Abstract
Proteins are indispensable for maintaining a healthy diet and performing crucial functions in a multitude of physiological processes. The growth of the global population and the emergence of environmental concerns have significantly increased the demand for protein-rich foods such as meat and dairy products, exerting considerable pressure on global food supplies. Single-cell proteins (SCP) have emerged as a promising alternative source, characterized by their high protein content and essential amino acids, lipids, carbohydrates, nucleic acids, inorganic salts, vitamins, and trace elements. SCP offers several advantages over the traditional animal and plant proteins. These include shorter production cycles, the use of diverse raw material sources, high energy efficiency, and minimal environmental impact. This review is primarily concerned with the microbial species employed in SCP production, utilization of non-food renewable materials as a source of feedstock, and application of rational and non-rational metabolic engineering strategies to increase SCP biomass and protein content. Moreover, the current applications, production shortages, and safety concerns associated with SCP are discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Proteins are fundamental building blocks of life and play vital roles in biological reactions, molecular transport, and signal transduction (Rezaei et al. 2016). As projected by the United Nations projections, the global population is expected to reach 10 billion by 2050, necessitating a 50%-80% increase in global food demand (Rischer et al. 2020). To maintain the current level of animal-based protein consumption, the world must produce 1.25 billion tons of meat and dairy products annually (Ritala et al., 2017; Ciani et al. 2021). As living standards improve, the consumer demand for high-quality protein and meat products has increased. However, the expansion of agricultural and livestock production has been constrained by limitations in land availability and environmental concerns. Furthermore, the conversion of plant-to-animal protein is an inefficient process, with a low nitrogen absorption rate from fertilizers and inherent nutrient loss (Berners-Lee et al., 2018). It is therefore evident that relying on traditional agricultural practices alone is unsustainable and inadequate for meeting the growing demand for proteins.
To address future protein shortages, it is necessary to explore alternative sources of protein. A joint report by the Boston Consulting Group and Blue Horizon forecasts that, by 2035, technological advancements and supportive regulations could significantly increase alternative protein consumption from 2% in 2020 to 22%, creating a substantial market valued at approximately $300 billion (Morrison 2022). Approximately one-tenth of the global consumption of meat, eggs, dairy products, and seafood is estimated to be derived from alternative proteins by 2035 (Qin et al. 2022). Four principal categories of alternative proteins are globally recognized: plant, microbial, cell-based, and insect proteins, each derived from distinct raw materials (Rischer et al. 2020; Cunha et al. 2023). These alternative proteins offer more efficient production methods, reducing environmental pollution and resource depletion and mitigating the inherent food safety risks associated with traditional meat products (Choi et al., 2012).
Single-cell proteins (SCP), also known as microbial proteins, refer to the microbial biomass used for protein supply. Protein content is a critical parameter for selecting suitable microorganisms for SCP production. The most commonly used microorganisms include yeast, fungi, algae, and bacteria (Suman. 2015). These organisms typically contain 30%–60% protein by dry cell weight (DCW) (Jach et al. 2022), which is higher than the protein content found in soy, fish, meat, and whole milk (Salazar-López et al., 2022). The concept of SCP gained prominence in the 1960s and the 1970s, a period marked by global food shortages and the pressing need to address protein deficits. During this period, the surplus in global oil production led to the use of oil-based substrates for yeast cultivation, which further drove the development of SCP (Ugalde and Castrillo., 2002). Furthermore, innovative raw materials have been investigated to achieve efficient and cost-effective production of microbial biomass. The term SCP was formally adopted in 1967 during the inaugural international SCP conference at the Massachusetts Institute of Technology (MIT) under the auspices of the United Nations Protein Advisory Group, confirming the potential role of SCP in addressing global food and feed challenges (Frazer 1967). However, concurrent increases in energy prices and global food production have resulted in a temporary decline in interest in the production of edible microbial biomass. As the global population continues to grow and the impact of climate change intensifies, there has been a resurgence of interest in SCP as a novel source of nutrients and proteins (Ciani et al. 2021; Choi et al., 2012). Advancements in synthetic biology have provided a promising avenue for the development of SCPs, as they have enabled the engineering of microbial chassis cells to enhance protein content and diversify protein types using renewable carbon sources. Furthermore, advancements in large-scale microbial fermentation technologies and post-processing methods have established a robust foundation for the future scalability of SCP production (Graham and Ledesma-Amaro. 2023). Consequently, whereas the initial focus of SCP research was on the selection of appropriate strains, current efforts are oriented toward sustainability and the capacity to address global protein supply challenges.
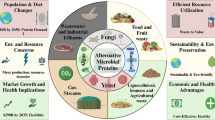
SCP are distinguished by a cytoplasmic mass that comprises carbohydrates, proteins, fats, nucleic acids, vitamins, and inorganic compounds. Compared with traditional animal and plant proteins, certain SCP achieve yields in bioreactors that can reach several kilograms per liter per hour, which is a significant improvement over traditional agriculture by several orders of magnitude (Ciani et al. 2021). SCP production requires less water and land, is ecofriendly, and does not compete with human food supplies or agricultural land. It is a sustainable protein source that benefits human health and the ecosystem (Bourdichon et al. 2012; Suman 2015; Ciani et al. 2021). SCP has been employed in a multitude of applications, not only as animal feed but also in the development of diverse food products suitable for human consumption (Liu et al. 2022; Qin et al. 2022) (Fig. 1). The replacement of 20% of global beef consumption with fungal proteins by 2050 could cause an annual reduction of over 50% in deforestation (Humpenöder et al. 2022). The use of yeast and its related fermentation products in food production has been a long-standing practice. The most commonly used yeasts for SCP production include Saccharomyces cerevisiae, Kluyveromyces marxianus, and Candida utilis. Methylotrophic yeasts, such as C. utilis, can produce biomass and proteins from methanol as a carbon source. This process has been successfully scaled up by companies such as Phillips Petroleum, with biomass concentrations reaching 130 g/L and yields exceeding 10 g/L/h (Rashad and W., 1990; Johnson 2013). Given these considerations, it seems reasonable to posit that SCP may play an important role as an alternative protein source.
SCP is an important protein supplement with considerable potential for diverse applications. This review begins by introducing the categories and characteristics of the SCP strains. It then explores non-food renewable materials suitable for SCP production before discussing the applications of rational and non-rational engineering strategies to enhance strain biomass and protein content in SCP. Finally, relevant case studies are examined. Future trends in SCP development are projected and pertinent challenges are addressed.
2 Microorganisms used for SCP
Given the accelerated growth of the global alternative protein market, microbial proteins produced through fermentation have garnered attention because of their advantages, including minimal resource consumption, high production efficiency, environmental sustainability, and comprehensive nutrition. The most commonly used microorganisms for microbial protein production include yeast, filamentous fungi, algae, and bacteria (Table 1).
2.1 Yeast
Yeast was one of the earliest microorganisms to be used by humans, with historical evidence indicating its use in the production of fermented beverages such as wine and bread (Patrick et al. 2004). Yeast demonstrates robust growth across a broad pH range (2.5–8.5) and temperature range (2–45℃). Yeast typically requires oxygen for optimal growth and its fermentation process can be efficiently scaled up for industrial applications (Jach et al. 2022). Yeast protein is devoid of any essential amino acids, lacks allergenic components, and is free of off-flavors. The protein content of yeast biomass is typically comparable to or higher than that of meat and soybeans, and exceeds that of milk protein. Moreover, the standardized protein efficiency ratio for casein indicates that SCP yeast is comparable to or more efficacious than traditional protein sources (Jach et al. 2022). Most yeast species contain relatively low quantities of nucleic acids, ranging from 5 to 8%. This is advantageous for reducing uric acid accumulation in humans. The long-standing use of yeast in food and beverage applications has contributed to its widespread use. Moreover, yeast can use a range of agricultural industrial by-products, including sugarcane bagasse and potato residues, enhancing the environmental sustainability of yeast production (Jach et al. 2022).
Saccharomyces cerevisiae (S. cerevisiae) is the most commonly used yeast species in industrial models for biotechnological research. It is generally recognized as safe (GRAS) and has applications in the food, feed, and pharmaceutical industries (Belda et al. 2019; Dunuweera et al., 2021). For example, Marmite®, a Unilever PLC product, has been consumed globally for over a century. Derived from brewing by-products, specifically S. cerevisiae extract, Marmite® contains 34% protein of dry cell weight (DCW) and various B vitamins, making it an appropriate dietary supplement for individuals with increased vitamin B requirements (Smetana et al. 2015). S. cerevisiae has a protein content between 30 and 55% DCW, high substrate tolerance, and is rich in trace minerals and bioactive compounds, rendering it an attractive feed ingredient that can partially replace products, such as fish meal and soybean meal (Jach et al. 2022).
Another commonly used yeast strain in industrial biotechnology is Yarrowia lipolytica(Y. lipolytica), which has a protein content ranging from 30 to 50% DCW and a lipid content exceeding 40%. Y. lipolytica has been designated as Generally Recognized As Safe (GRAS) by the Food and Drug Administration (FDA), and is regarded as a valuable protein source due to its high essential amino acid content. Since 2010, the dried and heat-killed biomass of Y. lipolytica cultivated in biofuel waste has been approved as a feed additive (Berge et al. 2013; Czech et al. 2018). In 2019, the European Food Safety Authority (EFSA) approved the use of dried and inactivated Y. lipolytica cells as a novel food ingredient, extending its application as a dietary supplement for individuals aged three and above (JUNCKER, 2019). In 2020, EFSA included selenium-rich Y. lipolytica biomass in the list of authorized novel foods owing to its high selenium protein content (Turck et al., 2019). Furthermore, the protein biomass of this yeast represents a valuable source of B-complex vitamins, including vitamin B12 (Jach et al. 2018).
Methylotrophic yeasts include Pichia pastoris (P. pastoris), Ogataea polymorpha, and Candida boidinii. These yeasts use methanol as a carbon source via the xylulose monophosphate (XuMP) pathway (Yongjin al., 2020). P. pastoris is a prominent industrial methanol yeast that has obtained GRAS certification from the FDA and is extensively used for the production of recombinant proteins (Ahmad et al. 2014; Zhu et al. 2018). Methanolic yeast exhibits a high protein content and is rich in essential amino acids and vitamins. This process effectively converts methanol into high-value SCP, reducing carbon emissions and promoting sustainable development. However, the intricate methanol metabolic pathway and toxicity of intracellular formaldehyde contribute to low methanol utilization efficiency in natural methanol-producing yeasts. Using a combination of elevated fermentation temperatures and adaptive evolution, Meng et al. enhanced the methanol utilization efficiency of P. pastoris to 43% (Meng et al. 2023).
