Abstract
Fruit color influences fruit quality and commodity value. Most longan (Dimocarpus longan Lour.) varieties have a yellowish-brown or grayish-yellow pericarp, and the discovery of red pericarp (RP) longan expanded the color varieties of longan fruit. Our previous research showed that the red pericarp of RP fruit was mainly caused by anthocyanin accumulation; however, its underlying regulatory mechanism remains unknown. Herein, DlMYB113, an R2R3-MYB transcription factor, was discovered by examining differentially expressed genes in two longan cultivars. Dimocarpus longan MYB (v-myb avian myeloblastosis viral) 113 (DlMYB113) expression was significantly higher in the pericarp and leaves of RP longan than in ‘Shixia’ longan. Sequence alignment analysis revealed two amino acid substitutions in the R3 domain between DlMYB113rp in RP longan and DlMYB113sx in ‘Shixia’ longan. Transient expression of DlMYB113rp significantly increased anthocyanin accumulation in tobacco leaves, whereas DlMYB113sx had negligible effect. Meanwhile, DlMYB113 overexpression promotes anthocyanin accumulation in Arabidopsis and longan calli. Site-directed mutation detection revealed divergence in DlMYB113 function when the R3 repeat 197-position base T was replaced with G, and the 317- and 318-position AT bases were replaced with GA. Our findings indicate that DlMYB113 can regulate anthocyanin production in RP longan, and three mutations in its nucleic acid sequence lead to anthocyanin accumulation, thereby developing molecular markers associated with the anthocyanin accumulation trait in RP longan. This study will facilitate early screening of longan hybrids with desirable fruit color and be significant for breeding new characteristic varieties.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Longan (Dimocarpus longan Lour.), native to southern China, is an evergreen fruit tree from Sapindaceae with significant nutritional and medicinal value (Shi et al. 2015; Zhu et al. 2019). Fruit appearance and color are essential indicators for evaluating and determining fruit quality and influencing fruit commodity value. Most longan varieties have a yellowish-brown or grayish-yellow pericarp; the discovery of red pericarp (RP) longan, a wild species from Southeast Asia, expanded the information on fruit color types and provided candidate materials for breeding new longan varieties. We previously demonstrated that the red color of the longan pericarp was mainly driven by anthocyanin accumulation, with cyanin-3-O glucoside being the primary anthocyanin component (Yi et al. 2021).
Anthocyanin is a pigment that affects fruit color formation. Several genes are required for anthocyanin production via the phenylpropanoid pathway. These genes are classified as beginning biosynthetic genes (BBGs), early biosynthetic genes (EBGs) and late biosynthetic genes (LBGs) (Liu et al. 2014). The MBW (MYB-bHLH-WD40) complex primarily regulates these structural genes (Xu et al. 2015; Araguirang and Richter 2022). R2R3-MYBs in the MBW complex have been extensively studied in various fruit trees (Martin and Paz-Ares 1997; Stracke et al. 2001; Allan et al. 2008). Moreover, anthocyanin synthesis in apples is linked to the promoter region of Malus domestica MYB (v-myb avian myeloblastosis viral) 17 (MdMYB17)-specific anthocyanin synthesis (Wang et al. 2022a, b). MdMYB90-like directly activates anthocyanin biosynthesis genes, Malus domestica chalcone synthase (MdCHS) and Malus domestica Uridine diphosphate (UDP)-glucose: flavonoid 3-O-glycosyl-transferase (MdUFGT) (Sun et al. 2021). Heterologous expression of PSMYB10.2 from plum and v-myb avian myeloblastosis viral 10 (MYB10) and MYB110 from kiwifruit can activate genes involved in tobacco anthocyanin synthesis (Fang et al. 2021a, b; Wang et al. 2022a, b). Additionally, LcMYB1 can activate Litchi chinensis Glutathione S- transferase (LcGST) expression and promote anthocyanin accumulation in vitro (Hu et al. 2016). However, whether and how the longan MYB protein regulates anthocyanin production is unclear.
Genetic mutations cause variations in anthocyanin accumulation phenotypes, which are caused by two kinds of mutations: coding and promoter region variations. Variation in the gene coding region frequently alters the three-dimensional structure of the protein, weakening or even eliminating the activity of proteins encoded by structural gene or the ability to bind to the important regulatory elements on the promoter of downstream structural genes of anthocyanin synthesis. Analysis of red and white strawberry alleles in octoploid strawberry (Fragaria × ananassa) revealed that in white strawberry, the coding region of FaMYB10 was inserted with an 8 bp ACTTATAC sequence, resulting in premature protein translation termination (Wang et al. 2020). Mutation of the G/A single nucleotide in the R3-conserved domain of turnip (Brassica rapa subsp. rapa) Brassica rapa MYB (v-myb avian myeloblastosis viral) 1a (BrPAP1a) leads to amino acid substitution, ultimately producing an all-white turnip mutant (Yang et al. 2021). Glutathione-S-transferase (GST) is an anthocyanin transporter that transports anthocyanins into vacuoles for storage and gives color to plants (Zhao et al. 2020). BSA-seq was performed on F2 populations of peaches (Prunus persica) with different flower colors; the 2 bp insertion or 5 bp deletion of PpGST in the third exon altered the reading frame and rendered the protein inactive (Lu et al. 2021).
To further elucidate the regulatory mechanism of anthocyanin biosynthesis in RP longan, an R2R3-MYB transcription factor, DlMYB113, that promotes anthocyanin accumulation was identified using transcriptomic and qPCR analyses. Subsequently, its subcellular localization was investigated, and its role in anthocyanin production was confirmed through transient transformation and overexpression. For molecular-assisted breeding, molecular markers associated with the accumulation trait of RP longan anthocyanin were developed. This study deepens our understanding of the mechanism of anthocyanin accumulation in the pericarp of RP longan, as well as the regulatory mechanism of pericarp coloration.
