Abstract
Pear (Pyrus bretschneideri), a valuable widely cultivated fruit, faces significant economic losses due to black spot disease caused by Alternaria alternate (Fr.) Keissl. Trihelix transcription factors (TFs) are crucial in regulating plant defense and autoimmunity. This study aimed to analyze the trihelix transcription factor (GT) genes within pear through genome-wide identification, phylogenetic, gene structure, synteny, and cis-acting elements analyses. Among the 31 trihelix genes, 28 were on 12 known chromosomes, while the remaining 3 were located on unknown chromosomes. These genes were categorized into five clades: SIP1, GTγ, GT1, GT2 and SH4, containing 7, 2, 9, 11 and 2 genes, respectively. Synteny analysis indicated eight duplicated gene pairs. Based on the expression pattern of PbGT genes in seven tissues from the database, the PbGT genes of the GT2 clade were selected for further investigation. The quantitative reverse transcriptase–polymerase chain reaction confirmed that PbrGT5, PbrGT6, PbrGT15 and PbrGT16 correlated with black spot disease resistance. Notably, the salicylic acid (SA) treatment significantly upregulated the expression levels of PbrGT10, PbrGT13, PbrGT15 and PbrGT23. Among these, PbrGT15 showed the highest induction to both SA and black spot infection. Subcellular localization demonstrated that PbrGT15 functions as a nuclear protein. Virus-induced gene silencing of PbrGT15 increased pear plants' susceptibility to black spot disease, indicating its pivotal role in enhancing resistance. These results indicated that PbrGT15 positively regulated black spot disease resistance in pears.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants face various biotic and abiotic stresses during their growth. Biotic stress primarily arising from pathogenic sources (bacteria, fungi and viruses) adversely impacts yield and quality (Bhar et al. 2022). Signal transduction interacting with biotic stress has been shown to bolster stress tolerance (Peck et al. 2020). Plants integrate diverse signals using transcription factors (TFs) from various families, including v-myb avian myeloblastosis viral (MYBs) and dehydration responsive element binding proteins (DREBs) (Zandalinas et al. 2020). TFs play a crucial role in plant growth, development and stress responses (Osorio et al. 2012). Over 60 TF families have been classified based on their DNA-binding domains (Riechmann et al. 2000). The trihelix TFs derive their name from their helix-loop-helix-loop-helix structure in the DNA-binding domain and are also known as trihelix transcription factor (GT) due to their strong affinity for GT elements (Kaplan-Levy et al. 2012; Qin et al. 2014).
The GT family, relatively understudied since the 1990s, has recently regained interest. It encompasses genes from various plants, with Arabidopsis and Oryza sativa each containing 30 and 31 characterized GT genes, respectively. The GT genes have been divided into five clades: GT1, GT2, SH4, GTγ and SIP1 (Kaplan-Levy et al. 2012). The name of each branch is based on the name of the first gene that has been identified (Gao et al. 2009). Additionally, 63 GT genes are also found in soybean (Song et al. 2016), 20 in chrysanthemum (Song et al. 2016), 59 in maize (Du et al. 2016), 36 in tomato (Yu et al. 2015) and 94 in wheat (Xiao et al. 2019). These GT genes (Osorio et al. 2012) are involved in plant development, including flowers (Brewer et al. 2004), leaves (Li et al. 2008), stomata (Yoo et al. 2010), embryogenesis (Willmann et al. 2011) and seeds (Lin et al. 2007), respond to stresses. For instance, the GT2-like 1 (GTL1) factor in Arabidopsis regulates bacteria-triggered immunity (Völz et al. 2018). Paired GT factors ASR3/AITF1 in Arabidopsis regulate plant defense and autoimmunity (Wang et al. 2023). In Arabidopsis, the GT3a and GT3b of GT1 clade genes reportedly responded to salt and pathogen stress, mirroring findings in soybean (Ayadi et al. 2004; Park et al. 2004). Furthermore, OsRML1, a rice GT factor, displays increased expression in seedlings post-fungal blast infection (Wang et al. 2004). Notably, GhGT23 and GhGT26 in cotton improve salinity tolerance (Li et al. 2022; Li et al. 2023), while GhGT3b_A04 enhances resistance to Verticillium dahliae in cotton (Mao et al. 2023). The maize GT factor ZmGT3b is associated with the defense response against Fusarium graminearum (Zhang et al. 2021).