2.2 Filamentous fungus
Filamentous fungi are a rich source of nutrients, comprising approximately 45% protein, 25% fiber, 13% fat, and 10% carbohydrates in dry cell weight (DCW), along with a variety of vitamins and minerals (Matassa et al. 2016). Compared with the standards set forth by the Food and Agriculture Organization of the United Nations (FAO), the amino acid profile of filamentous fungi is advantageous. These organisms typically exhibit elevated levels of threonine and lysine while displaying relatively lower methionine content (Ugalde and Castrillo 2002). One of the distinctive advantages of filamentous fungi as microbial protein producers is their capacity to flourish on a diverse array of carbon sources, including fruit by-products, spent grains from beer production, and agricultural residues (Ugalde and Castrillo 2002). This versatility not only enhances their efficiency in protein production but also offers a sustainable solution for the management of waste and by-products from food production and processing. Filamentous fungi are therefore of value not only for their nutritional output but also for their role in the promotion of circular economy principles through the conversion of waste materials into high-quality proteins. The optimal temperature range for fungal growth is 20–30 °C, with a pH range of 5.5–8.0 being conducive to growth. Oxygen is a vital component of the fungal metabolism and growth. The most commonly used filamentous fungi for protein production include Fusarium venenatum, Aspergillus oryzae, and Paecilomyces varioti. For example, fungal proteins derived from F. venenatum are employed as suitable substitutes for chicken breast tissue in chicken nuggets, whereas A. oryzae contributes to burger patty production (Gamarra-Castillo et al. 2022; Hashempour-Baltork et al. 2023). F. venenatum is extensively used for the production of fungal protein. For example, the Marlow Foods SCP product Quorn™ contains approximately 50% protein as determined by DCW (Smetana et al. 2015). This product is primarily used in the production of sausage patties and ready-to-eat burgers. Furthermore, fungal proteins are a source of B vitamins and beneficial secondary metabolites, including β-carotene and ergosterol. The growth of fungi is slower than that of bacteria, particularly for large edible fungi. Most fungi contain nucleic acids at concentrations between 7 and 10% (Li et al., 2023). However, during the cultivation process, toxins produced by some fungi represent a significant concern when using fungi as strains to produce microbial proteins (Anupama & Ravindra, 2000).
2.3 Bacteria
Bacteria have a longstanding history of SCP, with a typical protein content in the DCW range of 50%-80% (Matassa et al. 2016; Khumchai et al., 2022). The essential amino acid content of the bacterial SCP is likely to align closely with the recommendations that the FAO has set forth. Notably, the methionine content in bacterial proteins can reach as high as 3.0%, which is higher than the typical methionine levels found in algal or fungal SCP (Ritala et al., 2017). However, bacterial SCP usually has a relatively high nucleic acid content, ranging from 8 to 12% (Nasseri et al., 2011), which may necessitate additional processing steps to reduce nucleic acid levels to a level suitable for human consumption. Bacterial SCP are produced under both aerobic and anaerobic conditions. Bacteria can thrive in a broad range of temperatures, typically between 15 °C and 45 °C, and require a suitable pH environment, between 6 and 8. Bacterial SCP are distinguished by their rapid growth, with cell mass doubling times ranging from 20 min to 2 h, contingent on the specific strain and cultivation conditions. They can use a diverse range of substrates, including carbohydrates such as starch and sugars, methane, CO2, and hydrogen (Pander et al. 2020; Adeoye et al. 2021). The bacteria most commonly employed in SCP production include purple photosynthetic, hydrogen-oxidizing, and methane-used bacteria (FU et al. 2023).
Hydrogen-oxidizing bacteria (HOB), also known as hydrogen–oxygen mixed gas bacteria, use H2 and O2 as electron donors and acceptors, rapidly fix CO2, and assimilate nitrogen for protein synthesis (Matassa et al. 2016). The principal advantage of HOB in microbial protein production is its short production cycle, high protein content (up to 75% of DCW), and amino acid profile, which closely resembles that of high-quality animal protein. As a substantial component of SCP production, HOB uses H2 and O2 as raw materials generated by renewable energy, offering a potential solution for industries with considerable carbon dioxide emissions to mitigate their environmental impact (Yu, 2018). Furthermore, the HOB SCP contains poly β-hydroxybutyrate, which has been identified as a potential inhibitor of pathogenic bacteria in aquaculture and exhibits probiotic properties (Qin et al. 2022).
Purple phototrophic bacteria (PPB) are facultative anaerobic photosynthetic bacteria capable of metabolic synthesis under both light and dark conditions, using autotrophic and heterotrophic pathways (Capson-Tojo et al. 2020). SCP derived from PPB has a protein content of up to 60% DCW and is enriched with essential amino acids, carotenoids, vitamins, and polyhydroxyalkanoates (PHA), rendering it highly nutritious (Hülsen et al. 2022). The protein powder produced by PPB is predominantly used as a substitute for fish meal in aquaculture feed. During anaerobic fermentation, PPB employs the Calvin-Benson-Bassham (CBB) cycle to fix CO₂; it absorbs near-infrared light through bacteriochlorophylls (BChls) and visible light through its abundant carotenoids (Berg 2011). PPB has been employed in secondary and tertiary wastewater treatments, demonstrating its efficacy in the removal of organic matter, nitrogen, and phosphorus from a range of wastewater sources (Delamare-Deboutteville et al. 2019). However, PPB growth requires adequate illumination, and the cell yield per unit volume frequently remains relatively low.
Clostridium autoethanogenum (CA) is an anaerobic chemolithoautotrophic organism that uses CO or CO₂ as a carbon source via the Wood-Ljungdahl pathway and H₂ as a reducing agent to produce fuel ethanol, SCP, and other products (Xing-fa et al. 2020). CA has a protein content exceeding 80% of DCW, with an essential amino acid composition similar to that of fish meal, and is rich in trace elements without anti-nutritional factors (Norman et al. 2018). CA production not only yields a substantial quantity of high-quality protein but also significantly facilitates the recycling of industrial waste gases. Beijing Shougang Technology Co., Ltd., in collaboration with the Feed Research Institute of the Chinese Academy of Agricultural Sciences, has successfully industrialized the production of clean energy, including ethanol and microbial proteins, using CA fermentation technology (Liu et al. 2020). The primary raw material was CO from steel industry gas, the nitrogen source was ammonia water, and other trace elements were added to the medium. This results in an annual industrial production capacity of tens of thousands of tons of biosynthesized SCP (Wan et al. 2024). This approach effectively mitigated carbon emissions and reduced environmental pollution.
Methanotrophic bacteria use methane as both an energy and carbon source, and have been employed for SCP production for decades (Linder 2019). They demonstrated a high growth rate, with a protein content exceeding 70% DCW. Methanotrophic bacteria play a pivotal role in reducing atmospheric methane (CH4) levels and mitigating the greenhouse effect (Gao et al. 2024). Several SCP products derived from methane are commercially available. The Danish company UniBio and the American company Calysta have developed fermentation technologies that employ aerobic methanotrophic bacteria via the RuMP pathway to convert methane into SCP on a large scale. Methylococcus capsulatus is a promising source of protein (Guo et al. 2022). UniBio has developed UniProtein® through continuous fermentation, resulting in a protein content of 72% of the DCW. The product is distinguished by free-flowing reddish-brown granules with a methane conversion rate of 0.7 g DCW/g, and is devoid of toxins and heavy metals (Øverland et al. 2010). The protein produced by Calysta's M. capsulatus is marketed as FeedKind® and displays an extremely high nutritional value, with a protein content exceeding 80% DCW and an annual production capacity of 20,000 tons. M. capsulatus is renowned for its nutritional quality, balanced amino acid profile, and high digestibility and absorption rates, rendering it an exemplary source of protein. Research indicates that M. capsulatus protein can directly substitute for a portion of high-quality fishmeal in aquaculture feed and can sometimes replace white fishmeal (Zhang et al. 2022).
Besides microorganisms that have been the subject of much discussion, several bacteria that have been designated as Generally Recognized As Safe (GRAS) have been identified as having significant potential as SCP producers because of their safe profiles and efficient protein production capabilities. Bacillus subtilis is an aerobic gram-positive soil bacterium that is widely used for heterologous protein production (Earl et al., 2008; Dijl and Hecker, 2013). It is renowned for its high protein yield, which typically ranges from 40 to 60% of DCW. This strain is preferred for SCP production because of its rapid growth and brief fermentation period, typically approximately 48 h (Dijl and Hecker, 2013; Su et al., 2020). B. subtilis is a versatile organism capable of utilizing a variety of substrates, including agricultural and industrial byproducts, making it an efficient and sustainable candidate for SCP production. Corynebacterium glutamicum is another significant GRAS strain, primarily recognized for its use in amino acid production, but is also a valuable SCP producer. The protein content of this strain is approximately 50%–70% in DCW (Liu et al., 2016). Its metabolic flexibility and genetic tractability render it a promising candidate for metabolic engineering, enabling the production of SCP with tailored amino acid profiles suitable for specific nutritional requirements (Gopinath et al. 2011). Lactobacillus species, which are commonly used in the food industry, can produce SCP with a protein content ranging from 20 to 40% DCW. Moreover, these bacteria can produce beneficial metabolites, such as lactic acid and vitamins, enhancing their utility in the food and feed industries (Dempsey and Corr, 2022).
2.4 Microalgae
Microalgae are photosynthetic microorganisms that use CO₂ as a carbon source and convert it into glyceraldehyde-3-phosphate through the Calvin-Benson-Bassham (CBB) cycle to derive energy from sunlight (Ritala et al., 2017). They serve a variety of purposes as SCP, benefiting from advantages such as straightforward cultivation, utilization of solar energy, rapid growth rates, and high protein content (Yao et al. 2019). Microalgae typically contain approximately 60%-70% crude protein of DCW and have long been considered an effective means of addressing the "protein gap," making them an important source of microbial protein. Microalgae are well known for their highly digestible protein content and balanced amino acid profiles. Additionally, they are rich sources of vitamins, minerals, and polyunsaturated fatty acids (Niccolai et al. 2019). Besides their nutritional value, microalgae have been demonstrated to exhibit a range of bioactivities, including anti-inflammatory, antibacterial, antioxidant, lipid-lowering, and anticancer properties. This makes them a highly attractive subject of study in both nutrition and pharmaceutical fields (Bishop & Zubeck, 2012). Furthermore, microalgae exhibit relatively low nucleic acid levels, typically between 3 and 8% (Nasseri et al., 2011), suggesting their potential for the production of functional high-protein foods (Pereira et al. 2022). Microalgae demonstrate robust growth in a salinity range of 4%-36%, with optimal growth occurring at temperatures of 10–36 °C. They flourished in a neutral environment with a pH of 7.5–8.5. Furthermore, microalgae require light for photosynthesis, with an optimal light intensity of 8,000–15,000 lx. The most commonly used microalgal protein sources, including Chlorella, Chlamydomonas, Nannochloropsis, and Spirulina, are predominantly used in supplement formulations. The global market for autotrophic microalgae is approximately 30,000 tons, with an increasing number of applications in foods based on Spirulina platensis and Chlorella proteins. Recently, Tetraselmis chuii and Chlamydomonas reinhardtii have been approved as new food resources (Wan et al. 2024). Despite considerable interest among researchers in the biotechnological applications of autotrophic microalgae, commercial-scale production of these organisms remains limited. This is primarily due to the requirement for sufficient light and the substantial energy consumption associated with the process. Therefore, it is imperative to optimize production costs and improve production efficiency if microalgal protein production becomes a commercially viable proposition.
3 Non-food raw materials for SCP production
SCP production has traditionally involved the use of conventional substrates, including starch, molasses, and fruits. However, the cited sources do not provide sufficient evidence to support this claim. (Ugalde and Castrillo 2002; Tang et al. 2023). However, these substrates compete with human food and land resources, and are expensive. Therefore, it is necessary to develop non-food substrates, including agricultural and industrial byproducts, kitchen waste, and one-carbon substrates, for SCP production.
