Abstract
Plants have evolved a complex and elaborate signaling network to respond appropriately to the pathogen invasion by regulating expression of defensive genes through certain transcription factors. The APETALA2/ethylene response factor (AP2/ERF) family members have been determined as key regulators in growth, development, and stress responses in plants. Moreover, a growing body of evidence has demonstrated the critical roles of AP2/ERFs in plant disease resistance. In this review, we describe recent advances for the function of AP2/ERFs in defense responses against microbial pathogens. We summarize that AP2/ERFs are involved in plant disease resistance by acting downstream of mitogen activated protein kinase (MAPK) cascades, and regulating expression of genes associated with hormonal signaling pathways, biosynthesis of secondary metabolites, and formation of physical barriers in an MAPK-dependent or -independent manner. The present review provides a multidimensional perspective on the functions of AP2/ERFs in plant disease resistance, which will facilitate the understanding and future investigation on the roles of AP2/ERFs in plant immunity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants have developed elaborate and complex response mechanisms to cope with challenges of various pathogens (e.g. fungi, bacteria, and viruses). Pathogen-associated molecular pattern (PAMP)-triggered immunity (PTI) and effector-triggered immunity (ETI) are the two layers of immune systems that play critical roles in defense responses (Dangl, et al. 2013; Ngou, et al. 2022). PAMPs are perceived and recognized by pattern recognition receptors (PRRs) localized at the cell surface which initiate PTI, the first layer of plant immune system (Bernoux, et al. 2022; Yuan, et al. 2023). To survive and colonize host plants, pathogens often secrete virulence effectors into host cells to suppress PTI. Plants counteract the dampened PTI using resistance (R) genes which are often intracellular proteins with nucleotide-binding domain leucine-rich repeat (NLR) domains that directly or indirectly detect pathogen effectors and activate ETI (Jones, et al. 2016; Ngou, et al. 2022) (Table 1).
These two layers of immune system are commonly associated with various plant immune responses, such as accumulation of salicylic acid (SA), reactive oxygen species (ROS) production, and the expression of various defense-related genes (Greenberg and Yao 2004; van der Hoorn and Kamoun 2008). Generally, in the early stage of PTI, cellular responses include the activation of mitogen-activated protein kinase (MAPK), cytoplasmic Ca2+ influx, apoplastic alkalinization, stomata closure, membrane depolarization, production of ROS, and genome-wide transcriptional reprogramming (Bigeard, et al. 2015; Zhang, et al. 2023a, b, c). While in the late stage of PTI, cellular responses include biosynthesis of secondary metabolites and defense hormones SA and ethylene (ET), and deposition of callose and lignin in plant cell walls (Clay, et al. 2009; Zhang, et al. 2023a, b, c).
Transcription factors (TFs) are proteins that bind specifically to cis-regulatory elements in the promoter regions of genes, resulting in activation or suppression of gene expression (Pireyre and Burow 2015; Sohn, et al. 2006; Xing, et al. 2017). TF family APETALA2/ethylene response factors (AP2/ERFs) is a conserved superfamily widely involved in plant growth and development, including light acclimation, leaf senescence, flower pedicel abscission, and fruit ripening (Dey and Corina Vlot 2015; Koyama, et al. 2013; Nakano, et al. 2014; Vogel, et al. 2014; Zhai, et al. 2022). Moreover, AP2/ERFs also play key roles in regulating plant responses to various biotic and abiotic stresses, including salinity, drought, temperature, and multiple pathogens (e.g. viruses, fungi, and bacteria) (Qi, et al. 2023; Xie, et al. 2019). AP2/ERF superfamily is traditionally considered to specifically exist in plants, until its orthologs have been identified in genomes of microbes, such as the cyanobacterium Trichodesmium erythraeum, the virus Enterobacteria phage Rb49 (Magnani, et al. 2004; Wessler 2005). To date, a huge number of genes encoding AP2/ERF proteins have been identified in multiple plant species, including Solanum lycopersicum (146 genes) (Pirrello, et al. 2012), Malus domestica (259 genes) (Girardi, et al. 2013), sweet orange (Citrus sinensis, 108 genes) (Ito, et al. 2014), and others. Different types of duplication events, such as tandem and segmental duplication, or whole-genome duplication may have contributed to AP2/ERFs expansion across plant kingdom (Zhang, et al. 2021a, b, c; Zhuang, et al. 2009). The duplication and expansion enable functional differentiation of AP2/ERF family members to regulate diverse cellular processes, one of the key functions being defense responses (Feng, et al. 2020; Gao, et al. 2020).
A rising body of evidence shows that AP2/ERFs are closely involved in defense responses against various pathogens in plants (Reboledo, et al. 2022; Zang, et al. 2021). For example, overexpressing NtERF5 increases expression levels of pathogenesis-related (PR) genes and confers resistance to tobacco mosaic virus in tobacco (Nicotiana tobacum), while overexpressing Pti4/5/6 confer increased resistance to Erysiphe orontii, and Pseudomonas syringae pv tomato in Arabidopsis (Fischer and Dröge-Laser 2004; Gu, et al. 2002). Thus, this review will focus on the roles of AP2/ERF family in plant immune responses, and summarize AP2/ERFs as direct targets of MAPK signaling and how they regulate genes associated with defense hormonal signaling pathways, synthesis of antimicrobial secondary metabolites and physical barriers.
Features, classification, and binding specificity of AP2/ERF family
Member of AP2/ERF family encode a conserved APETALA2 (AP2)/Ethylene Responsive Element Binding Factor (EREB) domain, which contains 60–70 amino acid residues and is responsible for recognizing and binding to cis-regulatory elements in target genes (Nakano, et al. 2006; Okamuro, et al. 1997; Xie, et al. 2022). More specifically, arginine and tryptophan residues within the β-sheet of the AP2/EREB domain are pivotal for DNA binding (Aiese Cigliano, et al. 2013). In plants, the first AP2/EREB domain-containing protein was identified in model plant Arabidopsis thaliana (Jofuku, et al. 1994).
