Abstract
The C-terminal binding proteins (CtBPs), CtBP1 and CtBP2, are transcriptional co-repressor that interacts with multiple transcriptional factors to modulate the stability of chromatin. CtBP proteins were identified with overexpression in the high-grade serous ovarian carcinoma (HGSOC). However, little is known about CtBP proteins’ regulatory roles in genomic stability and DNA repair in HGSOC. In this study, we combined whole-transcriptome analysis with multiple research methods to investigate the role of CtBP1/2 in genomic stability. Several key functional pathways were significantly enriched through whole transcription profile analysis of CtBP1/2 knockdown SKOV3 cells, including DNA damage repair, apoptosis, and cell cycle. CtBP1/2 knockdown induced cancer cell apoptosis, increased genetic instability, and enhanced the sensitivity to DNA damage agents, such as γ-irradiation and chemotherapy drug (Carboplatin and etoposide). The results of DNA fiber assay revealed that CtBP1/2 contribute differentially to the integrity of DNA replication track and stability of DNA replication recovery. CtBP1 protects the integrity of stalled forks under metabolic stress condition during prolonged periods of replication, whereas CtBP2 acts a dominant role in stability of DNA replication recovery. Furthermore, CtBP1/2 knockdown shifted the DSBs repair pathway from homologous recombination (HR) to non-homologous end joining (NHEJ) and activated DNA-PK in SKOV3 cells. Interesting, blast through TCGA tumor cases, patients with CtBP2 genetic alternation had a significantly longer overall survival time than unaltered patients. Together, these results revealed that CtBP1/2 play a different regulatory role in genomic stability and DSBs repair pathway bias in serous ovarian cancer cells. It is possible to generate novel potential targeted therapy strategy and translational application for serous ovarian carcinoma patients with a predictable better clinical outcome.
Similar content being viewed by others
Introduction
Serous ovarian cancer is a gynecological tumor that is more common in women, and high-grade serous ovarian cancer (HGSOC) accounts for roughly 70% of ovarian cancer deaths [1, 2]. As an aggressive tumor type of ovarian cancer, HGSOC had a higher relapse rate (~25%) and poorer overall 5-year survival rate (31%) [3, 4]. The genetic alterations of HGSOC were well characterized, including copying number gains and losses, mutations and deletions [5], and were associated with a homologous recombination (HR) defect and increased sensitivity to DNA damage agents [5, 6]. Intracellular signaling is activated by genetic alterations to detect mispatch and regulate cell cycle progression and promote the repair of DNA lesions by DNA damage response (DDR). Double-strand breaks (DSBs) are among the most lethal types of DNA lesions in mammalian cells, and they are primarily repaired by the non-homologous end joining (NHEJ) or HR pathways. Pathways for DNA repair disruption, whether through chemotherapy drugs or other clinical cancer treatment methods, had been identified as an effective therapeutic strategy for HGSOC [7, 8].
The C-terminal binding protein (CtBP) family proteins consisted of two isoforms, CtBP1 and CtBP2, that shared a highly conserved protein structure (78% sequence homolog) and performed some similar functions in human cells, such as carbohydrate metabolism and epigenetic regulatory multiple transcription factors [9]. CtBP proteins had a distinct intracellular distribution that was determined by a C-terminal PDZ binding motif (CtBP1) and a unique N-terminal nuclear localization domain (CtBP2). CtBP1 was found in both the cytoplasm and the nucleus, whereas CtBP2 was exclusively resident in the nucleus [10]. CtBP1 interacted with human adenovirus E1A via a PLDLS motif and would be functional as a tumor suppressor [11, 12]. CtBP proteins had been found to be overexpressed in a variety of solid tumors, including breast, ovarian, prostate, colon, and gastric cancer, and had been linked to poor patient survival in clinical outcome [13, 14]. CtBP2 had been identified as a novel oncogene in serous ovarian cancer [15, 16], and overexpression of CtBP2 had been linked to abnormal proliferation and a lower survival rate [13]. CtBP proteins suppressed the expression of death receptors D4/5 and determine the fate of serous ovarian cancer cells [17]. CtBP proteins inhibited apoptosis by decreasing the expression of pro-apoptotic genes [18,19,20]. Up to now, little is known about the contribution of CtBP proteins in genomic stability in serous ovarian cancer.
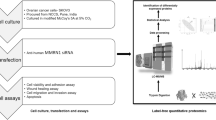
In this study, we combined whole-transcriptome analysis with multiple DNA damage repair research methods to investigate the role of CtBP1/2 in genomic stability in serous ovarian cancer cells. Several enriched key functional pathways, including DDR and apoptosis, had been thoroughly investigated in CtBP1/2 knockdown serous ovarian cancer cells utilizing transcription profile enrichment analysis. CtBP proteins had been studied for their role in genomic stability and DNA repair.
Results
Established CtBP1/2 stable knockdown in serous ovarian cancer cells
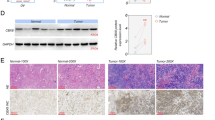
CtBP1/2 was found to be abnormally overexpressed in several ovarian cancer cell lines, including MCAS, SKOV3, RMG1, and RMUGL (Fig. 1A), and a very weak signal was found in human normal ovarian epithelium (HOSE). It is thought that CtBP1/2 overexpression contributes to the abnormality of ovarian cancer cells. The lentivirus shRNA constructs successfully generated scramble control and CtBP1/2 knockdown (CtBP1 KD, CtBP2 KD) SKOV3 cells. To avoid off-target effects, each target gene was initially screened with five different shRNA constructs, and then selected two independent clones for the next step functional assay, which were validated by Western blot with clearly CtBP knockdown (Fig. 1B). Similar to our previous findings [21], specific single knockdown of the CtBP1 or CtBP2 gene did not significantly induce isoform gene compensatory expression. Transiently dual CtBP1/2 knockdown had been achieved by a 48-h swap of CtBP1/2 siRNA interference in CtBP1/2 stable knockdown cancer cells, and validated by western blot analysis (Fig. 1B).