3.1 Byproducts of the brewing industry
The solid waste produced by the brewing industry, known as distillery yeast sludge, contains significant quantities of proteins and essential amino acids, rendering it an optimal substrate for SCP production. Valentino et al. investigated the potential of various microorganisms isolated from brewer distillery yeast sludge to produce SCP via the fermentation of dried sludge (Valentino 2015). The highest crude protein content (33.7% DCW) was observed in the S. cerevisiae strains inoculated with brewer's sludge, which was significantly higher than that in the non-inoculated strains (25.1%). Furthermore, essential amino acids such as methionine and tryptophan increased considerably, suggesting the potential of brewer distillery yeast sludge as a carbon source for SCP production. In another study, Chai et al. sterilized and disinfected the spent grain of a brewer by fermenting it with Rhizopus oligosporus. Following a three-day microbial fermentation, researchers employed microwave-assisted three-phase partitioning bio-separation to extract and separate microbial proteins, obtaining 200 g of protein from one kilogram of feedstock (Chai and Chen 2024). Moreover, the brewing industry generates considerable quantities of solid byproducts and wastewater that can be aerobically treated to facilitate SCP production. In a related study, Lee et al. (2014) investigated the enrichment of diazotrophic microbial communities in brewery wastewater and optimized the conditions for producing SCP, achieving a protein content exceeding 55% DCW. This approach addresses both the solid waste and wastewater issues prevalent in the brewing industry.
3.2 Byproducts of agro-processing industries
The processing of potato starch generates a considerable amount of waste material, which presents a significant environmental challenge. The high water content of potato pulp, typically exceeding 80%, limits its use in animal feed owing to its low nutritional value. Liu, B.N. et al. investigated a novel approach involving the mixed fermentation of potato residue and wastewater, devoid of additional nitrogen sources, to address solid and liquid pollution in the starch industry (Liu et al. 2013). The researchers identified four key microorganisms involved in this process: Curacaobacter, Pseudoalteromonas, Paenibacillus, and Bacillus. The resulting SCP products contained 46.09% protein. The potential for industrial applications was demonstrated by scaling up to a 150 m3 fermentation system. Moreover, the utilization of waste materials derived from date syrup and cheese byproducts as substrates has demonstrated considerable potential for the generation of high-quality SCP (Yadav et al. 2014; Al-Farsi et al. 2019).
Lignocellulosic biomass, derived from plants through photosynthesis, primarily comprises cellulose, hemicellulose, and lignin. Although it is employed in several industries, including papermaking, construction, textiles, and wood processing, its full potential remains largely untapped (Liu 2010). The bioconversion of lignocellulosic biomass is of paramount importance for the two principal strategies for SCP production: enzymatic hydrolysis followed by yeast fermentation or simultaneous saccharification and fermentation using cellulose and hemicellulose-degrading bacteria and yeast (Zhang et al. 2006). Yeast species have been demonstrated to be effective in utilizing lignocellulosic hydrolysates, which represent the most abundant bioresources for bioproducts. Wu et al. used lignocellulosic hydrolysates and xylose to generate SCP with Candida intermedia FL023, using peptone and yeast extract as nitrogen sources, and attained a crude protein content of 484.2 g/kg DCW (Wu et al. 2018). Silva et al. (2003) optimized the nutritional conditions for culturing Paecilomyces variotii in eucalyptus hemicellulose hydrolysate by adding ammonium sulfate and sodium phosphate, resulting in a cell concentration of 12.06 g/L (Silva et al., 2003).
These studies underscore the potential of a range of agro-processing byproducts and lignocellulosic biomass as sustainable substrates for SCP production, advancing both environmental sustainability and economic viability across diverse industrial sectors.
3.3 Byproducts of livestock and fisheries
The byproduct of fishmeal processing—water—serves as a substrate for SCP production. In a separate study, Kam et al. employed a combination of Lactobacillus acidophilus and Aspergillus niger to transform stick water into SCP. Both microorganisms demonstrated effective utilization, with L. acidophilus reaching a biomass of 7.29 g/L and A. niger reaching 5.20 g/L (Kam et al. 2012). Undigested poultry litter (UPL) and shrimp shell waste have been investigated as potential alternative raw materials for SCP production (Jalasutram et al. 2012; Wu et al. 2018). The resulting SCP products can be used as supplements in animal feed, offering advantages, such as cost reduction and mitigation of waste-related environmental issues.
3.4 One-carbon compounds
One-carbon (C1) compounds, including methanol, methane, and carbon dioxide, are ubiquitous in both natural and industrial processes; however, they are frequently under-used. The conversion of these C1 compounds into high-value SCPs not only reduces carbon emissions and mitigates the greenhouse effect but also promotes sustainable development (Ritala et al., 2017; FU et al. 2023).
Methanol, a low-cost and abundant raw material, is widely used as a starting material for SCP production. Analysis of the cell composition of Methylobacterium organophilum revealed a high crude protein content and the presence of essential amino acids, indicating its potential for use in SCP biomass. Ana et al. cultivated M. organophilum using methanol as a carbon source, attaining a maximum DCW of approximately 5 g/L with an initial methanol concentration of 12 g/L and a methanol conversion rate of 0.42 g DCW/g (Simões et al. 2022). P. pastoris, which is renowned for its natural methanol assimilation capability, was optimized for methanol-based SCP synthesis via adaptive laboratory evolution (ALE). Meng et al. successfully enhanced methanol utilization efficiency and temperature tolerance in P. pastoris, achieving high biomass production (63.37 g DCW/L), a methanol conversion rate of 0.43 g DCW/g, and a protein content of 0.506 g/g DCW in pilot-scale fed-batch cultivation at 33 °C (Meng et al. 2023a).
Methane (CH4), a potent greenhouse gas emitted from natural and anthropogenic sources (Jain, 2019), has been harnessed for SCP production using M. capsulatus in large-scale fermentation processes by companies such as UniBio and Calysta (Øverland et al. 2010; Zhang et al. 2022). Advances in urban sewage and garbage treatment technologies have enabled the use of methane-oxidizing bacteria to convert methane into SCP. Zha et al. demonstrated the successful growth of a mixed strain comprising 56.26% Methylomonas and 24.60% Methylobacter on pasteurized anaerobic digestion (AD) supernatant and biogas from sewage sludge, achieving promising dry cell weight yields (0.66 gDCW/gCH4 and 11.54 gDCW/gNH4+) with a protein content exceeding 41% in the DCW (Zha et al. 2021). Benyamin et al. employed biogas as a substrate for methane-oxidizing bacteria, resulting in high biomass yields (0.87 gDCW/gCH4) through the addition of nitrogen from pasteurized centrifuged digestate or electrochemically extracted ammonium from digestate (Khoshnevisan et al., 2019).
Carbon dioxide (CO₂) is a primary greenhouse gas and a key target for energy conservation and emission reduction. Autotrophic bacteria capable of CO₂ fixation are particularly well-suited for the sustainable production of SCP. Photosynthetic cyanobacteria, which use light as an energy source, demonstrate considerable potential for carbon dioxide fixation in SCP (Hülsen et al. 2022). They offer notable advantages over plant-based approaches, including rapid growth and versatile application potentials.
4 Engineering strategies for enhancing SCP production
Microbial biomass yield and protein content are of paramount importance for influencing SCP production. An increase in microbial biomass yield has been demonstrated to enhance SCP productivity, whereas an improvement in protein content has been shown to elevate the nutritional quality of SCP. Consequently, considerable effort has been invested in the pursuit of increasing microbial biomass and protein content to enhance SCP production.
4.1 Strategies to enhance microbial biomass yield
Microbial growth is a critical factor influencing the production of SCP, as it directly affects the biomass yield and protein content. Efficient microbial growth is a prerequisite for a high biomass output, which is a necessary condition for cost-effective SCP production. Several factors influence microbial growth, including nutrient availability, presence of toxic compounds, substrate tolerance, cultivation conditions, and metabolic constraints (Reihani and Khosravi-Darani, 2019). Various strategies are being used to address these challenges and enhance microbial growth to improve the protein content in SCP production (Balagurunathan et al., 2022) (Fig. 2).
4.1.1 Optimization of culture conditions and fermentation processes
The production of microbial biomass has been enhanced by optimizing culture conditions (Tesfaw and Assefa, 2014; Hezarjaribi et al., 2016; Kerckhof et al. 2021; Vethathirri et al. 2021; Hu et al. 2022) and improving fermentation processes to achieve high-protein yields (Reihani and Khosravi-Darani, 2019). For example, optimization of the culture medium for S. cerevisiae using the Taguchi method, which adjusts the concentrations of ammonium sulfate, iron sulfate, glycine, and glucose based on signal-to-noise ratio analysis, has been demonstrated to significantly improve SCP production. Before optimization, the yield was 7.69–7.91 log colony-forming unit (CFU) mL−1 with a protein content of 34% DCW. Following optimization, the yield increased to 8.84 log and the protein content reached 44.6% (Hezarjaribi et al. 2016). Research on the optimization of microbial biomass production through co-culture methods has demonstrated notable improvements in both the biomass yield and protein content of SCP. For instance, the co-culturing of white-rot fungi (Ganoderma lucidum) with yeast (Candida utilis) resulted in a notable increase in protein content from 8.75% to 16.23%. Similarly, the combination of Trichoderma reesei and A. niger resulted in a 91% increase in biomass yield, from 11.2 g/L to 21.4 g/L (Tesfaw and Assefa, 2014). Subsequent research demonstrated that the co-cultivation of Xanthobacter variabilis and Shinella sp. NM-101 resulted in an 11.4-fold increase in bacterial growth, along with a 24% increase in protein content, 28% increase in essential amino acids, and 26% increase in overall biomass yield compared to monocultures (Hu et al. 2022). The co-cultivation of distinct microorganisms facilitates the collaborative utilization of carbon sources, breakdown of complex substrates, and removal of deleterious metabolic byproducts.
The yield and quality of SCP production are significantly influenced by fermentation parameters, including carbon and nitrogen sources, inoculum size, age, aeration, temperature, and pH. These parameters play a crucial role in optimizing microbial growth and metabolic activity, which affects the yield and quality of SCP production. It has been demonstrated that adjusting these parameters is crucial for maximizing biomass yield, enhancing productivity, reducing production costs, and improving SCP quality (Reihani and Khosravi-Darani, 2019).
4.1.2 Rational metabolic engineering approaches
Metabolic engineering, encompassing both rational and non-rational engineering methodologies, is of paramount importance for enhancing SCP production. Modifying microbial metabolic pathways can enhance substrate utilization efficiency and increase nutrient conversion efficiency, improving overall biomass and protein yield (Jang et al., 2012).
Rational engineering approaches involve modification of transcription factors, transporters, and metabolic pathways to achieve the desired phenotype (Hara et al. 2017; Gupta et al. 2020). For example, optimization of transporter proteins in yeast, such as glucose transporters (Hxt1p and Hxt7p), sucrose transporters (Agt1p), cellodextrin transporters (CDT-1), xylose transporters (Hxt7p), and galactose transporters (Gal2p), has been demonstrated to significantly enhance substrate utilization efficiency (Hara et al., 2017). The overexpression of Hxt7p resulted in enhanced glucose consumption and lactate production, whereas the introduction of CDT-1 from Neurospora crassa led to a 30%–50% increase in sugar utilization rates and a 20%-40% increase in ethanol yield (Hara et al., 2017). These transporter engineering strategies optimize the bioproduction processes in yeast, demonstrating their potential for industrial applications in SCP production. Furthermore, global transcription machinery engineering (gTME) has yielded notable advancements in microbial biomass and product yields (Tan et al. 2016; El-Rotail et al., 2017). For instance, the implementation of gTME has demonstrated a remarkable enhancement in the biomass and ethanol production capacity of S. cerevisiae. The mutants exhibited a notable enhancement in DCW, reaching 2.01 g/L in 3% ethanol, compared to the control strain, which demonstrated a DCW of 1.39 g/L. Moreover, the most efficient mutant exhibited an ethanol yield of 15.72 g/L, representing a 60.3% increase over that of the control strain (9.814 g/L) (El-Rotail et al., 2017). Similarly, the Zymomonas mobilis mutant strain ZM4-mrpoD4 exhibited enhanced growth with a maximum cell density (OD600) of approximately 1.8 compared to the control's 1.2 under 9% ethanol stress, and increased ethanol production from 6.6–7.7 g/L to 13.0–14.1 g/L (Tan et al. 2016). Metabolically engineered S. cerevisiae strains were cultivated in ethylene glycol, isopropanol, and propionic acid to produce glucose. The protein content of these genetically modified strains reached approximately 50% of their dry cell weight (Tang et al. 2023).