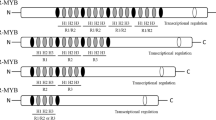
The AP2/ERF superfamily can be catagorized into four families: AP2, ERF, RAV (Related to Abscisic acid insensitive3/Viviparous1), and Soloist, mainly based on the number of AP2/EREB domains and the biological function of the members (Feng, et al. 2020; Nakano, et al. 2006). Among them, the members of RAV and ERF family usually have a single AP2/EREB domain, while AP2 family members usually contain two (Fig. 1). For RAV family members, they usually contain a single AP2/EREB domain and another DNA-binding domain, B3 domain (Fig. 1). In addition, depending on conserved amino acid residues of the DNA binding domains, the ERF family members are further classified into two subfamilies, ERF and the CBF/DREB (C-repeat-binding factor/dehydration-responsive element-binding protein) (Fig. 1) (Lata and Prasad 2011; Nakano, et al. 2006; Sakuma, et al. 2002). Specifically, in CBF/DREB proteins, the 14th valine and the 19th glutamic acid are conserved in the DNA binding domain, while these two positions are occupied by alanine and aspartic in the DNA binding domain of ERF proteins, respectively (Liu, et al. 1998; Sakuma, et al. 2002). These two positions are located on the β-sheet in the DNA binding domain, which determines the DNA binding affinity and specificity of the two subfamilies, resulting in regulation of different sets of stress-responsive genes (Liu, et al. 1998; Sakuma, et al. 2002). For instance, members of CBF/DREB subfamily usually regulate ABA, drought, heat, and cold-responsive genes (Agarwal, et al. 2006), while members of the ERF subfamily preferentially regulate expression of genes associated with ethylene response, biotic stress, and disease resistance (Table 1) (Amorim, et al. 2017; Ohme-Takagi and Shinshi 1995).
Schematic representation of domain structures of different types of AP2/ERFs. The position and amount of domains in different types of AP2/ERFs were illustrated. AP2, APETALA2; ERF, ethylene response factors; RAV, RELATED to ABSCISIC ACID INSENSITIVE 3/VIVIPAROUS 1; DREB, dehydration-responsive element-binding protein
As classical TFs, AP2/ERF proteins directly bind to the cis-acting elements within the promoter of targets, leading to activation or suppression of target genes. Different families of AP2/ERFs preferentially bind to different elements. For example, CBF/DREB subfamily members bind to DRE/CRE (Dehydration-Responsive or C-Repeat Element) motif with core sequence of A/GCCGAC on abiotic-stress responsive genes, while ERFs prefer to bind ERE (Ethylene-Response Element) motif with core sequence of AGCCGCC (named GCC-box) on the biotic-stress responsive genes (Xie, et al. 2019; Zhai, et al. 2022). Additionally, the B3 and AP2 domains of the RAV family members specifically bind to motifs with core sequence of CAACA and CACCTG, respectively (Kagaya, et al. 1999). However, some ERFs and CBF/DREBs in Arabidopsis bind both DRE/CRE and ERE elements, indicating they function in response to both biotic and abiotic stresses (Xie, et al. 2019). Additionally, multiple cis-elements have been reported to be recognized and bound by AP2/ERFs, such as HRPE (Hypoxia-Responsive Promoter Element), CE1 (Coupling Element 1), and others (Xie, et al. 2019). As aforementioned, the DNA-binding affinity of AP2/ERF proteins can be affected by certain amino acid residues of the AP2 domain, resulting in different DNA-binding specificity of different AP2/ERF proteins (Shoji, et al. 2013). Moreover, the binding affinity of AP2/ERFs can also be affected by flanking sequences of the cis-elements, leading to various binding specificity of AP2/ERFs to different target genes (Shoji, et al. 2013). The family specific preferential cis-element sequences were found to be conserved across a range of plant species including Zea mays (Liu, et al. 2013), Oryza sativa (Wan, et al. 2011), Triticum aestivum (Gao, et al. 2018), and Glycine max (Zhang, et al. 2009).
AP2/ERFs play vital roles in MAPK cascades-mediated plant disease resistance
Mitogen-activated protein kinase (MAPK)-mediated signaling pathways are conserved among all eukaryotes. In plants, MAPKs usually act downstream of sensors/receptors that recognize endogenous stimuli (e.g. peptide ligands) or exogenous stimuli (e.g. PAMPs and environmental factors) to coordinate plant growth, development, and immunity (Zhang, et al. 2018; Zhang and Zhang 2022). Activated MAPKs mediate the phosphorylation of various downstream substrates, such as protein kinases, TFs, structural proteins, and other enzymes to activate cellular responses (Zhang, et al. 2018; Zhang and Zhang 2022).
MAPK signaling pathways are among the early stage responses of PTI, and MAPK cascades have been widely demonstrated to be important signaling modules in plant disease resistance (Cheong, et al. 2003; Meng, et al. 2013; Zhang, et al. 2018; Zhang and Zhang 2022). To date, multiple AP2/ERF TFs have been determined to be substrates of MAPKs connecting plant defense responses (Table 1 and Fig. 2) (Bethke, et al. 2009; Cao, et al. 2018; Wang, et al. 2013). For example, a rice MAP kinase BWMK1 (blast and wounding-activated MAP kinase 1) phosphorylates and promotes binding affinity of OsEREBP1 (rice ethylene-responsive element-binding protein 1) to the GCC-box within the promoter of the PR genes, resulting in enhanced expression of these PRs and disease resistance (Cheong, et al. 2003). Additionally, two different groups reported that MPK3/MPK6, two partially redundant MAPKs in Arabidopsis, phosphorylate ERF6 and ERF72 to enhance disease resistance to Botrytis cinerea via different pathways (Li, et al. 2022a, b; Meng, et al. 2013). Specifically, MPK3/MPK6 phosphorylate ERF6 to increase its protein stability and DNA binding affinity to enhance the expression of defensive genes, including PDF1.1 and PDF1.2, while MPK3/MPK6 phosphorylate ERF72 to promote its transactivation activity, leading to increased accumulation of camalexin and elevated immunity (Li, et al. 2022a, b; Meng, et al. 2013). Moreover, MAPK kinase 4 (GmMKK4)-GmMPK6 module phosphorylates GmERF113 to promote its protein stability and transcriptional activity, resulting in increased expression of defensive genes (e.g. GmPR1 and GmPR10-1) and enhanced immune responses to Phytophthora sojae in Glycine max (Gao, et al. 2022).
