 to PZa; F72F16003120002 to PZa and MS in 2016; U278.2021/AI.259.RP to PZa in 2021), Regione Autonoma della Sardegna, Italy (F72I15000920002 to PZa in 2015), POR FESR Sardegna, Italy (F31G17000050002 to PZa in 2018), Instituto de Salud Carlos III, Spain (PI16/01126 to MAA and JMU in 2017; PI19/00163 to MA in 2020; PI18/01484 and PI21/01766 to SA in 2019 and 2021), Fundación Mutua Madrileña, Spain (AP174232020 to SA in 2020), Gobierno de Navarra Salud, Spain (58/2017 to MAA and JMU in 2017; 55/2018 to MA and CB in 2018), FEDER/Ministerio de Ciencia, Innovación y Universidades-Agencia Estatal de Investigación, Spain (Grants PID2019-104878RB-100 and PI02019-104265RB-100 to MAA and CB in 2019), Strategic Euroregion Collaborations, Aquitaine, Euskadi, Navarra, Spain (2020/101 to CB, MA, JMU and MAA in 2020), Asociación Española Contra el Cáncer (AECC) post-doctoral fellowship (to MA).
Author information
Authors and Affiliations
Contributions
ELo: contributed to samples preparation and processing, contributed to the genome-wide methylation experiments, performed the bioinformatics analyses of the genome-wide methylation experiments, interpreted the results of the genome-wide methylation analysis, designed ddPCR methylation assays, interpreted the results of the ddPCR analysis, wrote the manuscript draft, prepared tables and figures, made critical revision of the manuscript for important intellectual content. CZ: contributed to establishing the pipeline for the refined biomarkers selection writing an in-house algorithm, made critical revision of the manuscript for important intellectual content. AT: contributed to establishing the pipeline for the refined biomarkers selection writing an in-house algorithm, made critical revision of the manuscript for important intellectual content. LM: contributed to samples preparation and processing, contributed to the genome-wide methylation experiments. MC: contributed to the collection and retrieval of samples and clinical data, contributed to samples preparation and processing, made critical revision of the manuscript for important intellectual content. AP: performed methylation analyses by ddPCR, supervised the activities of the study, made critical revision of the manuscript for important intellectual content. CS: performed methylation analyses by ddPCR. AFV-B: prepared tables and figures, made critical revision of the manuscript for important intellectual content. PZi: contributed to the collection and retrieval of samples and clinical data. MP: contributed to the collection and retrieval of samples and clinical data. ELa: contributed to the collection and retrieval of samples and clinical data. LF: contributed to the collection and retrieval of samples and clinical data. MR: contributed to the collection and retrieval of samples and clinical data. JC: contributed to the collection and retrieval of samples and clinical data. IA: contributed to the collection and retrieval of samples and clinical data. JMU: contributed to the collection and retrieval of samples and clinical data. MA: contributed to the collection and retrieval of samples and clinical data. CB: contributed to the collection and retrieval of samples and clinical data, contributed to samples preparation and processing, supervised the activities of the study, made critical revision of the manuscript for important intellectual content. EF: performed methylation analyses by ddPCR, supervised the activities of the study, made critical revision of the manuscript for important intellectual content. AC-G: contributed to the collection and retrieval of samples and clinical data, supervised the activities of the study, made critical revision of the manuscript for important intellectual content. MAA: contributed to the collection and retrieval of samples and clinical data, supervised the activities of the study, made critical revision of the manuscript for important intellectual content. SA: performed the bioinformatics analyses of the genome-wide methylation experiments, interpreted the results of the genome-wide methylation analysis, supervised the activities of the study, made critical revision of the manuscript for important intellectual content. MS: conceived and designed the study, contributed to the collection and retrieval of samples and clinical data, supervised the activities of the study, made critical revision of the manuscript for important intellectual content. PZa: conceived and designed the study, contributed to samples preparation and processing, contributed to the genome-wide methylation experiments, interpreted the results of the genome-wide methylation analysis, designed ddPCR methylation assays, interpreted the results of the ddPCR analysis, supervised the activities of the study, wrote the manuscript draft, made critical revision of the manuscript for important intellectual content. All authors critically reviewed and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests. PZa, MS, ELo, CZ, AT, SA, MAA and AC-G have filed a patent (application number: 102021000021455, 06/08/2021), of which they have transferred ownership to the University of Cagliari.
Ethics approval and consent to participate
All the analysed biological samples were obtained with written informed consent from participants prior to inclusion in the study. The study protocol conforms to the ethical guidelines of the 1975 Declaration of Helsinki as reflected in a priori approval by the relative institution’s human research committees: the Independent Ethics Committee of the University Hospital of Cagliari (protocol # PG/2021/1527), the Ethics Committee of Romagna (CEROM) (protocol # IRST-B050) and the Ethics Committees of the Navarra University Hospital Complex, Pamplona, Spain (protocol # 2016/91).
Consent to publish
Not applicable.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Loi, E., Zavattari, C., Tommasi, A. et al. HOXD8 hypermethylation as a fully sensitive and specific biomarker for biliary tract cancer detectable in tissue and bile samples. Br J Cancer 126, 1783–1794 (2022). https://doi.org/10.1038/s41416-022-01738-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-022-01738-1
- Springer Nature Limited