The optimization of carbon fixation and nitrogen assimilation pathways is a critical strategy for enhancing microbial biomass. For example, in Methylophilus methylotrophus, overexpression of genes involved in carbon fixation, such as rbcL (encoding the RuBisCO large subunit) and prkA (encoding phosphoribulokinase), can enhance the flow of carbon into biomass production. Concurrently, overexpression of genes involved in nitrogen assimilation, such as gdhA (encoding glutamate dehydrogenase) and glnA (encoding glutamine synthetase), has been shown to enhance the availability of amino acids essential for protein synthesis. These modifications have increased protein content by up to 25% and biomass yield by approximately 20%, contributing to a higher SCP output (Windass et al., 1980).
Another crucial strategy is to minimize byproduct formation and carbon loss. In F. venenatum, research has concentrated on the inhibition of the ethanol synthesis and gluconeogenesis pathways, which are the primary routes for byproduct formation. Deletion of the genes FvPDC6 (pyruvate decarboxylase), which is involved in ethanol production, and FvPCK (phosphoenolpyruvate carboxykinase), which is involved in gluconeogenesis, whereas the overexpression of FvPYC (pyruvate carboxylase) to enhance carbon fixation and optimize the fermentation medium with the addition of ZnSO, resulted in an improvement in the carbon conversion ratio by up to 73% and an increase in protein content by 25%, reaching a protein concentration of 61. This equates to 9% DCW. Furthermore, the synthesis rate increased by 57% and CO₂ emissions were reduced by 39%, thus rendering the process both efficient and environmentally friendly (Tong et al., 2024).
4.1.3 Non-rational metabolic engineering approaches
Besides these rational engineering techniques, non-rational engineering approaches, including ALE, chemical mutagenesis, and genome engineering, have been employed for biomass accumulation by improving cell growth and substrate tolerance (Pham et al., 2017; Bennett et al. 2021; Meng et al. 2023). For example, a pH-sensing riboswitch system with a self-regulating genetic program enables bacteria to evolve autonomously, resulting in strains with enhanced survival rates ranging from 20 to 40% in the presence of various organic acids (fumaric acid, citric acid, L-malic acid, itaconic acid, lactic acid, and succinic acid) compared to the control strain T10 (Pham et al. 2017). The combination of ALE with chemical mutagenesis using N-methyl-N'-nitro-N-nitrosoguanidine (NTG) has resulted in notable improvements in the tolerance of mutant strains to methanol. The resulting strains, which exhibited mutations in the 30S ribosomal subunit proteins, particularly in rpsQ and rpsL, demonstrated growth rate improvements of 38% with 2 M methanol, 57% with 3 M methanol, and 10% with 1 M methanol. In addition, they exhibited a two- to three-fold improvement in methanol utilization at 60 mM (Bennett et al. 2021). Moreover, recent developments in methanol-used yeast P. pastoris have demonstrated considerable potential for sustainable SCP production. By employing the ALE approach and implementing genetic modifications, the engineered strain HTX-33 demonstrated a biomass yield of 63.37 g DCW/L, a methanol conversion rate of 0.43 g DCW/g, and a protein content of 0.506 g/g DCW at 33 °C. Compared with the parent strain X-33, which exhibited a protein content of 0.364 g/g DCW and a total nitrogen content of 0.058 g/g DCW at 30 °C, these findings indicate a 39% increase in protein content and a 25.86% increase in total nitrogen (Gao et al., 2023; Meng et al. 2023). These improvements highlight the potential of genetically modified P. pastoris strains for efficient SCP production.
4.2 Strategies to enhance protein content of microbial biomass
Besides increasing microbial biomass to enhance protein yield, current metabolic engineering strategies focus on directly boosting protein production. These strategies encompass an array of approaches, including enhancement of mRNA stability, reduction of protein degradation, promotion of protein synthesis, and facilitation of protein folding and refolding (Balagurunathan et al., 2022; see Fig. 2).
4.2.1 Increasing mRNA stability
The reduction of mRNA degradation can be achieved through genetic engineering, which allows for the development of RNase-deficient microbes, optimization of mRNA sequence elements, and regulation of RNA-binding proteins. These strategies collectively enhance mRNA stability and protein synthesis, increasing the SCP production in microbial systems. The degradation of mRNA is influenced by several factors, including the sequence of mRNA, activity of RNA-binding proteins, and presence of ribonucleases (Hui et al. 2014; Roux et al., 2022). To reduce total mRNA degradation, a direct method involves the creation of ribonuclease (RNase)-deficient microbes through genetic engineering techniques, such as CRISPR-Cas9-mediated knockout or inhibition of key RNase gene expression, such as the RNase E gene. These modifications have been demonstrated to significantly extend mRNA half-life by decreasing mRNA decay rates, improving mRNA stability, and increasing overall mRNA availability for translation, consequently enhancing protein synthesis (Roux et al., 2022). For example, E. coli strains, such as BL21 Star™(DE3), with the rne-131 allele for truncated RNase E have been demonstrated to be effective in stabilizing mRNA and enhancing heterologous protein synthesis. The use of RNase-deficient strains has been shown to increase protein yields. This is evidenced by strategies such as MazF induction, which has been shown to boost protein levels by a factor of 3–11 (Venturelli et al. 2017; Wu et al. 2020).
It is crucial to optimize the sequence elements of mRNA to ensure stability. Modifications to the 5' untranslated region (UTR) through the removal of destabilizing sequences and addition of stabilizing elements protect mRNA from degradation, enhance ribosome binding, and improve heterologous protein expression. For example, incorporating hairpin structures derived from ompA resulted in an increase in the mRNA half-life from three minutes to 5–6 min. Further addition of an AU-rich element extends this to 12.9 min (Viegas et al. 2018). The introduction of synthetic REP sequences at the 3′-terminus of prokaryotic cells prevents 3′-5′ exonuclease degradation, significantly improving heterologous protein expression. For example, the activity of cyclodextrin glucosyltransferase (CGTase) increased from 210.6 to 291.5 U/mL, whereas glucosamine-6-phosphate N-acetyltransferase 1 (GNA1) activity rose from 524.8 to 890.7 U/mg (Deng et al. 2019). Furthermore, differential mRNA decay within bacterial operons, such as the tatABC operon, highlights the influence of stabilizing RNA elements on protein synthesis. Stabilizing tatA mRNA over tatBC has been demonstrated to results in 25-fold higher TatA protein expression (Dar and Sorek, 2018). Another effective strategy involves the regulation of RNA-binding proteins. Proteins that stabilize mRNA, such as HuR, bind to mRNA and protect it from degradation. Conversely, destabilizing proteins, such as tristetraprolin, reduce mRNA stability by increasing mRNA decay rates. Therefore, mRNA stability can be enhanced by reducing the expression or activity of destabilizing proteins (Mukherjee et al., 2011).
4.2.2 Promoting protein synthesis
The construction of an efficient, high-expression system through codon optimization, increase in gene copy number, and transcriptional regulation is of paramount importance for achieving high-protein yields. Codon optimization enhances translation efficiency by adjusting codon usage frequencies, whereas increasing the gene copy number involves the integration of multiple copies of the target gene into plasmids or the genome. For example, codon optimization has been employed to replace rare codons in the native T. emersonii α-amylase gene with codons preferred by S. cerevisiae, resulting in a 1.6-fold increase in protein yield. Similarly, codon optimization of the T. emersonii glucoamylase gene resulted in a 3.3-fold increase in the protein yield (Cripwell et al., 2019). The use of high-copy number 2μ-based YEp plasmids, which can maintain 10–40 copies, has demonstrated a significant enhancement in protein expression, with yields of up to 5 g/L for human albumin and albumin fusion proteins (Chen et al.). 2012; Da Silva and Srikrishnan 2012). Transcriptional regulation uses strong promoters, inducible promoters, synthetic promoters, and optimized terminator sequences to enhance transcriptional and translational efficiency. Strong constitutive promoters in S. cerevisiae include pTDH3, pPGK1, pADH1, and pTPI1 as well as inducible promoters such as pGAL1, pGAL7, pGAL10, pPHO5, and pMET25 (Zhao et al., 2024). The GAP1 promoter system, with a medium-switching strategy, yielded approximately 40 g/L DCW and 1 mg/L Gap1 protein, increasing target protein production by approximately five-fold compared with the PMA1 promoter system (Debailleul et al., 2013). With the advent of synthetic biology, high-activity synthetic promoters with broad activity ranges have been developed (LaFleur et al., 2022). Synthetic promoters have been used to enhance the production of green fluorescent proteins and β-glucosidase in S. cerevisiae (Deng et al., 2021; Wu et al. 2023). The use of the inducible MET25 promoter in methionine-free media resulted in a significant enhancement in the expression of target proteins, with an 18- to 70-fold increase observed during the 16 to 24-h period compared to conditions with methionine (Solow et al., 2005). Moreover, the implementation of these strategies integratedly has been demonstrated to significantly enhance heterologous protein production in S. cerevisiae, providing effective solutions for industrial-scale applications (Zhao et al., 2024).
4.2.3 Enhancing protein folding and refolding
Enhancement of protein folding and refolding represents an effective strategy for increasing protein production, whereby reduction of protein degradation due to misfolding is achieved. Proteasomes or autophagy pathways are responsible for the degradation of properly folded proteins, increasing the net protein yield (Díaz-Villanueva et al., 2015). Molecular chaperones such as Hsp70 and Hsp90 facilitate the correct folding of proteins by binding to unfolded proteins and preventing their aggregation (Taldone et al. 2014). Overexpression of the chaperone protein BiP or disulfide isomerase Pdi1p has been demonstrated to enhance the production of β-glucosidase, endoglucanase, and α-amylase (Tang et al., 2015). Significant improvements in protein production efficiency have also been reported (Kim et al. 2020). The co-expression of chaperones and foldases has demonstrated a significant enhancement in protein yields through the optimization of refolding conditions. For example, the helper plasmid pTUM4, which overexpresses four periplasmic folding catalysts (DsbA, DsbC, FkpA, and SurA), has been demonstrated to significantly enhance the folding efficiency of recombinant proteins, increasing the yield of soluble proteins. Specifically, the use of pTUM4 in the production of human plasma retinol-binding protein (RBP) resulted in a four-fold increase in the yield of correctly folded RBP, reaching 0.6 mg/L/OD (Schlapschy et al. 2006). Furthermore, the expression of genes involved in the unfolded protein response (UPR) pathways, including HAC1, ATF6, PERK, and IRE1, has been shown to enhance folding capacity and mitigate folding stress induced by misfolded proteins (Smith et al. 2011; Chambers & Marciniak, 2014). Overexpression of HAC1 enhances the production of a range of recombinant proteins, including xylanase and α-amylase (Valkonen et al., 2003; Bao et al., 2020). Moreover, chemical chaperones such as 4-phenylbutyrate (4-PBA) and tauroursodeoxycholic acid have been shown to stabilize protein folding and reduce aggregation, representing a potential strategy for improving protein content (Lindquist and Kelly, 2011).