The pear, a Rosaceae family fruit, holds significant economic value with a rich cultivation history. Black spot, a devastating disease affecting pear cultivation and management globally, spreads via wind and rain, significantly impacting leaves, fruit and new shoots. This disease prompts premature leaf drop and fruit decay, thereby diminishing storage value, which consequently affects fruit yield and quality. However, limited studies have delved into the pear trihelix family, leaving the involvement of these genes in pear black spot uncertain. This study focuses on genome-wide identification, phylogenetic analysis, gene structure, synteny analysis, and cis-acting elements of 31 GT genes in Pyrus bretschneideri. Transcriptome data was utilized to examine relative GT gene expression in tissues, particularly assessing the expression levels of GT2 clade genes in different resistant varieties (R-01 and S-01). Furthermore, this research elucidates the role of PbrGT15 in conferring black spot disease resistance using virus-induced gene silencing (VIGS).
Materials and methods
Plant materials and treatment
Leaves of P. bretschneideri (‘Xiangmianli’, R-01) and P. sinkiangensis (‘Hongjujuli’, S-01) from an orchard (Baima, Nanjing, China) were pierced with sterile needles and inoculated with Alternaria alternata at the pinholes. RNA was extracted from these leaves after 0, 3, 12, 48 and 72 h. Subsequently, 45-day-old P. betulaefolia seedlings underwent salicylic acid (SA) treatment or VIGS at 25°C under a 16-h day/8-h night cycle. The plants were sprayed with 50 mL of 0.2 mM SA solution or water distilled. The true leaf of each plant was sampled with three biological replicates at 0, 6, 12, 24, 48 and 72 h, and stored at −80°C.
Identification of PbrGT genes in pear
A protein database of Chinese white pear was established (Wu et al. 2013). The trihelix protein sequences were downloaded from the TAIR database (http://www.arabidopsis.org) and the rice genome (http://rice.plantbiology.msu.edu/). A local BLASTP with an e-value of le-5 and a 50% homology threshold was used to screen the pear local protein database. Subsequently, the Hidden Markov Model (HMM) was constructed using HMMER 3.0 (http://hmmer.org/) based on the trihelix protein sequence alignment results of Arabidopsis and rice, then searched in the local database. The hits containing the PF13837 (Myb/SANT-LIKE domain) were compared with the BLASTP results, and the overlapping genes were considered to be the PbrGT genes.
The ID, location, length, theoretical isoelectric point (pI) and molecular weight (MW) of PbrGT genes were determined using ExPASy (http://web.expasy.org/compute_pi/).
Phylogenetic analysis
Arabidopsis, rice and pear trihelix protein sequences were aligned with ClustalX 2.0 (Thompson et al. 1997). The unrooted trees were constructed with MEGA 7.0 using the neighbor-joining (NJ) method, repeated 1000 times with the bootstrap test.
Synteny analysis of PbrGT genes
The syntenic chains and duplication mode of the trihelix gene family were identified using MCScanX (Qiao et al. 2015). The figure was generated using TBtools. The nonsynonymous (Ka) and synonymous (Ks) substitution ratio was calculated using the KaKs_Calculator 2.0 (Wang et al. 2010).
Gene structure construction, motif and cis-acting element identification
The exon/intron structure was obtained from GSDS 2.0 (http://gsds.gao-lab.org/) (Hu et al. 2015). The conserved protein motifs were analyzed using MEME (http://meme-suite.org/tools/meme) (Bailey et al. 2006).
The cis-acting elements in the 2 kbp upstream region of the genes were explored using the PlantCARE database (https://bioinformatics.psb.ugent.be/webtools/plantcare/html/). The graphic was output using TBtools.
Tissue-specific expression of PbrGT genes based on transcriptome data
Using RNA-seq data of P. bretschneideri ‘Dangshansuli’ pear (Li et al. 2019), the expression of PbrGT genes was examined in seven pear tissues: leaf, fruit, petal, sepal, ovary, stem and bud, obtained from P. bretschneideri ‘Dangshansuli’ grown naturally. TBtools displayed the expression pattern using the value log2 (RPKM + 1) (Reads Per Kilobase of exon model per Million mapped reads) (Chen et al. 2020).
Expression pattern analysis by quantitative reverse transcriptase–polymerase chain reaction
The RNA samples were extracted using the Total RNA Kit (Foregene, Chengdu, China). Reverse transcription utilized the HiScript III RT SuperMix (+ gDNA wiper) (Vazyme, Nanjing, China). Primers, designed via Primer Premier 6.0 (Table S1), were employed. Quantitative reverse transcriptase–polymerase chain reaction (qRT‒PCR) was conducted in a 20 μL SYBR Green reaction (Synergy Brands) on a Roche LightCycler 480 instrument (Roche, Basel, Switzerland) with the following protocol: 45 cycles of 95°C for 10 s, 60°C for 30 s, and 72°C for 20 s. The TUB gene was considered as the reference (Chen et al. 2015), and the expression profiles were calculated via the 2−ΔΔCt value (Livak and Schmittgen 2001). Each sample was repeated thrice.
