Abstract
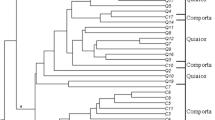
Wild Cicer germplasm is known to havegenes for disease resistance, stresstolerance and other important traits, andhence could be exploited for improvingcultivated genotypes. However, only fewCicer species are interfertile and itis essential to overcome crossabilitybarriers to utilize the germplasm moreeffectively. Genetic diversity analysis ofCicer species can give importantclues in understanding speciesrelationships and may assist in developingand planning breeding strategies. Weselected 6 annual and 7 perennial wildspecies, which were amplified using 15 ISSRprimers and UPGMA and AMOVA were used toevaluate the genetic diversity. On anaverage, 6.6 polymorphic bands per primerwere observed. Cluster analysis using theUPGMA algorithm indicated three majorgroups of species at the similarity valueof 0.60 with many subclusters. Theclustering pattern was in agreement withthe data based on crossability, seedstorage protein, isozyme, allozyme and RAPDmarker analysis. The among-populationcomponent of annual and perennial groupscalculated using AMOVA accounted for39.00%. Our results suggested that wildannuals of Cicer were notmonophyletic in nature.
Similar content being viewed by others
References
Aggarwal, R.K., D.S. Brar, S. Nandi, N. Huang & G.S. Khush, 1999. Phylogenetic relationships among Oryza species revealed by AFLP markers. Theor Appl Genet 98(8): 1320-1328.
Ahmad, F., 1988. Interspecific hybridisation and genetic relationships among the annual Cicer L. species. Ph.D thesis University of Saskatchewan, Saskatoon, Canada.
Ahmad, F., 1999. Random amplified polymorphic DNA (RAPD) analysis reveals genetic relationships among the annual Cicer species. Theor Appl Genet 98: 657-663.
Ahmad, F. & A.E. Slinkard, 1992. Genetic relationships in the genus Cicer L. as revealed by polyacrylamide gel electrophoresis of seed storage proteins. Theor Appl Genet 84: 688-692.
Ahmad, F., A.E. Slinkard & G.J. Scoler, 1987. The cytogenetic relationship between Cicer judaicum Boiss. and Cicer chroassannicum (Bge) M. Pop. Genome 29: 883-886.
Anonymous, 1995. Kabuli Chickpea Improvement. Annu Rep Germplasm Program: Legumes. ICARDA, Aleppo, Syria pp. 59-132.
Barret, B.A. & K.K. Kidwell, 1998. AFLP based genetic diversity assessment among wheat cultivars from the pacific Northwest. Crop Sci 38: 1261-1271.
Blair, M.W., O. Panaud & S.R. McCouch, 1999. Inter simple sequence repeat (ISSR) amplification for analysis of microsatellite motif frequency and fingerprinting in rice (Oryza sativa L.) Theor Appl Genet 98: 780-792.
Brummer, E.C., J.H. Bouton & G. Kochert, 1995. Analysis of annual Medicago species using RAPD markers. Genome 38: 362-367.
Campos, L.P., J.V. Raelson & W.F. Grant, 1994. Genome relationships among Lotus species based on random amplified polymorphic DNA (RAPD). Theor Appl Genet 88: 417-422.
Chan, K.F. & M. Sun, 1997. Genetic diversity and relationships detected by isozyme and RAPD analysis of crop and wild species of Amaranthus. Theor Appl Genet 95: 865-873.
Davierwala, A.P., K.V. Chowdari, A. Shivkumar, A.P.K. Reddy, P.K. Ranjekar & V.S. Gupta, 2001. Genetic diversity evaluation of Indian elite rice varieties using molecular markers. Genetica (in press).
Doyle, J.J. & J.L. Doyle, 1987. A rapid DNA isolation procedure for small amount of fresh leaf tissue. Phytochem Bull 19: 11-15.
Fahima, T., G.L. Sun, A. Beharav, T. Krugman, A. Beiles & E. Nevo, 1999. RAPD polymorphism of wild emmer wheat populations, Triticum dicoccoides in Israel. Theor Appl Genet 98: 434-447.
Felsenstein, J., 1985. Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39: 783-791.
Garcia, G.M., H.T. Stalker, E. Shroeder & G. Kochert, 1996. Identification of RAPD, SCAR and RFLP markers tightly linked to nematode resistance genes introgressed from Arachis cardenasii into Arachis hypogaea. Genome 39: 836-845.
Gotz, G., M. Nenno, K. Weising, D. Zink, W. Nagl & G. Kahl, 1998. Chromosomal localization and distribution of Simple sequence repeats and the Arabidopsis-type telomere sequence in the genome of Cicer arietinum L. Chromosome Research 6: 97-104.