AP2/ERF TFs are involved in plant disease-resistance through multiple signaling pathways. In plant defence responses, pathogen-secreted molecules are perceived by membrane-localized PRRs, which subsequently induce cellular responses, such as MAPK signaling cascades. AP2/ERFs could serve as substrates of MAPKs (labeled in orange) and are involved in MAPK signaling-mediated plant defence responses. Moreover, AP2/ERFs serve as TFs to regulate genes involved in hormonal signaling pathways, secondary metabolites biosynthesis, and formation of physical barriers (cuticle and cell wall). AP2/ERFs are typical transcription factors, they usually modulate cellular responses at transcription level in the nucleus, as indicated by the red-labeled processes. Molecules labeled in blue indicate that they interact with and enhance the DNA-binding activity or transactivition activity of corresponding TFs
AP2/ERFs regulate plant disease resistance as transcriptional activators or repressors
As typical TFs, AP2/ERFs might act as transcriptional activator or repressor to regulate plant disease resistance by binding to corresponding targets. For instance, AP2/ERF transgenic plants develop immunity to a variety of pathogens by activating expression of multiple defense genes (Liu, et al. 2023; Zhu, et al. 2022). Specifically, overexpressing GmERF113 in soybean (cultivar Dongnong 50) activates expression of GmPR1 and GmPR10-1 to enhance resistances to infection of Phytophthora sojae (Zhao, et al. 2017). Similarly, overexpressing OsERF83 in rice significantly inhibits the lesion formation induced by fungal pathogen Magnaporthe oryzae, suggesting OsERF83 plays a positive role in regulating resistance against rice blast (Tezuka, et al. 2019).
In some cases, AP2/ERFs act as transcriptional repressor to participate in plant disease resistance. For instance, AtERF9 is a transcription repressor binding to the GCC-box of PATHGEN-INDUCIBE PLANT DEFENSIN (PDF1.2) gene, and knocking out AtERF9 significantly promotes AtPDF1.2 expression, resulting in enhanced resistance to Botrytis cinerea (Maruyama, et al. 2013). Moreover, some AP2/ERF proteins contain a conserved C-terminal (L/F)DLN(L/F)xP motif, also known as EAR (ERF-associated amphiphilic repression) motif, and function to inhibit target gene expression (Fujimoto, et al. 2000). In yeast cells, EAR type StERF3 from tomato could bind to the promoter of HIS3 gene via the GCC-box element (Tian, et al. 2015). In tomato, silencing StERF3 promotes the expression of defensive genes (PR1, NPR1, and WRKY1), resulting in enhanced immunity to Phytophthora infestans (Tian, et al. 2015).
AP2/ERFs integrates hormonal signaling to regulate plant disease resistance
SA, jasmonic acid (JA), and ET are the three major phytohormones that are extensively involved in plant defense responses. SA is mainly responsible for immune responses against biotrophic pathogens, while JA and ET signaling are synergistically regulated and are closely associated with immunity against necrotrophic pathogens and herbivores (Li, et al. 2019a, b; Peng, et al. 2021; Zhang, et al. 2023a, b, c). Moreover, SA and JA/ET usually regulate disease resistance in an antagonistic manner, but new evidence has proved that they may also function synergistically to regulate plant immunity against virus infection in rice (Li, et al. 2019a, b; Zhang, et al. 2023a, b, c).
SA is the most important defense hormone in plant immune system. SA regulates the immune responses either in local infection site or in systemic organs through the master regulator NPR1 in its signaling pathway (Peng, et al. 2021; Zhang and Li 2019). AP2/ERFs have been reported to regulate plant resistance by regulating SA biosynthesis or signaling pathway (Table 1 and Fig. 2) (Hawku, et al. 2021; Wang, et al. 2020). For instance, MdERF11 positively regulates defense responses against Botryosphaeria dothidea by promoting SA biosynthesis in apple (Malus doemstica) (Wang, et al. 2020). ERF11 is induced upon treatment of SA and Pst DC3000, and disruption of ERF11 suppresses SA-mediated defense responses, leading to decreased resistance to Pst DC3000 (Zheng, et al. 2019). Further analysis shows that ERF11 could activate the expression of BT4 (BTB and TAZ domain protein 4), which encodes a positive regulator in SA-mediated immunity, by binding to its promoter through GCC-box (Zheng, et al. 2019). These data indicate that ERF11 acts upstream of BT4 to regulate SA response and disease resistance (Zheng, et al. 2019).
In addition to SA, AP2/ERFs are also involved in regulating disease resistance through JA/ET signaling pathway (Table 1 and Fig. 2). For instance, constitutive expression of ERF5 or ERF6 promotes expression of JA/ET-responsive genes, leading to enhanced immunity to Botrytis cinerea, a typical necrotrophic pathogen (Moffat, et al. 2012). Another AP2/ERF protein ORA59 is reported to promote JA/ET response by directly binding to the two GCC-boxes on the promoter of PDF1.2 to elevate immune response to Botrytis cinerea (Huang, et al. 2022; Zarei, et al. 2011). In contrast to ORA59 trans-activating PDF1.2, AtERF9 is reported to repress PDF1.2 expression by targeting to the GCC-box within its promoter, suggesting different AP2/ERFs might co-regulate the same target in either positive or negative manner (Maruyama, et al. 2013).
More importantly, AP2/ERFs are among key factors that mediate the crosstalk between signaling pathways of SA and JA/ET (Fig. 2). Analysis revealed that GCC-box motifs are present in the promoters of various JA/ET-responsive genes, and those motifs are critical for SA-induced repression of JA/ET-responsive genes (Van der Does, et al. 2013; Zander, et al. 2014). ORA59 plays key roles in JA/ET-mediated defense responses (Huang, et al. 2022; Zarei, et al. 2011), and SA could down-regulate both the transcription and protein accumulation of ORA59 (Van der Does, et al. 2013; Zander, et al. 2014). Thus, ORA59 stands at the middle of SA and JA/ET signaling to modulate the SA-mediated repression of JA/ET response. Moreover, ectopic expression of MdERF100 promotes expression of PR1 and PDF1.2, two marker genes of SA- and JA-signaling pathway, respectively, resulting in enhanced immune response to powdery mildew in Arabidopsis (Zhang, et al. 2021a, b, c).
In some cases, AP2/ERFs regulate disease resistance by modulating the biosynthesis of SA and JA simultaneously. For example, OsERF3 positively regulates resistance to herbivores by increasing the accumulation of SA, JA, and ET in rice (Lu, et al. 2011). In addition, SlERF2 also promotes accumulation of SA and JA to increase immune response to fungal pathogen Stemphylium lycopersici in tomato (Yang, et al. 2021a, b).