Subcellular localization of PbrGT15
The coding sequence (CDS) without a stop codon of PbrGT15 was cloned. The fusion plasmid 35S:PbrGT15:GFP was transfected into Agrobacterium tumefaciens GV3101 using freeze‒thawing, then cultured for approximately 24 h at 28°C. The bacterial suspensions were pressure-injected into tobacco leaves and observed after 48–72 h incubation at 25°C using a confocal laser scanning microscope (LSM800, Zeiss, Jena, Germany), with subsequent image capture.
VIGS
PbrGT15 DNA fragment was cloned and inserted into the pTRV2 vector for PbrGT15 silencing in pear (Gao and Shan 2013). GV3101 cells harboring pTRV1, pTRV2 and PbrGT15-TRV were cultured overnight at 28°C, then harvested (5500 g, 5 min). The OD600 value was then adjusted to 0.8 with infiltration buffer MMA (10 mM MgCl2, 10 mM 2-(4 morpholino)-ethane sulfonic acid, 150 μM acetosyringone, pH 5.7). A 1:1 ratio mixture of pTRV1 and PbrGT15-TRV, and pTRV1 and pTRV2, was used for seedling infection. The infected seedlings were cultured under a 16 h light/8 h dark cycle for 20 days after a dark incubation period. Subsequently, isolated leaves were inoculated with A. alternata and cultured in the dark for six days, followed by measuring the spot diameter. Electrolyte leakage (EL) was determined as described previously (Dahro et al. 2016). The enzyme activity, including phenylalanine ammonia-lyase (PAL), chitinase (CHI), catalase (CAT) and polyphenol oxidase (PPO) were measured using their respective test kits (Jiancheng, Nanjing, China).
Statistical analysis
Data were summarized using Microsoft Excel. GraphPad Prism 6 was used for data analysis and column chart creation.
Results
Identification and analysis of PbrGT genes in pear
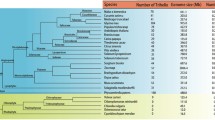
From the pear protein databases, 31 GT genes were identified and labelled PbrGT1 to PbrGT31 based on their chromosomal locations (Table 1). Among these, 28 PbrGT genes are distributed across 12 distinct chromosomes, while the locations of other three PbrGT genes remain unknown. Notably, PbrGT13 encoded the smallest protein (92 amino acids), whereas PbrGT7/8/31 encoded the largest proteins, each comprising 624 amino acids. The pI of the PbrGT genes range from 5.11 to 9.57 and their MW ranged between 16.06 and 102.97 kDa.
Phylogenetic tree construction
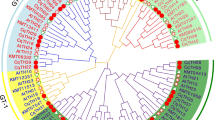
To understand the phylogenetic relationships among GT proteins in Arabidopsis, rice and pear, we constructed an NJ phylogenetic tree using 30 amino acid sequences from Arabidopsis, 31 from rice (detailed in Table S2) and 30 from pear. Building upon prior research on Arabidopsis and rice (Kaplan-Levy et al. 2012), we categorized the 92 sequences into five distinct clades, encompassing 7, 2, 9, 11 and 2 pear GT proteins, respectively (Fig. 1).
Phylogenetic tree of trihelix transcription factor (GT) genes. The pear, Arabidopsis and rice GT genes are marked with green circles, red triangles and blue diamonds, respectively. The groups are represented by different colors: GT-1 (blue), GT-2 (green), GTγ (yellow), SIP1 (purple) and SH4 (pink). PbrGT, pear trihelix transcription factor family gene
Synteny analysis
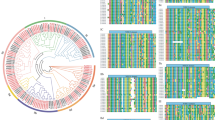
The pear genome exhibited diverse gene duplication events: 18 genes resulted from whole-genome duplication or segmental duplication (WGD or segmental), 10 from dispersed duplication and 3 from tandem duplication. Among these duplications, eight pairs were categorized as WGD or segmental genes (Table S3; Fig. 2). Notably, all duplicated genes demonstrated Ka/Ks values below 1, indicating strong purifying selection.
Gene structure, motif and cis-acting element analysis
The PbrGT proteins within the same branch showed analogous motif compositions and exon/intron structures, implying potential functional similarities. Using the MEME search tool, ten motifs were identified. Figure 3a shows the organizational patterns of these motifs within the PbrGT proteins. Motifs 1 and 6 were conserved across most PbrGT proteins, while some comprised only a single motif and others incorporated up to eight motifs. Detailed motif representations are shown in Figure S1. From the PlantCARE database, 37 cis-acting elements were identified. Figure 3b shows the further analysis of 15 stress, hormone-related and plant growth and development-related elements. Except for PbrGT12, all PbrGT genes incorporated cis-acting elements associated with various hormones, including abscisic acid (ABA), gibberellin (GA), SA, and methyl jasmonate (MeJA). Regarding gene structure, seven PbrGT genes (PbrGT1/2/4/11/13/14/17) lacked introns, while 10 genes (PbrGT3/5/6/10/12/15/18/19/23/26) had a single intron. Additionally, four genes (PbrGT7/8/22/31) featured 17 introns, while the remaining genes contained between 2 and 7 introns, as shown in Fig. 3c.