Materials and methods
Plant materials
RP longan and the main cultivar ‘Shixia’ (SX) longan were used as experimental materials. The mature leaves and pericarps of the two cultivars were harvested 95 d post-anthesis (DPA) for transcriptome sequencing analysis. Pericarp samples were collected during five developmental stages of RP (15, 35, 55, 75 and 95 DPA). RP mesocarp and seeds were sampled at 95 DPA. Furthermore, all samples were collected and preserved at −80°C.
Complementary DNA (cDNA) library construction and RNA sequencing
RP and SX longan pericarp (SAMN16132431 and SAMN16132432) and leaf samples were used as transcriptome sequencing samples. Transcriptome sequencing was performed by Guangzhou Gene Denovo Biotechnology Co. Ltd. (Guangzhou, China). The Hisat2 (Kim et al. 2015) was used for sequence alignment on the ‘JDB’ longan genome (Genome assembly ASM2298485v1), the StringTie (Pertea et al. 2015) assembler was used to assemble and quantify the reads. The EdgeR Bioconductor package (Robinson et al. 2010) was used to identify differentially expressed genes (DEGs). The KEGG and Swiss-Prot databases were used for annotation, TBtools and R packages clusterProfiler for analysis and visualization (Gasteiger et al. 2001; Xie et al. 2011; Chen et al. 2020; Wu et al. 2021).
Multiple sequence alignment and phylogenetic analysis
Multiple sequence alignment of MYB amino acid sequences was performed using MAFFT software (v.7.487) (Katoh and Standley 2013). A maximum likelihood (ML) phylogenetic tree was constructed using FastTree 2.1 software with a resample value of 1,000, in which JTT (Jones et al. 1992) was the best substitution model. The sequences used for the phylogenetic tree included AtMYB75 (AT1G56650), AtMYB090 (At1g66390), AtMYB113 (At1g66370), AtMYB114 (At1g66380), CsRUBY (AFB73913), AcMYB10 (MN602037), AcMYB75 (APZ74276.1), LcMYB1 (APP94121.1), MdMYB10 (ACQ45201.1), MdMYB1 (ADQ27443.1), MdMYB110a (JN711473), VvMYBA1 (BAD18977), VvMYBA2 (BAD18978), PyMYB10 (ALN66630.1), PcMYB10 (AFC88545.1), FaMYB10 (ABX79947.1), PbMYB10 (EU153577), PdmMYB10 (EU153580), EjMYB10 (EU153572), and CoMYB10 (EU153571).
Quantitative real-time PCR (qPCR) analysis
DEG expression levels in the biosynthetic pathway of anthocyanin were analyzed using qPCR. The gene-specific primers were designed using Primer 5.0 and are listed in Table S1. The reaction system was constructed using SYBR Green Master Mix (Thermo Fisher Scientific, Waltham, MA, USA). qPCR was conducted on the LightCyclerR480 system (Roche, Switzerland). The expression levels were calculated using the 2−ΔΔCT method.
Total
After weighing 0.25 g of powder from each sample, total anthocyanins were extracted in the dark at 4°C for 24 h using 10 mL of methanol: hydrochloric acid (99:1, v/v) (Tuan et al. 2015). The samples were centrifuged at 10,000 rpm for 10 min. A 1 mL supernatant was mixed with 1 mL KCL buffer (pH 1.0), and a 1 mL supernatant plus NaAc buffer (pH 4.5) was incubated at 4°C for 15 min. The absorbance was measured at 510 and 700 nm using a spectrophotometer. Three measurements were taken for each biological replicate. The relative anthocyanin content was normalized using the following formula reported by Romero et al. (2008) and Fang et al. (2021a).
Subcellular localization analysis
Subcellular localization was determined using fluorescent protein fusions after transient expression in onion epidermal cells (Nebenführ 2014). Dimocarpus longan MYB (v-myb avian myeloblastosis viral) 113rp (DlMYB113rp) and DlMYB113sx were cloned and inserted into the pCAMBIA2300_35S:GFP vector. A fresh onion was cut into 1 cm2, and the inner epidermal cells were placed in Murashige and Skoog (MS) medium for 2 d at 28°C in the dark. After 8 min of immersion in resuspended Agrobacterium tumefaciens, the cells were placed in a dark culture at 28°C for 24–72 h, and GFP expression was measured using LSM 800 laser confocal microscopy (Zeiss, Germany). A 35S:GFP vector was used as a control.
Transient expression of DlMYB113 in tobacco leaves
Transient expression assays were performed in young Nicotiana tabacum leaves cultivated in a greenhouse, as described previously (Fang et al. 2021). The target gene-containing plant expression vector was transformed into A. tumefaciens and cultured in darkness at 28°C for 2 d. GV3101 bacteria were resuspended in an infiltration buffer containing 10 mM MgCl2 and 100 μM acetosyringone, cultivated for 2 h in the dark, and injected into tobacco leaves.
Generation of DlMYB113-overexpressing plants
Flower-soaking transformation yielded transgenic overexpression lines of DlMYB113rp and DlMYB113sx driven by the 35S promoter (Zhang et al. 2006). Fresh A. tumefaciens solution (20 μL) was inoculated into 150 mL of liquid YEP medium with corresponding antibiotics, and the strain was shaken at 28°C to an OD600 of approximately 1.0. The bacteria were centrifuged at room temperature for 10 min at 6000 rpm, and the supernatant was discarded. The bacteria were suspended in a 5% sucrose solution containing 0.05% Sillwet-77. Arabidopsis thaliana inflorescences were suspended for 30 s, incubated in the dark for 24 h, and then returned to normal conditions until seeds were harvested.
Generation of transgenic longan calli
The target gene-containing plant expression vector was transformed into A. tumefaciens GV3101, which was then incubated with longan calli for 30 min and transferred to agar-solidified MS medium and cultured for 2 d in the dark. The calli were subsequently transferred to a screening medium containing 100 μM kanamycin and 500 mg/mL Timentin.
Statistical analysis
Data are means ± SD of 3 biological replicates, and the asterisks indicate statistically significant differences (*P < 0.05, **P < 0.01, and ***P < 0.001). Bars with different letters indicate significant differences at P < 0.05 by ANOVA with Tukey’s multiple comparison test.