The overexpression of CtBP1/2 was analyzed by western blot in ovarian cancer cell lines (A). CtBP1/2 stable knockdown in SKOV3 cells generated by lentivirus containing CtBP1/2-targeting shRNA constructs and siRNA transient double knockdown (KD) (B). The Venn diagram showed the overlapped significant changed genes among CtBP1KD, CtBP2 KD, and CtBP1/2 double KD (C). The enriched functional pathways were summarized in CtBP1KD vs. control (D), CtBP2 KD vs. control (E), and CtBP1/2 double KD vs. control (F).
Transcriptome analysis discovered key functions regulated by CtBP1/2 proteins
Normalized and filtered through comparison, 355 genes (upregulated 182 genes; downregulated 173 genes; CtBP1 KD vs. control, Fig. S1A), 805 genes (upregulated 784 genes; downregulated 21 genes; CtBP2 KD vs. control, Fig. S1B) and 4297 genes (upregulated 1364 genes; downregulated 2933 genes; CtBP1/2 DKD vs. control, Fig. S1C) had been identified as significantly changed genes and computationally clustered, respectively. A Venn diagram demonstrated the overlapped significantly changed genes among three knockdown groups (Fig. 1C). Different groups shared a variety of common genes, including 56 genes (CtBP1 KD vs. CtBP2-KD) (Table S1), 91 genes (CtBP1 KD vs CtBP1/2 DKD), 108 genes (CtBP2 KD vs CtBP1/2 DKD), and 11 genes were overlapped for three groups. Key functional pathways were enriched and characterized in CtBP1 KD, CtBP2 KD, and CtBP1/2 DKD cells using GSEA. Three key pathways were negatively enriched in CtBP1-KD cells, including epithelial to mesenchymal transition in colorectal cancer, MAPK signaling pathway, and cell cycle, while focal adhesion-PI3K-Akt-mTOR-signaling and integrin-mediated cell adhesion were positively enriched (Fig. 1D). Positively enriched pathways in CtBP2-KD cells included G1 to S cell cycle control, MAPK signaling pathway, cell cycle, DDR, and focal adhesion-PI3K-Akt-mTOR-signaling pathway; negatively enriched pathways included TGF-beta signaling pathway, ErbB signaling pathway, apoptosis, and epithelial to mesenchymal transition in colorectal cancer (Fig. 1E). Positively enriched pathways in CtBP1/2-DKD cells included rapamycin (TOR) signaling and apoptosis target; negatively enriched pathways included focal adhesion-PI3K-Akt-mTOR-signaling pathway, tumor suppression activity of SMARCB1, Wnt signaling pathway, photodynamic therapy-induced HIF-1 survival signaling, and glycolysis and gluconeogenesis, and metabolic reprogramming in the colon cancer (Fig. 1E). Cell cycle and focal adhesion-PI3K-Akt-mTOR signaling pathways were common enriched pathways between CtBP1/2-KD cells. Overexpression of CtBP1/2 was thought to play regulatory roles in the cell cycle and adhesion of serous ovarian cancer cells. The apoptosis pathway, in particular, had been highlighted in the enriched functional pathways of CtBP1/2-DKD and CtBP2-KD cells.
CtBP1/2 knockdown impaired cellular functions of serous ovarian cancer cell
Because the epithelial to mesenchymal transition and adhesion-related pathways were enriched in the CtBP1-KD/CtBP2-KD cells, the ability of cell proliferation, adhesion and migration were evaluated. CtBP protein knockdown significantly reduced cell proliferation (Fig. S2A), clone formation in vitro agar gel (Fig. S2B), and adhesion to extracellular matrix proteins collagen I and collagen IV (Fig. S2C), as measured by the scratch wound and transwell assays (Fig. S2D&E). CtBP2-KD had a greater impact on cellular functions, according to these findings. CtBP2 inhibition is proposed as an appropriate therapeutic target for serous ovarian cancer therapy and as an effective treatment strategy for serous ovarian cancer.
CtBP1/2 knockdown triggered apoptosis in serous ovarian cancer cells
Because the cell cycle signaling pathway was significantly positively enriched in CtBP1-KD/CtBP2-KD cells (Fig. 1D, E), cell cycle-related genes were gathered for further analysis (Fig. 2A). The heatmap depicts the 14 genes involved in cell cycle regulation that were significantly upregulated in CtBP1-KD and CtBP2-KD cells (Fig. 2B). Knockdown CtBP1 or CtBP2 disrupted the cell cycle phases, significantly increased the portion of sub-G1 phase (CtBP1-KD, 44.3%; CtBP2-KD, 44.0%), and decreased the portion of G1 phase and G2/M phase (Fig. 2C, D). It is proposed that CtBP1 or CtBP2 knockdown induce apoptosis in serous ovarian cancer cells, and that CtBP1/2-DKD accelerate the process of apoptosis while failing to generate stable double knockdown cells. In CtBP1-KD/ CtBP2-KD cells, the phosphorylation level of Cdc25A-Y15 increased while the expression level of Cdk2 decreased (Fig. 2E), both of which were indicators of cell cycle progression and enhanced apoptosis [22].
The enrichment plot of cell cycle-related genes was identified by GSEA in CtBP1 KD (left) and CtBP2 KD (right), compared to control cells (A). Heat maps significantly comparing the expression of genes that involved in cell cycle regulation in CtBP1/2 KD and control groups (B). The comparison between CtBP1/2 KD and control groups in cell cycle by flow cytometry (C, D). The cell cycle profile also was created by measuring total DNA content with PI. Data were representing each phase of cell cycle and quantitative analysis the percentage generated by three independent experiments. Western blot analyzed the changed pattern of cell cycle key regulators between CtBP1/2 knockdown and control (E). All the data represented the average of triplicates values generated by three independent experiments. *P < 0.05; **P < 0.001; ***P < 0.0001.
