Abstract
Autism Spectrum Disorder (ASD) presents a significant challenge due to its complex genetic basis and associated comorbidities. Among the genes implicated in ASD, SHANK3 has been identified as a critical player, affecting synaptic structure and function. This review examines the role of SHANK3 in ASD, highlighting the genetic diversity and the systemic nature of the disorder. Utilizing animal models, studies have uncovered autism-like behaviours and synaptic dysfunctions linked to SHANK3 deficiency, suggesting potential therapeutic targets. Furthermore, the review delves into the specific gene families associated with ASD, emphasizing the dynamic regulation between translation and transcription processes and the impact of mutations on synaptic translation and proteins. Molecular changes in SHANK3-deficient animal models reveal alterations in protein composition, localization, and transcription, particularly affecting the striatum and involving essential proteins and signalling pathways. Therapeutic strategies, including pharmaceutical compounds and genetic restoration, show promise in addressing the neuropsychiatric symptoms and physiological abnormalities observed in SHANK3-deficient mice. This research not only advances our understanding of ASD's neurobiological basis but also underscores the potential of targeted interventions to mitigate symptoms and improve the quality of life for individuals affected by ASD and related disorders.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Autism Spectrum Disorder (ASD) is a complex neurodevelopmental disorder characterized by repetitive and characteristic patterns of behaviour and difficulties with social communication and interaction [1, 2]. The genetic underpinnings of ASD involve a multiplicity of loci, with over 15 identified contributing to its development [3]. This genetic diversity points towards a heterogeneous nature of ASD, complicating the pursuit of a singular genetic explanation [4].
A key element in ASD research involves examining its frequent comorbidities—epilepsy, sleep disorders, and gastrointestinal abnormalities—as well as mitochondrial dysfunction, which is one of the critical pathologies of autism spectrum disorder, illuminating the disorder's systemic characteristics [5,6,7]. This multi-system involvement necessitates a holistic approach to both understanding and addressing ASD.
The genetic underpinnings of ASD involve a multiplicity of loci, with over 15 identified contributing to its development (Table 1). For example, genes involved in presynaptic vesicle cycling and transport, such as STXBP1 and SYN1, play critical roles in synaptic function and neurotransmitter release [8]. Mutations in genes regulating cytoskeleton dynamics, like ARHGEF6, can impact neuronal structure and connectivity [3, 9]. Additionally, genes involved in cell adhesion and trans-synaptic signaling, such as NRXN1 and NLGN3, are essential for synapse formation and maintenance [9].
Among the myriad genes implicated in ASD, SHANK3 stands out due to its critical role in synaptic function and its strong association with neurobehavioral disorders such as Phelan-McDermid Syndrome (PMS) [12, 13]. SHANK3 encodes a scaffolding protein essential for synaptic function, interacting with other synaptic proteins like SAPAP and Homer to form complexes that regulate synaptic signaling and plasticity (Fig. 1) [14]. Disruptions in SHANK3 and its associated proteins can lead to synaptic dysfunction, a hallmark of ASD [8]. Research utilizing Shank3-deficient mice has uncovered autism-like behaviors and synaptic dysfunctions, suggesting the potential of modulating actin regulators as a therapeutic strategy (Fig. 1) [15].
Schematic of SHANK3 protein domains and their interactions in the synaptic post-dendritic density. This diagrammatic representation illustrates the SHANK3 protein topology within neuronal synapses’ postsynaptic density (PSD). The N-terminal region of SHANK3 is depicted anchoring to the actin-based cytoskeleton via β-fodrin α. The protein consists of multiple domains: an Ankyrin repeat domain (Ank6), an SH3 domain, a Postsynaptic Density 95, Discs Large, Zonula Occludens-1 (PDZ) domain, a proline-rich region, and a C-terminal Sterile Alpha Motif (SAM). These domains facilitate the SHANK3 protein's interaction with a variety of synaptic proteins, including somatostatin receptor 2 (sst2), the calcium-independent receptor of alpha latrotoxin (CIRL), NMDA receptor (NMDA-R), metabotropic glutamate receptor (mGluR), PSD95, SAPAP/GKAP, and cortactin [24, 33]
Despite significant progress in understanding the genetic underpinnings of ASD, challenges remain in translating these findings into effective treatments [9]. By addressing gaps in existing research, the findings are expected to advance our understanding of ASD pathophysiology and contribute to the development of targeted therapeutic interventions. The study's main objectives are to investigate the genetic and molecular mechanisms by which SHANK3 mutations contribute to ASD, evaluate the efficacy of different therapeutic strategies for SHANK3 deficiency, and highlight the translational potential of these treatments for clinical application in ASD management.
2 Gene families associated with ASD
Recent findings underscore a nuanced interplay between translation and transcription processes implicated in ASD pathophysiology. Mutations in translation pathways, including mTOR, ERK, and FMRP-eIF4E-CYFIP, result in abnormally elevated synaptic translation and protein levels [16]. ASD-associated genes often regulate synaptic structure and function, including encoding scaffold proteins, neurotransmitter receptors, and related proteins. Recent advances in ASD research have identified differentially expressed genes in the brain tissue of ASD patients. These genes are often linked to alterations in synaptic function and neuronal communication. For instance, studies have shown that the expression of genes involved in synaptic transmission, neuronal development, and immune response are significantly altered in individuals with ASD [17]. This underscores the connection between genetic expression patterns and ASD-related symptoms, providing potential pathways for therapeutic intervention.