4.2.4 Reducing protein degradation
The most direct method for enhancing the protein yield is to inhibit protein degradation via protease knockout. The study deleted eight protease genes (nprE, aprE, epr, bpr, mpr, nprB, vpr, and wprA) were identified in Bacillus subtilis, resulting in second-generation protease-deficient strains. The engineered strain PD8, which was created by eliminating eight detrimental proteases, demonstrated notable improvements in methyl parathion hydrolase (MPH) production (79.9 U/mL in PD8 vs. 32.5 U/mL in the WB800-control strain) and chlorothalonil hydrolytic dehalogenase (Chd) production (13.4 U/L in PD8 vs. 7.4 U/L in WB800) (Zhao et al., 2019). Another study investigated the potential of protease-deficient S. cerevisiae strains to produce human-compatible glycoproteins. Disruption of the PEP4 and PRB1 genes in the resulting protease-deficient strain YAB101-4 resulted in a tenfold increase in human interferon-β (hIFN-β) secretion compared to the unmodified strains (Tomimoto et al., 2013). These studies demonstrate considerable potential for improving foreign protein production through the use of protease knockout strategies.
The ubiquitin–proteasome system and autophagy pathways play crucial roles in the degradation of misfolded proteins, maintaining proteostasis (Varshavsky, 2017). Spatial compartmentalization mechanisms such as juxtanuclear quality control (JUNQ) and insoluble protein deposition (IPOD) sequester misfolded proteins, facilitating proper folding or degradation (Kaganovich et al. 2008; Wolff et al. 2014). Despite the limited attention these mechanisms have received in the context of protein production, their optimization has the potential to enhance the protein content in SCP production.
5 Industrial application of SCP
The high-protein content and nutritional richness of SCP, coupled with its sustainable production capabilities, make it a suitable candidate for a plethora of industrial applications, including food, animal feed, and feedstock for microbial fermentation (Onyeaka et al., 2022; Fact.MR, 2023).
5.1 Food applications
SCP is a widely used ingredient in the food industry, largely because of its high-protein content and nutritional value. This provides a sustainable alternative to traditional protein sources. One notable application of SCP is the production of high-protein foods and beverages, which are especially advantageous for individuals who require a high-protein diet, such as vegetarians and athletes. SCP has been incorporated into a variety of food products, including protein bars, drinks, and powders, and is a rich source of essential amino acids (Onyeaka et al. 2022; Fact.MR 2023). Solar Foods produce Solein®, a protein-rich product with a content of 65%-70% DCW, derived from Xanthobacter tagetidis using electricity and carbon dioxide. Solein is used in a variety of food products, including protein shakes, bread, and pasta (Solar Foods n.d.). Furthermore, SCP serves as a principal component in the production of meat substitutes, imitating the texture and flavor of meat while reducing its environmental impact. Examples of such products include plant-based burgers, vegetarian sausages, and chicken substitutes (Bratosin et al. 2021). Nature's Fynd offers Fy Protein™, a high-protein ingredient (> 50%) derived from fungal microbes, which is used in the production of products such as meatless breakfast patties and dairy-free cream cheese (Nature's Fynd n.d.). Similarly, Quorn employs F. venenatum fermentation technology to produce mycoproteins, which are incorporated into a diverse array of vegetarian and plant-based foods, including Quorn Meatless Grounds and Quorn Meatless Nuggets (Quorn n.d.). Approximately 50% of DCW comprises mycoproteins. Furthermore, SCP is employed in the production of fermented foods, such as yogurt and tofu, to enhance their nutritional value and flavor while extending their shelf life. For example, Perfect Day employs microbial fermentation technology to create Perfect Day Dairy Protein™, a protein substitute incorporated into products such as Brave Robot ice cream and Modern Kitchen cream cheese. This enhances the nutritional value and flavor of these products (Perfect Day n.d.).
SCP is a valuable source of essential amino acids, vitamins, and other nutrients, making it an optimal ingredient for the production of dietary supplements, functional foods, and bioactive compounds for therapeutic treatments. These products benefit from the rich protein content and essential nutrients of SCP, which are crucial for maintaining a balanced diet and preventing nutrient deficiencies (Onyeaka et al., 2022; Fact.MR, 2023). The fermentation process yielded Blue Origins®, an SCP product derived from microalgae. This product is rich in omega-3 fatty acids and is used in dietary supplements and functional foods to promote cardiovascular health (Fermentalg n.d.). Solazyme developed AlgaVia®, an SCP that is rich in protein and essential nutrients. It is used in a range of nutraceutical products to enhance their health benefits (TerraVia n.d.). Therefore, SCP is a promising alternative protein source for the food industry.
5.2 Feed applications
SCP has been extensively used as a valuable alternative to traditional protein sources such as soybean meal and fishmeal in animal feed applications, including poultry, aquaculture, and livestock feed. This is because of its high-protein content, balanced amino acid profile, and sustainable production methods. In poultry feed, incorporation of SCP has been demonstrated to enhance growth rates and improve overall health. This is attributed to its contribution to improved feed efficiency and the production of higher-quality meat (Onyeaka et al., 2022; Fact.MR, 2023). For example, Unibio employs M. capsulatus to manufacture UniProtein®, an SCP with a protein content of 70% of the DCW. This product is derived from methane and is used in poultry feed, offering a sustainable and high-quality protein alternative (Unibio n.d.). SCP provides the essential nutrients required to support the growth and health of fish and shrimp, conferring significant benefits to aquaculture through the use of SCP-based aquaculture feed. FeedKind® by Calysta is produced through the fermentation of M. capsulatus using natural gas, resulting in an SCP with a protein content of 80% of the DCW. It is frequently used as an aquaculture feedstuff and offers notable sustainability benefits (Calysta n.d.). Furthermore, SCP is used in livestock feed to enhance protein intake and facilitate the growth of cattle, swine, and other farm animals. The utilization of SCP in livestock feed mitigates the environmental impact of animal husbandry, while ensuring a consistent and reliable supply of high-quality protein. KnipBio produces KnipBio Meal (KBM), an SCP derived from M. extorquens, which is used in livestock feed to enhance feed conversion rates and promote healthy animal growth (KnipBio n.d.). These applications in animal feed illustrate the potential for SCP to provide environmental and economic benefits to the livestock, poultry, and aquaculture industries.
5.3 Feedstock applications
SCP is a rich source of protein, making it a frequently used feedstock for microbial fermentation for the production of a range of valuable products, including bioplastics and biofuels, which have high protein and carbohydrate contents. For example, NovoNutrients produce NovoMeal, an SCP derived from industrial carbon dioxide emissions. NovoMeal™ is an exemplary fermentation feedstock for the production of sustainable protein ingredients that are highly digestible and exhibit superior amino acid profiles compared with numerous conventional protein sources. These proteins are employed in animal feed, plant-based foods, and aquaculture feed, underscoring the versatility and environmental benefits of utilizing SCP in fermentation processes (NovoNutrients n.d.). Mango Materials is another company that employs methane-derived SCP to manufacture polyhydroxyalkanoates (PHA), a biopolymer used in the production of biodegradable plastic products, including packaging, agricultural films, and consumer goods (Mango Materials n.d.). Furthermore, SCP has been successfully employed for biofuel production. For example, LanzaTech employs carbon-rich industrial emissions to cultivate C. autoethanogenum, yielding SCP with 80% DCW protein content. Subsequently, SCP is converted into bioethanol and other valuable products via fermentation (Ma et al., 2022). This process not only results in a reduction in greenhouse gas emissions but also produces a renewable biofuel that can be used as an alternative to traditional gasoline. Algenol produces biodiesel from SCP derived from the microalga Euglena gracilis, which has a protein content exceeding 70% DCW. The process exploits the high lipid content of microalgae to create a sustainable alternative fuel source while simultaneously producing a nutrient-rich protein byproduct (Algenol n.d.). SCP proves to be the ideal feedstock for bioplastics, biofuels, and efficient production of valuable chemicals.
6 Shortages and safety concerns of SCP production
SCP production offers substantial environmental and nutritional advantages by transforming agricultural and industrial waste into high-protein biomass, thus advancing sustainability and mitigating the environmental consequences associated with conventional protein sources (Ritala et al., 2017). Microorganisms, including algae, fungi, bacteria, and yeast, facilitate the efficient production of SCP with rapid growth rates and high yields of proteins enriched with essential amino acids, vitamins, and minerals (Aidoo et al. 2023). Nevertheless, the production and consumption of SCP have certain shortcomings and safety concerns (Flight et al. 2024).
A noteworthy concern is the elevated nucleic acid content of SCP, particularly in bacteria and yeasts. This can cause uric acid accumulation in humans, which may lead to adverse health outcomes such as gout and kidney stones (Yang et al. 1979; Ravindra, 2000). To mitigate these risks, processes such as thermal treatment, chemical extraction, and enzymatic degradation can be used to reduce nucleic acid levels. Heat treatment can denature nucleic acids, rendering them less bioavailable, thus reducing the risk of uric acid accumulation (Onyeaka et al., 2022).
Certain microorganisms involved in SCP production can produce harmful endotoxins and mycotoxins, posing a toxic risk. To address this issue, the selection of non-toxigenic strains and the application of genetic modifications can help to minimize the presence of these toxins (Ravindra, 2000; Hadi and Brightwell 2021). It is of paramount importance to implement robust regulatory frameworks and quality control measures to guarantee that SCP products comply with requisite safety standards. For example, the European Union regulations on novel foods and the Food and Drug Administration (FDA) guidelines for food safety set rigorous standards for the production and quality control of SCP products (Council, 2015).
The digestibility and bioavailability of SCPs are of great consequence to human health, necessitating a comprehensive evaluation through clinical studies to ensure their suitability for consumption. For example, research on the digestibility of algal proteins has demonstrated that processing techniques can enhance bioavailability and nutritional quality (Junaid et al. 2020; Bratosin et al. 2021).
It is imperative to address consumer neophobia and potential allergic reactions to foster the acceptance of SCP as an alternative protein source. Consumers’ acceptance of SCP products is a crucial factor in determining their success (Hung et al. 2023). The public's perception of SCPs can be shaped by several factors, including awareness of their environmental benefits, trust in their safety and nutritional value, and cultural attitudes toward novel food sources (Siddiqui et al. 2022). Effective communication strategies, transparency in production processes, and educational campaigns are essential to foster consumer trust and acceptance. Implementing public education campaigns and transparent communication on the safety and benefits of SCP can facilitate the establishment of consumer trust. Moreover, allergenicity testing is crucial for identifying and mitigating potential allergic reactions (Siddiqui et al. 2022; Hung et al. 2023).
In conclusion, whereas SCP production offers a sustainable solution with substantial benefits, it is imperative to address safety and risk factors through optimized processes and rigorous regulatory oversight. This information is crucial for establishing SCP as a safe and viable alternative protein source for the global food system. By employing advanced biotechnological methods, adhering to strict regulatory standards, and engaging in comprehensive safety evaluations, SCP can contribute to sustainable protein production and food security.
