Abstract
Assimilatory nitrate reduction and methane (CH4) oxidation by bacteria play important roles in carbon (C) and nitrogen (N) biogeochemical cycles. Here, an investigation of enzymatic assimilatory nitrate reduction and CH4 oxidation by Methylobacter sp. YHQ from the wetlands is presented, specifically concentrating on N and oxygen (O) isotope fractionation with various initial nitrate and oxygen concentrations. The N enrichment factors (15 ε assimilation) increased from 4.2 ± 0.7‰ to 6.9 ±1.3‰ and the O isotope enrichment factors (18 ε assimilation) increased from 2.7 ± 0.9‰ to 4.7 ± 0.8‰ during nitrate assimilation when initial nitrate concentrations increased from 0.9 mM to 2 mM. Similar 18 ε and 15 ε values were observed at different oxygen concentrations. The values of 18 ε and 15 ε provided vital parameters for the assessment of assimilatory nitrate reduction via the Rayleigh equation approach. The ratios of O and N isotope enrichment factors (18 ε:15 ε)assimilation ranged from 0.64 ± 0.15 to 0.74 ± 0.18 during nitrate assimilation by Methylobacter sp. YHQ with Nas, which were different from (18 ε:15 ε)assimilation for assimilatory eukaryotic nitrate reductase (eukNR) from literature data. Thus, N and O isotope fractionation could be useful tools to distinguish eukNR from Nas during nitrate assimilation. Additionally, the rates of CH4 oxidation and nitrate reduction were evaluated with a reaction-based kinetic model, and it quantitatively described the enzymatic reactions of nitrate assimilation. Combining dual N-O isotope analysis with kinetic modeling provides new insights into the microbially driven C-N interactions.
Graphical Abstract

Highlights
• Effects of nitrate and oxygen concentrations on isotope fractionation were revealed.
• The (18 ε:15 ε)assimilation ratios induced by freshwater bacteria were 0.64 ± 0.15–0.74 ± 0.18.
• Dual N-O isotopes are useful approaches for distinguishing Nas from eukNR.
• The rates of CH4 oxidation and nitrate reduction can be predicted using a kinetic model.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Enzymatic nitrate reduction is a major pathway that controls biogeochemical N cycling, which involves dissimilatory and assimilatory reduction (Rivett et al. 2008). During dissimilatory reduction, nitrate is utilized as an electron acceptor for metabolic energy generation and respiration. During assimilation, nitrate is utilized as an N source into the biomass for growth. Nitrate assimilation occurs widely in bacteria, fungus, algae, and plants (Crawford and Arst Jr 1993; Lin and Stewart 1997). Assimilatory nitrate reduction contributes to the N flux in marine and freshwater ecosystems of 2000 and 330 Tg.yr-1, respectively (Kuypers et al. 2018).
N and O isotopes are useful tools for differentiating sources, and evaluating biogeochemical transformation pathways of contaminant nitrate (Li et al. 2023). Because heavier isotopes (e.g., 15N and 18O) react slower than lighter ones (e.g., 14N and 16O) during assimilatory nitrate reduction, the remaining nitrate becomes progressively enriched with heavy isotopes (Granger et al. 2004). N and O isotope fractionation is unaffected by physical processes (e.g., dilution). Thus, isotope fractionation analysis provides the benefit of assessing the extent of nitrate reduction over traditional concentration measurements (Chen et al. 2020). Numerous experimental factors (e.g., light intensity, nitrate/iron concentrations, and growth rate) on N isotope fractionation during nitrate assimilation by different marine phytoplankton have been studied (Granger et al. 2004; Joseph et al. 2003; Pennock et al. 1996; Wada and Hattori 1978). Granger et al. (2004) and Wada and Hattori (1978) suggested that nitrate concentration was an important factor controlling N isotope fractionation, whereas Pennock et al. (1996) reported that N isotope fractionation was independent of nitrate concentrations during assimilatory nitrate reduction by marine diatoms. Meanwhile, microbial nitrate reduction, including nitrate uptake rates and enzymatic reduction rates of bacteria, may increase with elevated oxygen concentrations (Zhang et al. 2020). Changes in nitrate and oxygen concentrations can greatly affect nitrate transfer processes before bond breakage and may further affect isotope fractionation (Granger et al. 2004). The effects of nitrate and oxygen concentrations on isotope fractionation during nitrate assimilation by freshwater bacteria remain unclear.
The ratios of O and N isotope enrichment factors (18 ε:15 ε) have long been utilized as an invaluable tool for estimating global N budget (Brandes and Devol 1997). Dual isotopic tracers provide complementary information on co-occurring biogeochemical N transformations that cannot be unveiled by single-isotope measurements alone (Chen et al. 2023). Assimilatory nitrate reductase includes a prokaryotic nitrate reductase (Nas) for bacteria and eukaryotic nitrate reductase (eukNR) for fungi, algae, and plants (Stolz and Basu 2002). Various studies of N-O isotope fractionation during nitrate assimilation by marine and freshwater eukaryotes with eukNR under various environmental conditions have been reported (Granger et al. 2010; Granger et al. 2004; Karsh et al. 2014; Treibergs and Granger 2017). In culture experiments, the (18 ε:15 ε)assimilation ratios are close to 1.0, during nitrate assimilation with eukNR by marine eukaryotes (e.g., marine phytoplankton Thalassiosira weissflogii) or freshwater eukaryotes (e.g., freshwater diatoms Chlorella vulgaris and Chlorella pyrenoidosa, fungus Aspergillus niger, yeast Pichia angusta, and plant Arabidopsis thaliana) (Granger et al. 2010; Granger et al. 2004; Karsh et al. 2012; Treibergs and Granger 2017). A distinct (18 ε:15 ε)assimilation ratio of approximately 2.0 was found during nitrate assimilation by the marine bacterium Alteromonas macleodii Jul88 with Nas (Granger et al. 2010). The (18 ε:15 ε)assimilation ratios during nitrate assimilation by freshwater bacteria with Nas are still unknown.
While dual isotopes provide complementary information on co-occurring biogeochemical N transformations, it is still a challenge for quantitatively assessing the elementary reaction steps involved in assimilatory nitrate reduction. Kinetic modeling is a powerful tool to reveal multi-mechanisms of dominant reactions (Chen et al. 2023). The contributions for chemical and microbial processes during microbially driven nitrate reduction accompanied with Fe(II) oxidation can be evaluated using kinetic modeling (Liu et al. 2019). According to our understanding of assimilatory nitrate reduction, the equations containing elementary reactants can be established. The rate constants of all the dominant reactions and characteristics of microbial nitrate assimilation and substrate consumption can be obtained via the kinetic model, and the rates of assimilatory nitrate reduction and substrate consumption can be further predicted. Quantitative descriptions of nitrate assimilation are still lacking.