AP2/ERFs regulate plant disease resistance by manipulating biosynthesis of secondary metabolites
Plant metabolites are traditionally categorized into two classes, primary and secondary. Primary metabolites are widely present in all plants and are necessary for plant growth and development, while secondary metabolites are involved in multiple physiological and biochemical processes, such as adaptation to harsh environments and defense responses (De Geyter, et al. 2012; Zhan, et al. 2022). Based on the core structures, secondary metabolites can be classified into three major groups: nitrogen/sulfur-containing compounds (e.g. alkaloids, glucosinolates, and cyanogenic glycosides), terpenoids/isoprenoids, and phenolics (Marone, et al. 2022). Secondary metabolites contribute to plant defencse responses via multiple pathways, including serving as antioxidants to scavenge ROS, being toxic to invaders (e.g. microbial pathogens, herbivores, and competing plant species), triggering the expression of defense-related genes, and accumulation of other metabolites (Zhan, et al. 2022).
TFs have been shown to be extensively associated with regulation of the secondary metabolites synthesis in response to biotic stresses, and AP2/ERFs are among them (Table 1 and Fig. 2) (Imano, et al. 2021; Nakayasu, et al. 2018; Shoji, et al. 2010). Resveratrol is a major stilbene-type phytoalexin characterized with antimicrobial activity in plants and medical properties in humans (Hart 1981; Jang et al. 1997). VqMYB35 directly binds to the promoters and activates the expression of stilbene synthase-encoding genes, and VqERF114 could interact with VqMYB35 and enhance its transactivation activity, leading to increased stilbene biosynthesis in Vitis quinquangularis (Wang and Wang 2019). Nicotine is a type of alkaloids that mainly produced in Nicotiana species, and it is involved in immune responses against insect and herbivore invasion (Zenkner, et al. 2019). JA and methyl JA (MeJA) promote nicotine formation by inducing the expression of putrescine N-methyltransferase-encoding gene (NtPMT1a), which catalyzes the first committed step in nicotine pyrrolidine ring formation (Baldwin, et al. 1994; Sachan and Falcone 2002). A set of AP2/ERFs, including NtERF1, NtERF32, and NtERF121, activate the expression of NtPMT1a by binding specifically to its promoter, leading to elevated accumulation of nicotine and total alkaloid (Sears, et al. 2014). In addition to nicotine, JA/ET treatment also induced the biosynthesis of hydroxycinnamic acid amides (HCAAs), another type of antimicrobial metabolites that are associated with immune responses against necrotrophic pathogens (Campos, et al. 2014). Agmatine coumaryl transferease (ACT) is a key enzyme in HCAAs biosynthesis (Muroi, et al. 2009). Further investigations show that ORA59, a critical regulator in JA/ET-mediated defense signaling pathways, activates the ACT expression by binding to the GCC-box within its promoter, resulting in increased accumulation of HCAAs (Li, et al. 2018).
Additionally, AP2/ERFs have also been reported to regulate terpenoid indole alkaloids (TIAs) biosynthesis in both Ophiorrhiza pumila (Udomsom, et al. 2016) and Catharanthus roseus (Paul, et al. 2020, 2017), steroidal glycoalkaloids (SGAs) synthesis in both Solanum lycopersicum and Solanum tuberosum (Cárdenas, et al. 2016; Nakayasu, et al. 2018; Thagun, et al. 2016), and artemisinin production in Artemisia annua (Lu, et al. 2013; Yu, et al. 2012). These results confirm the critical roles of AP2/ERFs in modulating metabolism of secondary metabolites to regulate plant immunity.
AP2/ERFs regulates plant disease resistance by mediating biosynthesis of physical barriers
In plant defense systems, plant cell wall is one of the key barriers that mirobes need to break to achieve successful colonization in host plant tissues. Plant cell wall functions in several facets to prevent pathogen invasion. First, cell wall is a physical barrier, as colonization inside host cells require breakdown of wall matrix by microbial-secreted cell wall-degrading enzymes (Miedes, et al. 2014). Upon cell wall breakdown, the cell wall-docked antimicrobial compounds are released and trigger downstream immune responses (Miedes, et al. 2014; Yang, et al. 2021a, b). Additionally, impaired cell wall integrity results in release of signaling molecules (e.g. damage-associate molecular patterns, DAMPs) and activation of specific immune responses, including SA and JA signaling pathways (Cantu, et al. 2008; Chowdhury, et al. 2019; Engelsdorf, et al. 2019).
Lignin, one of the key constituents of plant cell walls (CWs), is closely involved in determining the mechanical strength, antioxidation, and hydrophobicity of CWs (Cesarino 2019; Vanholme, et al. 2019). Upon pathogen invasion, the accumulated lignin offers a fundamental barrier to exclude invaders outside the host cells. It also limits the substance exchange between host and microbes, such as restricting mirobe-secreted toxins and enzymes entering into host cells, as well as the nutrients transportation from hosts to invaders (Dong and Lin 2021; Ma, et al. 2018). Thus, lignin has become a critical target for developing strategies to control plant diseases. Multiple investigations have reported the critical function of AP2/ERFs in regulating plant defense responses by manipulating lignin biosynthesis (Table 1 and Fig. 2). For example, overexpressing GbERF1-like significantly increases the transcripts of lignin biosynthetic genes, resulting increased lignin contents and enhanced resistance to Verticillium dahliae in upland cotton (Gossypium hirsutum) (Guo, et al. 2016). Two different groups report that ERF114 regulates disease resistance by manipulating lignin biosynthesis in both apple (Malus domestica) and Arabidopsis (Li, et al. 2022a, b; Liu, et al. 2023). Specifically, AtERF114 activates lignin binsynthetic gene AtPAL1 expression by directly binding to its promoter, leading to increased accumulation of lignin and elevated immunity in Arabidopsis (Li, et al. 2022a, b). PEROXIDASEs (PRXs) catalyze the lignin formation through oxidative polymerization of three monolignol precursors (Marjamaa, et al. 2009). In apple, MdERF114 activates MdPRX63 expression by directly binding to the GCC-box in its promoter, leading to increased lignin accumulation in apple roots and promoted immune responses against Fusarium solani (Liu, et al. 2023). Moreover, other TFs like MdWRKY75 and MdMYB8 promote the lignin formation by either regulating the expression or transcriptional activity of MdERF114, resulting in enhanced lignin deposition and disease resistance (Liu, et al. 2023).