Availability of data and materials
No datasets were generated or analysed during the current study.
References
Adeoye A, Akegbejo-Samsons Y, Fawole FJ, Olatunji PO, Muller N, Wan AHL, Davies SJ. From waste to feed: Dietary utilisation of bacterial protein from fermentation of agricultural wastes in African catfish (Clarias gariepinus) production and health. Aquaculture. 2021;531:1–9.
Ahmad M, Hirz M, Pichler H, Schwab H. Protein expression in Pichia pastoris: recent achievements and perspectives for heterologous protein production. Appl Microbiol Biotechnol. 2014;98:5301–17.
Aidoo R, Kwofie EM, Adewale P, Lam E, Ngadi M. Overview of single cell protein: production pathway, sustainability outlook, and digital twin potentials. Trends Food Sci Technol. 2023;138:577–98.
Al-Farsi M, Bakir AA, Marzouqi HA, Thomas R. Production of single cell protein from date waste. In by-Products Palm Trees Their Appl. 2019;2:303–12.
Algenol: Advanced Algal Biofuels and Biochemicals. n.d. from https://www.algenol.com/advanced-algal-biofuels-biochemicals.
Anupama RP. Value-added food: single cell protein. Biotechnol Adv. 2000;18:459–79.
Balagurunathan B, Ling H, Choi WJ, Chang MW. Potential use of microbial engineering in single-cell protein production. Curr Opin Biotechnol. 2022;76:102740.
Bao C, Li J, Chen H, Sun Y, Wang G, Chen G, Zhang S. Expression and function of an Hac1-regulated multi-copy xylanase gene in Saccharomyces cerevisiae. Sci Rep. 2020;10:11686.
Belda I, Ruiz J, Santos A, Van Wyk N, Pretorius IS. Saccharomyces cerevisiae. Trends Genet. 2019;35:956–7.
Bennett RK, Gregory GJ, Gonzalez JE, Har JRG, Antoniewicz MR, Papoutsakis ET. Improving the methanol tolerance of an Escherichia coli methylotroph via adaptive laboratory evolution enhances synthetic methanol utilization. Front Microbiol. 2021;12:638426.
Berg IA. Ecological aspects of the distribution of different autotrophic CO2 fixation pathways. Appl Environ Microbiol. 2011;77:1925–36.
Berge GM, Hatlen B, Odom JM, Ruyter B. Physical treatment of high EPAYarrowia lipolyticabiomass increases the availability of n-3 highly unsaturated fatty acids when fed to Atlantic salmon. Aquac Nutr. 2013;19:110–21.
Berners-Lee, Kennelly C, Watson R, Hewitt CN. Current global food production is sufficient to meethuman nutritional needs in 2050 provided there isradical societal adaptation. Elem-Sci Anthrop. 2018;6:237–46.
Bishop WM, Zubeck HM. Evaluation of microalgae for use as nutraceuticals and nutritional supplements. J Nutr Food Sci. 2012;02:147–59.
Bourdichon F, Casaregola S, Farrokh C, Frisvad JC, Gerds ML, Hammes WP, Harnett J, Huys G, Laulund S, Ouwehand A, et al. Food fermentations: microorganisms with technological beneficial use. Int J Food Microbiol. 2012;154:87–97.
Bratosin BC, Darjan S, Vodnar DC. Single cell protein: a potential substitute in human and animal nutrition. Sustainability. 2021;13:1–25.
Calysta. FeedKind®: Sustainable protein for aquaculture. n.d. from: https://www.vbdata.cn/intelDetail/114249.
Capson-Tojo G, Batstone DJ, Grassino M, Vlaeminck SE, Puyol D, Verstraete W, Kleerebezem R, Oehmen A, Ghimire A, Pikaar I, et al. Purple phototrophic bacteria for resource recovery: challenges and opportunities. Biotechnol Adv. 2020;43: 107567.
Chai KF, Chen WN. Recovery of antioxidative protein hydrolysates with functional properties from fermented brewer’s spent grain via microwave-assisted three phase partitioning. Innov Food Sci Emerg Technol. 2024;91:103551–9.
Chambers JE, Marciniak SJ. Cellular mechanisms of endoplasmic reticulum stress signaling in health and disease. Protein misfolding and ER stress. Am J Physiol-Cell Physiol. 2014;307:657–70.
Chen Y, Partow S, Scalcinati G, Siewers V, Nielsen J. Enhancing the copy number of episomal plasmids in Saccharomyces cerevisiae for improved protein production. FEMS Yeast Res. 2012;12:598–607.
Choi KR, Jung SY, Lee SY. From sustainable feedstocks to microbial foods. Nat Microbiol. 2012;9:1167–75.
Ciani M, Lippolis A, Fava F, Rodolfi L, Niccolai A, Tredici MR. Microbes: Food for the Future. Foods. 2021;10:971.
Council EPa. Regulation (EU) 2015/2283 of the European Parliament and of the Council of 25 November 2015 on novel foods. Official J Eur Union. 2015. http://data.europa.eu/eli/reg/2015/2283/2021-03-27.
Cripwell RA, Rose SH, Viljoen-Bloom M, van Zyl WH. Improved raw starch amylase production by Saccharomyces cerevisiae using codon optimisation strategies. FEMS Yeast Res. 2019;19:127.
Cunha N, Andrade V, Ruivo P, Pinto P. Effects of insect consumption on human health: a systematic review of human studies. Nutrients. 2023;15:1–23.
Czech A, Smolczyk A, Ognik K, Wlazło Ł, Nowakowicz-Dębek B, Kiesz M. Effect of dietary supplementation with Yarrowia lipolytica or Saccharomyces cerevisiae yeast and probiotic additives on haematological parameters and the gut microbiota in piglets. Res Vet Sci. 2018;119:221–7.
Dar D, Sorek R. Extensive reshaping of bacterial operons by programmed mRNA decay. PLoS Genet. 2018;14:e1007354.
Da Silva NA, Srikrishnan S. Introduction and expression of genes for metabolic engineering applications in Saccharomyces cerevisiae. FEMS Yeast Res. 2012;12:197–214.
Debailleul F, Trubbia C, Frederickx N, Lauwers E, Merhi A, Ruysschaert J-M, André B, Govaerts C. Nitrogen catabolite repressible GAP1 promoter, a new tool for efficient recombinant protein production in S. cerevisiae. Microbial Cell Fact. 2013;12:1–10.
Delamare-Deboutteville J, Batstone DJ, Kawasaki M, Stegman S, Salini M, Tabrett S, Smullen R, Barnes AC, Hülsen T. Mixed culture purple phototrophic bacteria is an effective fishmeal replacement in aquaculture. Water Res x. 2019;4:100031–42.
Dempsey E, Corr SC. Lactobacillus spp. for gastrointestinal health: current and future perspectives. Front Immunol. 2022;13: 840245.
Deng C, Lv X, Li J, Liu Y, Du G, Amaro RL, Liu L. Synthetic repetitive extragenic palindromic (REP) sequence as an efficient mRNA stabilizer for protein production and metabolic engineering in prokaryotic cells. Biotechnol Bioeng. 2019;116:5–18.
Deng J, Wu Y, Zheng Z, Chen N, Luo X, Tang H, Keasling JD. A synthetic promoter system for well-controlled protein expression with different carbon sources in Saccharomyces cerevisiae. Microbial Cell Fact. 2021;20:1–10.
Díaz-Villanueva JF, Díaz-Molina R, García-González V. Protein folding and mechanisms of proteostasis. Int J Mol Sci. 2015;16:17193–230.
Dijl JMv, Hecker M. Bacillus subtilis: from soil bacterium to supersecreting cell factory. Microb Cell Fact. 2013;12:1–6.
Dunuweera AN, Nikagolla DN, Ranganathan K, Ahmad A. Fruit Waste Substrates to Produce Single-Cell Proteins as Alternative Human Food Supplements and Animal Feeds Using Baker’s Yeast (Saccharomyces cerevisiae). J Food Qual. 2015;2021:1–6.
Earl AM, Losick R, Kolter R. Ecology and genomics of Bacillus subtilis. Trends Microbiol. 2008;16:269–75.
El-Rotail AA, Zhang L, Li Y, Liu SP, Shi GY. A novel constructed SPT15 mutagenesis library of Saccharomyces cerevisiae by using gTME technique for enhanced ethanol production. AMB Express. 2017;7:1–12.
Fact. MR. Single cell protein market. 2023. https://www.cellofuel.com/.
Fermentalg. Innovative solutions in algal biotechnology. n.d. https://www.fermentalg.com/innovative-solutions-in-algal-biotechnology.
Flight MH, Tait J, Chronopoulos T, Betancor M, Wischhusen P, Burton E, O’Neill HM, Van Der Heul K, Hays J, Rowe P. Analysing responsible innovation along a value chain-A single-cell protein case study. Eng Biol. 2024;8:16–29.
Frazer AC. Single cell proteins. Nature. 1967;216:743.
Fu X, Qiao W, Shi S. Microbial production of single cell proteins from single carbon substrates: a review. Food Science. 2023;44:1–11.
Gamarra-Castillo O, Echeverry-Montaña N, Marbello-Santrich A, Hernández-Carrión M, Restrepo S. Meat substitute development from fungal protein (Aspergillus oryzae). Foods. 2022;11:1–15.
Gao Z, Guo S, Chen Y, Chen H, Fu R, Song Q, Li S, Lou W, Fan D, Li Y. A novel nutritional induction strategy flexibly switching the biosynthesis of food-like products from methane by a methanotrophic bacterium. Green Chem. 2024;26:7048–58.
Gao L, Meng J, Dai W, Zhang Z, Dong H, Yuan Q, Zhang W, Liu S, Wu X. Deciphering cell wall sensors enabling the construction of robust P. pastoris for single-cell protein production. Biotechnol Biofuels Bioprod. 2023;7:16178–92.
Gopinath V, Murali A, Dhar KS, Nampoothiri KM. Corynebacterium glutamicum as a potent biocatalyst for the bioconversion of pentose sugars to value-added products. Appl Microbiol Biotechnol. 2011;93:95–106.
Gour Suman MN, Singh A, Bhatnagar P. Single cell protein production: a review. Int J Curr Microbiol App Sci. 2015;4:251–63.
Graham AE, Ledesma-Amaro R. The microbial food revolution. Nature. Communications. 2023;14:2231.
Guo S, Nguyen DTN, Chau THT, Fei Q, Lee EY. Systems metabolic engineering of Methanotrophic Bacteria for biological conversion of methane to value-added compounds. Adv Biochem Eng Biotechnol. 2022;180:91–126.
Gupta JK, Rai P, Jain KK, Srivastava S. Overexpression of bicarbonate transporters in the marine cyanobacterium Synechococcus sp. PCC 7002 increases growth rate and glycogen accumulation. Biotechnol Biofuels. 2020;13:1–12.
Hadi J, Brightwell G. Safety of alternative proteins: technological, environmental and regulatory aspects of cultured meat, plant-based meat, insect protein and single-cell protein. Foods. 2021;10: 1226.
Hara KY, Kobayashi J, Yamada R, Sasaki D, Kuriya Y, Hirono-Hara Y, Ishii J, Araki M, Kondo A. Transporter engineering in biomass utilization by yeast. FEMS Yeast Research. 2017;17: fox061.