In the present study, Methylobacter sp. YHQ isolated from a freshwater wetland was chosen as a typical strain for assimilatory nitrate reduction and aerobic methane (CH4) oxidation with widespread distribution in freshwater ecosystems (Hao et al. 2020). Microbial aerobic CH4 oxidation serves as a vital CH4 removal pathway in wetlands and other ecosystems (He et al. 2019). During assimilatory nitrate reduction and aerobic CH4 oxidation process, CH4 serves as the carbon source and electron donor, whereas nitrate serves as the nitrogen source for methanotrophs. The overall goals were: (1) to systematically investigate the effect of initial nitrate and oxygen concentrations on N-O isotope fractionation during nitrate assimilation accompanying aerobic CH4 oxidation in laboratory experiments, (2) to test whether assimilatory prokaryotic Nas can be distinguished from eukaryotic nitrate reductase eukNR by N-O isotopes, and (3) to develop a kinetic model for finely describing nitrate assimilation and CH4 oxidation.
2 Materials and methods
2.1 Culturing
The methanotrophic strain, Methylobacter sp. YHQ was cultured in a modified medium of nitrate mineral salts (Puri et al. 2015), which contains 0.14 g L-1 CaCl2·6H2O, 0.2 g L-1 MgSO4·7H2O, and 1.0 g L-1 KNO3 at pH 6.8. The strains were then transferred into 125 mL serum bottles with 24 mL medium. The bottles were sealed with butyl rubber stoppers and aluminum caps and autoclaved at 121℃ for 20 min. A 5% phosphate solution containing 5.44 g L-1 KH2PO4 and 5.68 g L-1 Na2HPO4, and a 0.2% trace element solution were supplemented. The concentrations of trace metals are listed in Table S1. CH4 (25%) in the headspace (approximately 9.60 mM) was spiked into bottles as the substrate, and the strains were cultured in the dark at 30 ℃ for three days until the exponential growth phase was reached.
2.2 CH4 monooxygenase and nitrate reductase measurements
The 16S rRNA gene sequence of the Methylobacter sp. YHQ was uploaded to the NCBI website (No. MT242555). Sequences of closely related methanotroph taxa were retrieved from the GenBank database using BLASTN (McGinnis and Madden 2004). A neighbor-joining phylogenetic tree was built using MEGA 7.0. Furthermore, the key functional genes of assimilatory and dissimilatory nitrate reductase were measured to determine the nitrate reductase type of strain YHQ. Genomic DNA of strain YHQ was extracted by TIANamp Bacteria DNA Kit (Tiangen, Beijing, China). Qualitative and quantitative detection of DNA extraction effect was performed by agarose gel electrophoresis and a Nanodrop ND-100 spectrophotometer. Subsequently, the DNA template (1 μL) was mixed with 10 μL of Sangon Biotech® Taq PCR Mix (2×), 1 μL primers (10 μM), and 8 μL RNase-free dH2O. Primers for the particulate CH4 monooxygenase (pmoA) gene (Lin et al. 2005), soluble CH4 monooxygenase (mmoX) gene (Hutchens et al. 2004), periplasmic nitrate reductase (napA) gene (Flanagan et al. 1999), and membrane-bound nitrate reductase (narG) gene (Pastorelli et al. 2013) are listed in Table S2. Specific primers for the assimilatory nitrate reductase (nasA) gene were designed using conserved regions to target homologous genes in Methylobacter spp. Shewanella oneidensis MR-1 with periplasmic nitrate reductase (nap) and Klebsiella pneumoniae L17 with membrane-bound nitrate reductase (nar) genes were used as positive controls (Gao et al. 2009; Liu et al. 2014). The nitrate reductase genomes of S. oneidensis MR-1 contain napA (Gao et al. 2009), whereas that of K. pneumoniae L17 include narG (Liu et al. 2014). The strains were used as positive controls to demonstrate that the dissimilatory nitrate reductase gene can be detected under our experimental conditions when the strain contains nap or nar. Culture conditions of S. oneidensis MR-1 and K. pneumoniae L17 and PCR reaction conditions of genes for CH4 monooxygenase and nitrate reductase were shown in the Text S1 of Supplementary material. The PCR products of nasA were sequenced by SanGon Biotech (Shanghai, China). The acquired sequences were blasted on the NCBI and KEGG websites to verify functions.
2.3 Experimental procedures
Using modified NMS without nitrate, the strain YHQ was centrifuged (8,000 ×g, 5 min) to separate it from the medium. Different experiments were performed under varying concentrations of nitrate and air in 125 mL serum bottles with 24 mL modified NMS without nitrate. Three treatments with 25% CH4 in the headspace were conducted with 4% inoculum under the conditions of 2 mM nitrate, 75% air; 0.9 mM nitrate, 75% air; 2 mM nitrate, 40% air, and 35% helium. By comparing the treatments (2 mM nitrate / 25% methane / 75% air) and (2 mM nitrate / 25% methane / 40% air), the effect of initial oxygen concentrations on isotope fractionation can be revealed. By comparing the treatments (2 mM nitrate / 25% methane / 75% air) and (0.9 mM nitrate / 25% methane / 75% air), the effect of initial nitrate concentrations on isotope fractionation can be determined. Different initial dissolved oxygen concentrations were achieved by adjusting air volume in the culture system. For the treatment (2 mM nitrate / 25% methane / 75% air), the same volume (25 mL) of air was removed before injecting CH4 (25 mL, 25%) using a sterile syringe. For the treatment (2 mM nitrate / 25% methane / 40% air), 60 mL of the air was withdrawn and reinjected with 25 mL (25%) of methane and 35 mL (35%) of helium to balance the gas pressure. Helium is an inert gas and is not utilized by bacteria (Wang et al. 2023). All the experiments were performed under static and dark conditions at 30 °C in duplicate, and experiments in the absence of cells were conducted as a negative control. A destructive sampling method was used in this study, which means that serum bottles were prepared for sampling at each given time intervals, and these serum bottles were used only once in the experiments. Headspace (0.2 mL) was drawn into syringes and injected into a gas chromatograph (Agilent 7890 B, USA) to measure concentrations of CH4 and CO2 (Hao et al. 2020). The possibility of CH4 dissolution in the medium can be ignored (Xiao et al. 2020). The dissolved oxygen was determined by a microelectrode (Unisense, Denmark). The OD600 values (optical density at 600 nm wavelength) reflect the bacterial density, which were determined by a UV-vis spectrophotometer (UV1102II, Techcomp Ltd., China). After filtering through 0.22 μm filters, nitrate concentrations of samples were determined by ion chromatography (Dionex ICS-600, USA) (Chen et al. 2023). CH4 oxidation and nitrate reduction rates were defined as V= ΔC/Δt, where ΔC is the change in CH4 or nitrate concentration over the time interval Δt.