CtBP1/2 knockdown exacerbated the DNA damage response to irradiation and chemotherapy drugs in serous ovarian cancer cells
DDR-signaling pathway was significantly positive enriched (NES:1.2074, FDR q:0.034) pathways in the CtBP2-KD cell (Fig. 3A). 11 key genes involved in DDR were significantly changed, with ten genes upregulated (BAX, CCNE1, CDKN1B, GAPP45A, NFKB1, NFKB2, TCF7, TCF7L1, and TGFB1) and one gene downregulated (CCNG2). CtBP1-KD cells showed a similar change pattern without significant enrichment (Fig. 3B). Key regulatory genes, NFKB1, NFKB2, TCF7, and TCF7L1, were significantly upregulated, which are involved the DDR and oxidative stress through WNT/beta-catenin signaling pathway [23, 24]. There was no significant difference in the ratios of tail DNA between CtBP1/2-KD and control cells. The ratio of tail DNA was significantly increased in CtBP1-KD and CtBP2-KD cells after γ-irradiation (6 and 12 Gy) treatment (Fig. 3C). CtBP2-KD cells were more sensitive to γ-irradiation and generated the highest ratio of tail DNA (6 Gy, 35.8%, P < 0.001; 12 Gy, 45.8%, P < 0.001).
The upregulated genes in CtBP2 KD vs. control were enriched in DNA damage response as assessed by GSEA (A). Heat maps significantly compared expression of genes that involved in DNA damage response regulation in CtBP1/2 KD and control groups (B). Comet assay analyzed the DNA damage response the after g-irradiation treatment (6 and 12 Gy) on ovarian cancer cells. The ratio of tail DNA was as quantitative analysis (C). Immunofluorescence of γH2AX and RPA32 phosphorylation detected in control and CtBP1/2 KD cells after cell treated with carboplatin and etoposide (D). Nuclear DNA was counterstained with DAPI. Quantitative analysis foci formation per cell in each group was performed by Image J and presented as γH2AX foci/cell (E) and RPA32 foci/cell (F). Rest represented without stimulation. For each sample, 100 cells were blinded and scored independently by two markers. *P < 0.05; **P < 0.01; ***P < 0.001.
The γH2AX and RPA32 phosphorylation foci signals were employed as readout indicators to evaluate the impact of CtBP1/2 knockdown on the DNA damage and genomic stability in cancer cells, which were typical markers for DNA damage [25], genomic stability, and cell survival [26]. Under rest conditions (without any treatment), the γH2AX phosphorylation foci number per cell was significantly increased in CtBP1-KD and CtBP2-KD cells (Fig. 3D, E). The RPA32 phosphorylation foci signal was almost negative in these cells, with no statistically significant difference (Fig. 3F). The intensity of γH2AX and RPA32 phosphorylation foci signal became brighter after treated with carboplatin or etoposide (Fig. 3D) and the number of foci per cell increased significantly, especially in CtBP2-KD cells (Fig. 3E, F). CtBP1/2 knockdown was thought to increase genetic instability and activate DNA-dependent protein kinase (DNA-PK) in the DDR induced by carboplatin or etoposide treatment.
CtBP1/2 knockdown shortened the DNA replication fork and increased instability
The DNA fiber assay was used to evaluate the impact of CtBPs knockdown on genomic stability. The retention of IdU label was measured (Σ IdU), with or without HU treatment, as described in the “Methods” section, to reflect the stability of stalled forks (Fig. 4A). Without HU treatment, the median IdU tract length of CtBP1-KD/CtBP2-KD cells had significantly shorter (1.7793 kb, CtBP1-KD; 5.3212 kb, CtBP2-KD, respectively, P < 0.0001, two-tailed Mann–Whitney U test) than control cells (7.0396 kb) (Fig. 4B, C, Table 1A). HU treatment resulted in a significantly shorter median IdU tract length (1.5592 kb, CtBP1-KD) than the control (4.5338 kb, P < 0.0001, two-tailed Mann–Whitney U test), despite the fact that (Fig. 4B, D, Table 1B), although HU treatment resulted in a significantly shorter median IdU tract length in the CtBP1-KD/CtBP2-KD and control cells (Table S2). It is proposed that CtBP1/2 contribute to maintain the integrity of DNA replication track, while CtBP1 play an important role in the protection of stalled forks integrity under metabolic stress condition during prolonged periods of replication.
Schematic of single DNA fiber analysis experimental design. Green tracts, IdU; red tracts, CldU. Examples of various types of tracts are shown (A). The retention of IdU label was measured, with or without HU treatment, and reflected the stability of stalled forks (Σ IdU). The lengths of CldU tract formed after breakage have been measured (ΣCIdU), with or without HU exposure, to evaluate the impact of CtBP1/2 knockdown on the DNA replication recover ability in serous ovarian cancer cells. Demonstration of representative images of ldU/CldU tract, with or without HU treatment, in the control, CtBP1 KD, and CtBP1 KD groups (B). IdU tract length distributions from DNA fibers from control and CtBP1/2 KD cells in the presence (replication stalling) (C) or absence (unperturbed replication) of HU (D). CIdU tract length distributions from DNA fibers from control and CtBP1/2 KD cells in the presence (replication stalling) (E) or absence (unperturbed replication) of HU (F). Distribution curves of the ratio of CldU/IdU tract lengths with (G) or without HU in cells (H). Median tract lengths and cumulative distributions were given in parentheses here and in subsequent figures. Error bars represented the standard error of the mean (SEM).
Simultaneously, the lengths of CldU tract formed after breakage were measured, with or without HU exposure, to evaluate the impact of CtBP1/2 knockdown on the DNA replication recover ability in serous ovarian cancer cells. Without HU exposure, CtBP1/2 knockdown significantly decreased the median CIdU tract length (3.1818 kb, control) (CtBP1-KD, 1.2872 kb, P < 0.0001; CtBP2-KD, 1.7793 kb, P < 0.0001, two-tailed Mann–Whitney U test) (Fig. 4E, Table 1C). CtBP1/2 knockdown was strongly suggested to increase the instability of DNA replication recovery and shorten the CldU tract lengths. Similar to the IdU response pattern, the replication fork recovery rate was significantly slowered with HU treatment, and the length of CldU track was significantly decreased (Table S3). The HU-induced stress of deoxyribonucleoside triphosphate pool did not alter the significantly different pattern of CldU tracks between the CtBP1-KD/CtBP2-KD and control cells (Fig. 4F, Table 1D).