Hashempour-Baltork F, Jannat B, Dadgarnejad M, Mirza Alizadeh A, Khosravi-Darani K, Hosseini H. Mycoprotein as chicken meat substitute in nugget formulation: physicochemical and sensorial characterization. Food Sci Nutr. 2023;11:4289–95.
Hezarjaribi M, Ardestani F, Ghorbani HR. Single cell protein production by Saccharomyces cerevisiae using an optimized culture medium composition in a batch submerged bioprocess. Applied biochemistry and biotechnology. 2016;179:1336–45.
Hu X, Vandamme P, Boon N. Co-cultivation enhanced microbial protein production based on autotrophic nitrogen-fixing hydrogen-oxidizing bacteria. Chem Eng J. 2022;429: 132535.
Hui MP, Foley PL, Belasco JG. Messenger RNA degradation in bacterial cells. Annu Rev Genet. 2014;48:537–59.
Hülsen T, Barnes AC, Batstone DJ, Capson-Tojo G. Creating value from purple phototrophic bacteria via single-cell protein production. Curr Opin Biotechnol. 2022;76:102726–41.
Humpenöder F, Bodirsky BL, Weindl I, Lotze-Campen H, Linder T, Popp A. Projected environmental benefits of replacing beef with microbial protein. Nature. 2022;605:90–6.
Hung Y, Van der Stricht H, Verbeke W. Consumer acceptance and nutritional expectations of microalgae protein products: insights from a cross-European study. In Proceedings. 2023;91:87.
Jach ME, Masłyk M, Juda M, Sajnaga E, Malm A. Vitamin B12-Enriched Yarrowia lipolytica Biomass Obtained from Biofuel Waste. Waste and Biomass Valorization. 2018;11:1711–6.
Jach ME, Serefko A, Ziaja M, Kieliszek M. Yeast protein as an easily accessible food source. Metabolites. 2022;12:1–8.
Jain S. Global potential of biogas. 2019. In Available at: https://www.worldbiogasassociation.org/global-potential-of-biogas/.
Jalasutram V, Kataram S, Gandu B, Anupoju GR. Single cell protein production from digested and undigested poultry litter by Candida utilis: optimization of process parameters using response surface methodology. Clean Technol Environ Policy. 2012;15:265–73.
Jang Y-S, Park JM, Choi S, Choi YJ, Cho JH, Lee SY. Engineering of microorganisms for the production of biofuels and perspectives based on systems metabolic engineering approaches. Biotechnol Adv. 2012;30:989–1000.
Johnson EA. Biotechnology of non-Saccharomyces yeasts–the ascomycetes. Appl Microbiol Biotechnol. 2013;97:503–17.
Junaid F, Khawaja LA, Ali S. Single cell proteins as a potential meat substitute: a critical review. World J Pharm Res. 2020;9:141–61.
Juncker JC. Commission implementing regulation (EU) 2019_760. Official Journal of the European Union. 2019;125:13–5.
Kaganovich D, Kopito R, Frydman J. Misfolded proteins partition between two distinct quality control compartments. Nature. 2008;454:1088–95.
Kam S, Kenari AA, Younesi H. Production of Single Cell Protein in Stickwater byLactobacillus acidophilusandAspergillus niger. J Aquat Food Prod Technol. 2012;21:403–17.
Kerckhof F-M, Sakarika M, Van Giel M, Muys M, Vermeir P, De Vrieze J, Vlaeminck SE, Rabaey K, Boon N. From biogas and hydrogen to microbial protein through co-cultivation of methane and hydrogen oxidizing bacteria. Frontiers in Bioengineering and Biotechnology. 2021;9:733753.
Khoshnevisan B, Tsapekos P, Zhang Y, Valverde-Pérez B, Angelidaki I. Urban biowaste valorization by coupling anaerobic digestion and single cell protein production. Bioresource Technol. 2019;290:121743.
Khumchai J, Wongchai A, On-uma R, Sabour A, Alshiekheid M, Narayanan M, Karuppusamy I, Pugazhendhi A, Brindhadevi K, Lan Chi NT. A viable bioremediation strategy for treating paper and pulp industry effluents and assessing the prospect of resulted bacterial biomass as single cell protein (SCP) using indigenous bacterial species. Chemosphere. 2022;304:135246.
Kim K, Choe D, Lee D-H, Cho B-K. Engineering biology to construct microbial chassis for the production of difficult-to-express proteins. Int J Mol Sci. 2020;21: 990.
KnipBio. KnipBio Meal: Premium Single Cell Protein for Aquaculture. n.d. from: https://www.knipbio.com/knipbio-meal-premium-single-cell-protein-aquaculture.
LaFleur TL, Hossain A, Salis HM. Automated model-predictive design of synthetic promoters to control transcriptional profiles in bacteria. Nat Commun. 2022;13:5159.
Lee JZ, Logan A, Terry S, Spear JR. Microbial response to single-cell protein production and brewery wastewater treatment. Microb Biotechnol. 2014;8:65–76.
Li R, Fan X, Jiang Y, Wang R, Guo R, Zhang Y, Fu S. From anaerobic digestion to single cell protein synthesis: A promising route beyond biogas utilization. Water Res. 2023;243:120417.
Linder T. Making the case for edible microorganisms as an integral part of a more sustainable and resilient food production system. Food Security. 2019;11:265–78.
Lindquist SL, Kelly JW. Chemical and biological approaches for adapting proteostasis to ameliorate protein misfolding and aggregation diseases–progress and prognosis. Cold Spring Harb Perspect Biol. 2011;3:a004507.
Liu Y, Deng M, Chen J. Microbial alternative protein biomanufacturing: advances and perspectives. Journal of Chinese Institute of Food Science and Technology. 2022;22:1–5.
Liu B, Song J, Li Y, Niu J, Wang Z, Yang Q. Towards industrially feasible treatment of potato starch processing waste by mixed cultures. Appl Biochem Biotechnol. 2013;171:1001–10.
Liu S. Woody biomass: Niche position as a source of sustainable renewable chemicals and energy and kinetics of hot-water extraction/hydrolysis. Biotechnol Adv. 2010;28:563–82.
Liu X, Zhang F, Meng Z, Sun Q , T S, Liu C. The production of bacterial proteins from ethanol mash by fermentation of iron and steel industry tail gas. Chem Eng Manage. 2020;11:143–45.
Liu X, Zhang W, Zhao Z, Dai X, Yang Y, Bai Z. Protein secretion in Corynebacterium glutamicum. Crit Rev Biotechnol. 2016;37:541–51.
Ma S, Liang X, Chen P, Wang J, Gu X, Qin Y, Blecker C, Xue M. A new single-cell protein from Clostridium autoethanogenum as a functional protein for largemouth bass (Micropterus salmoides). Anim Nutr. 2022;10:99–110.
Mango Materials. Converting methane into biodegradable polymers. n.d. from: https://www.mangomaterials.com/biodegradable-polymers.
Matassa S, Boon N, Pikaar I, Verstraete W. Microbial protein: future sustainable food supply route with low environmental footprint. Microb Biotechnol. 2016;9:568–75.
Meng J, Liu S, Gao L, Hong K, Liu S, Wu X. Economical production of Pichia pastoris single cell protein from methanol at industrial pilot scale. Microb Cell Fact. 2023;2023(22):1–16.
Morrison O. New wave of cleaner, healthier, and tastier alternative proteins, report predicts. In Investing in the alternative protein sector has one of the biggest impacts on decarbonization when assessed in terms of the market value of avoided CO2e emissions per dollar invested in mitigation efforts, claims a new report. 2022. https://www.foodnavigator.com/article/2022/07/11/new-wave-of-cleaner-healthier-and-tastier-alternative-proteins-report-predicts.
Mukherjee N, Corcoran DL, Nusbaum JD, Reid DW, Georgiev S, Hafner M, Ascano M, Tuschl T, Ohler U, Keene JD. Integrative regulatory mapping indicates that the RNA-binding protein HuR couples pre-mRNA processing and mRNA stability. Molecular Cell. 2011;43:327–39.
Nasseri A, Rasoul-Ami S, Morowvat MH, Ghasemi Y. Single Cell Protein-Production and Process. Am J Food Technol. 2011;6:103–16.
Nature's Fynd. Fy protein: a versatile fungal protein. n.d. from:https://www.naturesfynd.com/fy-protein.
Niccolai A, Chini Zittelli G, Rodolfi L, Biondi N, Tredici MR. Microalgae of interest as food source: biochemical composition and digestibility. Algal Res. 2019;42:101617.
Norman ROJ, Millat T, Winzer K, Minton NP, Hodgman C. Progress towards platform chemical production using Clostridium autoethanogenum. Biochem Soc Trans. 2018;46:523–35.
NovoNutrients. NovoMeal™: Sustainable Protein for Industrial Applications. n.d. from: https://www.novonutrients.com/products/novomeal.
Onyeaka H, Anumudu CK, Okpe C, Okafor A, Ihenetu F, Miri T, Odeyemi O, Anyogu A. Single cell protein for foods and feeds: a review of trends. 2022;16:e187428582206160.
Øverland M, Tauson A-H, Shearer K, Skrede A. Evaluation of methane-utilising bacteria products as feed ingredients for monogastric animals. Arch Anim Nutr. 2010;64:171–89.
Pander B, Mortimer Z, Woods C, McGregor C, Dempster A, Thomas L, Maliepaard J, Mansfield R, Rowe P, Krabben P. Hydrogen oxidising bacteria for production of single-cell protein and other food and feed ingredients. Engineering Biology. 2020;4:21–4.
Patrick E, McGovern JZ, Tang J, Zhang Z, Gretchen R, Robert AM, Alberto N, Eric DB, Michael PR, Wang C, Cheng G, Zhao Z, Wang C. Fermented beverages of pre- and proto-historic China. Proc Natl Acad Sci USA. 2004;101:17593–8.
Pereira AG, Fraga-Corral M, Garcia-Oliveira P, Otero P, Soria-Lopez A, Cassani L, Cao H, Xiao J, Prieto MA, Simal-Gandara J. Single-cell proteins obtained by circular economy intended as a feed ingredient in aquaculture. Foods. 2022;11:2831–53.
Perfect Day. Animal-Free Dairy Proteins. n.d. From:https://www.perfectday.com/animal-free-dairy-proteins.
Pham HL, Wong A, Chua N, Teo WS, Yew WS, Chang MW. Engineering a riboswitch-based genetic platform for the self-directed evolution of acid-tolerant phenotypes. Nature communications. 2017;8:411.
Qin S, Kaikai L, Tuo Q, Xuetuan W. Application and development of microbial proteins as high quality alternative protein resources—a review. Futre Food Science. 2022;2:96–106.
Quorn. Quorn products: sustainable protein for a healthier planet. n.d. From https://www.quorn.us/products.
Rashad MM, Moharib SA, Jwanny WE. Yeast conversion of mango waste or methanol to single cell protein and other metabolites. Biol Wastes. 1990;32:277–84.
Ravindra P. Value-added food: Single cell protein. Biotechnology advances. 2000;18:459–79.
Reihani SFS, Khosravi-Darani K. Influencing factors on single-cell protein production by submerged fermentation: a review. Electron J Biotechnol. 2019;37:34–40.
Rezaei R, Wu Z, Hou Y, Bazer FW, Wu G. Amino acids and mammary gland development: nutritional implications for milk production and neonatal growth. J Anim Sci Biotechnol. 2016;7:20.
Rischer H, Szilvay GR, Oksman-Caldentey KM. Cellular agriculture-industrial biotechnology for food and materials. Curr Opin Biotechnol. 2020;61:128–34.