2.4 Isotope analysis
N and O isotopic compositions (δ 15N and δ 18O) were determined by the denitrifyer method (Sigman et al. 2001). Nitrate was quantitatively converted to N2O gas by Pseudomonas chlororaphis subsp. aureofaciens ATCC 13985. Without N2O reductase, these bacteria do not transform N2O into N2. The δ 15N and δ 18O values were measured using an IsoPrime continuous flow isotope ratio mass spectrometer (CF-IRMS) equipped with trace gas preconcentrator unit (IsoPrime, GV, UK) and a Gilson GX271 autosampler. For calibration, the international reference materials IAEA-N3, USGS-35, USGS-34, and USGS-32 as well as two laboratory reference materials were utilized. The standard deviations (SDs) of replicate measurements for reference materials and samples were 0.3‰ for δ 15N and 0.5‰ for δ 18O. R reactant and R product are the isotope ratios of the reactant and product, respectively. The magnitude of isotope fractionation is represented as isotope fractionation factor (α). The δ 15N and δ 18O values of the residual nitrate were matched to Rayleigh fractionation equation to determine the α values (where α = R reactant / R product) (modified from Mariotti et al. 1981). For convenience, the α values are converted to enrichment factors (ε) for N isotopes (15 ε) and ε values for O isotopes (18 ε) (where 15 ε = [15 α–1] × 1000; 18 ε = [18 α–1] × 1000) (Mariotti et al. 1981). The value and uncertainty of ε were determined by Isoplot (version 4.15) (Chen et al. 2020).
2.5 Kinetic models
The program KinTek Explorer 6.5.1 was used to fit kinetic models to the kinetic data in our experiments (Johnson et al. 2009) and was established for description of the experimental data of nitrate assimilation and CH4 oxidation under various experimental conditions.
3 Results
3.1 Identification of genes encoding for assimilatory nitrate reduction and CH4 oxidation in Methylobacter sp. YHQ
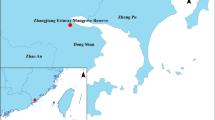
The strain in this study was isolated from wetland soil in the Yellow River Delta of China and identified as Methylobacter sp. YHQ in our previous study (Hao et al. 2020). The 16S rRNA gene sequence analysis for pure culture representatives placed strain YHQ in a clade with Methylobacter marinus A45 from the closely-related species Methylobacter tundripaludum SV96 and Methylobacter psychrophilus Z-0021 (Fig. S1).
CH4 monooxygenase enzymes (MMOs) are responsible for the catalysis of CH4 oxidation and convert CH4 to methanol, which is the first step of CH4 oxidation (Koo and Rosenzweig 2021). Two types of MMOs have been identified, particulate CH4 monooxygenase (pMMO) and soluble CH4 monooxygenase (sMMO) (Koo and Rosenzweig 2021). Analysis of CH4 monooxygenase (pmoA and mmoX) genes of Methylobacter sp. YHQ (Fig. 1a) indicated that Methylobacter sp. YHQ had the pmoA gene (Fig. 1a, lane 1) rather than the mmoX. Negative controls were prepared using the respective primers without a DNA template (Fig. 1a, lanes 2 and 4). Analyses of the nasA, napA, and narG genes of Methylobacter sp. YHQ (Fig. 1b) suggested that Methylobacter sp. YHQ contained nasA (Fig. 1b, lane 1) rather than napA or narG genes. S. oneidensis MR-1 (Fig. 1b, lane 4) and K. pneumoniae L17 (Fig. 1b, lane 7) were used as the positive controls. The nitrate reductase genes of S. oneidensis MR-1 were shown to be napA, whereas the nitrate reductase genes of K. pneumoniae L17 were shown to be narG (Gao et al. 2009; Liu et al. 2014). Negative controls (Fig. 1b, lanes 2, 5, and 8) were prepared using the respective primers without the DNA template. Correspondingly, the amplified sequences of Methylobacter sp. YHQ by nasA primers had the highest score with nasA, which codes for the assimilatory nitrate reductase catalytic subunit (nasA EC:1.7.99.-) by BLAST analysis on the KEGG website. Therefore, we conclude that nitrate reductase in Methylobacter sp. YHQ is Nas, and nitrate reduction by Methylobacter sp. YHQ is assimilatory nitrate reduction.
a Analysis of CH4 monooxygenase (pmoA and mmoX) genes of Methylobacter sp. YHQ by agarose gel electrophoresis of DNA fragments generated in PCR reactions. Reactions were performed with total DNA isolated from Methylobacter sp. YHQ (lanes 1 and 3). Negative controls (lanes 2 and 4) were operated using respective primer without DNA template. b Analysis of nitrate reductase (nasA, napA, and narG) genes of Methylobacter sp. YHQ by agarose gel electrophoresis of DNA fragments generated in PCR reactions. Reactions were performed with total DNA isolated from Methylobacter sp. YHQ (lanes 1, 3 and 6), Shewanella oneidensis MR-1 (lane 4), and Klebsiella pneumoniae L17 (lane 7). Negative controls (lanes 2, 5, and 8) were operated using respective primer without DNA template. Lane M: marker
3.2 Kinetics and isotope fractionation under different nitrate concentrations
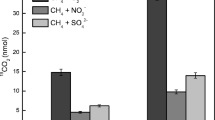
Effects of the nitrate concentration on kinetics and isotope fractionation of assimilatory nitrate reduction and CH4 oxidation were investigated under a CH4 concentration of 25% (9.6 mM). The average rate of assimilatory nitrate reduction was 0.35 mM/day with 2 mM initial nitrate, and 0.23 mM/day with 0.9 mM initial nitrate during the first four days, whereas nitrate reduction was not observed in the no-cells control (Fig. 2a). Nitrite and ammonium were not detected throughout the process. Changes in nitrate concentration affect CH4 oxidation. The rates of CH4 oxidation with 2 and 0.9 mM initial nitrate were 1.1 and 1.0 mM/day, respectively (Fig. 2b), indicating that CH4 oxidation occurred simultaneously with assimilatory nitrate reduction. Dissolved oxygen was consumed from 0.221 to 0.08 mM with 2 mM nitrate and from 0.212 to 0.06 mM with 0.9 mM nitrate (Fig. 2d), indicating that oxic conditions gradually changed to hypoxic conditions. The concentrations of CO2 generated by Methylobacter sp. YHQ, under the condition of 2 mM and 0.9 mM initial nitrate in five days, were 1.46 and 1.42 mM, respectively (Fig. 2c). Much more CH4 was consumed than CO2 was produced. The ratios of CH4 oxidized to CO2 produced were 3.7:1 and 3.5:1 under the conditions of 2 mM and 0.9 mM initial nitrate, respectively. Methanotrophs utilize CH4 as an electron donor and a carbon source. Methylobacter sp. YHQ incorporated 73% and 72% CH4-derived carbon into the biomass under the condition of 2 mM and 0.9 mM initial nitrate, respectively. The OD600 value increased to 0.30 and 0.40 with 0.9 mM and 2 mM nitrate during 5 d incubations, respectively (Fig. S2), indicating that higher nitrate concentrations are conducive to bacterial growth.