Without HU treatment, the CldU tracts length were longer that the IdU track’s in control cells (median CldU/IdU = 1.725), whereas the CldU/IdU ratio was lower than 1.0 in CtBP1-KD cells (median CldU/IdU = 0.6774, P < 0.0001) and greater than 1.0 in CtBP2-KD cells (median CldU/IdU = 1.161, P < 0.0001, Fig. 4G). Although HU treatment reduced the CldU/IdU ratio of CtBP1/2 KD cells to less than 1.0 (CtBP1 KD, median = 0.7052, P < 0.0001; CtBP2 KD, median = 0.8738, P < 0.05; control, median = 1.174), it had no effect on the difference pattern between the CtBP1/2 KD and control cells (Fig. 4H). It was implied that CtBP1/2 protect against the degradation of stalled replication forks with opposite directions for the leading and lagging strands and CtBP2 appears to play a dominant protective role in serous ovarian cancer cells.
CtBP1/2 knockdown synergistically activated DNA-PK in DNA damage response
DNA-PK activity was necessary for the NHEJ pathway [27]. Two DNA-PK selective inhibitors, KU-0060648 and NU7441, were employed as probe to investigate the regulatory role of DNA-PK during DDR in the CtBP1/2 KD ovarian cancer cells. KU-0060648 was a dual inhibitor of DNA-PK and PI3K that could increase the sensitivity of cancer cells to DNA damage induced by cytotoxic drug [27], such as etoposide. NU7441 was an ATP-selective competitive inhibitor of DNA-PK and had no inhibitory effect on the DNA-PK-related enzymes ATM and ATR even at 100 μM concentration [28]. To evaluate the contribution of DNA-PK in CtBP1/2 KD induced DDR, cytotoxic assay and fluorescence foci assay were employed to quickly explore the optimal dosage of specific or non-specific DNA-PK inhibitors for CtBP1/2 KD cells.
NU7441 treatment significantly decreased survival and differentiated the cytotoxic response among CtBP1-KD (0.5 μM, 68.3 ± 5.80%, P < 0.05; 2 μM, 22.1 ± 1.48%, P < 0.001; 5.0 μM, 24.3 ± 5.97%, N.S), CtBP2-KD cells (0.5 μM, 68.4 ± 4.50%, P < 0.05; 2 μM, 35.6 ± 3.30%, P < 0.05; 5.0 μM, 32.1 ± 5.89%, N.S) and control cells (0.5 μM, 82.5 ± 4.08%; 2 μM, 45.8 ± 3.87%; 5.0 μM, 27.9 ± 3.70%), respectively (Fig. 5A). There was no significant difference in cytotoxic response between the CtBPs knockdown and control groups when treated with a higher dosage (5 μM) of NU7441. The dosage (2 μM) of NU7441 was chosen for foci assay. KU-0060648 exhibits differential growth inhibitory effects but not profoundly cytotoxic in multiple human cancer cell lines, and it works as a dual DNA-PK (IC50 = 8.6 nM) and PI3Ks inhibitor (IC50 = 0.59 μM for PI3Kγ) [27]. Although KU-0060648 (2 μM) treatment induced cytotoxicity in CtBP1-KD (35.5 ± 9.00%, P = 0.4934), CtBP2-KD (44.4 ± 4.80%, P = 0.8374) and control cells (43.0 ± 4.40%), there was no significant difference between different groups (Fig. 5B). The dosage (2.0 μM) of KU-0060648 was intended to completely block the DNA-PK and PI3Ks contribution in CtBP1/2-KD cells and is employed to evaluate the DDR via the foci assay.
The cell survival comparison between CtBP1/2 KD and control the response to DNA-PK inhibitor NU7441 (A) and KU treatment (B). Immunofluorescence of gH2AX and RPA32 phosphorylation detected in control and CtBP1/2 KD cells after cell treated with ENA-PK inhibitor NU7441 and KU000487. Nuclear DNA was counterstained with DAPI (C). Quantitative analysis foci formation per cell in each group was performed by Image J and presented as γH2AX foci/cell (D) and RPA32 foci/cell (E). *P < 0.05; **P < 0.001. The contribution of CtBP1/2 in the regulation of balance shifted between homology-dependent vs. homology-independent repair model in ovarian cancer cells (F). The SeeSaw 2.0 reporter was employed to detect the balance shift between homology-dependent vs. homology-independent repair among CtBP1/2 knockdown and control cells in rest condition, treated with carboplatin and etoposide. To measure the deviation from the balance between NHEJ and HR, the ratio between green vs. red cells in each condition was calculated. To facilitate comparing experiments, this ratio was normalized shRNA control. Those skewed the balance towards an increase in homology-independent repair has a fold-increase of over 1, while those with an increase in HR have a fold-decrease of less than one. Data represent a minimum of three sets of duplicated experiments. Western blot analyzed the key regulators between CtBP1/2 knockdown and control (G). Rest represents without stimulation.
Nuclear γH2AX and RPA32 phosphorylation foci number/cell were calculated and compared between CtBP1-KD/CtBP2-KD and control cells after 24-h treatment KU-0060648/NU7441 (2 μM) (Fig. 5C). The γH2AX foci number/cell significantly increased in CtBP1/2-KD cells (P < 0.001, Fig. 5D). KU-0060648 treatment increased the γH2AX and RPΑ32 foci number slightly but not significantly (Fig. 5C–E). NU7441 significantly increased the number of γH2AX and RPΑ32 phosphorylation foci in CtBP1/2-KD cells than control cells (Fig. 5C–E). KU-0060648 had the potential to increase the rate of homology-directed repair while inhibiting the NHEJ repair shift [27]. In light of these findings, it is proposed that DNA-PK be activated for the initial DNA repair and significantly increase the sensitivity to the specific inhibitor of DNA-PK in the CtBP1/2 knockdown serous ovarian cancer cells.