Additionally, clinical assessments and genetic research have elucidated that ASD encompasses diverse neurobehavioral disorders, each characterised by distinct genetic foundations (Table 1). For example, Fragile X Syndrome is caused by mutations in the FMR1 gene and often presents with intellectual disability and autistic behaviours [18]. Rett Syndrome, primarily affecting females, is associated with mutations in the MECP2 gene and is characterised by severe cognitive and physical impairments [19]. Tuberous Sclerosis Complex, caused by mutations in the TSC1 or TSC2 genes, leads to the formation of benign tumours in multiple organs and is often associated with epilepsy and autism [20]. These examples highlight the genetic heterogeneity underlying ASD, emphasising the need for personalised approaches in diagnosis and treatment.
Cell adhesion molecules (CAMs) are pivotal in brain development, facilitating initial synaptic contacts [21, 22]. Predominantly, CAMs within the synaptic cleft belong to families such as calcineurin, the immunoglobulin superfamily, integrins, and the complexes of neurotoxins with their neuroglial protein chaperones [23]. Genetic mutations in these neuronal CAMs have been linked to ASD or susceptibility to it. Unique and rare genetic variants in synaptic CAMs (e.g., CDH9, LGN3, NLGN4X, NRXN1, SHANK2, SHANK3) highlight the critical role of synaptic cell adhesion in cognitive and behavioural functions [24, 25]. The SHANK gene family, in particular, has garnered extensive study for its consistent phenotypic manifestations across mouse and mammalian models, offering a robust theoretical framework for understanding ASD's molecular underpinnings.
The SHANK3 gene is found on chromosome 22q13.3 in humans, chromosome 15E3 in mice and rats, and on 7q34 in rodents [26]. SHANK3 is a postsynaptic scaffolding protein that regulates synaptic development, function, and plasticity. As illustrated in Fig. 1, SHANK3 interacts with a variety of synaptic proteins, facilitating the formation of complexes that are essential for synaptic signaling and plasticity. SHANK3 is pivotal in assembling postsynaptic density signalling complexes at the macromolecular scale [26]. Strong associations between mutations in the SHANK gene and ASD have been established. Diverse SHANK3 isoforms from complex regulation and splicing support the theory that synaptic dysfunction underlies ASD pathophysiology. For instance, isoforms involved in synapse formation and maturation include those with Ankyrin repeats and SH3 domains, which are crucial for synaptic scaffolding and signalling [27, 28]. Isoforms with proline-rich regions interact with actin filaments and intracellular signalling pathways, playing a role in maintaining synaptic plasticity [29].
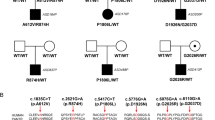
Human genetic research, including genome-wide association studies (GWAS) and targeted sequencing of ASD cohorts, has established a causal link between SHANK3 mutations and ASD. For instance, studies involving large cohorts of ASD patients and their families have identified several de novo mutations in SHANK3 [30]. These findings are corroborated by analyses of inherited mutations in families with multiple affected members, further strengthening the evidence for SHANK3's role in ASD. According to a meta-analysis, truncating mutations in the SHANK gene family are thought to be responsible for about 1% of all ASD cases [31]. Given the variety of etiologies for ASD, this is an unexpectedly high rate. Additionally, both asymptomatic parents and their children diagnosed with ASD have been found to carry SHANK3 mutations [8, 32]. This suggests that while SHANK3 mutations alone may not be sufficient to cause ASD, they contribute significantly to its development within a genetically susceptible environment.
3 Molecular changes in animal models of Shank 3 deficiency
SHANK3-deficient murine animal models exhibit increased repetitive behaviour, inappropriate social behaviour, heightened anxiety, decreased neuronal physiology, and altered PSD levels of HOMER [34], DLGAPs, NMDARs [14], AMPARs [32], and other HOMER proteins [35]. These phenotypes closely mimic the symptoms observed in humans with ASD [36,37,38,39]. This resemblance underscores the utility of these animal models in studying the underlying mechanisms of ASD and developing potential treatments.
Disruptions in SHANK3 result in significant alterations in brain structure and function, observed at the molecular level as changes in protein composition, localization, or transcription. Studies have shown that SHANK3 disruption impacts numerous proteins, including DLG4, HOMER1, and GAIA1, which are intricately linked with actin-related mechanisms and the broader cytoskeletal architecture or fall under the specific categorization of glutamate receptor subclasses [26]. These changes were primarily observed in SHANK3-deficient mouse models, such as those with exon-specific mutations (e.g., ex13-16|PDZ, ex21|PRO) and heterozygous mutant rodents [40].