Results
Fruit and leaf phenotype
As the picture in Fig. 1 shows, SX longan has a yellowish-brown pericarp, similar to that of most cultivars, whereas RP longan has a dark red pericarp, which is uncommon in longan cultivars. There was no anthocyanin accumulation in SX longan cotyledons, while there was some in RP longan cotyledons. However, the aril of RP longan was similar to that of ordinary longan and appeared milk-white due to a lack of pigment accumulation (Fig. 1a, b). Moreover, SX longan leaves were green with pale yellow veins (Fig. 1c), whereas RP longan leaves were dark red on the adaxial side and green on the axial side (Fig. 1d).
Transcriptome sequencing and analysis
RNA-seq data from pericarp and leaf samples of two different varieties were generated. All clean reads were mapped to longan’s reference genome, and most reads of RP (88.71%) and SX (93.78%) matched the reference genome. Principal component analysis (PCA) revealed that the twelve pericarp and leaf samples were then classified into four groups, each corresponding to two varieties, demonstrating that the gene expression data obtained in this study had good reproducibility (Fig. S1). In addition, 4985 DEGs were discovered in the transcriptomes of RP longan and SX longan leaf and pericarp (Fig. S2).
DEGs were annotated using gene ontology (GO) terms to help classify their function. Biological process was more closely related to the secondary metabolic activity occurring between the pericarp and leaf of RP and SX longans (Figs. S3 and S4). DEGs were analyzed using the eggNOG-mapper and TBtools analytic tools to identify key metabolic pathways in RP longan. The KEGG pathways (P < 0.05) are shown in Figs. S5 and S6. The pericarp upregulated DEGs were significantly enriched in cytochrome P450, phenylpropanoid biosynthesis, flavone and flavonol biosynthesis, and biosynthesis of other secondary metabolites pathway in RP longan. The upregulated DEGs in the leaves were significantly enriched in anthocyanin biosynthesis, phenylpropanoid biosynthesis, biosynthesis of other secondary metabolites, and cytochrome P450 pathway.
Based on the Swiss-Prot database annotation information, 25 anthocyanin-related structural genes were screened from the pericarp and leaf common DEGs, including two 4-coumarate CoA ligase (4CL), one Phenylalanine ammonialyase (PAL), two Chalcone synthase (CHS), one Flavonoid 3′,5′-hydroxylase (F3′5′H), six UFGT, and 13 GST (Fig. S7). In addition, 81 transcription factors were identified from DEGs, including 24 MYBs, 11 bHLH, 18 AP2/ERF, nine MADS-box, seven NACs, and 10 WRKYs (Fig. S8).
Function prediction and qPCR validation
As MYB transcription factor play important roles in anthocyanin, a phylogenetic tree of the MYB family was constructed between longan and other species. Seven DlMYBs were clustered into four subgroups of S4–S7 of Arabidopsis (Fig. S9), suggesting that these seven genes are involved in flavonoid biosynthesis. To further determine the potential function of these seven DlMYBs, a clustering analysis of anthocyanin-related MYBs from other species was performed. Only D.long035496.01 (named DlMYB113 according to the Swiss-Prot annotation) shared the most similarities with Litchi chinensis MYB (v-myb avian myeloblastosis viral) 1 (LcMYB1) of litchi (Fig. 2), which indicated that DlMYB113 may be related to anthocyanin accumulation in the longan pericarp. DlMYB113 expression levels were examined in different developmental stages of the pericarp and tissues (Fig. 3a, b, c). DlMYB113 expression levels increased first and then decreased during the pericarp pigmentation process. The expression level of DlMYB113 in different tissues (pericarp, pulp, seed and leaf) was used to confirm tissue specificity. DlMYB113 was only expressed in anthocyanin-containing tissues but not in anthocyanin-free tissues (Fig. 3c).
Cluster analysis of anthocyanin biosynthesis-related MYB genes in different species. Cluster analysis and functional prediction of MYB in different species. The phylogenetic tree relationship between DlMYB113 and MYB proteins from other plant species via Fasttree. The red letters indicate DlMYB113 protein
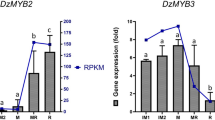
Characteristics of v-myb avian myeloblastosis viral (MYB) gene expression. (a) Total anthocyanin content in five developmental stages of fruits in red pericarp (RP) longan (G1–G5 stand for 15, 35, 55, 75 and 95 DPA, respectively). (b) The expression levels of Dimocarpus longan MYB (v-myb avian myeloblastosis viral) 113 (DlMYB113) in the pericarp during different developmental stages. (c) The expression levels of DlMYB113 in different tissues of RP longan
Sequence analysis and subcellular localization of DlMYB113
To analyze the function of DlMYB113, its coding sequence (CDS) region was cloned from the mature pericarp of RP and SX and homologously recombined into the pCAMBIA2300 expression vector, with 804 bp encoding 267 amino acids. Three SNPs were found in the conserved region of R3 in SX, resulting in amino acid variation, particularly the mutation of the third helix, which may cause a loss of MYB binding ability, which interacts specifically with DNA as a DNA recognition helix (Ogata et al. 1994). Phylogenetic analysis revealed that DlMYB113 shared more similarities with litchi LcMYB1 and Citrus sinensis MYB (v-myb avian myeloblastosis viral) (Citrus sinensis CsRUBY) Multiple sequence alignment showed that they both contained the conserved R2R3 domain (Fig. 4a). DlMYB113 from RP longan and SX longan was designated as DlMYB113rp (GenBank: OQ911501) and DlMYB113sx (GenBank: OQ911500), respectively.