Expression of PbrGTs in different tissues
The expression data for PbrGT genes across seven tissues were obtained from a previous study (Li et al. 2019). Utilizing the log2-transformed RPKM values, a heatmap (Fig. 4, Table S4) was generated. The majority of PbrGT genes displayed tissue-specific patterns: 16 genes were predominantly expressed in petal, calyx, stem, ovary and fruit; 18 in the bud and 15 in the leaf. A subset of 12 genes exhibited high expression across all tissues, while 10 genes consistently exhibited low expression in all tissues. Interestingly, PbrGT12, PbrGT22 and PbrGT23 had limited expression outside the bud, thus suggesting their potential role in bud development.
Expression of PbrGTs in different cultivars
A marked difference was found in the leaf inoculation spot diameter between R-01 and S-01 (Fig. 5a-c). This discrepancy indicated differing resistance levels, with R-01 exhibiting superior resistance compared to S-01.
Expression patterns of PbrGT genes in different cultivars. (a) Phenotype of R-01. Bars=2 cm. (b) Phenotype of S-01. Bars=2 cm. (c) Mean diameter of lesions (*p < 0.05; **p < 0.01; n = 12). (d)-(n) Relative expression level: PbrGT5, PbrGT6, PbrGT9, PbrGT10, PbrGT11, PbrGT13, PbrGT15, PbrGT16, PbrGT17, PbrGT21, PbrGT23. (* P < 0.05; ** P < 0.01; n = 3). PbrGT, pear trihelix transcription factor family gene; R, resistant variety; S, susceptible variety
The GTL1, a GT2 clade member, enhances SA metabolism (Kaplan-Levy et al. 2012) and positively regulates defense genes along with bacterial-triggered immunity (Völz et al. 2018). Therefore, the GT2 clade gene expression in R-01 and S-01 were analyzed using qRT‒PCR (Fig. 5d–n).
In the resistant R-01 variety, significant expression level alterations were observed in 11 genes, all displaying bimodal patterns. An initial peak emerged after 3 h, with PbrGT15 being notably higher in expression than in S-01. Subsequent peaks occurred at either after 48 or 72 h, wherein PbrGT5, PbrGT6, PbrGT15 and PbrGT16 displayed considerably elevated levels compared to S-01. These findings suggested that these four PbrGT genes were activated post-infection, crucial for reinforcing disease resistance. Conversely, expression levels of PbrGT9, PbrGT11, PbrGT13, PbrGT17 and PbrGT23 were significantly subdued compared to S-01, indicating potential susceptibility correlations. The consistent expression patterns of PbrGT10 and PbrGT21 between R-01 and S-01 suggested that they might not directly impact resistance.
Expression profiles of PbrGT genes under SA treatment
The study employed qRT‒PCR to assess GT2 clade gene expression post SA treatment. Significant expression variations were observed in 11 PbrGT genes (Fig. 6a–k), indicating a correlation between PbrGT genes and SA metabolism. PbrGT5 and PbrGT17 showed stable expression within 72 h post-SA spraying. Expression patterns for PbrGT6, PbrGT9 and PbrGT21 initially decreased after 6 or 12 h, then rose, and finally peaking values ~ 1‒2 times higher than initial levels. In contrast, PbrGT16 expression initially increased, peaking after 24 h before decreasing. The SA treatment notably induced the expression of PbrGT10, PbrGT13, PbrGT15 and PbrGT23, displaying a bimodal pattern, with peak values 4‒10 times their original levels.
Subcellular localization of PbrGT15 protein
The Plant-mPLoc tool deduced the subcellular localization of the PbrGT15 protein in the nucleus. Subsequently, the fusion protein PbrGT15::GFP was transiently expressed in tobacco leaves. As predicted, PbrGT15 was confirmed to be a nuclear-localized protein (Fig. 7).
Silencing PbrGT15 by VIGS led to reduced disease resistance
The qRT‒PCR analysis indicated sustained activation of PbrGT15 post-infection, a key gene in pear black spot resistance. PbrGT15 from the GT2 clade was further tested via VIGS. After 20 days, leaves from both VIGS-treated and control groups were collected. After six days post-inoculation, the experimental group displayed notably larger lesions than the control (Fig. 8a and b). The transcript abundance of PbrGT15 in silenced plants (TRV-3/5/6) was reduced by 60–80% compared to the control group (TRV) (Fig. 8c and d). EL in TRV-3/5/6 leaves was significantly higher than in TRV leaves (Fig. 8e). Additionally, the PAL, CHI, CAT and PPO activities were notably reduced (Figs. 8f–i). These results emphasize PbrGT15's role in enhancing black spot resistance. Moreover, the expression levels of defense-related genes, PbrPR1 and PbrNPR1, in the VIGS-treated plants were significantly lower than those in the control plants (Fig. 8j and k).