a Assimilatory nitrate reduction as a function of time in the presence of 2 mM nitrate / 75% air, 2 mM nitrate / 40% air, and 0.9 mM nitrate / 75% air; (b) the concentration of CH4 in the presence of 2 mM nitrate / 75% air, 2 mM nitrate / 40% air, and 0.9 mM nitrate / 75% air; (c) the concentration of CO2 in the presence of 2 mM nitrate / 75% air, 2 mM nitrate / 40% air, and 0.9 mM nitrate / 75% air; (d) the concentration of dissolved oxygen in the presence of 2 mM nitrate / 75% air, 2 mM nitrate / 40% air, and 0.9 mM nitrate / 75% air
The value of δ 15N in the remaining nitrate increased as assimilatory nitrate reduction proceeded and followed a Rayleigh fractionation model. With 67% nitrate reduction of 2 mM nitrate, the value of δ 15N of the residual nitrate increased from –0.9‰ to 8.4‰ (Fig. 3a), and the 15 ε assimilation value obtained from the experiments was 6.9 ± 1.3‰. For the 0.9 mM initial nitrate experiments, the value of δ 15N of the residual nitrate increased from –0.8‰ to 4.2‰ with 77% nitrate reduction, and the 15 ε assimilation value was 4.2 ± 0.7‰ (Fig. 3b). These results indicated that the values of 15 ε assimilation decreased with decreasing nitrate concentrations. The values of 18 ε assimilation were 4.7 ± 0.8‰ with 2 mM initial nitrate, and 2.7 ± 0.9‰ with 0.9 mM initial nitrate (Table 1), respectively, indicating that the 18 ε assimilation values decreased with decreasing nitrate concentrations as well as 15 ε assimilation. The δ 18O values in the remaining nitrate followed the Rayleigh equation trend. The ratios of (18 ε:15 ε)assimilation were 0.70 ± 0.05 for 2 mM nitrate and 0.64 ± 0.15 for 0.9 mM nitrate (Fig. 3c).With the δ 18O values of 24.3 ± 0.5‰ for nitrate at the beginning and the δ 18O values of 24.2 ± 0.5‰ at the end in the no-cell treatments, O isotope exchange between water and nitrate can be neglected (Chen et al. 2020).
a The N isotopic composition (δ 15N) vs. fraction of assimilatory nitrate reduction with 2 and 0.9 mM initial nitrate; (b) the O isotopic composition (δ 18O) vs. fraction of assimilatory nitrate reduction with 2 and 0.9 mM initial nitrate; and (c) the ratios of (18 ε:15 ε)assimilation with 2 and 0.9 initial nitrate
3.3 Kinetics and isotope fractionation with different oxygen concentrations
The rates of nitrate reduction were 0.35 and 0.24 mM/day under the conditions of 75% air and 40% air in the headspace with a dissolved oxygen concentration of 0.221 mM and 0.165 mM, respectively (Fig. 2a), indicating that the assimilatory nitrate reduction rate increased with higher oxygen concentration. Nitrite was not detected during the incubation period. The rates of CH4 oxidation were 1.1 and 0.87 mM/day under the conditions of 75% air and 40% air (Fig. 2b), respectively. The concentration of dissolved oxygen was reduced from 0.221 to 0.08 mM under the condition of 75% air, while it was reduced from 0.164 to 0.07 mM under the condition of 40% air (Fig. 2d). Therefore, Methylobacter sp. YHQ is sensitive to changes in oxygen concentration, which strongly affects oxygen bioavailability and CH4 utilization by methanotrophs (Mancinelli 1995). The CO2 concentrations in the headspace generated by Methylobacter sp. YHQ were 1.46 mM and 1.19 mM, respectively, in the presence of 75% air and 40% air (Fig. 2c). Based on the mass balance of C, the CH4 utilized by Methylobacter sp. YHQ is either converted to CO2 or transformed into biomass (van Grinsven et al. 2021). OD600 increased from 0.01 to 0.40 under the condition of 75% air while that increased from 0.01 to 0.24 under the condition of 40% air (Fig. S2), indicating that higher oxygen concentrations can promote bacterial growth.
Although the assimilatory nitrate reduction rates were quite different under the two oxygen headspace conditions, the values of 15 ε assimilation were similar under the conditions of 75% air (6.9 ± 1.3‰) and 40% air (6.5 ± 1.4‰) (Fig. 4a), and the values of 18 ε assimilation were also similar at 75% air (4.7 ± 0.8‰) and 40% air (4.8 ± 1.3‰) (Fig. 4b). The ratios of (18 ε:15 ε)assimilation were 0.70 ± 0.05 with 75% air and 0.74 ± 0.18 with 40% air, respectively (Fig. 4c).
(a) The N isotopic composition (δ 15N) vs. fraction of assimilatory nitrate reduction in the presence of 75% air and 40% air; (b) the O isotopic composition (δ 18O) vs. fraction of assimilatory nitrate reduction in the present of 75% air and 40% air; and (c) the ratios of (18 ε:15 ε)assimilation in the present of 75% air and 40% air
4 Discussion
4.1 Effects of nitrate and oxygen concentrations on isotope fractionation
Nitrate assimilation begins with nitrate uptake into the cell. Nitrate is then reduced to nitrite in a two-electron reaction by assimilatory nitrate reductase, and is finally converted to organism-level N (Luque-Almagro et al. 2011). Effect of nitrate concentrations on isotope fractionation is caused by nitrate-transfer limitation and the ratio of cellular nitrate transport efflux to flux (Fig. 5; Kritee et al. 2012). Specifically, intrinsic kinetic isotope fractionation is caused by an isotopically sensitive reaction step, which is N-O bond cleavage (Karsh et al. 2012). The nitrate-transfer processes prior to bond breaking are isotopically insensitive steps with minimal isotope fractionation (Chen et al. 2020). However, when the nitrate-transfer processes gradually turn into rate-limiting steps, the observed isotope fractionation is substantially less pronounced (Chen et al. 2023), which is the “masking effect”. The nitrate-transfer processes of microbial assimilatory nitrate reduction in this study included nitrate diffusion, nitrate uptake via the cell envelope, and binding to Nas. Thus, the 15 ε assimilation and 18 ε assimilation values were determined by the ratio of nitrate transport efflux to flux (Fig. 5; Grange et al. 2004).
Schematic of isotope fractionation during assimilatory nitrate reduction. 15 ε assimilation and 18 ε assimilation values were determined by the ratio of nitrate transport efflux (green arrow) to flux (blue arrow). The arrow thickness represents relative rates of flux and efflux. During high ratio of efflux to flux, the isotope fractionation was less masked by mass-transfer limitation and the values of 15 ε assimilation and 18 ε assimilation were higher (a); during low ratio of efflux to flux, the isotope fractionation was more masked by mass-transfer limitation and the values of 15 ε assimilation and 18 ε assimilation were lower (b)
The values of 15 ε assimilation decreased when nitrate concentration decreased (Table 1), which is consistent with previous studies (Kritee et al. 2012). At higher concentration, the ratio of nitrate molecules involved in the enzyme-substrate complexes to the free nitrate molecules decreased. Thus, more nitrate molecules may diffuse back into the solution. N isotope fractionation was less suppressed by nitrate-transfer limitation. When the nitrate concentration decreased, the bioavailable nitrate in the solution decreased, whereas the amount of nitrate consumption was still relatively high. Therefore, the ratio of the enzyme-substrate complexes to the free nitrate molecules increased, and fewer nitrate molecules may return to the solution (Kritee et al. 2012). As a result, isotope fractionation was substantially masked by nitrate-transfer limitations and became less pronounced.