Specifically, research often focuses on the striatum, the most studied and significantly affected brain region in these models. The levels of many proteins, including HOMER1 [40], HOMER2 [41], HOMER-proteins in general [42], DLG [36], DLG4 [43], which mediate protein–protein interactions, were lowered in the striatum of mutant mice. Additionally, reduced levels of various glutamate receptor subunits were observed, predominantly in the postsynaptic fractions of the striatum, notably including AMPAR subunits (GRIA1, GRIA2, and GRIA4) [44], NMDAR-subunits (GRIN1, GRIN2A, and GRIN2B) [45, 46], GRM5 [47], and GRIK [48]. Another group of proteins, such as DOCK4 or TRIO, linked to actin or the cytoskeleton and were diminished in the striatal synapses of SHANK3-deficient mice [49]. Furthermore, reductions in HOMER1 protein levels have been identified across various brain regions, including the hippocampus [24], cortical synapses [50], and prefrontal areas [51]. This decrease aligns with the molecular disturbances frequently noted in SHANK3 deficiency, which disrupts the signalling mediated by metabotropic glutamate receptors (e.g., GRM5) [32, 49]. It also leads to diminished phosphorylation of targets within SYN1 and CREB [52], as well as in the PI3K/AKT/mTOR [53] and the MAPK/ERK pathways [54], impacting synaptic MAPK phosphorylation [34]. Additionally, a decline in crucial elements of the RAC1-dependent signalling pathway contributes to the phosphorylation and subsequent inactivation of CFL1. This process triggers an increase in actin depolymerisation due to the activation of CFL1 [14], further complicating the synaptic alterations observed in SHANK3 deficiency.
Epigenetic regulation also plays a crucial role in gene expression in vivo. In SHANK3-deficient animals, only the prefrontal cortex showed a significant decrease in H3 acetylation and an increase in H3K9m2 di-methylation at the Arc promoter, linked to HDAC2, EHMT1, and EHMT2 dysregulation [15]. Moreover, targeting histone demethylase LSD1 has ameliorated autism-like deficits in Shank3-deficient mice, further underscoring the therapeutic potential of epigenetic interventions [55]. Such epigenetic dysregulation underscores a potential therapeutic avenue, given that epigenetic interventions can normalise the expression of numerous genes found to be dysregulated.
While the focus has predominantly been on neurological alterations, recent findings also report the dysregulation of cytokine levels in SHANK3-deficient mice [25], indicating the gastrointestinal system as an additional extracerebral system affected by SHANK3 deficiency, alongside the previously noted systems. This is evidenced by the observed upregulation of TJP1 (ZO-1) in the small intestine. Such an increase may potentially influence the functional integrity of the intestinal barrier [56]. This involvement is further supported by the observed upregulation of TJP1 in the small intestine, which may affect the intestinal barrier's functional integrity, and increased levels of bacterial lipopolysaccharide in the liver, suggesting enhanced gastrointestinal leakage in these mice. The comprehensive investigation into SHANK3 deficiency reveals significant neurological and extra-neural disruptions, highlighting the complexity of its pathophysiology and underscoring the potential of epigenetic therapies in addressing the broad spectrum of resultant alterations.
4 Treatment strategies for SHANK3 deficiency
4.1 Pharmaceutical compounds
Numerous chemical substances have been suggested as potential treatments for SHANK3-associated neuropsychiatric disease in light of those above molecular and physiological abnormalities in SHANK3-deficient mice. For example, it has been proposed that modulating glutamate receptors may be advantageous since AMPAR, NMDAR, or GRM5 hypofunction could result in an imbalance between excitement and inhibition, potentially responsible for specific ASD-like symptoms in SHANK3-mutant mice [50]. The research team targeted this molecular loop involving the modulation of AMPAR, NMDAR, and GRM5 receptors. They developed a tailored drug, AMPAR-PAM, a positive allosteric modulator (PAM) of AMPAR via compound CX546. This drug has normalised social deficits in ex13-16|PDZ mice [57]. These findings suggest that AMPAR-PAM could be a viable candidate for clinical trials aimed at treating social and behavioural symptoms of ASD in humans. However, social behaviour was unaffected in another investigation [58]. Additionally, during social interactions, AMPAR-PAM reduced increased repetitive behaviour and partially repaired changes in synaptic transmission in the anterior cingulate cortex [28, 59]. Ex13-16|PDZ mouse hypoactivity was unaffected by CX546 [60].
Furthermore, insulin-like growth factor-1 (IGF-1) has shown promise in treating Phelan-McDermid syndrome (PMS), a condition caused by SHANK3 haploinsufficiency. A clinical trial demonstrated that IGF-1 treatment improved social withdrawal and repetitive behaviours in PMS patients, highlighting its potential for broader application in SHANK3 deficiency-related conditions [61]. This suggests that IGF-1 could be repurposed for treating similar symptoms in ASD, emphasising the need for further clinical investigations to establish its efficacy and safety in a broader ASD population. Additionally, NMDAR activation through D-cycloserine, which acts as a partial agonist at the glycine modulatory site of the NMDAR, enhanced social interaction in ex13-16|PDZ mutant mice without affecting their repetitive behaviour or hypoactivity [57, 62].
The potential for these pharmaceutical compounds to transition from animal models to human treatments underscores the importance of conducting rigorous clinical trials. These trials should evaluate these compounds' safety, efficacy, and optimal dosing in individuals with SHANK3 mutations. Success in these trials could lead to the development of targeted pharmacological therapies that significantly improve the quality of life for those affected by ASD, offering a hopeful outlook for effective management and intervention in this complex disorder.