Silencing of PbrGT15. (a) Inoculated leaf phenotype. (b) Trypan blue-dyed leaves. (c) Expression of PbrGT15. (d) Mean diameter of lesions. (e) EL (%) value. f PAL enzyme activity. g CHI enzyme activity. h CAT enzyme activity. i PPO enzyme activity. j Relative expression of PbrPR1. k Relative expression of PbrNPR1. (*, P < 0.05; **, P < 0.01; n = 3–12). TRV, tobacco rattle virus; EL, electrolyte leakage; PAL, phenylalanine ammonia-lyase; CHI, chitinase; CAT, catalase; PPO, polyphenol oxidase; PbrPR1, pear pathogenesis-related protein 1; PbrNPR1, nonexpressor of pathogenesis-related protein 1
Discussion
Biotic stress reduces yield or quality in plants, prompting the activation of defense systems, one of which involves integrating various signals through transcription networks using TFs (Zandalinas et al. 2020). The trihelix TFs are pivotal in regulating developmental processes and stress responses. However, the pear trihelix gene family remains understudied.
This study identified and systematically named 31 pear members based on chromosomal positions, following naming conventions similar to the GT family in Arabidopsis and rice. The count of pear GT genes closely resembled that of Arabidopsis and rice. Phylogenetic analysis categorized PbrGTs into five subfamilies: GT1, GT2, SH4, SIP1 and GTγ (Gao et al. 2009). Gene duplications in pear involved whole-genome duplication, segmental duplication, dispersed duplication and tandem duplication. WGD or segmental duplication primarily contributed to PbrGT gene amplification. Eight pairs of duplicated genes were identified, all originating from WGD or segmental genes. Analysis of Ka/Ks values indicated that the GT genes underwent rigorous purifying selection.
Phylogenetically clustered genes often share similar motifs, implying the conservation of these motifs' importance in specific group functionalities. Promoter cis-elements are pivotal in stress responses (Fujita et al. 2006). The analysis revealed numerous cis-acting elements associated with hormones like ABA, auxin, GA, MeJA and SA within the GT gene family. Following SA treatment, several genes showed significant expression changes, as confirmed by qRT‒PCR.
From the RNA-seq dataset analysis unveiled diverse expression patterns among GT genes across different tissues. A total of 12 genes displayed consistently high ubiquitous expression levels across all tissues, while 10 genes were consistently low-expressed. Moreover, a subset of 15 genes demonstrated increased expression in leaves, thereby suggesting a potential association with leaf functionality.
Multiple trihelix TFs, such as ASR3/AITF1 (Wang et al. 2023), GT3a, GT3b in Arabidopsis (Ayadi et al. 2004; Park et al. 2004), OsRML1 in rice (Wang et al. 2004), GhGT3b_A04 in cotton, and ZmGT3b in maize (Zhang et al. 2021), play roles in defense responses against biotic stress. GTL1, a member of the GT2 subfamily in Arabidopsis, is integral to the MPK4 signaling cascade, essential for plant immunity and SA homeostasis (Völz et al. 2018). SA, a pivotal plant hormone for pathogen resistance (Wang et al. 2020; Malamy et al. 1990; Gaffney et al. 1993), induces systemic acquired resistance (SAR) associated with upregulated pathogenesis-related (PR) genes (Zhou and Zhang 2020). Genes encoding PR proteins and enzymes are upregulated during pathogen invasion (Kachroo et al. 2020; Jia et al. 2023). Endogenous SA acts as a signaling molecule inducing SAR, while externally applied SA enhances plant resistance to various biotic and abiotic stresses (Vlot et al. 2009). Based on the constructed phylogenetic tree, 11 GT genes were identified within the GT2 subfamily. PbrGT5, PbrGT6, PbrGT15 and PbrGT16 showed markedly elevated expression compared to the susceptible cultivar S-01, indicating their rapid activation post-infection and crucial role in disease resistance. Furthermore, SA treatment significantly induced the expression of PbrGT10, PbrGT13, PbrGT15 and PbrGT23, suggesting their swift activation following SA exposure.
The qRT‒PCR findings indicated a swift response of the PbrGT15 gene in disease-resistant pear varieties after SA treatment. Furthermore, its membership in the pear GT2 subfamily suggests its potential role in regulating black spot resistance. This prompted a detailed analysis of PbrGT15. The VIGS results further confirmed its proactive role in enhancing pear resistance to black spot disease. Subsequent qRT‒PCR analyses demonstrated higher PbrGT15 expression in resistant varieties compared to susceptible ones. Moreover, since disease resistance reduced post-gene silencing, it highlighted a positive correlation between PbrGT15 and black spot resistance.