Sequence alignment and subcellular localization of DlMYB113. (a) Sequence alignment of DlMYB113, LcMYB1, and CsRUBY. The R2 and R3 domains are indicated using black lines, and the red star indicates the amino acid substitution site of DlMYB113 in the conserved domains of red pericarp (RP) longan and ‘Shixia’ (SX) longan. (b) Subcellular localization of DlMYB113 in onion epidermal cells. pCAMBIA2300-35s: GFP was used as a positive control
To investigate DlMYB113 subcellular localization, recombinant (35S:DlMYB113:GFP) and control vectors were agroinfiltrated into onion epidermal cells. The control vector’s GFP fluorescence was distributed throughout the cell, whereas 35S:DlMYB113:GFP vectors displayed high fluorescence signals in the nuclei of onion epidermal cells. Therefore, DlMYB113 was likely localized and functioned in the nucleus (Fig. 4b).
Differences in DlMYB113 sequence cause variation in its function
To further validate the different function of two types DlMYB113, Agrobacterium tumefaciens GV3101 containing pCAMBIA2300:35S:DlMYB113 was transiently transformed into tobacco leaves. Transient overexpression of DlMYB113rp promoted anthocyanin accumulation in tobacco leaves while the difference in anthocyanin content between DlMYB113sx and the control was not significant (Fig. 5a, b). These results indicated that DlMYB113rp may regulate anthocyanin biosynthesis; however, DlMYB113sx did not significantly increase anthocyanin content. The inability of DlMYB113sx to bind to downstream target genes in tobacco may be related to the base mutation in its R3 conserved region, which may also explain why anthocyanins do not accumulate in the pericarp of SX longan.
Morphological characteristics and related gene expression of Dimocarpus longan MYB (v-myb avian myeloblastosis viral) 113 (DlMYB113) in tobacco leaves, Arabidopsis and Dimocarpus longan Lour. (longan) calli. (a) Phenotypic characteristics of DlMYB113 transiently transformed tobacco. pCAMBIA2300-35S was used as a positive control. (b) Total anthocyanin content in transformed leaves of tobacco. (c) Phenotypic characteristics of transgenic Arabidopsis. A, B and C represent Arabidopsis flower organs transformed into empty 35S:DlMYB113sx and 35S:DlMYB113rp vectors, respectively. EV, empty vector. (d) The expression levels of Dimocarpus longan MYB (v-myb avian myeloblastosis viral) 113 (DlMYB113) in transgenic Arabidopsis. (e) Expression of structural genes involved in anthocyanin biosynthesis in Arabidopsis overexpressing DlMYB113. (f) Phenotypic characteristics of transgenic longan calli. (g) Expression of structural genes involved in anthocyanin biosynthesis in longan calli overexpressing DlMYB113
Subsequently, the DlMYB113rp and DlMYB113sx genes were evaluated using overexpression driven by the 35S promoter in Arabidopsis. The 35S:DlMYB113rp transformants accumulated significantly more anthocyanins in petals, petioles and the anterior ends of the filaments and calyx than the control 35S:DlMYB113sx (Fig. 5c). DlMYB113 was highly expressed in 35S:DlMYB113rp and 35S:DlMYB113sx plants (Fig. 5d). Additionally, DlMYB113 overexpression upregulated the expression of structural genes involved in anthocyanin biosynthesis, including Arabidopsis thaliana chalcone synthase (AtCHS), Arabidopsis thaliana chalcone isomerase (AtCHI), Arabidopsis thaliana flavanone 3-hydroxylase (AtF3H), Arabidopsis thaliana dihydroflavonol-4-reductase (AtDFR), Arabidopsis thaliana anthocyanidin synthase (AtANS) and Arabidopsis thaliana uridine diphosphate (UDP)-glucose: flavonoid 3-O-glycosyl-transferase (AtUFGT) (Fig. 5e). Therefore, DlMYB113 can regulate anthocyanin accumulation, and the difference in sequence could be the key to anthocyanin accumulation in the RP longan pericarp.
The transformation of the longan calli confirmed that anthocyanin accumulated in RP longan due to the difference in the DlMYB113 sequence, and DlMYB113 was the key transcription factor regulating RP longan coloration (Fig. 5f). DlMYB113rp overexpression significantly increased the expression of anthocyanin biosynthesis-related structural genes (Fig. 5g), which was similar to DlMYB113rp overexpression in Arabidopsis.
Base substitution in the conserved R3 domain resulted in function variation of DlMYB113
DlMYB113’s functional characterization varies due to sequence variation in the coding region. A point mutation sequence was constructed and ligated to the plant binary expression vector pCAMBIA2300:35S to test whether amino acid substitutions in the R3 conserved structural domain of DlMYB113 were responsible for the functional variation in its coding region. DlMYB113rp base T at position 197 was substituted with G (amino acid at position 66 was replaced from methionine to arginine) and named DlMYB113−197. The bases AT at positions 317 and 318 in DlMYB113rp were replaced by GA to AT (amino acid AT position 106 was replaced from arginine to histidine) and named DlMYB113−317. Simultaneously, a double-mutant vector named DlMYB113−197/317 was constructed. DlMYB113−197 was more capable of promoting anthocyanin accumulation in tobacco leaves than DlMYB113−317, whereas DlMYB113−197/317 was consistent with DlMYB113sx, with little anthocyanin accumulation (Fig. 6a). qPCR revealed that DlMYB113−197, DlMYB113−317 and DlMYB113−197/317 treatment showed varying degrees of significant downregulation of anthocyanin structural genes compared to DlMYB113rp-treated leaves and DlMYB113−197/317 was the most significant, followed by DlMYB113−317 (Fig. 6b). Interestingly, the trend of anthocyanin structural gene expression in tobacco leaves with point mutations was consistent with the trend of total anthocyanin content (Fig. 6c).