4.2 Genetic restoration
Given ASD's neurodevelopmental nature, exploring strategies to mitigate behavioural and synaptic irregularities in adulthood is crucial. Despite the lack of observed enhancements in anxiety or motor coordination deficits, reinstating SHANK3 expression in adult mice within the cKI ex13-16|PDZ murine model markedly improved recurrent self-injurious grooming and social interaction shortfalls [41]. This finding highlights the promise of therapeutic interventions post-development for ASD. Moreover, genetic rectification reinstated typical synaptic protein composition and spine morphology and corrected neurotransmission deficiencies. Interventions in early postnatal stages and germline adjustments successfully reversed the irreversible behavioural anomalies at maturity [63]. Similarly, early genetic interventions reduced exacerbated repetitive behaviours, social challenges, and hypoactivity in Ex13|PDZ mice [40]. Another study on adult mice's genetic rescue of Shank3 by Cre demonstrated improvements in avoidance behaviour, motor function, and synaptic transmission but did not address hypoactivity [64]. This suggests that the observed improvements were explicitly due to the Cre-mediated rescue of SHANK3 expression, as hypoactivity might be influenced by other factors not corrected by the Cre-transgene alone. Moreover, local neuronal activation is also seen as an active therapeutic strategy. For instance, activation of prefrontal regions has been shown to partially correct alterations in synaptic transmission and spike density [65]. However, this attempt did not improve avoidance, repetitive, or hypoactive behaviours [66].
Dietary therapy is also an active area in the treatment of ASD. Recognizing the potential connection between zinc deficiency and an elevated risk of ASD [67], researchers explored the impact of zinc supplementation in the SHANK3-deficient ex13-16|PDZ model. Oral zinc supplementation (150 ppm) for six weeks completely normalised phenotypes such as social cognition, repetitive behaviours and anxiety-like behaviours while only partially alleviating hyperactivity in these mice [68]. A study by Lee et al. (2022) explored long-term maternal zinc supplementation (150 ppm) as a novel approach to mitigate ASD-like behaviours in ex13-16|PDZ SHANK3 mutant mice [60]. This intervention successfully alleviated social behavioural deficits, reduced repetitive behaviours, and diminished anxiety levels. It also restored striatal NMDAR-mediated synaptic transmission and presynaptic short-term plasticity, though AMPAR-mediated transmission continued to be compromised. Intriguingly, while maternal zinc supplementation adversely affected AMPAR- and NMDAR-mediated transmission in wild-type animals, it did not influence their behaviour [30, 69].
In addition to zinc supplementation, the ketogenic diet (KD) has emerged as another potential dietary intervention for ASD. KD is a high-fat, low-carbohydrate diet that has been shown to improve various neurological conditions, including epilepsy and ASD. Studies have indicated that KD may enhance mitochondrial function and reduce oxidative stress, potentially leading to improved social behaviours and reduced repetitive behaviours in ASD patients [70]. A modified ketogenic, gluten-free diet with supplemental medium-chain triglycerides (MCTs) also showed improvements in social affect scores among children with ASD [71].
The translational potential of genetic restoration strategies for SHANK3 mutations is significant, with emerging research highlighting promising avenues for treating ASD in humans. Studies have shown that early genetic intervention in SHANK3-related ASD can lead to substantial improvements in social interactions, repetitive behaviours, and other ASD-like phenotypes in mouse models [41, 72]. The application of gene-editing technologies, such as CRISPR-Cas9, offers the potential for precise genetic corrections in patients with SHANK3 mutations. For example, Jaramillo et al. (2020) demonstrated that early restoration of SHANK3 expression in mice prevented core ASD-like behavioural phenotypes, including deficits in social interaction and increased repetitive behaviours [40]. These findings underscore the importance of early intervention and the potential efficacy of gene therapy for ASD. By leveraging advanced gene-editing tools and conducting thorough clinical evaluations, there is potential to develop targeted treatments that significantly improve the quality of life for individuals with ASD, offering hope for effective management and intervention of this complex disorder.
5 Conclusions
This comprehensive examination of SHANK3's role in ASD and related conditions like PMDS highlights the multifaceted genetic and molecular underpinnings of these disorders. The research underscores the critical role of SHANK3 in synaptic structure, function, and plasticity, with its diverse isoforms contributing to various aspects of neuronal development and signalling. The insights gained from studying SHANK3-deficient animal models have been pivotal in replicating ASD-like behaviours and synaptic dysfunctions, thus providing valuable models for testing potential therapeutic interventions.
Pharmaceutical interventions targeting glutamate receptors, such as AMPAR-PAM and NMDAR activation through D-cycloserine, have demonstrated mixed results in SHANK3-deficient models. While these drugs have shown promise in normalising social deficits and reducing repetitive behaviours, their effects on hypoactivity and other symptoms have been inconsistent. These findings indicate that while modulation of glutamatergic signalling can address some ASD-related symptoms, a more comprehensive approach may be necessary to achieve broader therapeutic efficacy.