CtBP1/2 knockdown regulated NHEJ/HR pathway shift in DSBs repair
To explore the impact of CtBPs knockdown in DSBs repair, the SSR 2.0 reporter was used to evaluate the NHEJ/HR pathway shift in shRNA-mediated CtBP1/2 KD serous ovarian cancer cells, which is a genetic encoding sensor that specifically measures the shift of NHEJ or HR pathway by I-SceI-induced DSBs repair [29] (Fig. S3A). Under rest condition, the NHEJ/HR ratio in CtBP1/2 KD cells was identified to be greater than 1.0 when normalized with scrambled shRNA control (1.21 ± 0.17, CtBP1-KD; 1.59 ± 0.18, CtBP2-KD) (Fig. S3B). As a result, CtBP1/2 KD shifted the balance to NHEJ pathway (Fig. 5F). Carboplatin induces high-cytotoxicity DNA lesions in cells, which is mainly repaired by HR pathway [30]. Carboplatin or etoposide treatment induced a different repair pathway balance shift in CtBP1/2 KD cells. CtBP2-KD cells dramatically shifted to NHEJ pathway (NHEJ/HR ratio = 6.69 ± 1.69) and CtBP1-KD slightly inverted to HR pathway (NHEJ/HR ratio = 0.96 ± 0.07) after carboplatin (200 μM, 24 h) treatment (Fig. S3C). Etoposide bound to topoisomerase II (TOP2) and induces DSBs, which were primarily repaired by NHEJ pathway, while BRCA1 promoted the removal of TOP2-cleavage complexes from DSBs [31]. Treated with etoposide (50 μM, 24 h), CtBP1-KD cells shifted slightly from HR to NHEJ pathway (NHEJ/HR ratio = 1.28 ± 0.33), whereas CtBP2-KD cells remained in the NHEJ pathway (NHEJ/HR ratio = 1.058 ± 0.11) (Fig. S3D, Fig. 5F).
Furthermore, western blot analysis revealed that the phosphorylation levels of Chk1-Ser345 and Chk2-Thr68 were significant lower in CtBP1/2 KD cells than the control cells (Fig. 5G). It was hypothesized that the activity of Chk1 and Chk2 would be suppressed in the CtBP1/2 KD cells. As a key molecule of DDR-signaling downstream and DNA damage marker, phosphorylation level of Chk1-Ser345 was crucial for Chk1 activation and necessary for HR repair [32]. When exposed to ionizing radiation or UV, phosphorylate level of Chk2-Thr68 increased, causing the pathway switch from the NHEJ to error-free HR. DNA-PK controlled the Chk2-Thr68 phosphorylation level and regulated the Chk2–BRCA1 pathway to reverse pathway shift and ensured chromosomal stability [33]. We proposed that CtBP2 would play a dominant role in maintaining genetics stability and regulate NHEJ/HR pathway shift in DSBs repair.
Patients with CtBP2 genetic alterations significantly extended the overall survival time
To validate the hypothesis that disrupting CtBP2’s function may inhibit the abnormal growth of cancer cells, we utilized the TCGA via cBioportal to explore the potential correlation of CtBP1/2 genetic alterations with patient’s overall survival time in serous ovarian cancer patients. There are total of 153 (~10.0%, Table S4) patients associated with CtBP1/2 genes genetic alternation among 1680 cases, 78 (5%) in CtBP1 and 75 (5%) in CtBP2, respectively. Interesting, patients with CtBP2’s genetic alternation were significantly associated with longer overall survival time (median time: 61.67 months, log rank test, P value: 3.198e−4, n = 72) than unaltered patients (median time: 44.48 months, n = 1561) in ovarian serous cystadenocarcinoma patients during 10-year survival period (Fig. S4B). While CtBP1-related alterations did not significantly impact the overall survival time (median time: 52.0 months, log rank test, P value: 0.358, q value: 0.613, n = 78) than unaltered patients (median time: 44.55 months, n = 1555, Fig. S4A). These genetic alternations of CtBP1/2 were summarized, including P308H missense mutation and LYAR fusion in CtBP1, and K434Nfs*33 frameshift deletion and KCNMA1-CtBP2 gained fusion merged in CtBP2 (Table S4), including fusion, amplification and homo deletion, missense mutation, and copy-number alterations, which are primarily distributed in ovarian cancer and ovarian epithelial cancer. Recently, genomics features of several ovarian cancer cell lines were well characterized and identified typical genetic alternations in these cancer cells, which are broadly used in research, including SKOV3 and OVSAHO cells. More interesting, the typical genes deletions of NOTCH1, CDKN2A and TP53 in SKOV3 cells [34] were significantly associated with CtBP2’s alternation in the CtBP2 altered patients’ profile (Table S5). It was more convincible and clinical relevant that CtBP2’s genetic alternation would disrupt its function and attenuate it abnormal impact in serous ovarian cancer.
Discussion
In this study, multiple research techniques, including shRNA-based stable CtBP1/2 knockdown, transcription profiles analysis and DNA Fiber, et al., were applied to investigate the contribution of CtBP1/2 in apoptosis, genomic stability, and DDR of serous ovarian cancer cells. Initially, the study purpose was to investigate the contribution of the CtBP proteins play in the HGSOC. According to previously findings, overexpression of CtBP2 was linked with abnormal proliferation, epigenetically silencing BRCA1 function [21], and poorer survival rate in ovarian cancer patients [13]. CtBP1/2 were found to be overexpressed in multiple subtypes of ovarian cancer cells by western blot analysis (Fig. 1A). The SKOV3 cell was chosen as model to investigate the role of CtBP1/2 in HGSOC by integrated shRNA-based stable knockdown and transcription profiles analysis (Fig. 1B, C). The most significantly enriched functional pathways in CtBP1/2 KD cells were mainly concentrated on cell cycle regulation, adhesion, DDR, and apoptosis, according to enrichment analysis for significant change genes (Fig. 1D–E). The abilities of proliferation (Fig. S2A), cell migration and clone-forming in vitro (Fig. S2B–E) were significantly reduced in stable CtBP1/2 KD cells. Single CtBP1-KD or CtBP2-KD induced a high portion of apoptosis (Fig. 2C, D), whereas CtBP1/2 DKD accelerated the apoptosis. CtBP proteins had been identified as anti-apoptotic proteins and negatively regulator through several program cell death genes, such as caspase-3, BIK and D3R, in human cancer cells [17, 35, 36]. In our results, TGF-b was found to be significantly upregulated in the CtBP1/2 KD cells (Figs. 2B and 3B). CtBP1/2 KD increased the metabolic stress and DNA replication instability, and could activate HIPK2, a serine /threonine kinase in the TGF-β signaling pathway, by checkpoint kinase ATM under stress and leading to apoptosis in cancer cells, which directly or indirectly (via JNK1) phosphorylates the sites of Ser422 and Ser428 in CtBP1 and CtBP2 [36,37,38], respectively. It may create a feedback loop, exaggerating and hastening the apoptosis in CtBP DKD cells. CtBP proteins were thought to act as apoptosis suppressers and interact with TGF-β signaling pathway in serous ovarian cancer cells. It could be partially explained that genetic alterations in CtBP2 producing abnormally functional protein and inducing tumor apoptosis, thereby extending the patients’ survival time.