Phenotype and gene expression of DlMYB113rp site-directed mutagenesis in tobacco. (a) Transient phenotype of tobacco with DlMYB113 site-directed mutations. The control groups in the model are DlMYB113rp and DlMYB113sx, and the point mutation sites are the bases indicated using orange arrows. (b) Tobacco anthocyanin biosynthesis-related gene expression levels in different point mutation types. (c) Total anthocyanin content of tobacco with different point mutation types. NtCHS, Nicotiana tabacum chalcone synthase; NtCHI, Nicotiana tabacum chalcone isomerase; NtF3H, Nicotiana tabacum flavanone 3-hydroxylase; NtF3′H, Nicotiana tabacum flavonoid 3'-hydroxylase; NtDFR, Nicotiana tabacum dihydroflavonol-4-reductase; NtANS, Nicotiana tabacum anthocyanidin synthase; NtUFGT, Nicotiana tabacum uridine diphosphate (UDP)-glucose: flavonoid 3-O-glycosyl-transferase
Development of functional markers for red pericarp trait screening
It is critical to develop screening markers for RP longan resource breeding based on DlMYB113 mutation variations. Therefore, a myb113arms functional marker was developed in this study based on the double mutation site (AT/GA) of DlMYB113’s R3 conserved domain (Fig. 7a and Table S1). According to the Tetra-primer ARMS-PCR primer design strategy, two bands with 716 and 253 bp in RP longan and 716 and 506 bp in SX longan could be amplified. Using the myb113arms functional marker to perform the Tetra-primer ARMS-PCR in 22 germplasm resources revealed that only the anthocyanin-accumulating RP longan could amplify a specific band of 253 bp, whereas the remaining 20 varieties could only amplify a band of 253 bp, as in SX longan (Fig. 7b). Using primer characteristics, the true hybrids of two longan strains (F1–7 and F1–8) were identified, revealing three band types of 716, 506, and 253 bp (Fig. 7c). This finding provides a functional marker for identifying favorable progeny in subsequent RP longan cross-breeding.
The development and utilization of myb113arms marker for screening various cultivars. (a) Primer design strategy for the functional marker myb113arms. Two allele-specific amplicons were generated using two pairs of primers: one (Fa and Rb) producing an amplicon representing the GA allele and the other (Fb and Ra) producing an amplicon representing the AT allele. (b) Tetra-primer ARMS-PCR was used to verify the functional markers of 22 longan germplasm resources (the full names of the varieties are listed in Table S2). Red arrows indicate the specific band of red pericarp (RP) longan. (c) Tetra-primer ARMS-PCR was used to identify the hybrid offspring of RP longan and ‘Shixia’ (SX) longan. Red arrows indicate the specific band of RP longan. Fa, Forward primer a; Ra, Reverse primer a; Fb, Forward primer b; Rb, Reverse primer b; DLMYB113sx, Dimocarpus longan MYB (v-myb avian myeloblastosis viral) 113sx
Discussion
Longan is an important tropical and subtropical fruit tree. Although there are numerous longan varieties, most cultivated varieties have yellowish-brown or grayish-yellow peels. Longan’s relatively single and dull color severely limits its diversity and market potential. The discovery of RP longan resources provided the foundation for breeding new longan varieties with bright colors. Longan is a woody fruit tree with a long breeding cycle. Elucidating the regulatory mechanism of anthocyanin synthesis in RP longan and developing red trait screening markers can facilitate early RP progeny screening and improve breeding efficiency.
The red color of red longan fruit is generated by anthocyanin accumulation, and cyanidin 3-O glucoside is the main anthocyanin component in the pericarp (Yi et al. 2021). BBGs, EBGs and LBGs regulate anthocyanin production, and most structural genes are tissue-specific (Brugliera et al. 1999; Polashock et al. 2002; Zhao et al. 2012; Gu et al. 2019). Various genes involved in the anthocyanin biosynthesis pathway were detected as DEGs between RP longan and SX longan. Two 4CL, one PAL, two CHS, one F3′5′H, six UFGT, and 13 GST genes were identified (Fig. S4). These differentially structured genes in the anthocyanin synthesis pathway provide important support for elucidating the transcriptional regulatory mechanism of anthocyanin biosynthesis in RP longan.
The MBW complex regulates anthocyanin production primarily in plants, and the former research indicated that MYB transcription factors may be involved in anthocyanin accumulation in RP longan (Yi et al. 2021). Many MYB transcription factors regulate anthocyanin biosynthesis by modulating structural gene transcription (Zhou et al. 2019). LcMYB1 expression was positively correlated with anthocyanin content (Lai et al. 2014). Moreover, LcMYB5 expression is relatively consistent in the pericarp during the coloring period of rapid anthocyanin accumulation (Lai et al. 2019). Furthermore, Arabidopsis thaliana AtMYB113, eggplant SmMYB113, and potato StMYB113 could bind to downstream target genes to promote anthocyanin production. In the present study, sequence analysis revealed that DlMYB113 differed between RP longan and SX longan (Fig. 4a). Three base mutations in R3’s conserved domain caused the substitution of two amino acids, one within the loop and one in the third helix. The transient transformation of tobacco leaves, Arabidopsis and longan calli indicated that the longan pericarp coloring was caused by a sequence variation in DlMYB113. All these results confirmed the powerful function of DLMYB113 in anthocyanin accumulation of RP longan.
Point mutations can produce new alleles and genetic functions, resulting in variations in the morphological structure of organisms (such as shape, size and color), which can provide important donor materials for molecular design breeding. Mutations in the R2 and R3 domains alter the promoter target specificity and DNA binding affinity of MYB proteins (Williams and Grotewold 1997; Heppel et al. 2013). In MYB domain studies, the third helix of the R2 and R3 repeats is considered the recognition helix and binds directly to a specific cis-regulatory element (Ogata et al. 1995; Tominaga et al. 2007). However, the first two helices of the R2 and R3 repeats play a crucial role in DNA binding affinity (Jia et al. 2014). The analysis of PpMYB10.1 and PpMYB10.2 showed that the change in two amino acids, Arg/Lys66 and Gly/Arg93, located in the R3 domain’s second helix and loop of the R3 domain, was the cause of anthocyanin accumulation and that the mutation of the R3 domain recognition helix was not the main factor affecting anthocyanin accumulation (Zhou et al. 2018). In this study, transcriptome analysis was used to identify a potential transcription factor, DlMYB113, that regulates anthocyanin biosynthesis. Point mutations reveal that the DlMYB113 sequence variation is one of the factors driving pericarp pigmentation; however, further research is warranted.