Genetic restoration strategies have shown significant potential, with studies demonstrating that reinstating SHANK3 expression in adult mice can improve social interactions, reduce self-injurious behaviours, and correct synaptic protein composition and spine morphology. Early postnatal interventions and germline adjustments have also successfully reversed behavioural anomalies that are otherwise irreversible at maturity. These results highlight the promise of gene therapy approaches in mitigating the long-term impacts of SHANK3 mutations. Dietary interventions, particularly zinc supplementation and ketogenic diets, have emerged as promising avenues for addressing ASD-related symptoms. Zinc supplementation has effectively normalised social cognition, reduced repetitive behaviours, and alleviated anxiety-like behaviours in SHANK3-deficient models. The ketogenic diet has been associated with improvements in social behaviours and reductions in oxidative stress, further underscoring the potential of dietary modifications in managing ASD.
The translational potential of these treatments is significant, suggesting that targeted genetic, pharmaceutical, and dietary interventions could offer viable therapeutic options for individuals with ASD. The following steps in this research should focus on conducting rigorous clinical trials to validate these preclinical findings and ensure the safety and efficacy of these interventions in humans. Moreover, a holistic approach that integrates genetic, molecular, and environmental factors will be crucial for developing personalised treatment protocols that address the diverse needs of individuals with ASD and related disorders.
Despite the promising advancements in understanding the role of SHANK3 mutations in ASD and related disorders, several limitations must be acknowledged. The reliance on animal models, primarily mice, to study SHANK3 mutations and their effects poses significant translational challenges, as these models may not fully capture the complexity of human ASD, which involves intricate genetic and environmental interactions. Additionally, ASD is highly heterogeneous, with numerous genes implicated in its pathogenesis. Focusing predominantly on SHANK3 mutations might overlook other critical genetic and epigenetic factors that contribute to the disorder, potentially limiting the applicability of findings to the broader ASD population. The genetic and phenotypic variability among individuals with ASD further complicates the development of universal treatments, as variations in SHANK3 mutations and their interactions with other genetic and environmental factors can lead to differing clinical presentations. Translating promising preclinical findings into effective human treatments remains a significant hurdle, with the safety, efficacy, and long-term effects of potential therapies, such as gene-editing techniques and pharmaceutical compounds, requiring rigorous testing through well-designed clinical trials. Furthermore, the role of epigenetic modifications and environmental factors in ASD is not fully understood, and studies primarily focusing on genetic aspects might miss crucial insights into how these factors interact with genetic predispositions. Developing targeted therapies must also consider potential off-target effects and long-term safety, as the complexity of SHANK3's interactions within the synaptic environment poses risks for unintended consequences that could affect other critical neurological functions.
In conclusion, this research advances our understanding of ASD's neurobiological basis and paves the way for innovative therapeutic strategies. By leveraging genetic and molecular insights, there is a promising potential to develop targeted treatments that can significantly improve the quality of life for individuals affected by ASD, ultimately offering hope for effective management and intervention in this complex disorder.
Data availability
No datasets were generated or analysed during the current study.
References
Widiger T. Diagnostic and statistical manual of mental disorders (DSM). Psychology. 2011. https://doi.org/10.1093/obo/9780199828340-0022.
Akhter S, Hussain AHME, Shefa J, et al. Prevalence of autism spectrum disorder (ASD) among the children aged 18–36 months in a rural community of Bangladesh: a cross sectional study. Research. 2018;7:424. https://doi.org/10.12688/f1000research.13563.1.
Jeste SS, Geschwind DH. Disentangling the heterogeneity of autism spectrum disorder through genetic findings. Nat Rev Neurol. 2014;10:74–81. https://doi.org/10.1038/nrneurol.2013.278.
Maestrini E, Pagnamenta AT, Lamb JA, et al. High-density SNP association study and copy number variation analysis of the AUTS1 and auts5 loci implicate the IMMP2L–DOCK4 gene region in autism susceptibility. Mol Psychiatry. 2009;15:954–68. https://doi.org/10.1038/mp.2009.34.
Kates WR, Burnette CP, Eliez S, et al. Neuroanatomic variation in monozygotic twin pairs discordant for the narrow phenotype for autism. Am J Psychiatry. 2004;161:539–46. https://doi.org/10.1176/appi.ajp.161.3.539.
Bauman ML. Medical comorbidities in autism: challenges to diagnosis and treatment. Neurotherapeutics. 2010;7:320–7. https://doi.org/10.1016/j.nurt.2010.06.001.
Khachadourian V, Mahjani B, Sandin S, et al. Comorbidities in autism spectrum disorder and their etiologies. 2022; https://doi.org/10.1101/2022.03.10.22272202.
Durand CM, Betancur C, Boeckers TM, et al. Mutations in the gene encoding the synaptic scaffolding protein Shank3 are associated with autism spectrum disorders. Nat Genet. 2006;39:25–7. https://doi.org/10.1038/ng1933.
Rodriguez-Gomez DA, Garcia-Guaqueta DP, Charry-Sánchez JD, et al. A systematic review of common genetic variation and biological pathways in autism spectrum disorder. BMC Neurosci. 2021. https://doi.org/10.1186/s12868-021-00662-z.
Li X, Zou H, Brown WT. Genes associated with autism spectrum disorder. Brain Res Bull. 2012;88(6):543–552. https://doi.org/10.1016/j.brainresbull.2012.05.017.