Conclusion
This study analyzed the expression of pear GT genes across different cultivars post-SA treatment, and utilized VIGS to knock out specific genes, confirming that PbrGT15 was pivotal in enhancing pear black spot resistance.
Availability of data and materials
Date will be shared upon request by the readers.
References
Ayadi M, Delaporte V, Li YF, Zhou DX. Analysis of GT-3a identifies a distinct subgroup of trihelix DNA-binding transcription factors in Arabidopsis. FEBS Lett. 2004;562:147–54. https://doi.org/10.1016/S0014-5793(04)00222-4.
Bailey TL, Williams N, Misleh C, Li WW. MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res. 2006;34:W369–73. https://doi.org/10.1093/nar/gkl198.
Bhar A, Chakraborty A, Roy A. Plant responses to biotic stress: old memories matter. Plants. 2022;11:84. https://doi.org/10.3390/plants11010084.
Brewer PB, Howles PA, Dorian K, Griffith ME, Ishida T, Kaplan-Levy RN, et al. PETAL LOSS, a trihelix transcription factor gene, regulates perianth architecture in the Arabidopsis flower. Development. 2004;131:4035–45. https://doi.org/10.1242/dev.01279.
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, et al. TBtools: An integrative Toolkit developed for interactive analyses of big biological data. Mol Plant. 2020;13:1194–202. https://doi.org/10.1016/j.molp.2020.06.009.
Chen J, Li X, Wang D, Li L, Zhou H, Liu Z, et al. Fabrice MR. Identification and testing of reference genes for gene expression analysis in pollen of Pyrus bretschneideri. Sci Hortic-Amsterdam. 2015;190:43–56. https://doi.org/10.1016/j.scienta.2015.04.010.
Dahro B, Wang F, Peng T, Liu JH. PtrA/NINV, an alkaline/neutral invertase gene of Poncirus trifoliata, confers enhanced tolerance to multiple abiotic stresses by modulating ROS levels and maintaining photosynthetic efficiency. BMC Plant Biol. 2016;76:1–18. https://doi.org/10.1186/s12870-016-0761-0.
Du H, Huang M, Liu L. The genome wide analysis of GT transcription factors that respond to drought and waterlogging stresses in maize. Euphytica. 2016;208:113–22. https://doi.org/10.1007/s10681-015-1599-5.
Fujita M, Fujita Y, Noutoshi Y, Takahashi F, Narusaka Y, Yamaguchi-Shinozaki K, et al. Crosstalk between abiotic and biotic stress responses: a current view from the points of convergence in the stress signaling networks. Curr Opin Plant Biol. 2006;9:436–42. https://doi.org/10.1016/j.pbi.2006.05.014.
Gaffney T, Friedrich L, Vernooij B, Negrotto D, Nye G, Uknes S, et al. Requirement of salicylic acid for the induction of systemic acquired resistance. Science. 1993;261:754–6. https://doi.org/10.1126/science.261.5122.75.
Gao MJ, Lydiate DJ, Li X, Lui H, Gjetvaj B, Hegedus D-D, et al. Repression of seed maturation genes by a trihelix transcriptional repressor in Arabidopsis seedlings. Plant Cell. 2009;21:54–71. https://doi.org/10.1105/tpc.108.061309.
Gao XQ, Shan LB. Functional genomic analysis of cotton genes with agrobacterium-mediated virus-induced gene silencing. Methods Mol Biol. 2013;975:157–65. https://doi.org/10.1007/978-1-62703-278-0_12.
Hu B, Jin J, Guo A-Y, Zhang H, Luo J, Gao G. GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics. 2015;31:1296–7. https://doi.org/10.1093/bioinformatics/btu817.
Jia XQ, Wang L, Zhao HY, Zhang YB, Chen ZX, Xu L, et al. The origin and evolution of salicylic acid signaling and biosynthesis in plants. Mol Plant. 2023;16:245–59. https://doi.org/10.1016/j.molp.2022.12.002.
Kachroo P, Liu H, Kachroo A. Salicylic acid: transport and long-distance immune signaling. Curr Opin Virol. 2020;42:53–7. https://doi.org/10.1016/j.coviro.2020.05.008.
Kaplan-Levy RN, Brewer PB, Quon T, Smyth DR. The trihelix family of transcription factors–light, stress and development. Trends Plant Sci. 2012;17:163–71. https://doi.org/10.1016/j.tplants.2011.12.002.