In addition to the cell wall, the cuticle is another critical physical barrier that covers almost all the above ground organs of plants. The cuticle is composed of cutin and waxes, and plays critical roles in regulating epidermal permeability and nonstomatal water loss (Arya, et al. 2021; Sieber, et al. 2000). Similar with the function of cell wall, it not only defends against insects, fungi, and bacteria as physical barrier, but also acts as a chemical deterrent and activator of plant immune responses (Arya, et al. 2021). SHINE (SHN) transcription factors (TFs) belong to group V ERF subfamily and have been found to regulate the formation of the cuticle (Aharoni, et al. 2004; Riechmann and Meyerowitz 1998). Overexpressing SlSHN3 in tomato promotes the accumulation of cutin monomers, resulting in a more permeable cuticle, which contributes to enhanced resistance to Botrytis cinerea and Xanthomonas campestris pv. vesicatoria (Bessire, et al. 2007; Buxdorf, et al. 2014). Moreover, application of cutin monomers extracted from SlSHN3 overexpressing plants alters the expression of disease-related genes (e.g. PR1a and AOS), thereby restricting the development of disease symptom in wild-type plants (Buxdorf, et al. 2014). In addition, GhWIN2 (WAX INDUCER 2), a cotton (Gossypium hirsutum) homolog of SlSHN3, activates the cuticle biosynthetic genes and promotes cuticle formation in cotton (Li, et al. 2019a, b). However, silencing GhWIN2 promotes resistance to Verticillium dahliae, probably because of the increased content of SA in cotton (Li, et al. 2019a, b), consistent with previous notions that AP2/ERFs integrate multiple pathways to regulate disease resistance in plants.
Conclusions and future perspectives
The AP2/ERFs are critical stress-responsive TFs that are tightly associated with immune responses in plants. Here, we mainly summarize the functions of AP2/ERFs in plant defensive responses against microorganisms, such as fungi, bacteria, and viruses. Multiple AP2/ERF-encoding genes are induced by pathogens or defence-related phytohormones (e.g. SA, JA, or ET). Upon pathogen invasion, AP2/ERFs may serve as substrates of MAPKs to associate with MAPK signaling cascade-mediated defensive pathways (Table 1 and Fig. 2). Moreover, AP2/ERFs function as TFs to reprogram expression of defence-related genes involved in hormonal signaling pathways, secondary metabolism, and formation of physical barriers to regulate plant disease resistance (Tabel 1 and Fig. 2). Importantly, some AP2/ERFs may function as a signal hub or mediate the crosstalk between different signaling pathways. For example, while ORA59 serves as a master regulator in JA/ET-mediated defence response (Huang, et al. 2022; Zarei, et al. 2011), it can also be regulated by SA at both transcriptional and protein levels (Van der Does, et al. 2013; Zander, et al. 2014). Moreover, it is also directly involved in regulating biosynthesis of HCAAs, an important antimicrobial metabolite in plant defence system (Li, et al. 2018). Therefore, novel AP2/ERF family members that regulate plant disease resistance in multiple facets, as well as the underlying mechanism, need to be explored in the future researches.
At the current stage, with the rapid development of modern technologies, such as next generation sequencing, multi-omics analysis, the gene-editing system clustered regularly interspaced short palindromic repeats/Cas9 (CRISPR/Cas9), genome-wide association studies (GWAS), and others, the exploration and dissection of AP2/ERFs regulatory network will broaden our understanding of their roles in plant disease resistance. It is worth to mention that, in addition to transcriptional regulation, post-translational modification (PTM) of proteins are also tightly associated with the regulation of plant immunity (Gough and Sadanandom 2021; Withers and Dong 2017; Yin, et al. 2019). PTMs are highly specific alterations of protein structures, and most of them are rapid and reversible in plant cells (Jensen 2004). To date, more than 300 types of PTMs have been identified in plants, such as acetylation, ubiquitination, methylation, sumoylation, phosphorylation, glycosylation, and so on (Jensen 2004). PTMs usually affect the protein conformation, localization, stability, activity, and protein–protein interactions (Jensen 2004; Withers and Dong 2017). For AP2/ERFs, multiple reports have demonstrated their functions in plant immunity. For example, MAPKs target and phosphorylate AP2/ERFs to regulate their protein stability, DNA binding activity to regulate AP2/ERF-mediated plant disease resistance (Gao, et al. 2022; Li, et al. 2022a, b). In addition, a BTB/POZ domain protein GmBTB/POZ promotes the ubiquitination and degradation of AP2/ERF-like TF GmAP2, a negative regulator of plant resistance to Phytophthora sojae, resulting in enhanced resistance to Phytophthora sojae in soybean (Glycine max) (Zhang, et al. 2021a, b, c). Therefore, identification of PTMs of AP2/ERFs using multi-omics analysis will facilitate the dissection of molecular mechanisms underlying AP2/ERFs-regulated plant immunity. With the identification of novel AP2/ERFs, as well as the uncovering of underlying mechanism, AP2/ERFs can be potential candidates for disease resistance breeding in many crops.
Availability of data and materials
Not applicable.