Ross PJ et al. Modeling neuronal consequences of autism-associated gene regulatory variants with human induced pluripotent stem cells. Molecular Autism. 2020;11(1). https://doi.org/10.1186/s13229-020-00333-6.
Monteiro P, Feng G. Shank proteins: roles at the synapse and in autism spectrum disorder. Nat Rev Neurosci. 2017;18:147–57. https://doi.org/10.1038/nrn.2016.183.
Durand CM, Perroy J, Loll F, et al. Shank3 mutations identified in autism lead to modification of dendritic spine morphology via an actin-dependent mechanism. Mol Psychiatry. 2011;17:71–84. https://doi.org/10.1038/mp.2011.57.
Duffney LJ, Zhong P, Wei J, et al. Autism-like deficits in Shank3-deficient mice are rescued by targeting actin regulators. Cell Rep. 2015;11:1400–13. https://doi.org/10.1016/j.celrep.2015.04.064.
Qin L, Ma K, Wang Z-J, et al. Social deficits in Shank3-deficient mouse models of autism are rescued by histone deacetylase (HDAC) inhibition. Nat Neurosci. 2018;21:564–75. https://doi.org/10.1038/s41593-018-0110-8.
Jiang C-C, Lin L-S, Long S, et al. Signalling pathways in autism spectrum disorder: mechanisms and therapeutic implications. Signal Transduct Target Ther. 2022. https://doi.org/10.1038/s41392-022-01081-0.
Gandal MJ, Haney JR, Parikshak NN, et al. Shared molecular neuropathology across major psychiatric disorders parallels polygenic overlap. Science. 2018;359:693–7. https://doi.org/10.1126/science.aad6469.
Garber KB, Visootsak J, Warren ST. Fragile X syndrome. Eur J Hum Genet. 2008;16:666–72. https://doi.org/10.1038/ejhg.2008.61.
Amir RE, Van den Veyver IB, Wan M, et al. Rett syndrome is caused by mutations in X-linked MECP2 encoding methyl-CPG-binding protein 2. Nat Genet. 1999;23:185–8. https://doi.org/10.1038/13810.
Curatolo P, Bombardieri R, Jozwiak S. Tuberous sclerosis. The Lancet. 2008;372:657–68. https://doi.org/10.1016/s0140-6736(08)61279-9.
Jang S, Oh D, Lee Y, et al. Synaptic adhesion molecule IGSF11 regulates synaptic transmission and plasticity. Nat Neurosci. 2015;19:84–93. https://doi.org/10.1038/nn.4176.
Xing G, Li M, Sun Y, et al. Neurexin-Neuroligin 1 regulates synaptic morphology and functions via the wave regulatory complex in drosophila neuromuscular junction. eLife. 2018. https://doi.org/10.7554/elife.30457.
Joseph RM, Korzeniewski SJ, Allred EN, et al. Extremely low gestational age and very low birthweight for gestational age are risk factors for autism spectrum disorder in a large cohort study of 10-year-old children born at 23–27 weeks’ gestation. Am J Obstet Gynecol. 2017. https://doi.org/10.1016/j.ajog.2016.11.1009.
Wang X, McCoy PA, Rodriguiz RM, et al. Synaptic dysfunction and abnormal behaviors in mice lacking major isoforms of Shank3. Hum Mol Genet. 2011;20:3093–108. https://doi.org/10.1093/hmg/ddr212.
Tabouy L, Getselter D, Ziv O, et al. Dysbiosis of microbiome and probiotic treatment in a genetic model of autism spectrum disorders. Brain Behav Immun. 2018;73:310–9. https://doi.org/10.1016/j.bbi.2018.05.015.
Brose N. Faculty opinions recommendation of mutations in the gene encoding the synaptic scaffolding protein Shank3 are associated with autism spectrum disorders. Fac Opin Post-Publication Peer Rev Biomed Lit. 2007. https://doi.org/10.3410/f.1087538.540595.
Hollander E, Uzunova G. Faculty opinions recommendation of abnormal striatal development underlies the early onset of behavioral deficits in shank3b-/- mice. Fac Opin Post-Publication Peer Rev Biomed Lit. 2019. https://doi.org/10.3410/f.736889716.793567605.
Jiang Y, Ehlers MD. Modeling autism by shank gene mutations in mice. Neuron. 2013;78:8–27. https://doi.org/10.1016/j.neuron.2013.03.016.
Edmonson C, Ziats MN, Rennert OM. Altered glial marker expression in autistic post-mortem prefrontal cortex and cerebellum. Molecular Autism. 2014. https://doi.org/10.1186/2040-2392-5-3.
Vyas Y, Lee K, Jung Y, Montgomery JM. Influence of maternal zinc supplementation on the development of autism-associated behavioural and synaptic deficits in offspring Shank3-knockout mice. Mol Brain. 2020. https://doi.org/10.1186/s13041-020-00650-0.
Leblond CS, Nava C, Polge A, et al. Meta-analysis of shank mutations in autism spectrum disorders: a gradient of severity in cognitive impairments. PLoS Genet. 2014. https://doi.org/10.1371/journal.pgen.1004580.
Wang X, Bey AL, Katz BM, et al. Altered mglur5-homer scaffolds and corticostriatal connectivity in a Shank3 complete knockout model of autism. Nat Commun. 2016. https://doi.org/10.1038/ncomms11459.
