Abstract
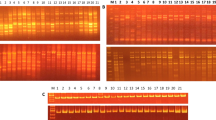
The cluster analysis of ten hop (Humulus lupulus L.) varieties and the comparison of amplified DNA polymorphism were assessed using RAPD, STS, ISSR and AFLP molecular methods. Thirty-five RAPD primers generated an average of 13.4 products with 5.7 polymorphic products (42.3% of polymorphism). Ten STS primer combinations generated an average of 9.3 products with 6.6 polymorphic products (71% of polymorphism). Seven ISSR microsatellite polymorphic primers and seven primer combinations generated an average of 12.9 products with 4.2 polymorphic products (32.6% of polymorphism) in agarose gels and 86.2 products with 24.3 polymorphic products (28.3% of polymorphism) in polyacrylamide gels. A total of 56 AFLP primer combinations generated an average of 91 products with 52.5 polymorphic products (57.6% of polymorphism). All molecular methods accurately distinguished all tested varieties, except for Osvald’s clones (31, 72 and 114), which were only distinguished by AFLP. The similarities between varieties were revealed by cluster analyses. These analyses were very similar for all molecular methods. Pairwise values of JCS (Jaccard’s similarity coefficient) ranged from 0.02 to 1.0 and the mean JCS values ranged from 0.395 (AFLP) to 0.462 (ISSR). The highest correlation coefficient was between ISSR and RAPD (0.966) and the lowest was between AFLP and STS (0.867). STS had the highest correlation of chemical data with molecular methods (0.592) and ISSR had the lowest (0.435).
Similar content being viewed by others
References
Abbott, M.S. & M.J. Fedele, 1994. A DNA-based identification procedure for hop leaf tissue.J Inst Br 100: 283–285.
Araki, S., Y. Tsuchiya, T. Masachika, T. Tamaki & K. Shinotsuka, 1998.Identification of hop cultivars by DNA marker analysis. J Amer Soc Br Chem 56: 93–98.
Blair, M.M.,O. Panaud & S.R. McCouch, 1999. Inter-simple sequence repeat (ISSR) amplification for analysis of microsatellite motif frequency and fingerprinting in rice (Oryza sativa L.). Theor Appl Genet 98: 780–792.
Bohn, M., H.F. Utz & A.E. Melchinger, 1999. Genetic similarities among winter wheat cultivars determined on the basis of RFLPs, AFLPs and SSRs and their use for predicting progeny variance. Crop Sci 39: 228–237.
Brady, J.L., N.S. Scott & M.R. Thomas, 1996. DNA typing of hops (Humulus lupulus) through application of RAPD and microsatellite marker sequences converted to sequence tagged sites (STS). Euphytica 91: 277–284.
Cervera, M.-T., J.A. Cabezas, J.C. Sancha F. Martínez de Toda & J.M. Martínez-Zapater, 1998. Application of AFLPs to the characterization of grapevine Vitis vinifera L. genetic resources. A case study with accessions from Rioja (Spain). Theor Appl Genet 97: 51–59.
Godwin, I.D., E.A.B. Aitken & L.W. Smith, 1997. Application of inter simple sequence repeat (ISSR)markers to plant genetics. Electrophoresis 18: 1524–1528.
Jaccard, P., 1908. Nouvelles recherches sur ladistribution florale. Bull Soc Vaud Sci Nat 44: 223–270.
Jakše, J., J. Šuštar-Vozliĉ & B. Javornik,1994. Identification of hop cultivars by RAPD markers. Proc Intl Colloquium on the Impact of Plant Biotechnology on Agriculture. Rogla, Slovenia, pp. 147–151.
Hartl, L. & S. Seefelder, 1998. Diversity of selected hopcultivars detected by fluorescent AFLPs. Theo Appl Genet 96: 112–116.
Krajl, D., J. Zupanec, D. Vasilj, S. Kralj & J. Pšeniĉnik, 1991. Variability of essential oils of hops, Humulus lupulus L. J Inst Br 97: 197–206.
Mantel, N.A., 1967. The detection of disease clustering and a generalized regression approach. CancRes 27: 209–220.
Murakami, A., 1998. The practical application of PCR for the verification of hop variety.MBAA Tech Q 35: 185–188.
Neve, R.A., 1991. Hops. Chapman and Hall, London.
Parsons, B.J.,H.J. Newbury, M.T. Jackson & B.V. Ford-Lloyd, 1997. Contrasting genetic diversity relationships are revealed in rice (Oryza sativa L.) using different marker types. Mol Breed 3: 115–125.
Patzak, J., P. Oriniaková, J. Matoušek & P. Svoboda, 1999. Czech hop characterization using RAPD method and genetic distance analysis of selected genotypes. Pl Prod 45: 165–172.
Peacock, V.E. & P. McCarty, 1992. Varietal identification of hops and hoppellets. MBAA Techn Quart 29: 81–85.
Pillay, M. & S.T. Kenny, 1996. Random amplifiedpolymorphic DNA (RAPD) markers in hop, Humulus lupulus: level of genetic variability and segregation in progeny. Theor Appl Genet 92: 334–339.
Polley, A., E. Seigner & M.W. Ganal, 1997. Identification of sex in hop (Humuluslupulus) using molecular markers. Genome 40: 357–361.
Powell, W., M. Morgante, C. Andre, M. Hanafey, J. Vogel, S. Tingey & A. Rafalski, 1996. The comparison of RFLP, RAPD, AFLP and SSR (microsatellite) markers for gemplasm analysis. Molec Breed 2: 225–238.
Rogers, D.G. & T.T. Tanimoto, 1960. A computerprogram for classifying plants. Science 132: 1115–1118.
Saghai-Maroof, M.A., K.M. Soliman, R.A. Jorgensen & R.W. Allard, 1984. Ribosomal DNA spacer-length polymorphism in barley: Mendelian inheritance, chromosomal location, and population dynamics. Proc Natl Acad Sci USA 81: 8014–8018.
Sensi, E., R. Vignani, W. Rohde & S. Biricolti, 1996. Characterization of genetic biodiversity with Vitis vinifera L. Sangiovese and Colorino genotypes by AFLP and ISTR DNA marker technology. Vitis 35: 183–188.
Šuštar-Vozliĉ, J. & B. Javornik,1999. Genetic relationships in cultivars of hop, Humulus lupulus L., determined by RAPD analysis. Plant Breed 118: 175–181.
Tsuchiya, Y., S. Araki, M. Takashio & T. Tamaki, 1997. Identification of hop varieties using specificprimers derived from RAPD markers. J Ferment Bioeng 84: 103–107.
Vejl, P., 1997. Identification ofgenotypes in hop (Humulus lupulus L.) by RAPD analysis using program GelManager forWindows. Pl Prod 43: 325–333.
Williams, J.G.K., A.R. Kubelik, K.J. Livak, J.A. Rafalski & S.V. Tingey, 1990. DNA polymorphismamplified by arbitrary primers are useful as genetic markers. Nucl Acids Res 18: 6531–6535.
Zabeau, M. & P. Vos, 1993. Selective restriction fragment amplification. A general method for DNA fingerprinting. European patent application no. 924026297. Publication no. 0534858.
Zietjiewicz, E., A. Rafalski & D. Labuda, 1994. Genomefingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genomics 20: 176–183.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Patzak, J. Comparison of RAPD, STS, ISSR and AFLP molecular methods used for assessment of genetic diversity in hop (Humulus lupulus L.). Euphytica 121, 9–18 (2001). https://doi.org/10.1023/A:1012099123877
Issue Date:
DOI: https://doi.org/10.1023/A:1012099123877