Conclusions
This study compared the published transcriptomes of RP longan and SX longan pericarp with transcriptomic data on the mature leaves of RP longan and SX longan. The structural and regulatory genes involved in anthocyanin accumulation in the pericarp were investigated. Experiments using heterologous transformation further demonstrate how DlMYB113rp, an important regulatory gene that regulates the coloring of RP longan pericarp, promotes anthocyanin accumulation in tobacco leaves, Arabidopsis and longan calli. This study provides a reference for investigating the factors that control longan fruit coloration and developing molecular markers for the anthocyanin accumulation trait.
Availability of data and materials
The RNA-Seq raw data have been deposited in the National Center for Biotechnology Information (NCBI) Sequence Read Archive (http://www.ncbi.nlm.nih.gov/sra/) under bioproject accession PRJNA663435, PRJNA943549 and PRJNA943551. All other data generated in this study are included in this published article and its supplementary information.
References
Allan AC, Hellens RP, Laing WA. MYB transcription factors that colour our fruit. Trends Plant Sci. 2008;13:99–102. https://doi.org/10.1016/j.tplants.2007.11.012.
Araguirang GE, Richter AS. Activation of anthocyanin biosynthesis in high light - what is the initial signal? New Phytol. 2022;236:2037–43. https://doi.org/10.1111/nph.18488.
Brugliera F, Barri-Rewell G, Holton TA, Mason JG. Isolation and characterization of a flavonoid 3’-hydroxylase cDNA clone corresponding to the Ht1 locus of Petunia hybrida. Plant J. 1999;19:441–51. https://doi.org/10.1046/j.1365-313x.1999.00539.x.
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant. 2020;13:1194–202. https://doi.org/10.1016/j.molp.2020.06.009.
Fang ZZ, Zhou DR, Ye XF, Jiang CC, Pan SL. Identification of candidate anthocyanin-related genes by transcriptomic analysis of ‘Furongli’ plum (Prunus salicina Lindl.) during fruit ripening using RNA-Seq. Front Plant Sci. 2021a;7:1338. https://doi.org/10.3389/fpls.2016.01338.
Fang ZZ, Wang KL, Zhou DR, Lin YJ, Jiang CC, Pan SL, et al. Activation of PsMYB10.2 transcription causes anthocyanin accumulation in flesh of the red-fleshed mutant of ‘Sanyueli’ (Prunus salicina Lindl.). Front Plant Sci. 2021b;12:680469. https://doi.org/10.3389/fpls.2021.680469.
Gasteiger E, Jung E, Bairoch A. SWISS-PROT: connecting biomolecular knowledge via a protein database. Curr Issues Mol Biol. 2001;3:47–55. https://doi.org/10.21775/cimb.003.047.
Gu Z, Zhu J, Hao Q, Yuan YW, Duan YW, Men S, et al. A novel R2R3-MYB transcription factor contributes to petal blotch formation by regulating organ-specific expression of PsCHS in tree peony (Paeonia suffruticosa). Plant Cell Physiol. 2019;60:599–611. https://doi.org/10.1093/pcp/pcy232.
Heppel SC, Jaffé FW, Takos AM, Schellmann S, Rausch T, Walker AR, et al. Identification of key amino acids for the evolution of promoter target specificity of anthocyanin and proanthocyanidin regulating MYB factors. Plant Mol Biol. 2013;82:457–71. https://doi.org/10.1007/s11103-013-0074-8.
Hu B, Zhao J, Lai B, Qin Y, Wang H, Hu G. LcGST4 is an anthocyanin-related glutathione S-transferase gene in litchi chinensis Sonn. Plant Cell Rep. 2016;35:831–43. https://doi.org/10.1007/s00299-015-1924-4.
Jia L, Clegg MT, Jiang T. Evolutionary dynamics of the DNA-binding domains in putative R2R3-MYB genes identified from rice subspecies indica and japonica genomes. Plant Physiol. 2014;134:575–85. https://doi.org/10.1104/pp.103.027201.
Jones DT, Taylor WR, Thornton JM. The rapid generation of mutation data matrices from protein sequences. Bioinformatics. 1992;8:275–82. https://doi.org/10.1093/bioinformatics/8.3.275.
Katoh K, Standley DM. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol. 2013;30:772–80. https://doi.org/10.1093/molbev/mst010.
Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015;12:357–60. https://doi.org/10.1038/nmeth.3317.
Lai B, Li XJ, Hu B, Qin YH, Huang XM, Wang HC, et al. LcMYB1 is a key determinant of differential anthocyanin accumulation among genotypes, tissues, developmental phases and ABA and light stimuli in litchi chinensis. PLoS One. 2014;9:e86293. https://doi.org/10.1371/journal.pone.0086293.
Lai B, Du LN, Hu B, Wang D, Huang XM, Zhao JT, et al. Characterization of a novel litchi R2R3-MYB transcription factor that involves in anthocyanin biosynthesis and tissue acidification. BMC Plant Biol. 2019;19:62. https://doi.org/10.1186/s12870-019-1658-5.
Liu Z, Shi MZ, Xie DY. Regulation of anthocyanin biosynthesis in Arabidopsis thaliana red pap1-D cells metabolically programmed by auxins. Planta. 2014;239:765–81. https://doi.org/10.1007/s00425-013-2011-0.
Lu Z, Cao H, Pan L, Niu L, Wei B, Cui G, et al. Two loss-of-function alleles of the glutathione S-transferase (GST) gene cause anthocyanin deficiency in flower and fruit skin of peach (Prunus persica). Plant J. 2021;107:1320–31. https://doi.org/10.1111/tpj.15312.
Martin C, Paz-Ares J. MYB transcription factors in plants. Trends Genet. 1997;13:67–73. https://doi.org/10.1016/S0168-9525(96)10049-4.
Nebenführ A. Identifying subcellular protein localization with fluorescent protein fusions after transient expression in onion epidermal cells. Methods Mol Biol. 2014;1080:77–85. https://doi.org/10.1007/978-1-62703-643-6_6.