Uchino S, Waga C. SHANK3 as an autism spectrum disorder-associated gene. Brain Develop. 2013;35(2):106–10. https://doi.org/10.1016/j.braindev.2012.05.013.
Wang Z, Zhong P, Ma K, et al. Amelioration of autism-like social deficits by targeting histone methyltransferases EHMT1/2 in Shank3-deficient mice. Mol Psychiatry. 2019;25:2517–33. https://doi.org/10.1038/s41380-019-0351-2.
Sala C. Faculty opinions recommendation of altered mglur5-homer scaffolds and corticostriatal connectivity in a Shank3 complete knockout model of autism. Fac Opin Post-Publication Peer Rev Biomed Lit. 2016. https://doi.org/10.3410/f.726345298.793522936.
Peça J, Feliciano C, Ting JT, et al. Shank3 mutant mice display autistic-like behaviours and striatal dysfunction. Nature. 2011;472:437–42. https://doi.org/10.1038/nature09965.
Yan Z, Rein B. Mechanisms of synaptic transmission dysregulation in the prefrontal cortex: Pathophysiological implications. Mol Psychiatry. 2021;27:445–65. https://doi.org/10.1038/s41380-021-01092-3.
Yao F, Zhang K, Feng C, et al. Protein biomarkers of autism spectrum disorder identified by computational and experimental methods. Front Psych. 2021. https://doi.org/10.3389/fpsyt.2021.554621.
Yoo Y, Yoo T, Lee S, et al. Shank3 mice carrying the human Q321R mutation display enhanced self-grooming abnormal electroencephalogram patterns and suppressed neuronal excitability and seizure susceptibility. Front Mol Neurosci. 2019. https://doi.org/10.3389/fnmol.2019.00155.
Jaramillo TC, Speed HE, Xuan Z, et al. Novel Shank3 Mutant exhibits behaviors with face validity for autism and altered striatal and hippocampal function. Autism Res. 2016;10:42–65. https://doi.org/10.1002/aur.1664.
Mei Y, Monteiro P, Zhou Y, et al. Adult restoration of Shank3 expression rescues selective autistic-like phenotypes. Nature. 2016;530:481–4. https://doi.org/10.1038/nature16971.
Zhou Y, Kaiser T, Monteiro P, et al. Mice with Shank3 mutations associated with ASD and schizophrenia display both shared and distinct defects. Neuron. 2016;89:147–62. https://doi.org/10.1016/j.neuron.2015.11.023.
Song T-J, Lan X-Y, Wei M-P, et al. Altered behaviors and impaired synaptic function in a novel rat model with a complete Shank3 deletion. Front Cell Neurosci. 2019. https://doi.org/10.3389/fncel.2019.00111.
Zhu Z, Wang W, Gu C, et al (2022) The M1 muscarinic acetylcholine receptor regulates the surface expression of the AMPA receptor subunit GLUA2 via PICK1. 2022; https://doi.org/10.21203/rs.3.rs-1476657/v1.
Myers SJ, Yuan H, Kang J-Q, et al. Distinct roles of grin2a and grin2b variants in neurological conditions. Research. 2019;8:1940.
Krzystanek M, Asman M, Witecka J, et al. Selected single-nucleotide variants in GRIN1, GRIN2A, and GRIN2B encoding subunits of the NMDA receptor are not biomarkers of schizophrenia resistant to clozapine: exploratory study. Pharmacol Rep. 2020;73:309–15. https://doi.org/10.1007/s43440-020-00165-4.
Matosin N, Newell KA, Quidé Y, et al. Effects of common GRM5 genetic variants on cognition, hippocampal volume and MGLUR5 protein levels in schizophrenia. Brain Imaging Behav. 2017;12:509–17. https://doi.org/10.1007/s11682-017-9712-0.
Hu T-M, Wu C-L, Hsu S-H, et al. Ultrarare loss-of-function mutations in the genes encoding the ionotropic glutamate receptors of kainate subtypes associated with schizophrenia disrupt the interaction with PSD95. J Personaliz Med. 2022;12:783. https://doi.org/10.3390/jpm12050783.
Lord C, Brugha TS, Charman T, et al. Autism spectrum disorder. Nat Rev Dis Primers. 2020. https://doi.org/10.1038/s41572-019-0138-4.
Vicidomini C, Ponzoni L, Lim D, et al. Pharmacological enhancement of mglu5 receptors rescues behavioral deficits in Shank3 knock-out mice. Mol Psychiatry. 2016;22:689–702. https://doi.org/10.1038/mp.2016.30.
Harony-Nicolas H, Kay M, du Hoffmann J, et al. Oxytocin improves behavioral and electrophysiological deficits in a novel Shank3-deficient rat. eLife. 2017. https://doi.org/10.7554/elife.18904.
Amal H, Barak B, Bhat V, et al. Shank3 mutation in a mouse model of autism leads to changes in the S-nitroso-proteome and affects key proteins involved in vesicle release and synaptic function. Mol Psychiatry. 2018;25:1835–48. https://doi.org/10.1038/s41380-018-0113-6.