Li X, Qin G, Chen Z, Gu H, Qu L-J. A gain-of-function mutation of transcriptional factor PTL results in curly leaves, dwarfism and male sterility by affecting auxin homeostasis. Plant Mol Biol. 2008;66:315–27. https://doi.org/10.1007/s11103-007-9272-6.
Li Y, Hu Z, Dong Y, Xie Z. Overexpression of the cotton trihelix transcription factor GhGT23 in Arabidopsis mediates salt and drought stress tolerance by binding to GT and MYB promoter elements in stress-related genes. Front Plant Sci. 2023;14:1144650. https://doi.org/10.3389/fpls.2023.1144650.
Li Y, Hu Z, Dong Y, Xie Z. Trihelix Transcriptional factor GhGT26 of cotton enhances salinity tolerance in Arabidopsis. Plants. 2022;11:2694. https://doi.org/10.3390/plants11202694.
Li Q, Qiao X, Yin H, Zhou Y, Dong H, Qi K, et al. Unbiased subgenome evolution following a recent whole-genome duplication in pear (Pyrus bretschneideri Rehd.). Hortic Res-england. 2019;6:34.
Lin Z, Griffith ME, Li X, Zhu Z, Tan L, Fu Y, et al. Origin of seed shattering in rice (Oryza sativa L.). Planta. 2007;226:11–20.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods. 2001;25:402–8. https://doi.org/10.1006/meth.2001.1262.
Malamy J, Carr JP, Klessig DF, Raskin I. Salicylic acid: a likely endogenous signal in the resistance response of tobacco to viral infection. Science. 1990;250:1002–4. https://doi.org/10.1126/science.250.4983.1002.
Mao HM, Zhang WQ, Lv JY, Yang JX, Yang SX, Jia B, et al. Overexpression of cotton Trihelix transcription factor GhGT-3b_A04 enhances resistance to Verticillium dahliae and affects plant growth in Arabidopsis thaliana. J Plant Physiol. 2023;283: 153947. https://doi.org/10.1016/j.jplph.2023.153947.
Osorio MB, Bücker-Neto L, Castilhos G, Turchetto-Zolet AC, Wiebke-Strohm B, Bodanese-Zanettini MH, et al. Identification and in silico characterization of soybean trihelix-GT and bHLH transcription factors involved in stress responses. Genet Mol Biol. 2012;35:233–46. https://doi.org/10.1590/S1415-47572012000200005.
Park HC, Kim ML, Kang YH, Jeon JM, Yoo JH, Kim MC, et al. Pathogen-and NaCl-induced expression of the SCaM-4 promoter is mediated in part by a GT-1 box that interacts with a GT-1-like transcription factor. Plant Physiol. 2004;135:2150–61. https://doi.org/10.1104/pp.104.041442.
Peck S, Mittler R. Plant signaling in biotic and abiotic stress. J Exp Bot. 2020;71:1649–51. https://doi.org/10.1093/jxb/eraa051.
Qiao X, Li M, Li L, Yin H, Wu J, Zhang S. Genome-wide identification and comparative analysis of the heat shock transcription factor family in Chinese white pear (Pyrus bretschneideri) and five other Rosaceae species. BMC Plant Biol. 2015;15:1–16. https://doi.org/10.1186/s12870-014-0401-5.
Qin Y, Ma X, Yu G, Wang Q, Wang L, Kong L, et al. Evolutionary history of trihelix family and their functional diversification. DNA Res. 2014;21:499–510. https://doi.org/10.1093/dnares/dsu016.
Riechmann JL, Heard J, Martin G, Reuber L, Jiang C-Z, Keddie J, et al. Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science. 2000;290:2105–10. https://doi.org/10.1126/science.290.5499.210.
Song A, Wu D, Fan Q, Tian C, Chen S, Guan Z, et al. Transcriptome-wide identification and expression profiling analysis of chrysanthemum trihelix transcription factors. Int J Mol Sci. 2016;17:198. https://doi.org/10.3390/ijms17020198.
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997;25:4876–82. https://doi.org/10.1093/nar/25.24.4876.
Vlot A-C, Dempsey D-A, Klessig D-F. Salicylic acid, a multifaceted hormone to combat disease. Annu Rev Phytopathol. 2009;47:177–206. https://doi.org/10.1146/annurev.phyto.050908.135202.
Völz R, Kim SK, Mi J, Mariappan KG, Guo X, Bigeard J, et al. The Trihelix transcription factor GT2-like 1 (GTL1) promotes salicylic acid metabolism, and regulates bacterial-triggered immunity. Plos Genet. 2018;14: e1007708. https://doi.org/10.1371/journal.pgen.1007708.
Wang R, Hong G, Han B. Transcript abundance of rml1, encoding a putative GT1-like factor in rice, is up-regulated by Magnaporthe grisea and down-regulated by light. Gene. 2004;324:105–15. https://doi.org/10.1016/j.gene.2003.09.008.