Ogata K, Morikawa S, Nakamura H, Sekikawa A, Inoue T, Kanai H, et al. Solution structure of a specific DNA complex of the Myb DNA-binding domain with cooperative recognition helices. Cell. 1994;79:639–48. https://doi.org/10.1016/0092-8674(94)90549-5.
Ogata K, Morikawa S, Nakamura H, Hojo H, Yoshimura S, Zhang R, et al. Comparison of the free and DNA-complexed forms of the DMA-binding domain from c-Myb. Nat Struct Mol Biol. 1995;2:309–20. https://doi.org/10.1038/nsb0495-309.
Pertea M, Pertea GM, Antonescu CM, Chang TC, Mendell JT, Salzberg SL. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol. 2015;33:290–5. https://doi.org/10.1038/nbt.3122.
Polashock JJ, Griesbach RJ, Sullivan RF, Vorsa N. Cloning of a cDNA encoding the cranberry dihydroflavonol-4-reductase (DFR) and expression in transgenic tobacco. Plant Sci. 2002;163:241–51. https://doi.org/10.1016/S0168-9452(02)00087-0.
Robinson MD, McCarthy DJ, Smyth GK. EdgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26:139–40. https://doi.org/10.1093/bioinformatics/btp616.
Romero I, Teresa Sanchez-Ballesta M, Maldonado R, Isabel Escribano M, Merodio C. Anthocyanin, antioxidant activity and stress-induced gene expression in high CO2-treated table grapes stored at low temperature. J Plant Physiol. 2008;165:522–30. https://doi.org/10.1016/j.jplph.2006.12.011.
Shi SY, Li WC, Zhang HN, Liu LQ, Shu B, Liang QZ, et al. Application of extended Biologische Bundesantalt, Bundessortenamt and Chemische Industrie scale for phenological studies in longan (Dimocarpus longan): application of extended BBCH scale for phenological studies in longan. Ann Appl Biol. 2015;167:127–34. https://doi.org/10.1111/aab.12214.
Stracke R, Werber M, Weisshaar B. The R2R3-MYB gene family in Arabidopsis thaliana. Curr Opin Plant Biol. 2001;4:447–56. https://doi.org/10.1016/S1369-5266(00)00199-0.
Sun C, Wang C, Zhang W, Liu S, Wang W, Yu X, et al. The R2R3-type MYB transcription factor MdMYB90-like is responsible for the enhanced skin color of an apple bud sport mutant. Hortic Res-England. 2021;8:156. https://doi.org/10.1038/s41438-021-00590-3.
Tominaga R, Iwata M, Okada K, Wada T. Functional analysis of the epidermal-specific MYB genes CAPRICE and WEREWOLF in Arabidopsis. Plant Cell. 2007;19:2264–77. https://doi.org/10.1105/tpc.106.045732.
Tuan PA, Bai S, Yaegaki H, Tamura T, Hihara S, Moriguchi T, et al. The crucial role of PpMYB10.1 in anthocyanin accumulation in peach and relationships between its allelic type and skin color phenotype. BMC Plant Biol. 2015;15:280. https://doi.org/10.1186/s12870-015-0664-5.
Wang H, Zhang H, Yang Y, Li M, Zhang Y, Liu J, et al. The control of red color by a family of MYB transcription factors in octoploid strawberry (Fragaria × ananassa) fruits. Plant Biotechnol J. 2020;18:1169–84. https://doi.org/10.1111/pbi.13282.
Wang S, Li LX, Zhang Z, Fang Y, Li D, Chen XS, et al. Ethylene precisely regulates anthocyanin synthesis in apple via a module comprising MdEIL1, MdMYB1, and MdMYB17. Hortic Res. 2022a;9:uhac034. https://doi.org/10.1093/hr/uhac034/6532229.
Wang WQ, Moss SMA, Zeng L, Espley RV, Wang T, Lin-Wang K, et al. The red flesh of kiwifruit is differentially controlled by specific activation-repression systems. New Phytol. 2022b;235:630–45. https://doi.org/10.1111/nph.18122.
Williams CE, Grotewold E. Differences between plant and animal Myb domains are fundamental for DNA binding activity, and chimeric Myb domains have novel DNA binding specificities. J Biol Chem. 1997;272:563–71. https://doi.org/10.1074/jbc.272.1.563.
Wu T, Hu E, Xu S, Chen M, Guo P, Dai Z, et al. ClusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation (Camb). 2021;2:100141. https://doi.org/10.1016/j.xinn.2021.100141.
Xie C, Mao X, Huang J, Ding Y, Wu J, Dong S, et al. KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res. 2011;39:W316–22. https://doi.org/10.1093/nar/gkr483.
Xu W, Dubos C, Lepiniec L. Transcriptional control of flavonoid biosynthesis by MYB-bHLH-WDR complexes. Trends Plant Sci. 2015;20:176–85. https://doi.org/10.1016/j.tplants.2014.12.001.
Yang J, Song HD, Chen Y, Chen B, Kim M, Kim P, et al. A single amino acid substitution in the R2R3 conserved domain of the BrPAP1a transcription factor impairs anthocyanin production in turnip (Brassica rapa subsp. rapa). Plant Physiol Biochem. 2021;162:124–36. https://doi.org/10.1016/j.plaphy.2021.02.011.
Yi D, Zhang H, Lai B, Liu L, Pan X, Ma Z, et al. Integrative analysis of the coloring mechanism of red longan pericarp through metabolome and transcriptome analyses. J Agric Food Chem. 2021;69:1806–15. https://doi.org/10.1021/acs.jafc.0c05023.
Zhang X, Henriques R, Lin SS, Niu QW, Chua NH. Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat Protoc. 2006;1:641–6. https://doi.org/10.1038/nprot.2006.97.