Abbreviations
- ACT:
-
Agmatine coumaryl transferease
- AP2/ERF:
-
APETALA2/ethylene response factor
- CBF/DREB:
-
C-repeat-binding factor dehydration-responsive element-binding protein
- CE1:
-
Coupling Element 1
- CW:
-
Cell wall
- DRE/CRE:
-
Dehydration-Responsive or C-Repeat Element
- EAR:
-
ERF-associated amphiphilic repression
- EREB:
-
Ethylene Responsive Element Binding Factor
- ET:
-
Ethylene
- ETI:
-
Effector-triggered immunity
- HCAA:
-
Hydroxycinnamic acid amide
- HR:
-
Hypersensitivity reaction
- HRPE:
-
Hypoxia-Responsive Promoter Element
- MAPK:
-
Mitogen-activated protein kinase
- NLR:
-
Nucleotide-binding domain, leucine-rich-repeat containing receptor
- PAMP:
-
Pathogen-associated molecular patterns
- PDF1.2:
-
PATHGEN-INDUCIBE PLANT DEFENSIN
- PR:
-
Pathogenesis-related
- PRR:
-
Pattern recognition receptor
- PTI:
-
Pathogen-associated molecular patterns-triggered immunity
- ROS:
-
Reactive oxygen species
- SA:
-
Salicylic acid
- SGA:
-
Steroidal glycoalkaloid
- TIA:
-
Terpenoid indole alkaloid
References
Agarwal PK, Agarwal P, Reddy MK, Sopory SK (2006) Role of DREB transcription factors in abiotic and biotic stress tolerance in plants. Plant Cell Rep 25:1263–1274. https://doi.org/10.1007/s00299-006-0204-8
Aharoni A, Dixit S, Jetter R, Thoenes E, van Arkel G, Pereira A (2004) The SHINE clade of AP2 domain transcription factors activates wax biosynthesis, alters cuticle properties, and confers drought tolerance when overexpressed in Arabidopsis. Plant Cell 16:2463–2480. https://doi.org/10.1105/tpc.104.022897
Aiese Cigliano R, Sanseverino W, Cremona G, Ercolano MR, Conicella C, Consiglio FM (2013) Genome-wide analysis of histone modifiers in tomato: gaining an insight into their developmental roles. BMC Genomics 14:57. https://doi.org/10.1186/1471-2164-14-57
Amorim LLB, da Fonseca Dos Santos R, Neto JPB, Guida-Santos M, Crovella S, Benko-Iseppon AM (2017) Transcription factors involved in plant resistance to pathogens. Curr Protein Pept Sci 18:335–351. https://doi.org/10.2174/1389203717666160619185308
Arya GC, Sarkar S, Manasherova E, Aharoni A, Cohen H (2021) The plant cuticle: an ancient guardian barrier set against long-standing rivals. Front Plant Sci 12:663165. https://doi.org/10.3389/fpls.2021.663165
Baldwin IT, Schmelz EA, Ohnmeiss TE (1994) Wound-induced changes in root and shoot jasmonic acid pools correlate with induced nicotine synthesis inNicotiana sylvestris spegazzini and comes. J Chem Ecol 20:2139–2157. https://doi.org/10.1007/bf02066250
Bernoux M, Zetzsche H, Stuttmann J (2022) Connecting the dots between cell surface- and intracellular-triggered immune pathways in plants. Curr Opin Plant Biol 69:102276. https://doi.org/10.1016/j.pbi.2022.102276
Bessire M, Chassot C, Jacquat AC, Humphry M, Borel S, Petetot JM, Metraux JP, Nawrath C (2007) A permeable cuticle in Arabidopsis leads to a strong resistance to Botrytis cinerea. EMBO J 26:2158–2168. https://doi.org/10.1038/sj.emboj.7601658
Bethke G, Unthan T, Uhrig JF, Pöschl Y, Gust AA, Scheel D, Lee J (2009) Flg22 regulates the release of an ethylene response factor substrate from MAP kinase 6 in Arabidopsis thaliana via ethylene signaling. Proc Natl Acad Sci U S A 106:8067–8072. https://doi.org/10.1073/pnas.0810206106
Bigeard J, Colcombet J, Hirt H (2015) Signaling mechanisms in pattern-triggered immunity (PTI). Mol Plant 8:521–539. https://doi.org/10.1016/j.molp.2014.12.022
Buxdorf K, Rubinsky G, Barda O, Burdman S, Aharoni A, Levy M (2014) The transcription factor SlSHINE3 modulates defense responses in tomato plants. Plant Mol Biol 84:37–47. https://doi.org/10.1007/s11103-013-0117-1
Campos L, Lisón P, López-Gresa MP, Rodrigo I, Zacarés L, Conejero V, Bellés JM (2014) Transgenic tomato plants overexpressing tyramine N-hydroxycinnamoyl transferase exhibit elevated hydroxycinnamic acid amide levels and enhanced resistance to Pseudomonas syringae. Mol Plant Microbe Interact 27:1159–1169. https://doi.org/10.1094/mpmi-04-14-0104-r
Cantu D, Vicente AR, Labavitch JM, Bennett AB, Powell AL (2008) Strangers in the matrix: plant cell walls and pathogen susceptibility. Trends Plant Sci 13:610–617. https://doi.org/10.1016/j.tplants.2008.09.002
Cao FY, DeFalco TA, Moeder W, Li B, Gong Y, Liu XM, Taniguchi M, Lumba S, Toh S, Shan L, Ellis B, Desveaux D, Yoshioka K (2018) Arabidopsis Ethylene Response Factor 8 (ERF8) has dual functions in ABA signaling and immunity. BMC Plant Biol 18:211. https://doi.org/10.1186/s12870-018-1402-6
Cárdenas PD, Sonawane PD, Pollier J, Vanden Bossche R, Dewangan V, Weithorn E, Tal L, Meir S, Rogachev I, Malitsky S, Giri AP, Goossens A, Burdman S, Aharoni A (2016) GAME9 regulates the biosynthesis of steroidal alkaloids and upstream isoprenoids in the plant mevalonate pathway. Nat Commun 7:10654. https://doi.org/10.1038/ncomms10654
Cesarino I (2019) Structural features and regulation of lignin deposited upon biotic and abiotic stresses. Curr Opin Biotechnol 56:209–214. https://doi.org/10.1016/j.copbio.2018.12.012
Cheong YH, Moon BC, Kim JK, Kim CY, Kim MC, Kim IH, Park CY, Kim JC, Park BO, Koo SC, Yoon HW, Chung WS, Lim CO, Lee SY, Cho MJ (2003) BWMK1, a rice mitogen-activated protein kinase, locates in the nucleus and mediates pathogenesis-related gene expression by activation of a transcription factor. Plant Physiol 132:1961–1972. https://doi.org/10.1104/pp.103.023176