The changes in oxygen concentrations had a limited effect on 15 ε assimilation and 18 ε assimilation values, suggesting no extra masking effect on isotope fractionation. Higher oxygen concentrations may increase the rate of bond cleavage whereas the rate of mass transfer may increase as well (Kritee et al. 2012). Therefore, the ratio of nitrate transport efflux to flux remained unchanged, leading to similar values of 15 ε assimilation and 18 ε assimilation despite varying oxygen concentrations (Treibergs and Granger 2017).
4.2 Distinguishing Nas from eukNR during assimilatory nitrate reduction
The ratios of (18 ε:15 ε)assimilation were similar under the conditions of different initial nitrate or air concentrations. Similar ratios of (18 ε:15 ε)assimilation at different nitrate concentrations indicate that, although 15 ε assimilation or 18 ε assimilation declined with decreasing nitrate concentration, the kinetic isotope effect was suppressed to the same extent by mass-transfer limitations and varied concurrently. Hence, compared to the single isotope, the dual isotopes were significantly less impacted by mass-transfer limitations.
N-O isotopes are useful approaches for distinguishing Nas from eukNR during nitrate assimilation (Granger et al. 2010; Treibergs et al. 2007). In this study, the ratios of (18 ε:15 ε)assimilation induced by freshwater bacteria with Nas ranged from 0.64 ± 0.15 to 0.74 ± 0.18. The ratios of (18 ε:15 ε)assimilation induced by freshwater eukaryotes with eukNR were nearly 1.0 (Granger et al. 2010; Karsh et al. 2012; Treibergs and Granger 2017), implying that N isotope fractionation changed concomitantly with O. The eukaryotes tested in previous studies included freshwater diatoms, fungus, and plants (Fig. 6). Therefore, the (18 ε:15 ε)assimilation ratios induced by eukNR in previous studies were distinct from those induced by Nas in freshwater bacteria (Fig. 6). Binding to the enzymes belongs to nitrate transfer processes because molecules of nitrate need to go through a channel to reach the active site of Nas before bond breakage takes place (Chen et al. 2020). The structures of Nas and eukNR are different (Stolz and Basu 2002). EukNR is a member of sulfite oxidase family of mononuclear molybdenum enzymes, whereas Nas is a member of dimethylsulfoxide reductase family (Stolz and Basu 2002). EukNR contains the molybdenum cofactor, flavin adenine dinucleotide, and a b-type heme, whereas Nas comprises a molybdenum cofactor along with an Fe-S cluster (Stolz and Basu 2002). Different structures and binding affinities between Nas and eukNR may subsequently influence isotope fractionation and cause distinct (18 ε:15 ε)assimilation ratios. Therefore, N-O isotopes could be promising methods to distinguish eukNR from Nas during assimilatory nitrate reduction. Additionally, the ratios of (18 ε:15 ε)assimilation during nitrate assimilation by freshwater bacteria in this study were distinct from those observed in a common marine bacterium Alteromonas macleodii Jul88 in Granger et al. (2010). To date, this (18 ε:15 ε)assimilation ratio of 2.0 was the only N-O isotopic data during nitrate assimilation by marine bacteria. Further investigations are needed to explore the differences in (18 ε:15 ε)assimilation ratios between freshwater and marine systems.
Compilation of known ratios of (18 ε:15 ε)assimilation by freshwater eukaryote with eukNR and bacteria with Nas. Triangle above and inverted triangle below the box plots correspond to the maximum and minimum, respectively. Diamond represents the average values. The horizontal lines in the top, middle, and bottom correspond to the 75% percentile, median, and 25% percentile, respectively. The (18 ε:15 ε)assimilation ratios of Nas were obtained from this study. The (18 ε:15 ε)assimilation ratios of eukNR were literature data collected from Granger et al. (2010), Karsh et al. (2012), and Treibergs and Granger (2017)
4.3 Kinetic modeling of assimilatory nitrate reduction and CH4 oxidation
Since it is difficult to follow the kinetics of every elementary reaction contained in microbial assimilatory nitrate reduction and CH4 oxidation, simplified reactions that encompassed the primary reactions were utilized in this study. Similar kinetic models to quantitatively describe the elementary reactions during multi-reactants and multi-mechanisms were also used in our previous studies (e.g., Chen et al. 2023; Liu et al. 2019). Firstly, O2 in the air was converted into dissolved oxygen before dissolved oxygen was used by Methylobacter (R1). In the presence of dissolved oxygen, CH4 was presented as carbon source to generate more bacteria and Nas (R2). According to the previous discussion, the oxidized Nas (Nasox) was reduced to the reduced Nas (Nasred) by combining with CH4 oxidation (CH4 was the only electron donor in the system) (R3). Nitrate was assimilatorily reduced to nitrite (R4). The equations of elementary reactions in bacterial CH4 oxidation coupled with nitrate assimilation by Methylobacter are expressed as follows:
\({\mathrm O}_{2\left(\mathrm{Air}\right)}\xrightarrow{{\mathrm k}_1}{\mathrm O}_{2\left(\mathrm{Dis}\right)}\) | R1 |
\({\mathrm{CH}}_4+{\mathrm O}_2\xrightarrow{{\mathrm k}_2}{\mathrm{Nas}}_{\mathrm{ox}}\) | R2 |
\({\mathrm{CH}}_4+{\mathrm{Nas}}_{\mathrm{ox}}\xrightarrow{{\mathrm k}_3}{\mathrm{CO}}_2+2{\mathrm H}_2\mathrm O+{\mathrm{Nas}}_{\mathrm{red}}\) | R3 |
\(\mathrm{NO}_3^-+{\mathrm{Nas}}_{\mathrm{red}}\xrightarrow{\mathrm k4}\;\mathrm{NO}_2^-+{\mathrm{Nas}}_{\mathrm{ox}}\) | R4 |
Here, rate constants from R1 to R4 are denoted as k 1, k 2, k 3, and k 4. Based on R1-R4 and the kinetic data in the presence of 25% CH4, 75% air and 2 mM nitrate (shown in Figs. 2 and S2), the kinetic models during microbial nitrate assimilation accompanied with CH4 oxidation were established and the values of k 1-k 4 are presented in Table 2, Figs. 7a and S3.