CtBP proteins enhanced mitotic fidelity and genome stability through their metabolic activity in the nucleus [14]. However, it is unclear what role CtBP1/2 in DNA replication and DNA damage repair in serous ovarian cancer cells. In this study, DDR-signaling pathway was highlighted and positively enriched in CtBP2-KD cells rather than in CtBP1-KD cells (Fig. 1D, E). CtBP1/2 knockdown significantly increased the genomic instability and the sensitivity to γ-irradiation, especially in CtBP2 KD cells (Fig. 3C). It could be attributed to difference in intracellular distribution and metabolic protective functions of CtBP1/2. CtBP2 played dominant localized in nuclear and works as transcriptional corepressors [39, 40]. Furthermore, CtBP1/2 knockdown significantly disrupted the stability of the DNA replication fork and increased the instability of DNA replication recovery in serous ovarian cancer cells (Fig. 4C–F). Under the metabolic stress condition (HU treatment), CtBP1 had a greater protective effect than CtBP2’s (Fig. 4D, F).
Approximately 44% of HGSOC patients had HR deficiency (24). Defects in DNA repair were likely to affect the sensitivity of platinum in HGSOC [4]. DSBs were most of the cytotoxicity response to genotoxic stress. NHEJ and HR pathways mainly involved in the DNA repair of DSBs. NHEJ pathway was a highly error-prone DNA repair mechanism that operates throughout the cell cycle [41]. DNA-PK was a mediator of NHEJ and specific DNA-PK inhibitors were widely used in platinum-resistant cancer cell lines as a chemo-sensitization strategy [42]. In this study, CtBP1/2 knockdown increased genetic instability and activated DNA-PK during DNA breakage. The preference of DNA repair pathway shifted from HR to NHEJ under resting condition. Chemotherapy drugs (Carboplatin and Etoposide) treatment had an effect on the trend of NHEJ pathway shift (Fig. 5F, Fig. S3B–D). CtBP2 knockdown did not enhance the carboplatin cytotoxic effect in serous cancer cells (data not shown). A synergic effect of a specific DNA-PK inhibitor and CtBP2 KD would exist in serous ovarian cancer cells. Based on these findings, a novel potential treatment strategy, the combination of specific CtBP2 protein inhibitor and DNA-PK inhibitors, was proposed to target CtBP2 protein and DNA-PK in platinum-resistant serous ovarian cancer.
We investigated the correlation of CtBP1/2 genetic alterations with patient overall survival time in serous ovarian cancer patients using TCGA cases to validate the results that disruption of CtBP proteins function would lead to cancer cell apoptosis and increase DNA instability. Although CtBP1 and CtBP2 had similar levels of genomic alterations (around 5%), CtBP2’s genetically altered cases had significantly longer survival time (Fig. S4B) than unaltered cases. Previously, CtBP2 overexpression was linked with poorer survival rate in invasive ovarian cancer [13]. Oncogene amplification and overexpression are not the same thing. The amplification copy of a gene may be translocated into parental alleles or other chromosome(s) or extra-chromosome acentric elements [43]. Gene overexpression could be caused by a variety of epigenetic and genetic modifications [44, 45], including copy number increase and paternal gene overactivation, etc. Oncogene’s overexpression may not be associated with detectable amplification. For example, the oncogene MET was amplified with 5.1% cases while overexpressed in 13.1% cases of glioma [46]. Gained-fused KCNMA1-CtBP2 gene may destroy the function of KCNMA1, which was overexpressed in ovarian cancer cells and promoted proliferation, migration and attenuation of apoptosis [47], and induced apoptosis in ovarian cancer cells. It is possible that genetic alterations of CtBP2 destroy the abnormal activity of CtBP2 in serous ovarian cancer, resulting in a longer survival time for patients with the disease. More intriguing, some typical genetic alternations of SKOV3 cell [34], such as deletion of NOTCH1, CDKN2A, and TP53 genes, were significantly associated with CtBP2’s alternation in the CtBP2 altered patients’ group (Table S5). It was more convincible and clinical relevant that CtBP2’s genetic alternation would disrupt its function and attenuate it abnormal impact in serous ovarian cancer. Up to now, little is known about the exactly mechanism linking between CtBP2’s genetic alternation and significantly longer overall survival time. It will be an exciting new research direction for CtBP2 role in ovarian cancer, with greater potential translational application for ovarian cancer patients.
In summary, this study discovered that CtBP1/2 played differentially protector of genetic stability and DNA repair pathways in serous ovarian cancer cells. CtBP1/2 contributed to maintain the stability of DNA replication fork and DNA repair in serous ovarian cancer cells. CtBP2-related genetic alterations may destroy its abnormal activity and abolish the proliferation-promoting effect in serous ovarian cancer. Combined with these results, it will give a new perspective on the regulatory role of CtBP proteins in genetic stability and DNA repair pathway in ovarian cancer. Advance knowledge of ovarian cancer deregulatory mechanism will accelerate the translational application and development of novel promising clinical treatment strategy and solutions for serous ovarian cancer patients.
Materials and methods
Ovarian cancer cells culture
Human ovarian cancer cell lines (HOSE, SKOV3, MCAS, RMG1, and RMUGL) were purchased from ATCC and cultured in DMEM medium or RPMI 1640 supplemented with 10% fetal bovine serum, 0.1 mg/mL penicillin, and 0.1 mg/mL streptomycin.