Wang W, Withers J, Li H, Zwack P, Rusnac DV, Shi H, et al. Structural basis of salicylic acid perception by Arabidopsis NPR proteins. Nature. 2020;586:311–6. https://doi.org/10.1038/s41586-020-2596-y.
Wang Y, Tang M, Zhang Y, Huang ML, Wei L, Lin Y, et al. Coordinated regulation of plant defense and autoimmunity by paired trihelix transcription factors ASR3/AITF1 in Arabidopsis. New Phytol. 2023;237:914–29. https://doi.org/10.1111/nph.18562.
Wang D, Zhang Y, Zhang Z, Zhu J, Yu J. KaKs_Calculator 2.0: a toolkit incorporating gamma-series methods and sliding window strategies. Genom Proteom Bioinf. 2010;8:77–80.
Willmann MR, Mehalick AJ, Packer RL, Jenik PD. MicroRNAs regulate the timing of embryo maturation in Arabidopsis. Plant Physiol. 2011;155:1871–84. https://doi.org/10.1104/pp.110.171355.
Wu J, Wang Z, Shi Z, Zhang S, Ming R, Zhu S, et al. The genome of the pear (Pyrus bretschneideri Rehd.). Genome Res. 2013;23:396–408. https://doi.org/10.1101/gr.144311.112.
Xiao J, Hu R, Gu T, Han J, Qiu D, Su P, et al. Genome-wide identification and expression profiling of trihelix gene family under abiotic stresses in wheat. BMC Genomics. 2019;20:1–14. https://doi.org/10.1186/s12864-019-5632-2.
Yoo CY, Pence HE, Jin JB, Miura K, Gosney MJ, Hasegawa PM, et al. The Arabidopsis GTL1 transcription factor regulates water use efficiency and drought tolerance by modulating stomatal density via transrepression of SDD1. Plant Cell. 2010;22:4128–41. https://doi.org/10.1105/tpc.110.078691.
Yu C, Cai X, Ye Z, Li H. Genome-wide identification and expression profiling analysis of trihelix gene family in tomato. Biochem Bioph Res Co. 2015;468:653–9. https://doi.org/10.1016/j.bbrc.2015.11.010.
Zandalinas SI, Fritschi FB, Mittler R. Signal transduction networks during stress combination. J Exp Bot. 2020;71:1734–41. https://doi.org/10.1093/jxb/erz486.
Zhang QQ, Zhong T, E LZ, Xu ML, Dai WX, Sun SC, et al. GT Factor ZmGT-3b is associated with regulation of photosynthesis and defense response to Fusarium graminearum infection in maize seedling. Front Plant Sci. 2021;12:724133. https://doi.org/10.3389/fpls.2021.724133.
Zhou JM, Zhang YL. Plant immunity: danger perception and signaling. Cell. 2020;181:978–89. https://doi.org/10.1016/j.cell.2020.04.028.
Acknowledgements
Not applicable.
Funding
This work has been supported by the National Key Research and Development Program of China (2022YFD1200503), the Jiangsu Agriculture Science and Technology Innovation Fund (CX(22)3215), the Key Research and Development Program of Jiangsu Province (BE2023328), the National Science Foundation of China (32072538) and the Fundamental Research Funds for the Central Universities (JCQY201901), the Project is Funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions, the Earmarked Fund for China Agriculture Research System (CARS-28), College Students’ Innovative Entrepreneurial Training Plan Program (202210307233P) and the Agricultural Science & Technology Independent Innovation Fund of Jiangsu Province [CX(21)3024].
Author information
Authors and Affiliations
Contributions
SZ and XH designed experiments. ZZ and HX performed all bioinformatics analysis. ZZ performed all molecular biology experiments. ZZ, XX, XC and TL analyzed all data. SZ, XH and ZZ wrote the manuscript. All authors have read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests, no financial or nonfinancial interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1:
Figure S1. The ten motifs of pear trihelix transcription factor family gene (PbrGT) detected by the online tool MEME.
Additional file 2:
Table S1. Primer sequences designed using Primer Premier 6.
Additional file 3:
Table S2. Overview of the GT genes in Arabidopsis and rice.
Additional file 4:
Table S3. Ka/Ks and duplicate type analysis for the paralogous gene pairs in PbrGT genes.
Additional file 5:
Table S4. Transcriptome data of PbrGT genes in various tissues.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, Z., Xie, H., Xu, X. et al. Genome-wide identification of trihelix transcription factor family genes in pear (Pyrus bretschneideri) and functional characterization of PbrGT15 in black spot resistance. HORTIC. ADV. 1, 12 (2023). https://doi.org/10.1007/s44281-023-00018-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44281-023-00018-z