Model-fitted kinetics of (a) 2 mM nitrate / 75% air, (b) 2 mM nitrate / 40% air, and (c) 0.9 mM nitrate / 75% air. The shadow area is the 95% inference confidence intervals at the extremes of the parameter confidence limits using the chi2 test. The kinetic models during microbial nitrate assimilation accompanied with CH4 oxidation were established with the experimental data of (2 mM nitrate / 75% air) (a). The experiments under the conditions of (2 mM nitrate / 30% air) (b) and (0.9 mM nitrate / 75% air) (c) were conducted to assess the accurateness of the model
Assimilatory nitrate reduction and CH4 oxidation by Methylobacter sp. YHQ were quantitatively described by the model. To assess the accurateness of the model, the experiments under the conditions of (2 mM nitrate + 40% air) and (0.9 mM nitrate + 75% air) were conducted as well (Fig. 7b, c). The trend lines generated by the model in Fig. 7b-d and kinetic data of CH4, nitrate, and CO2 can be well matched. These results indicate that the model is mechanistically feasible to predict rate of CH4 oxidation and assimilatory nitrate reduction. Due to the fact that the developed model may give a quantitative depiction of microbial nitrate assimilation and CH4 oxidation, such a straightforward model may have the potential to be used to determine CH4 oxidation and assimilatory nitrate reduction rate in other analogous systems.
4.4 Environmental implications
Enzymatic nitrate transformation is a vital step of biogeochemical N cycling (He et al. 2021). Dual N and O isotope analysis is an invaluable approach for quantitative assessments of biochemical nitrate transformation. This study is the first to use N-O isotopes to elucidate the molecular mechanisms of assimilatory nitrate reduction by freshwater bacteria. N-O isotopes offer a more accurate assessment of reaction processes compared to single isotope analysis because they are less prone to being impacted by mass-transfer limitations. Future work on N and O isotope fractionation induced by other species of bacteria and/or with other carbon sources can provide more insight into the (18 ε:15 ε)assimilation ratios of assimilatory nitrate reduction with Nas.
Assimilatory nitrate reduction accompanied with aerobic CH4 oxidation by bacteria is ubiquitous in natural environments and is a vital link between global C and N cycles. Nitrate is a prevalent environmental pollutant in freshwater ecosystems because of the widespread applications of nitrogenous fertilizers and the extensive discharge of industrial wastewater (Visser et al. 2022). CH4 oxidation provides energy for their metabolism, which can be used for nitrate assimilatory reactions. CH4 is a non-CO2 greenhouse gas with a 28 to 34 times higher greenhouse effect than CO2 (Rodhe 1990). Aerobic CH4 oxidation plays an important role in fluctuating redox zones, e.g., wetland ecosystems and periodically flooded paddy soils (Yun et al. 2013). Wetlands are the biggest natural CH4 reservoirs, which emit 164 Tg.yr-1, and are important atmospheric CH4 sinks (Bridgham et al. 2013). Methanotrophs play a central role in mitigating CH4 emissions to the atmosphere (Reis et al. 2022). Members of Methylobacter are responsible for most of aerobic CH4 oxidation in freshwater wetlands (Smith et al. 2018). In oxic soils, the estimated 43%–90% of CH4 (40 Tg y−1) is consumed by aerobic methanotrophs (Chowdhury and Dick 2013). Methanotroph growth requires a large amount of N (Babel 1983), resulting in assimilatory nitrate reduction (Hao et al. 2020). N2O is one of the major products of nitrate dissimilation but does not occur during nitrate assimilation (Chen et al. 2021). Thus, acceleration of rate for assimilatory nitrate reduction and aerobic CH4 oxidation can simultaneously lower the N2O and CH4 concentrations. Apart from CH4, Methylobacter strains can utilize methanol as a carbon and energy substrate, and a few strains can utilize methylamine as well (Bowman 2006). Methanol and methylamine are common metabolic intermediate in environment. Methanol is formed during microbial degradation of organic matter under anoxic and oxic settings (Sun et al. 2002). Methylamine is generated during organic matter decarboxylation, anaerobic decomposition of proteins, and numerous industries (Ge et al. 2011). CH4 may be transferrable to methanol and methylamine of easily degradable organic carbon and this needs to be studied in the future work.
5 Conclusions
In this study, the impacts of various environmental factors on N and O isotope fractionation during assimilatory nitrate reduction and CH4 oxidation by freshwater bacteria, were thoroughly investigated. N-O isotopes can be applied as “fingerprints” which provide information on quantitative assessment of assimilatory nitrate reduction utilizing laboratory-based 15 ε values and the Rayleigh model. The 15 ε assimilation values with different initial nitrate and oxygen concentrations were provided, which can back up appropriate choice of 15 ε for quantification of nitrate reduction. Dual isotope plots were much less affected than those of single isotope and could provide more reliable insights into N transformation mechanisms. The ratios of (18 ε:15 ε)assimilation induced by freshwater bacteria with Nas were pronouncedly different from those induced by eukaryotes with eukNR. This study shows a high significance for distinguishing eukNR with Nas through natural abundance N and O isotopes and quantitatively describing microbial assimilatory nitrate reduction and CH4 oxidation through a kinetic model.
Availability of data and materials
The datasets used or analyzed during the current study are available from the corresponding author on reasonable request.
References
Bowman J (2006) The methanotrophs—the families Methylococcaceae and Methylocystaceae. Prokaryotes 5:266–289. https://doi.org/10.1007/0-387-30745-1_15
Brandes JA, Devol AH (1997) Isotopic fractionation of oxygen and nitrogen in coastal marine sediments. Geochim Cosmochim Acta 61:1793–1801. https://doi.org/10.1016/S0016-7037(97)00041-0
Bridgham SD, Cadillo-Quiroz H, Keller JK, Zhuang Q (2013) Methane emissions from wetlands: biogeochemical, microbial, and modeling perspectives from local to global scales. Glob Change Biol 19:1325–1346. https://doi.org/10.1111/gcb.12131
Chen G, Chen D, Li F, Liu T, Zhao Z, Cao F (2020) Dual nitrogen-oxygen isotopic analysis and kinetic model for enzymatic nitrate reduction coupled with Fe(II) oxidation by Pseudogulbenkiania sp. strain 2002. Chem Geol 534:119456. https://doi.org/10.1016/j.chemgeo.2019.119456