Statistical analysis
Data were presented as means ± SEM. Significance of differences for the values was determined using the Student’s t-test with Prism software (GraphPad Software, Inc. San Diego, CA). A P value less than 0.05 was considered a significant difference.
Detailed descriptions of the following techniques were available in the Supplementary Materials and Methods:
-
(1)
Established CtBP1/2 stable knockdown ovarian cancer cells
-
(2)
Western blotting
-
(3)
Whole transcript expression profiling analysis
-
(4)
Proliferation, drug treatment, wound scratch, and transwell assay
-
(5)
Colony-forming assays
-
(6)
Cell cycle analysis
-
(7)
DNA fiber assay
-
(8)
Immunofluorescence image analysis for DNA damage response
-
(9)
Comet assay for DNA Damage induced by Irradiation
-
(10)
NHEJ and HR repair pathways shift Assay
-
(11)
Explore CtBP1/2 gene with serous ovarian carcinoma cases through TCGA
References
Altekruse S, Kosary C, Krapcho M, Neyman N, Aminou R, Waldron W, et al. SEER cancer statistics review, 1975-2007. Bethesda, MD, USA: National Cancer Institue; 2010.
Seidman JD, Horkayne-Szakaly I, Haiba M, Boice CR, Kurman RJ, Ronnett BM. The histologic type and stage distribution of ovarian carcinomas of surface epithelial origin. Int J Gynecol Pathol. 2004;23:41–44.
Bowtell DD, Bohm S, Ahmed AA, Aspuria PJ, Bast RC Jr, Beral V, et al. Rethinking ovarian cancer II: reducing mortality from high-grade serous ovarian cancer. Nat Rev Cancer. 2015;15:668–79.
Mittempergher L. Genomic characterization of high-grade serous ovarian cancer: dissecting its molecular heterogeneity as a road towards effective therapeutic strategies. Curr Oncol Rep. 2016;18:44.
Bowtell DD. The genesis and evolution of high-grade serous ovarian cancer. Nat Rev Cancer. 2010;10:803–8.
Cancer Genome Atlas Research N. Integrated genomic analyses of ovarian carcinoma. Nature. 2011;474:609–15.
Hosoya N, Miyagawa K. Targeting DNA damage response in cancer therapy. Cancer Sci. 2014;105:370–88.
Lord CJ, Ashworth A. The DNA damage response and cancer therapy. Nature. 2012;481:287–94.
Spyer M, Allday MJ. The transcriptional co-repressor C-terminal binding protein (CtBP) associates with centrosomes during mitosis. Cell Cycle. 2006;5:530–7.
Bergman LM, Morris L, Darley M, Mirnezami AH, Gunatilake SC, Blaydes JP. Role of the unique N-terminal domain of CtBP2 in determining the subcellular localisation of CtBP family proteins. BMC Cell Biol. 2006;7:35.
Boyd JM, Subramanian T, Schaeper U, La Regina M, Bayley S, Chinnadurai G. A region in the C-terminus of adenovirus 2/5 E1a protein is required for association with a cellular phosphoprotein and important for the negative modulation of T24-ras mediated transformation, tumorigenesis and metastasis. EMBO J. 1993;12:469–78.
Schaeper U, Boyd JM, Verma S, Uhlmann E, Subramanian T, Chinnadurai G. Molecular cloning and characterization of a cellular phosphoprotein that interacts with a conserved C-terminal domain of adenovirus E1A involved in negative modulation of oncogenic transformation. Proc Natl Acad Sci USA. 1995;92:10467–71.
Barroilhet L, Yang J, Hasselblatt K, Paranal RM, Ng SK, Rauh-Hain JA, et al. C-terminal binding protein-2 regulates response of epithelial ovarian cancer cells to histone deacetylase inhibitors. Oncogene. 2013;32:3896–903.
Di LJ, Byun JS, Wong MM, Wakano C, Taylor T, Bilke S, et al. Genome-wide profiles of CtBP link metabolism with genome stability and epithelial reprogramming in breast cancer. Nat Commun. 2013;4:1449.
Chinnadurai G. CtBP, an unconventional transcriptional corepressor in development and oncogenesis. Mol Cell. 2002;9:213–24.
Chinnadurai G. The transcriptional corepressor CtBP: a foe of multiple tumor suppressors. Cancer Res. 2009;69:731–4.
Ding B, Yuan F, Damle PK, Litovchick L, Drapkin R, Grossman SR. CtBP determines ovarian cancer cell fate through repression of death receptors. Cell Death Dis. 2020;11:286.
Bergman LM, Blaydes JP. C-terminal binding proteins: emerging roles in cell survival and tumorigenesis. Apoptosis. 2006;11:879–88.
Grooteclaes M, Deveraux Q, Hildebrand J, Zhang Q, Goodman RH, Frisch SM. C-terminal-binding protein corepresses epithelial and proapoptotic gene expression programs. Proc Natl Acad Sci USA. 2003;100:4568–73.
Kovi RC, Paliwal S, Pande S, Grossman SR. An ARF/CtBP2 complex regulates BH3-only gene expression and p53-independent apoptosis. Cell Death Differ. 2010;17:513–21.
May T, Yang J, Shoni M, Liu S, He H, Gali R, et al. BRCA1 expression is epigenetically repressed in sporadic ovarian cancer cells by overexpression of C-terminal binding protein 2. Neoplasia. 2013;15:600–8.
Donzelli M, Draetta GF. Regulating mammalian checkpoints through Cdc25 inactivation. EMBO Rep. 2003;4:671–7.
Dose M, Emmanuel AO, Chaumeil J, Zhang J, Sun T, Germar K, et al. beta-Catenin induces T-cell transformation by promoting genomic instability. Proc Natl Acad Sci USA. 2014;111:391–6.
Janssens S, Tschopp J. Signals from within: the DNA-damage-induced NF-kappaB response. Cell Death Differ. 2006;13:773–84.
Sharma A, Singh K, Almasan A. Histone H2AX phosphorylation: a marker for DNA damage. Methods Mol Biol. 2012;920:613–26.