Gupta, V.S., W. Ramakrishna, S.R. Rawat & P.K. Ranjekar, 1994. (CAC)5 detects DNA fingerprints and sequences homologous to gene transcripts in rice. Biochem Genet 32: 1-8.
Helbaek, H., 1959. Domestication of food plants in the old world. Science 130: 365-372.
Ishii, T., T. Nakano, H. Maeda & O. Kamijima, 1996. Phylogenetic relationships in A-genome species of rice revealed by RAPD analysis. Genes Genet Syst 71: 195-201.
Ishii, T., T. Terachi & K. Tsunewaki, 1988. Restriction endonuclease analysis of chloroplast DNA from A-genome diploid species of rice. Jpn J Genet 63: 523-536.
Joshi, S.P., V.S. Gupta, R.K. Aggarwal, P.K. Ranjekar & D.S. Brar, 2000. Genetic diversity and phylogenetic relationship as revealed by inter-simple sequence repeat (ISSR) polymorphisms in the genus Oryza. Theor Appl Genet 100: 1311-1320.
Kazan, K. & F.J. Muehlbauer, 1991. Allozyme variation and phylogeny in annual species of Cicer (Leguminosae). Pl Syst Evol 175: 11-21.
Labdi, M., L.D. Robertson, K.B. Singh & A. Charrier, 1996. A genetic diversity and phylogenetic relationships among the annual Cicer species as revealed by isozyme polymorphism. Euphytica 88: 181-188.
Ladizinsky & Adler, 1976. Genetic relationships among the annual species of Cicer L. Theor Appl Genet 48: 197-203.
Lanner, C., T. Bryngelsson & M. Gustafsson, 1996. Genetic variability of RAPD markers at the intra-and inter-specific level in wild Brassica species with n = 9. Theor Appl Genet 93: 9-14.
Lanner-Herrera, C., M. Gustafsson, A.S. Falt & T. Bryngelsson, 1996. Diversity in natural populations of wild Brassica oleracea as estimated by isozyme and RAPD analysis. Genet Res Crop Evol 43: 13-23.
Matsuoka, Y. & K. Tsunewaki, 1999. Evolutionary dynamics of Ty1-copia group retrotransposons in grass shown by reverse transcriptase domain analysis. Mol Biol Evol 16: 208-217.
McCoy, T.J. & C.S. Echt, 1993. Potential of trispecies bridge crosses and random amplified polymorphic DNA markers for introgression of Medicago daghestanica and Medicago pironae germplasm into alfalfa (Medicago sativa). Genome 36: 594-601.
Nelson, R.J. & V.I. Yap, 1996. Winboot: A Program for Performing Boot Strap Analysis of Binary Data to determine the Confidence Limits of UPGMA Based Dendrograms. IRRI Discussion paper series No. 14 International Rice Research Institute, PO Box 933, Manila, Phillipines.
Ocampo, B., G. Venora, A. Errico, K.B. Singh & F. Saccardo, 1992. Karyotype analysis in the genus Cicer. J Genet Breed 46: 229-240.
Patil, P., P.L. Vrinten, G.J. Scoles & A.E. Slinkard, 1995. Variation in the ribosomal RNA units of the genera Lens and Cicer. Euphytica 83: 33-42.
Peakall, R., S. Gilmore, W. Keys, M. Morgante & A. Rafalski, 1998. Cross-species amplification of soybean (Glycine max) simple sequence repeats (SSRs) within the genus and other legume genera: implications for the transferrability of SSRs in plants. Mol Biol Evol 15: 1275-1287.
Plaschke, J., M.W. Ganal & M.S. Roder, 1995. Detection of genetic diversity in closely related bread wheat using microsatellite markers. Theor Appl Genet 91: 1001-1007.
Pundir, R.P.S. & L.J.G. Vander Maesen, 1983. Interspecific hybridisation in Cicer. Int Chickpea Newsl 8: 4-5.
Ratnaparkhe, M.B., V.S. Gupta, M.R. Ven Murthy & P.K. Ranjekar, 1995. Genetic fingerprinting of pigeonpea (Cajanus cajan (L.) Millsp.) and its wild relatives using RAPD markers. Theor Appl Genet 91: 893-898.
Ratnaparkhe, M.B., D.K. Santra, A. Tullu & F.J. Muehlbauer, 1998. Inheritance of Inter simple sequence repeat polymorphism and linkage with fusarium wilt resistance gene in chickpea. Theor Appl Genet 96: 348-353.
Rieseberg, L., 1996. Homology among RAPD fragments in interspecific comparisons. Molec Ecol 5: 99-105.
Robertson, L.D., B. Ocampo & K.B. Singh, 1997. Morphological variation in wild annual Cicer species in comparison to the cultigen. Euphytica 95: 309-319.