Chowdhury J, Coad BR, Little A (2019) Cell wall responses to biotrophic fungal pathogen invasion. Annual Plant Rev 2: 1001–1030. https://doi.org/10.1002/9781119312994.apr0634
Clay NK, Adio AM, Denoux C, Jander G, Ausubel FM (2009) Glucosinolate metabolites required for an Arabidopsis innate immune response. Science 323:95–101. https://doi.org/10.1126/science.1164627
Dangl JL, Horvath DM, Staskawicz BJ (2013) Pivoting the plant immune system from dissection to deployment. Science 341:746–751. https://doi.org/10.1126/science.1236011
De Geyter N, Gholami A, Goormachtig S, Goossens A (2012) Transcriptional machineries in jasmonate-elicited plant secondary metabolism. Trends Plant Sci 17:349–359. https://doi.org/10.1016/j.tplants.2012.03.001
Dey S, Corina Vlot A (2015) Ethylene responsive factors in the orchestration of stress responses in monocotyledonous plants. Front Plant Sci 6:640. https://doi.org/10.3389/fpls.2015.00640
Dong NQ, Lin HX (2021) Contribution of phenylpropanoid metabolism to plant development and plant-environment interactions. J Integr Plant Biol 63:180–209. https://doi.org/10.1111/jipb.13054
Engelsdorf T, Kjaer L, Gigli-Bisceglia N, Vaahtera L, Bauer S, Miedes E, Wormit A, James L, Chairam I, Molina A, Hamann T (2019) Functional characterization of genes mediating cell wall metabolism and responses to plant cell wall integrity impairment. BMC Plant Biol 19:320. https://doi.org/10.1186/s12870-019-1934-4
Feng K, Hou XL, Xing GM, Liu JX, Duan AQ, Xu ZS, Li MY, Zhuang J, Xiong AS (2020) Advances in AP2/ERF super-family transcription factors in plant. Crit Rev Biotechnol 40:750–776. https://doi.org/10.1080/07388551.2020.1768509
Fischer U, Dröge-Laser W (2004) Overexpression of NtERF5, a new member of the tobacco ethylene response transcription factor family enhances resistance to tobacco mosaic virus. Mol Plant Microbe Interact 17:1162–1171. https://doi.org/10.1094/mpmi.2004.17.10.1162
Fujimoto SY, Ohta M, Usui A, Shinshi H, Ohme-Takagi M (2000) Arabidopsis ethylene-responsive element binding factors act as transcriptional activators or repressors of GCC box-mediated gene expression. Plant Cell 12:393–404. https://doi.org/10.1105/tpc.12.3.393
Gao H, Jiang L, Du B, Ning B, Ding X, Zhang C, Song B, Liu S, Zhao M, Zhao Y, Rong T, Liu D, Wu J, Xu P, Zhang S (2022) GmMKK4-activated GmMPK6 stimulates GmERF113 to trigger resistance to Phytophthora sojae in soybean. Plant J 111:473–495. https://doi.org/10.1111/tpj.15809
Gao J, Zhang Y, Li Z, Liu M (2020) Role of ethylene response factors (ERFs) in fruit ripening. Food Qual Safety 4:15–20. https://doi.org/10.1093/fqsafe/fyz042
Gao T, Li GZ, Wang CR, Dong J, Yuan SS, Wang YH, Kang GZ (2018) Function of the ERFL1a transcription factor in wheat responses to water deficiency. Int J Mol Sci 19:1465. https://doi.org/10.3390/ijms19051465
Girardi CL, Rombaldi CV, Dal Cero J, Nobile PM, Laurens F, Bouzayen M, Quecini V (2013) Genome-wide analysis of the AP2/ERF superfamily in apple and transcriptional evidence of ERF involvement in scab pathogenesis. Sci Hortic 151:112–121. https://doi.org/10.1016/j.scienta.2012.12.017
Gough C, Sadanandom A (2021) Understanding and exploiting post-translational modifications for plant disease resistance. Biomolecules 11:1122. https://doi.org/10.3390/biom11081122
Greenberg JT, Yao N (2004) The role and regulation of programmed cell death in plant-pathogen interactions. Cell Microbiol 6:201–211. https://doi.org/10.1111/j.1462-5822.2004.00361.x
Gu YQ, Wildermuth MC, Chakravarthy S, Loh YT, Yang C, He X, Han Y, Martin GB (2002) Tomato transcription factors pti4, pti5, and pti6 activate defense responses when expressed in Arabidopsis. Plant Cell 14:817–831. https://doi.org/10.1105/tpc.000794
Guo W, Jin L, Miao Y, He X, Hu Q, Guo K, Zhu L, Zhang X (2016) An ethylene response-related factor, GbERF1-like, from Gossypium barbadense improves resistance to Verticillium dahliae via activating lignin synthesis. Plant Mol Biol 91:305–318. https://doi.org/10.1007/s11103-016-0467-6
Hawku MD, Goher F, Islam MA, Guo J, He F, Bai X, Yuan P, Kang Z, Guo J (2021) TaAP2–15, an AP2/ERF transcription factor, is positively involved in wheat resistance to Puccinia striiformis f. sp. tritici. Int J Mol Sci 22:2080. https://doi.org/10.3390/ijms22042080
Huang LJ, Zhang J, Lin Z, Yu P, Lu M, Li N (2022) The AP2/ERF transcription factor ORA59 regulates ethylene-induced phytoalexin synthesis through modulation of an acyltransferase gene expression. J Cell Physiol 1-11. https://doi.org/10.1002/jcp.30935
Imano S, Fushimi M, Camagna M, Tsuyama-Koike A, Mori H, Ashida A, Tanaka A, Sato I, Chiba S, Kawakita K, Ojika M, Takemoto D (2021) AP2/ERF transcription factor NbERF-IX-33 is involved in the regulation of phytoalexin production for the resistance of Nicotiana benthamiana to Phytophthora infestans. Front Plant Sci 12:821574. https://doi.org/10.3389/fpls.2021.821574
Ito TM, Polido PB, Rampim MC, Kaschuk G, Souza SG (2014) Genome-wide identification and phylogenetic analysis of the AP2/ERF gene superfamily in sweet orange (Citrus sinensis). Genet Mol Res 13:7839–7851. https://doi.org/10.4238/2014.september.26.22
Jensen ON (2004) Modification-specific proteomics: characterization of post-translational modifications by mass spectrometry. Curr Opin Chem Biol 8:33–41. https://doi.org/10.1016/j.cbpa.2003.12.009