Ashley AK, Shrivastav M, Nie J, Amerin C, Troksa K, Glanzer JG, et al. DNA-PK phosphorylation of RPA32 Ser4/Ser8 regulates replication stress checkpoint activation, fork restart, homologous recombination and mitotic catastrophe. DNA Repair. 2014;21:131–9.
Munck JM, Batey MA, Zhao Y, Jenkins H, Richardson CJ, Cano C, et al. Chemosensitization of cancer cells by KU-0060648, a dual inhibitor of DNA-PK and PI-3K. Mol Cancer Ther. 2012;11:1789–98.
Hardcastle IR, Cockcroft X, Curtin NJ, El-Murr MD, Leahy JJ, Stockley M, et al. Discovery of potent chromen-4-one inhibitors of the DNA-dependent protein kinase (DNA-PK) using a small-molecule library approach. J Med Chem. 2005;48:7829–46.
Gomez-Cabello D, Jimeno S, Fernandez-Avila MJ, Huertas P. New tools to study DNA double-strand break repair pathway choice. PLoS ONE. 2013;8:e77206.
McHugh PJ, Spanswick VJ, Hartley JA. Repair of DNA interstrand crosslinks: molecular mechanisms and clinical relevance. Lancet Oncol. 2001;2:483–90.
Sasanuma H, Tsuda M, Morimoto S, Saha LK, Rahman MM, Kiyooka Y, et al. BRCA1 ensures genome integrity by eliminating estrogen-induced pathological topoisomerase II-DNA complexes. Proc Natl Acad Sci USA. 2018;115:E10642–E10651.
Draga M, Madgett EB, Vandenberg CJ, du Plessis D, Kaufmann A, Werler P, et al. BRCA1 is required for maintenance of phospho-Chk1 and G2/M arrest during DNA cross-link repair in DT40 cells. Mol Cell Biol. 2015;35:3829–40.
Shang Z, Yu L, Lin YF, Matsunaga S, Shen CY, Chen BP. DNA-PKcs activates the Chk2-Brca1 pathway during mitosis to ensure chromosomal stability. Oncogenesis. 2014;3:e85.
Elias KM, Emori MM, Papp E, MacDuffie E, Konecny GE, Velculescu VE, et al. Beyond genomics: critical evaluation of cell line utility for ovarian cancer research. Gynecol Oncol. 2015;139:97–103.
Paliwal S, Pande S, Kovi RC, Sharpless NE, Bardeesy N, Grossman SR. Targeting of C-terminal binding protein (CtBP) by ARF results in p53-independent apoptosis. Mol Cell Biol. 2006;26:2360–72.
Wang SY, Iordanov M, Zhang Q. c-Jun NH2-terminal kinase promotes apoptosis by down-regulating the transcriptional co-repressor CtBP. J Biol Chem. 2006;281:34810–5.
Hofmann TG, Stollberg N, Schmitz ML, Will H. HIPK2 regulates transforming growth factor-beta-induced c-Jun NH(2)-terminal kinase activation and apoptosis in human hepatoma cells. Cancer Res. 2003;63:8271–7.
Zhang Q, Yoshimatsu Y, Hildebrand J, Frisch SM, Goodman RH. Homeodomain interacting protein kinase 2 promotes apoptosis by downregulating the transcriptional corepressor CtBP. Cell. 2003;115:177–86.
Bergman LM, Birts CN, Darley M, Gabrielli B, Blaydes JP. CtBPs promote cell survival through the maintenance of mitotic fidelity. Mol Cell Biol. 2009;29:4539–51.
Birts CN, Bergman LM, Blaydes JP. CtBPs promote mitotic fidelity through their activities in the cell nucleus. Oncogene. 2011;30:1272–80.
Cortizas EM, Zahn A, Hajjar ME, Patenaude AM, Di, Noia JM, et al. Alternative end-joining and classical nonhomologous end-joining pathways repair different types of double-strand breaks during class-switch recombination. J Immunol. 2013;191:5751–63.
Boeckman HJ, Trego KS, Turchi JJ. Cisplatin sensitizes cancer cells to ionizing radiation via inhibition of nonhomologous end joining. Mol Cancer Res. 2005;3:277–85.
Matsui A, Ihara T, Suda H, Mikami H, Semba K. Gene amplification: mechanisms and involvement in cancer. Biomol Concepts. 2013;4:567–82.
D’Angelo D, Esposito F, Fusco A. Epigenetic mechanisms leading to overexpression of HMGA proteins in human pituitary adenomas. Front Med. 2015;2:39.
Minarovits J, Banati F, Szenthe K, Niller HH. Epigenetic regulation. Adv Exp Med Biol. 2016;879:1–25.
Kwak Y, Kim SI, Park CK, Paek SH, Lee ST, Park SH. C-MET overexpression and amplification in gliomas. Int J Clin Exp Pathol. 2015;8:14932–8.
Ma SY, Wei P, Qu F. KCNMA1-AS1 attenuates apoptosis of epithelial ovarian cancer cells and serves as a risk factor for poor prognosis of epithelial ovarian cancer. Eur Rev Med Pharm Sci. 2019;23:4629–41.
Acknowledgements
This work was supported by the grant for outstanding talent from abroad by Chinese Academic of Science and startup support funding from “CAS Pioneer Hundred Talents Program” (E0241211H1) and State Key Laboratory of Phytochemistry and Plant Resources in West China, Kunming Institute of Botany, the Chinese Academy of Sciences (Y8677211K1, Y8690211Z1) to SL.
Author information
Authors and Affiliations
Contributions
S.L. and Y.H. designed the overall project study. Y.Z.C., Z.C.H., C.C., J.L., S.L., Y.H., and S.L. collected data, performed data analysis. S.L. and Y.H. interpreted and summarized the results, wrote and revised the manuscript. All authors have read and approved the final version of manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
He, Y., He, Z., Lin, J. et al. CtBP1/2 differentially regulate genomic stability and DNA repair pathway in high-grade serous ovarian cancer cell. Oncogenesis 10, 49 (2021). https://doi.org/10.1038/s41389-021-00344-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41389-021-00344-9
- Springer Nature Limited