Zhao ZC, Hu GB, Hu FC, Wang HC, Yang ZY, Lai B. The UDP glucose: flavonoid-3-O-glucosyltransferase (UFGT) gene regulates anthocyanin biosynthesis in litchi (Litchi chinesis Sonn.) during fruit coloration. Mol Biol Rep. 2012;39:6409–15. https://doi.org/10.1007/s11033-011-1303-3.
Zhao Y, Dong W, Zhu Y, Allan AC, Lin-Wang K, Xu C. PpGST1, an anthocyanin-related glutathione S-transferase gene, is essential for fruit coloration in peach. Plant Biotechnol J. 2020;18:1284–95. https://doi.org/10.1111/pbi.13291.
Zhou H, Liao L, Xu S, Ren F, Zhao J, Ogutu C, et al. Two amino acid changes in the R3 repeat cause functional divergence of two clustered MYB10 genes in peach. Plant Mol Biol. 2018;98:169–83. https://doi.org/10.1007/s11103-018-0773-2.
Zhou H, Lin-Wang K, Wang F, Espley RV, Ren F, Zhao J, et al. Activator-type R2R3-MYB genes induce a repressor-type R2R3-MYB gene to balance anthocyanin and proanthocyanidin accumulation. New Phytol. 2019;221:1919–34. https://doi.org/10.1111/nph.15486.
Zhu XR, Wang H, Sun J, Yang B, Duan XW, Jiang YM. Pericarp and seed of litchi and longan fruits: constituent, extraction, bioactive activity, and potential utilization. J Zhejiang Univ Sci B. 2019;20:503–12. https://doi.org/10.1631/jzus.B1900161.
Acknowledgements
Not applicable.
Funding
This work was supported by the projects from the Natural Science Foundation of Hainan Province (321MS0857), the National Natural Science Foundation of China (No.32272674), the Central Public-interest Scientific Institution Basal Research Fund for Chinese Academy of Tropical Agricultural Sciences (No. PZS2023022), the Guangdong Province "Modern Seed Industry" Research and Development Plan (No.2020B020220006), the Guangdong Modern Agricultural Industry Technical System (No.2022KJ123) and the Crop Germplasm Resources Protection (No.2130135).
Author information
Authors and Affiliations
Contributions
Y-H Huang, Y-Z Wei, and S-Y Shi conceived and designed the study; Y-H Huang and Y-Z Wei conducted experiments and analyzed data; L-Q Liu, D-B Yi, J-N Zhou, and X-W Hu provided the transcriptome analysis; Y-Y Lu, L-F Zhu, M-Y Chen and Y-C Wang generated transgenic plants; Y-H Huang wrote the manuscript; Y-Z Wei and S-Y Shi revised the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
Authors declare that they have no conflict of interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Fig. S1.
Principal component analysis (PCA) of transcriptome data of Dimocarpus longan Lour. (longan) tissue samples. (a) PCA analysis of pericarps transcriptome data. (b) PCA analysis of leaves transcriptome data. RP, red pericarp; RPL, the leaf of red pericarp longan; SX, ‘Shixia’; SXL, the leaf of ‘Shixia’ longan Fig. S2. Number of differentially expressed genes (DEGs) between RP vs SX and RPL vs SXL. RP, red pericarp; RPL, the leaf of red pericarp longan; SX, ‘Shixia’; SXL, the leaf of ‘Shixia’ longan Fig. S3. Gene ontology (GO) enrichment analysis of differentially expressed genes (DEGs) in the leaf of red pericarp longan (RPL) and the leaf of ‘Shixia’ longan (SXL). BP, biological process; CC, cellular component; MF, molecular function Fig. S4. Gene ontology (GO) enrichment analysis of differentially expressed genes (DEGs) in the leaf of red pericarp longan (RPL) and the leaf of ‘Shixia’ longan (SXL). BP, biological process; CC, cellular component; MF, molecular function Fig. S5. The function of differentially expressed genes (DEGs) is analyzed by KEGG in the leaf of red pericarp longan (RPL) and the leaf of ‘Shixia’ longan (SXL) Fig. S6 The function of differentially expressed genes (DEGs) is analyzed by KEGG in the leaf of red pericarp longan (RPL) and the leaf of ‘Shixia’ longan (SXL) Fig. S7. Heat map of structural genes related to anthocyanin biosynthesis. Red, white, blue, yellow, purple and gray represent 4-coumarate CoA ligase (4CL), Phenylalanine ammonialyase (PAL), Chalcone synthase (CHS), Uridine diphosphate (UDP)-glucose: flavonoid 3-O-glycosyl-transferase (UFGT), Flavonoid 3′,5′-hydroxylase (F3',5'H) and Glutathione S- transferase (GST) respectively Fig. S8. Heat map of related genes involved in the transcription regulation of anthocyanin biosynthesis. Blue, red, gray, yellow, white, green and purple represent v-myb avian myeloblastosis viral (MYB), basic Helix-Loop-Helix (bHLH), basic leucine zipper (bZIP), APETALA2/ethylene responsive factor (AP2/ERF), MCM, AG, DEFICIENS, SERUM response factor (MADS-box), the NAM/ATAF1/2/CUC2 (NAC) transcription factor (NAC) and WRKY (Named for its special heptapeptide conserved sequence WRKYGOK), respectively. RP, red pericarp; RPL, the leaf of red pericarp longan; SB, the pericarp of ‘Shixia’longan; SXL, the leaf of ‘Shixia’ longan Fig. S9. Cluster analysis and functional prediction of v-myb avian myeloblastosis viral (MYB) in longan and Arabidopsis. Orange, blue, green, and purple represent the four subgroups S4, S7, S5 and S6, respectively, associated with anthocyanin biosynthesis in Arabidopsis.
Additional file 2: Table S1.
Gene specific primers used in this study.
Additional file 3: Table S2.
The full names and abbreviations of 22 longan germplasm resources.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Huang, YH., Liu, LQ., Yi, DB. et al. DlMYB113 mutation affects anthocyanin accumulation in red pericarp longan (Dimocarpus longan Lour.). HORTIC. ADV. 1, 11 (2023). https://doi.org/10.1007/s44281-023-00014-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44281-023-00014-3