Sales, E., S.G. Nebauer, M. Mus & J. Segura, 2001. Population genetic study in the Balearic endemic plant species Digitalis minor (Scrophulariaceae) using RAPD markers. Amer J Bot 88: 1750-1759.
Sambrook, J., E.F. Fritsch & T. Maniatis, 1989. Molecular Cloning: A Laboratory Manual (2nd edn.), Cold Spring Harbor Laboratory Press. Cold Spring Harbor, New York.
Sant, V.J., A.G. Patankar, V.S. Gupta, N.D. Sarode, L.B. Mhase, M.N. Sainani, R.B. Deshmukh & P.K. Ranjekar, 1999. Potential of DNA markers in detecting divergence and in analyzing heterosis in Indian elite chickpea cultivars. Theor Appl Genet 98: 1217-1225.
Saxena, M.C. & K.B. Singh, 1987. The Chickpea. CAB international, Wallingford, UK.
Serret, M.D., S.M. Udupa & F. Weigand, 1997. Assessment of genetic diversity of cultivated chickpea using microsatellitederived RFLP markers: Implications for origin. Plant Breed 116: 573-578.
Sharma, P.C., B. Huttel, P. Winter, G. Kahl, R.C. Gardner & K. Weising, 1995. The potential of microsatellites for hybridization and PCR-based DNA fingerprinting of Chickpea and related species. Electrophoresis 16: 1755-1761.
Simon, C.J. & F.J. Muehlbauer, 1997. Construction of a chickpea linkage map and its comparison with maps of pea and lentil. J Hered 88: 115-119.
Singh, K.B., 1987. Chickpea breeding. In: M.C. Saxena & K.B. Singh (Eds.), The Chickpea, pp. 127-162. CAB Int Publ, UK.
Singh, K.B. & B. Ocampo, 1997. Exploitation of wild Cicer species for yield improvement in chickpea. Theor Appl Genet 95: 418-423.
Sneath, P.H.A. & R.R. Sokal, 1973. Numerical Taxonomy WHF. San Francisco, pp. 100-308.
Tayyar, R.I., A.J. Lukaszewiski & J.G. Waines, 1994. Chromosome banding patterns in the annual species of Cicer. Genome 37: 656-663.
Tayyar, R.I. & J.G. Waines, 1996. Genetic relationships among annual species of Cicer (Fabaceae) using isozyme variation. Theor Appl Genet 92; 245-254.
Thomson, J., L.N. Randall & O.L. Vodki, 1999. Identification of diverse soybean germplasm using RAPD markers. Crop Sci 38: 1348-1355.
Tuwafe, S., A.L. Kahler, A. Bor & M. Fergusson, 1988. Inheritance and geographical distribution of allozyme polymorphism in chickpea (Cicer arietinum L). J Heredity 79: 170-174.
Udupa, S.M., L.D. Robertson, F. Weigand, M. Baum & G. Kahl, 1999. Allelic variation at (TAA)n microsatellite loci in a world collection of chickpea (Cicer arietinum L.) germplasm. Mol Gen Genet 261: 354-363.
Vairinhos, F. & D.R. Murray, 1983. The seed proteins of chickpea comparative studies of Cicer arietinum, C. reticulatum and C. echinospermum (Leguminosae). Plant syst Evol 142: 15-22.
Van der Maesen, L.J.G., 1972. Cicer L. A monograph on the genus with special reference to the chickpea (Cicer arietinum L.), its ecology and cultivation. Thesis, Agricultural University Wageningen. Meded Landbouwhogeschool, Wageningen 72-10.
Van der Maesen, L.J.G., 1987. Origin, history and taxonomy of Chickpea. In: M.C. Saxena & K.B. Singh (Eds.), The Chickpea, pp. 11-34. CAB international/ICARDA, Wallingford, Oxon, UK.
Wang, Z.Y., G. Second & S.D. Tanksley, 1992. Polymorphism and phylogenetic relationships among species in the genus Oryza as determined by nuclear RFLPs. Theor Appl Genet 83: 565-581.
Wu, J., K.V. Krutovski & S.H. Strauss, 2000. Nuclear DNA diversity, population differentiation and phylogenetic relationships in the California closed-cone pines based on RAPD and allozyme markers. Genome 42: 893-908.
Yang, G.P., M.A. Saghai Maroof, C.G. MA, Q. Zhang & R.M. Biyashev, 1994. Comparative analysis of microsatellite DNA polymorphism in landraces and cultivars of rice. Mol Gen Genet 245: 187-194.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Rajesh, P., Sant, V., Gupta, V. et al. Genetic relationships among annual and perennial wild species of Cicer using inter simple sequence repeat (ISSR) polymorphism. Euphytica 129, 15–23 (2003). https://doi.org/10.1023/A:1021567821141
Issue Date:
DOI: https://doi.org/10.1023/A:1021567821141