Chen G, Zhao W, Yang Y, Chen D, Wang Y, Li F, Zhao Z, Cao F, Liu T (2021) Chemodenitrification by Fe(II) and nitrite: Effects of temperature and dual N-O isotope fractionation. Chem Geol 575:120258. https://doi.org/10.1016/j.chemgeo.2021.120258
Chen D, Cheng K, Liu T, Chen G, Kappler A, Li X, Zeng RJ, Yang Y, Yue F, Hu S, Cao F, Li F (2023) Novel insight into microbially mediated nitrate-reducing Fe(II) oxidation by Acidovorax sp. Strain BoFeN1 using dual N-O isotope fractionation. Environ Sci Technol 57:12546–12555. https://doi.org/10.1021/acs.est.3c02329
Chowdhury TR, Dick RP (2013) Ecology of aerobic methanotrophs in controlling methane fluxes from wetlands. Appl Soil Ecol 65:8–22. https://doi.org/10.1016/j.apsoil.2012.12.014
Crawford NM, Arst HN Jr (1993) The molecular genetics of nitrate assimilation in fungi and plants. Annu Rev Genet 27:115–146. https://doi.org/10.1146/annurev.ge.27.120193.000555
Flanagan DA, Gregory LG, Carter JP, Karakas-Sen A, Richardson DJ, Spiro S (1999) Detection of genes for periplasmic nitrate reductase in nitrate respiring bacteria and in community DNA. FEMS Microbiol Lett 177:263–270. https://doi.org/10.1111/j.1574-6968.1999.tb13742.x
Gao H, Yang ZK, Barua S, Reed SB, Romine MF, Nealson KH, Fredrickson JK, Tiedje JM, Zhou J (2009) Reduction of nitrate in Shewanella oneidensis depends on atypical NAP and NRF systems with NapB as a preferred electron transport protein from CymA to NapA. ISME J 3:966–976. https://doi.org/10.1038/ismej.2009.40
Ge X, Wexler AS, Clegg SL (2011) Atmospheric amines—Part 1 A review. Atmos Env 45:524–546. https://doi.org/10.1016/j.atmosenv.2010.10.012
Granger J, Sigman DM, Needoba JA, Harrison PJ (2004) Coupled nitrogen and oxygen isotope fractionation of nitrate during assimilation by cultures of marine phytoplankton. Limnol Oceanogr 49:1763–1773. https://doi.org/10.4319/lo.2004.49.5.1763
Granger J, Sigman D, Rohde M, Maldonado M, Tortell P (2010) N and O isotope effects during nitrate assimilation by unicellular prokaryotic and eukaryotic plankton cultures. Geochem Cosmochim Acta 74:1030–1040. https://doi.org/10.1016/j.gca.2009.10.044
Hao Q, Liu F, Zhang Y, Wang O, Xiao L (2020) Methylobacter accounts for strong aerobic methane oxidation in the Yellow River Delta with characteristics of a methane sink during the dry season. Sci Total Environ 704:135383. https://doi.org/10.1016/j.scitotenv.2019.135383
He Z, Wang J, Hu J, Yu H, Jetten MS, Liu H, Cai C, Liu Y, Ren H, Zhang X (2019) Regulation of coastal methane sinks by a structured gradient of microbial methane oxidizers. Environ Pollu 244:228–237. https://doi.org/10.1016/j.envpol.2018.10.057
He Y, Zhang Y, Zhang S, Liu Y (2021) Predicting nitrogen and oxygen kinetic isotope effects of nitrate reduction by periplasmic dissimilatory nitrate reductase. Geochim Cosmochim Acta 293:224–239. https://doi.org/10.1016/j.gca.2020.10.027
Hutchens E, Radajewski S, Dumont MG, McDonald IR, Murrell JC (2004) Analysis of methanotrophic bacteria in Movile Cave by stable isotope probing. Environ Microbiol 6:111–120. https://doi.org/10.1046/j.1462-2920.2003.00543.x
Johnson KA, Simpson ZB, Blom T (2009) Global kinetic explorer: a new computer program for dynamic simulation and fitting of kinetic data. Anal Biochem 387(1):20–29. https://doi.org/10.1016/j.ab.2008.12.024
Joseph AN, Waser NA, Harrison PJ, Calvert SE (2003) Nitrogen isotope fractionation in 12 species of marine phytoplankton during growth on nitrate. Mar Ecol Prog Series 255:81–91. https://doi.org/10.3354/meps255081
Karsh KL, Granger J, Kritee K, Sigman DM (2012) Eukaryotic assimilatory nitrate reductase fractionates N and O isotopes with a ratio near unity. Environ Sci Technol 46:5727–5735. https://doi.org/10.1021/es204593q
Karsh K, Trull T, Sigman D, Thompson P, Granger J (2014) The contributions of nitrate uptake and efflux to isotope fractionation during algal nitrate assimilation. Geochim Cosmochim Acta 132:391–412. https://doi.org/10.1016/j.gca.2013.09.030
Koo CW, Rosenzweig AC (2021) Biochemistry of aerobic biological methane oxidation. Chem Soc Rev 50:3424–3436. https://doi.org/10.1039/D0CS01291B
Kritee K, Sigman DM, Granger J, Ward BB, Jayakumar A, Deutsch C (2012) Reduced isotope fractionation by denitrification under conditions relevant to the ocean. Geochim Cosmochim Acta 92:243–259. https://doi.org/10.1016/j.gca.2012.05.020
Kuypers MM, Marchant HK, Kartal B (2018) The microbial nitrogen-cycling network. Nature Rev Microbiol 16:263. https://doi.org/10.1038/nrmicro.2018.9
Li S, Luo Z, Wang S, Nan Q, Ji G (2023) Denitrification fractionates N and O isotopes of nitrate following a ratio independent of carbon sources in freshwaters. Environ Microbiol 25(11):2404–2415. https://doi.org/10.1111/1462-2920.16468
Lin JT, Stewart V (1997) Nitrate assimilation by bacteria. In: Poole RK (ed) Advances in Microbial Physiology. Academic Press, pp 1–30
Lin JL, Joye SB, Scholten JCM, Schäfer H, McDonald IR, Murrell JC (2005) Analysis of methane monooxygenase genes in Mono Lake suggests that increased methane oxidation activity may correlate with a change in methanotroph community structure. Appl Environ Microbiol 71:6458–6462. https://doi.org/10.1128/AEM.71.10.6458-6462.2005
Liu T, Li X, Zhang W, Hu M, Li F (2014) Fe(III) oxides accelerate microbial nitrate reduction and electricity generation by Klebsiella pneumoniae L17. J Colloid Interface Sci 423:25–32. https://doi.org/10.1016/j.jcis.2014.02.026
Liu T, Chen D, Luo X, Li X, Li F (2019) Microbially mediated nitrate-reducing Fe (II) oxidation: Quantification of chemodenitrification and biological reactions. Geochim Cosmochim Acta 256:97–115. https://doi.org/10.1016/j.gca.2018.06.040
Luque-Almagro VM, Gates AJ, Moreno-Vivián C, Ferguson SJ, Richardson DJ, Roldán MD (2011) Bacterial nitrate assimilation: gene distribution and regulation. Biochem Soc Trans 39:1838–1843. https://doi.org/10.1042/BST20110688
Mancinelli RL (1995) The regulation of methane oxidation in soil. Annu Rev Microbiol 49:581–605. https://doi.org/10.1146/annurev.mi.49.100195.003053
Mariotti A, Germon J, Hubert P, Kaiser P, Letolle R, Tardieux A, Tardieux P (1981) Experimental determination of nitrogen kinetic isotope fractionation: some principles; illustration for the denitrification and nitrification processes. Plant Soil 62:413–430. https://doi.org/10.1007/BF02374138
McGinnis S, Madden TL (2004) BLAST: at the core of a powerful and diverse set of sequence analysis tools. Nucleic Acids Res 32:W20–W25. https://doi.org/10.1093/nar/gkh435