Bidinosti M, Botta P, Krüttner S, et al. CLK2 inhibition ameliorates autistic features associated with Shank3 deficiency. Science. 2016;351:1199–203. https://doi.org/10.1126/science.aad5487.
Han Q, Kim YH, Wang X, et al. Shank3 deficiency impairs heat hyperalgesia and TRPV1 signaling in primary sensory neurons. Neuron. 2016;92:1279–93. https://doi.org/10.1016/j.neuron.2016.11.007.
Rapanelli M, Williams JB, Ma K, et al. Targeting histone demethylase LSD1 for treatment of deficits in autism mouse models. Mol Psychiatry. 2022;27:3355–66. https://doi.org/10.1038/s41380-022-01508-8.
Sauer AK, Bockmann J, Steinestel K, et al. Altered intestinal morphology and microbiota composition in the autism spectrum disorders associated Shank3 Mouse model. Int J Mol Sci. 2019;20:2134. https://doi.org/10.3390/ijms20092134.
Guo B, Chen J, Chen Q, et al. Anterior cingulate cortex dysfunction underlies social deficits in Shank3 Mutant Mice. Nat Neurosci. 2019;22:1223–34. https://doi.org/10.1038/s41593-019-0445-9.
Rhine MA, Parrott JM, Schultz MN, et al. Hypothesis-driven investigations of diverse pharmacological targets in two mouse models of autism. Autism Res. 2019;12:401–21. https://doi.org/10.1002/aur.2066.
Han K, et al. SHANK3 overexpression causes manic-like behavior with unique pharmacogenetic properties. Nature. 2013;503(7474):72–7. https://doi.org/10.1038/nature12606.
Lee K, Mills Z, Cheung P, et al. The role of zinc and NMDA receptors in autism spectrum disorders. Pharmaceuticals. 2022;16:1. https://doi.org/10.3390/ph16010001.
Kolevzon A, Breen MS, Siper PM, et al. Clinical trial of insulin-like growth factor-1 in Phelan-McDermid syndrome. Molecular Autism. 2022. https://doi.org/10.1186/s13229-022-00493-7.
Won H, et al. Autistic-like social behaviour in Shank2-mutant mice improved by restoring NMDA receptor function. Nature. 2012;486(7402):261–5. https://doi.org/10.1038/nature11015.
Betancur C, Buxbaum JD. SHANK3 haploinsufficiency: a “common” but underdiagnosed highly penetrant monogenic cause of autism spectrum disorders. Molecular Autism. 2013;4(1):17. https://doi.org/10.1186/2040-2392-4-17.
Speed HE, Kouser M, Xuan Z, et al. Apparent genetic rescue of adult shank3 exon 21 insertion mutation mice tempered by appropriate control experiments. Eneuro. 2019. https://doi.org/10.1523/eneuro.0317-19.2019.
Torossian A, Saré RM, Loutaev I, Smith CB. Increased rates of cerebral protein synthesis in Shank3 knockout mice: implications for a link between synaptic protein deficit and dysregulated protein synthesis in autism spectrum disorder/intellectual disability. Neurobiol Dis. 2021;148: 105213. https://doi.org/10.1016/j.nbd.2020.105213.
Uchino S, Waga C. Shank3 as an autism spectrum disorder-associated gene. Brain Develop. 2013;35:106–10. https://doi.org/10.1016/j.braindev.2012.05.013.
Goyal DK, Neil J, Simmons SD, et al. Zinc deficiency in autism: A controlled study. Insights Biomed. 2019. https://doi.org/10.36648/2572-5610.4.3.63.
Fourie C, Vyas Y, Lee K, et al. Dietary zinc supplementation prevents autism related behaviors and striatal synaptic dysfunction in Shank3 exon 13–16 mutant mice. Front Cell Neurosci. 2018. https://doi.org/10.3389/fncel.2018.00374.
Voineagu I, Wang X, Johnston P, et al. Transcriptomic analysis of autistic brain reveals convergent molecular pathology. Nature. 2011;474:380–4. https://doi.org/10.1038/nature10110.
Varesio C, Grumi S, Zanaboni MP, et al. Ketogenic dietary therapies in patients with autism spectrum disorder: facts or fads? A scoping review and a proposal for a shared protocol. Nutrients. 2021;13:2057. https://doi.org/10.3390/nu13062057.
Karhu E, Zukerman R, Eshraghi RS, et al. Nutritional interventions for autism spectrum disorder. Nutr Rev. 2019;78:515–31. https://doi.org/10.1093/nutrit/nuz092.
Liu C, Wang Y, Deng J, et al. Social deficits and repetitive behaviors are improved by early postnatal low-dose VPA intervention in a novel shank3-deficient Zebrafish model. Front Neurosci. 2021. https://doi.org/10.3389/fnins.2021.682054.
Funding
The authors declare that no funds, grants, or other support were received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
The author Xingshen Li wrote the main manuscript text, prepared the table and the figure and reviewed the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable. This study did not involve human participants, human data or tissues, or animals. Therefore, no ethical approval was required.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, X. Unravelling the role of SHANK3 mutations in targeted therapies for autism spectrum disorders. Discov Psychol 4, 110 (2024). https://doi.org/10.1007/s44202-024-00223-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44202-024-00223-5