Ritala A, Häkkinen ST, Toivari M, Wiebe MG. Single cell protein—state-of-the-art, industrial landscape and patents 2001–2016. Frontiers in microbiology. 2017;8:300587.
Roux C, Etienne TA, Hajnsdorf E, Ropers D, Carpousis AJ, Cocaign-Bousquet M, Girbal L. The essential role of mRNA degradation in understanding and engineering E. coli metabolism. Biotechnology Advances. 2022;54: 107805.
Salazar-López NJ, Barco-Mendoza GA, Zuñiga-Martínez BS, Domínguez-Avila JA, Robles-Sánchez RM, Ochoa MAV, González-Aguilar GA. Single-Cell Protein Production as a Strategy to Reincorporate Food Waste and Agro By-Products Back into the Processing Chain. Bioeng. 2022;9:623–34.
Schlapschy M, Grimm S, Skerra A. A system for concomitant overexpression of four periplasmic folding catalysts to improve secretory protein production in Escherichia coli. Protein Eng Des Sel. 2006;19:385–90.
Siddiqui SA, Alvi T, Sameen A, Khan S, Blinov AV, Nagdalian AA, Mehdizadeh M, Adli DN, Onwezen M. Consumer acceptance of alternative proteins: A systematic review of current alternative protein sources and interventions adapted to increase their acceptability. Sustainability. 2022;14:15370.
Silva JBAe, Lima UA, Taqueda MES, Guaragna FG. Use of response surface methodology for selection ofnutrient levels for culturing Paecilomyces variotii ineucalyptus hemicellulosic hydrolyzate. Bioresourc Technol. 2003;87:45–50.
Simões ACP, Fernandes RP, Barreto MS, da Costa GBM, de Godoy MG, Freire DMG, Pereira N Jr. Growth of Methylobacterium organophilum in methanol for the simultaneous production of single-cell protein and metabolites of interest. Food Technology and Biotechnology. 2022;60:338–49.
Smetana S, Mathys A, Knoch A, Heinz V. Meat alternatives: life cycle assessment of most known meat substitutes. The International Journal of Life Cycle Assessment. 2015;20:1254–67.
Smith MH, Ploegh HL, Weissman JS. Road to ruin: targeting proteins for degradation in the endoplasmic reticulum. Science. 2011;334:1086–90.
Solar Foods. Solein: a microbial protein powder. n.d. from: https://www.solarfoods.fi/solein.
Solow SP, Sengbusch J, Laird MW. Heterologous protein production from the inducible MET25 promoter in Saccharomyces cerevisiae. Biotechnol Prog. 2005;21:617–20.
Su Y, Liu C, Fang H, Zhang D. Bacillus subtilis: a universal cell factory for industry, agriculture, biomaterials and medicine. Microb Cell Fact. 2020;19:173.
Taldone T, Patel HJ, Bolaender A, Patel MR, Chiosis G. Protein chaperones: a composition of matter review (2008–2013). Expert Opin Ther Pat. 2014;24:501–18.
Tan F, Wu B, Dai L, Qin H, Shui Z, Wang J, Zhu Q, Hu G, He M. Using global transcription machinery engineering (gTME) to improve ethanol tolerance of Zymomonas mobilis. Microb Cell Fact. 2016;15:1–9.
Tang H, Bao X, Shen Y, Song M, Wang S, Wang C, Hou J. Engineering protein folding and translocation improves heterologous protein secretion in Saccharomyces cerevisiae. Biotechnol Bioeng. 2015;112:1872–82.
Tang H, Wu L, Guo S, Cao W, Ma W, Wang X, Shen J, Wang M, Zhang Q, Huang M, et al. Metabolic engineering of yeast for the production of carbohydrate-derived foods and chemicals from C1–3 molecules. Nat Catal. 2023;7:21–34.
TerraVia. Algae-based Food and Nutrition Solutions. n.d. from: https://www.terravia.com/algae-based-food-and-nutrition-solutions.
Tesfaw A, Assefa F. Co-culture: a great promising method in single cell protein production. Biotechnol Mol Biol Rev. 2014;9:12–20.
Tomimoto K, Fujita Y, Iwaki T, Chiba Y, Jigami Y, Nakayama K-i, Nakajima Y, Abe H. Protease-deficient Saccharomyces cerevisiae strains for the synthesis of human-compatible glycoproteins. Bioscience, biotechnology, and biochemistry. 2013;77:2461–6.
Tong S, Chen W, Hong R, Chai M, Sun Y, Wang Q, Li D. Efficient Mycoprotein Production with Low CO2 Emissions through Metabolic Engineering and Fermentation Optimization of Fusarium. J Agric Food Chem. 2024;72:604–12.
Turck D, Castenmiller J, de Henauw S, Hirsch-Ernst KI, Kearney J, Maciuk A, Mangelsdorf I, McArdle HJ, Naska A, Pelaez C. Safety of Yarrowia lipolytica yeast biomass as a novel food pursuant to regulation (EU) 2015/2283. EFSA J. 2019;17:e05594.
Ugalde U, Castrillo JI. Single cell proteins from fungi and yeasts. Applied Mycology and Biotechnolog. 2002;2:123–49.
Unibio. Uniprotein®: Sustainable Protein for Animal Feed. n.d. from: https://www.unibio.dk/uniprotein-sustainable-protein-animal-feed.
Valentino MJG. Mycota of distillery yeast sludge as source of single cell protein. Mycosphere. 2015;6:241–7.
Valkonen M, Penttilä M, Saloheimo M. Effects of inactivation and constitutive expression of the unfolded-protein response pathway on protein production in the yeast Saccharomyces cerevisiae. Applied and environmental microbiology. 2003;69:2065–72.
Varshavsky A. The ubiquitin system, autophagy, and regulated protein degradation. Ann Rev Biochem. 2017;86:123–8.
Venturelli OS, Tei M, Bauer S, Chan LJG, Petzold CJ, Arkin AP. Programming mRNA decay to modulate synthetic circuit resource allocation. Nat Commun. 2017;8: 15128.
Vethathirri RS, Santillan E, Wuertz S. Microbial community-based protein production from wastewater for animal feed applications. Biores Technol. 2021;341: 125723.
Viegas SC, Apura P, Martínez-García E, de Lorenzo V, Arraiano CM. Modulating heterologous gene expression with portable mRNA-stabilizing 5′-UTR sequences. ACS Synth Biol. 2018;7:2177–88.
Wan S, Lai M, Gao X, Zhou M, Yang S, Li Q, Li F, Xia L, Tan Y. Recent progress in engineering Clostridium autoethanogenum to synthesize the biochemicals and biocommodities. Synthetic and Systems Biotechnology. 2024;9:19–25.
Windass JD, Worsey MJ, Pioli EM, Pioli D, Barth PT, Atherton KT, Dart EC, Byrom D, Powell K, Senior PJ. Improved conversion of methanol to single-cell protein by Methylophilus methylotrophus. Nature. 1980;287:396–401.
Wolff S, Weissman JS, Dillin A. Differential scales of protein quality control. Cell. 2014;157:52–64.
Wu J, Bao M, Duan X, Zhou P, Chen C, Gao J, Cheng S, Zhuang Q, Zhao Z. Developing a pathway-independent and full-autonomous global resource allocation strategy to dynamically switching phenotypic states. Nat Commun. 2020;11:5521.
Wu Y, Deng J, Zheng Z, Chen N, Luo X, Tang H. Engineering an efficient expression using heterologous GAL promoters and transcriptional activators in Saccharomyces cerevisiae. ACS Synth Biol. 2023;12:1859–67.
Wu J, Hu J, Zhao S, He M, Hu G, Ge X, Peng N. Selection of marine yeasts for the generation of single cell protein from prawn-shell waste. Appl Biochem Biotechnol. 2018;185:163–78.
Xing-fa L, Fang-qi Z, Zhi-peng M, Qing-kun S, Shu-huan T, Chong-yang L. The production of bacterial proteins from ethanol mash by fermentation of iron and steel industry tail gas. Chemical Engineering Management. 2020;11:143–5.
Yadav JSS, Bezawada J, Ajila CM, Yan S, Tyagi RD, Surampalli RY. Mixed culture of Kluyveromyces marxianus and Candida krusei for single-cell protein production and organic load removal from whey. Biores Technol. 2014;164:119–27.
Yang H-H, Thayer D, Yang S. Reduction of endogenous nucleic acid in a single-cell protein. Appl Environ Microbiol. 1979;38:143–7.
Yao S, Lyu S, An Y, Lu J, Gjermansen C, Schramm A. Microalgae-bacteria symbiosis in microalgal growth and biofuel production: a review. J Appl Microbiol. 2019;126:359–68.
Yongjin GJZ. Advances in methanol bio-transformation. Synthetic Biology Journal. 2020;1:158–73.
Yu J. Fixation of carbon dioxide by a hydrogen-oxidizing bacterium for value-added products. World J Microbiol Biotechnol. 2018;34:89.
Zha X, Tsapekos P, Zhu X, Khoshnevisan B, Lu X, Angelidaki I. Bioconversion of wastewater to single cell protein by methanotrophic bacteria. Biores Technol. 2021;320:124351–8.
Zhang Q, Liang H, Longshaw M, Wang J, Ge X, Zhu J, Li S, Ren M. Effects of replacing fishmeal with methanotroph (Methylococcus capsulatus, Bath) bacteria meal (FeedKind®) on growth and intestinal health status of juvenile largemouth bass (Micropterus salmoides). Fish Shellfish Immunol. 2022;122:298–305.
Zhang X, Zhao ZG, Liu Z. Research Progress of Biodegradation and Bioconversion of Wooden Biomass. Scientia Silvae Sinicae. 2006;42:85–91.
Zhao M, Ma J, Zhang L, Qi H. Engineering strategies for enhanced heterologous protein production by Saccharomyces cerevisiae. Microbial Cell Fact. 2024;23:32.
Zhao L, Ye B, Zhang Q, Cheng D, Zhou C, Cheng S, Yan X. Construction of second generation protease-deficient hosts of Bacillus subtilis for secretion of foreign proteins. Biotechnol Bioeng. 2019;116:2052–60.
Zhu W, Gong G, Pan J, Han S, Zhang W, Hu Y, Xie L. High level expression and purification of recombinant human serum albumin in Pichia pastoris. Protein Expr Purif. 2018;147:61–8.
Acknowledgements
Not applicable.
Funding
This work was supported by the National Key Research and Development Program of China (2021YFA0911000), the National Natural Science Foundation of China (32071416), the Key-Area Research and Development Program of Guangdong Province (2022B1111080005), the Shenzhen Key Laboratory for the Intelligent Microbial Manufacturing of Medicines (ZDSYS20210623091810032), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB0480000).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by Zhoukang Zhuang and Guangyu Wan. The first draft of the manuscript was written by Tao Yu and Hongting Tang. Zhoukang Zhuang and Guangyu Wan, Xiaocong Lu and Linhai Xie corrected and approved the final manuscript. All authors commented on previous versions of the manuscript. All authors read and approved the final manuscript. Acknowledging additional support from the China Merchants Investment Development Co., Ltd and the China Blue Chemical Ltd.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
All authors agreed with the content and that all gave explicit consent to submit.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhuang, Z., Wan, G., Lu, X. et al. Metabolic engineering for single-cell protein production from renewable feedstocks and its applications. Adv. Biotechnol. 2, 35 (2024). https://doi.org/10.1007/s44307-024-00042-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44307-024-00042-8