Pastorelli R, Piccolo R, Simoncini S, Landi S (2013) New primers for denaturing gradient gel electrophoresis analysis of nitrate-reducing bacterial community in soil. Pedosphere 23:340–349. https://doi.org/10.1016/S1002-0160(13)60025-9
Pennock JR, Velinsky DJ, Ludlam JM, Sharp JH, Fogel ML (1996) Isotopic fractionation of ammonium and nitrate during uptake by Skeletonema costatum: Implications for δ 15N dynamics under bloom conditions. Limnol Oceanogr 41:451–459. https://doi.org/10.4319/lo.1996.41.3.0451
Puri AW, Owen S, Chu F, Chavkin T, Beck DA, Kalyuzhnaya MG, Lidstrom ME (2015) Genetic tools for the industrially promising methanotroph Methylomicrobium buryatense. Appl Environ Microbiol 81:1775–1781. https://doi.org/10.1128/AEM.03795-14
Reis PCJ, Thottathil SD, Prairie YT (2022) The role of methanotrophy in the microbial carbon metabolism of temperate lakes. Nat Commun 13:43. https://doi.org/10.1038/s41467-021-27718-2
Rivett MO, Buss SR, Morgan P, Smith JWN, Bemment CD (2008) Nitrate attenuation in groundwater: a review of biogeochemical controlling processes. Water Res 42:4215–4232. https://doi.org/10.1016/j.watres.2008.07.020
Rodhe H (1990) A comparison of the contribution of various gases to the greenhouse effect. Science 248(4960):1217–1219. https://doi.org/10.1126/science.248.4960.1217
Sigman DM, Casciotti KL, Andreani M, Barford C, Galanter M, Böhlke J (2001) A bacterial method for the nitrogen isotopic analysis of nitrate in seawater and freshwater. Analyt Chem 73:4145–4153. https://doi.org/10.1021/ac010088e
Smith GJ, Angle JC, Solden LM, Borton MA, Morin TH, Daly RA, Johnston MD, Stefanik KC, Wolfe R, Gil B, Wrighton KC (2018) Members of the genus Methylobacter are inferred to account for the majority of aerobic methane oxidation in oxic soils from a freshwater wetland. MBio 9:e00815-18. https://doi.org/10.1128/mBio.00815-18
Stolz JF, Basu P (2002) Evolution of nitrate reductase: molecular and structural variations on a common function. ChemBioChem 3:198–206. https://doi.org/10.1002/1439-7633(20020301)3:2/3%3c198::AID-CBIC198%3e3.0.CO;2-C
Sun MY, Aller RC, Lee C, Wakeham SG (2002) Effects of oxygen and redox oscillation on degradation of cell-associated lipids in surficial marine sediments. Geochim Cosmochim Acta 66:2003–2012. https://doi.org/10.1016/S0016-7037(02)00830-X
Treibergs LA, Granger J (2017) Enzyme level N and O isotope effects of assimilatory and dissimilatory nitrate reduction. Limnol Oceanogr 62:272–288. https://doi.org/10.1002/lno.10393
van Grinsven S, Sinninghe Damsté JS, Harrison J, Polerecky L, Villanueva L (2021) Nitrate promotes the transfer of methane-derived carbon from the methanotroph Methylobacter sp. to the methylotroph Methylotenera sp. in eutrophic lake water. Limnol Oceanogr 66:878–891. https://doi.org/10.1002/lno.11648
Visser A, Wankel S, Frey C, Kappler A, Lehmann M, 2022. Unchanged nitrate and nitrite isotope fractionation during heterotrophic and Fe(II)-mixotrophic denitrification suggest a non-enzymatic link between denitrification and Fe (II) oxidation. Front Microbiol. 13. https://doi.org/10.3389/fmicb.2022.927475
Wada E, Hattori A (1978) Nitrogen isotope effects in the assimilation of inorganic nitrogenous compounds by marine diatoms. Geomicrobiol J 1:85–101. https://doi.org/10.1080/01490457809377725
Wang Y, Wu M, Lai CY et al (2023) Methane oxidation coupled to selenate reduction in a membrane bioreactor under oxygen-limiting conditions. Environ Sci Technol 57(51):21715–21726. https://doi.org/10.1021/acs.est.3c04958
Xiao L, Zheng S, Lichtfouse E, Luo M, Tan Y and Liu F. (2020) Carbon nanotubes accelerate acetoclastic methanogenesis: From pure cultures to anaerobic soils. Soil Biol Biochem. 150. https://doi.org/10.1016/j.soilbio.2020.107938
Yun J, Yu Z, Li K, Zhang H (2013) Diversity, abundance and vertical distribution of methane-oxidizing bacteria (methanotrophs) in the sediments of the Xianghai wetland, Songnen Plain, northeast China. J Soils Sediments 13:242–252. https://doi.org/10.1007/s11368-012-0610-1
Zhang RC, Chen C, Shao B, Wang W, Xu XJ, Zhou X, Xiang YN, Zhao L, Lee DJ, Ren NQ (2020) Heterotrophic sulfide-oxidizing nitrate-reducing bacteria enables the high performance of integrated autotrophic-heterotrophic denitrification (IAHD) process under high sulfide loading. Water Res 178:115848. https://doi.org/10.1016/j.watres.2020.115848
Acknowledgments
We are deeply indebted to Sebastian Kopf from University of Colorado Boulder for the constructive discussion and suggestions.
Funding
This work was funded by the National Natural Science Foundation of China (42125704, 42177035, 42107134, and U20A20109), National Key R&D Program of China (2023YFC3706900), Guangdong Basic and Applied Basic Research Foundation (2024A1515012421 and 2022A1515011518), Young Talent Project of GDAS (2023GDASQNRC-0203), GDAS' Project of Science and Technology Development (2024GDASZH-2024010102), and Guangdong Foundation for Program of Science and Technology Research (Grant No.2023B1212060044). Andreas Kappler acknowledges infrastructural support by the DFG under Germany’s Excellence Strategy, cluster of Excellence EXC2124, project ID 390838134.
Author information
Authors and Affiliations
Contributions
Tongxu Liu and Guojun Chen contributed to the study conception and design. Material preparation and data analysis were performed by Qinqin Hao, Guojun Chen, Fujun Yue, and Yang Yang. The original draft of the manuscript was written by Guojun Chen and Qinqin Hao. Tongxu Liu, Fangbai Li, and Fanghua Liu supervised the study. Tongxu Liu, Raymond Jianxiong Zeng, Xiaomin Li, Andreas Kappler, Fangbai Li, Shiwen Hu, Han Li, Dayi Qian, Baoguo Yang, and Kaster Sarkytkan reviewed and edited the article. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
Fangbai Li is an executive editor-in-chief and Tongxu Liu is an editorial board member of Carbon Research. They were not involved in the editorial review, or the decision to publish this article. All authors declare that there are no competing interests.
Additional information
Handling editor: Fengchang Wu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, G., Hao, Q., Zeng, R.J. et al. Unraveling the mechanism of assimilatory nitrate reduction and methane oxidation by Methylobacter sp. YHQ through dual N-O isotope analysis and kinetic modeling. Carbon Res. 3, 58 (2024). https://doi.org/10.1007/s44246-024-00143-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44246-024-00143-y