Jofuku KD, den Boer BG, Van Montagu M, Okamuro JK (1994) Control of Arabidopsis flower and seed development by the homeotic gene APETALA2. Plant Cell 6:1211–1225. https://doi.org/10.1105/tpc.6.9.1211
Jones JD, Vance RE, Dangl JL (2016) Intracellular innate immune surveillance devices in plants and animals. Science 354:aaf6395. https://doi.org/10.1126/science.aaf6395
Kagaya Y, Ohmiya K, Hattori T (1999) RAV1, a novel DNA-binding protein, binds to bipartite recognition sequence through two distinct DNA-binding domains uniquely found in higher plants. Nucleic Acids Res 27:470–478. https://doi.org/10.1093/nar/27.2.470
Koyama T, Nii H, Mitsuda N, Ohta M, Kitajima S, Ohme-Takagi M, Sato F (2013) A regulatory cascade involving class II Ethylene response factor transcriptional repressors operates in the progression of leaf senescence. Plant Physiol 162:991–1005. https://doi.org/10.1104/pp.113.218115
Lata C, Prasad M (2011) Role of DREBs in regulation of abiotic stress responses in plants. J Exp Bot 62:4731–4748. https://doi.org/10.1093/jxb/err210
Li J, Zhang K, Meng Y, Hu J, Ding M, Bian J, Yan M, Han J, Zhou M (2018) Jasmonic acid/ethylene signaling coordinates hydroxycinnamic acid amides biosynthesis through ORA59 transcription factor. Plant J 95:444–457. https://doi.org/10.1111/tpj.13960
Li N, Han X, Feng D, Yuan D, Huang LJ (2019) Signaling crosstalk between Salicylic acid and Ethylene/Jasmonate in plant defense: do we understand what they are whispering? Int J Mol Sci 20:671. https://doi.org/10.3390/ijms20030671
Li X, Liu N, Sun Y, Wang P, Ge X, Pei Y, Liu D, Ma X, Li F, Hou Y (2019b) The cotton GhWIN2 gene activates the cuticle biosynthesis pathway and influences the salicylic and jasmonic acid biosynthesis pathways. BMC Plant Biol 19:379. https://doi.org/10.1186/s12870-019-1888-6
Li Y, Liu K, Tong G, Xi C, Liu J, Zhao H, Wang Y, Ren D, Han S (2022a) MPK3/MPK6-mediated phosphorylation of ERF72 positively regulates resistance to Botrytis cinerea through directly and indirectly activating the transcription of camalexin biosynthesis enzymes. J Exp Bot 73:413–428. https://doi.org/10.1093/jxb/erab415
Li Z, Zhang Y, Ren J, Jia F, Zeng H, Li G, Yang X (2022b) Ethylene-responsive factor ERF114 mediates fungal pathogen effector PevD1-induced disease resistance in Arabidopsis thaliana. Mol Plant Pathol 23:819–831. https://doi.org/10.1111/mpp.13208
Liu Q, Kasuga M, Sakuma Y, Abe H, Miura S, Yamaguchi-Shinozaki K, Shinozaki K (1998) Two transcription factors, DREB1 and DREB2, with an EREBP/AP2 DNA binding domain separate two cellular signal transduction pathways in drought- and low-temperature-responsive gene expression, respectively, in Arabidopsis. Plant Cell 10:1391–1406. https://doi.org/10.1105/tpc.10.8.1391
Liu S, Wang X, Wang H, Xin H, Yang X, Yan J, Li J, Tran LS, Shinozaki K, Yamaguchi-Shinozaki K, Qin F (2013) Genome-wide analysis of ZmDREB genes and their association with natural variation in drought tolerance at seedling stage of Zea mays L. PLoS Genet 9:e1003790. https://doi.org/10.1371/journal.pgen.1003790
Liu Y, Liu Q, Li X, Zhang Z, Ai S, Liu C, Ma F, Li C (2023) MdERF114 enhances the resistance of apple roots to Fusarium solani by regulating the transcription of MdPRX63. Plant Physiol 192:2015. https://doi.org/10.1093/plphys/kiad057
Lu J, Ju H, Zhou G, Zhu C, Erb M, Wang X, Wang P, Lou Y (2011) An EAR-motif-containing ERF transcription factor affects herbivore-induced signaling, defense and resistance in rice. Plant J 68:583–596. https://doi.org/10.1111/j.1365-313x.2011.04709.x
Lu X, Zhang L, Zhang F, Jiang W, Shen Q, Zhang L, Lv Z, Wang G, Tang K (2013) AaORA, a trichome-specific AP2/ERF transcription factor of Artemisia annua, is a positive regulator in the artemisinin biosynthetic pathway and in disease resistance to Botrytis cinerea. New Phytol 198:1191–1202. https://doi.org/10.1111/nph.12207
Magnani E, Sjölander K, Hake S (2004) From endonucleases to transcription factors: evolution of the AP2 DNA binding domain in plants. Plant Cell 16:2265–2277. https://doi.org/10.1105/tpc.104.023135
Ma Q-H, Zhu H-H, Qiao M-Y (2018) Contribution of both lignin content and sinapyl monomer to disease resistance in tobacco. Plant Pathol 67:642–650. https://doi.org/10.1111/ppa.12767
Marjamaa K, Kukkola EM, Fagerstedt KV (2009) The role of xylem class III peroxidases in lignification. J Exp Bot 60:367–376. https://doi.org/10.1093/jxb/ern278
Marone D, Mastrangelo AM, Borrelli GM, Mores A, Laidò G, Russo MA, Ficco DBM (2022) Specialized metabolites: physiological and biochemical role in stress resistance, strategies to improve their accumulation, and new applications in crop breeding and management. Plant Physiol Biochem 172:48–55. https://doi.org/10.1016/j.plaphy.2021.12.037
Maruyama Y, Yamoto N, Suzuki Y, Chiba Y, Yamazaki K, Sato T, Yamaguchi J (2013) The Arabidopsis transcriptional repressor ERF9 participates in resistance against necrotrophic fungi. Plant Sci 213:79–87. https://doi.org/10.1016/j.plantsci.2013.08.008
Meng X, Xu J, He Y, Yang KY, Mordorski B, Liu Y, Zhang S (2013) Phosphorylation of an ERF transcription factor by Arabidopsis MPK3/MPK6 regulates plant defense gene induction and fungal resistance. Plant Cell 25:1126–1142. https://doi.org/10.1105/tpc.112.109074
Miedes E, Vanholme R, Boerjan W, Molina A (2014) The role of the secondary cell wall in plant resistance to pathogens. Front Plant Sci 5:358. https://doi.org/10.3389/fpls.2014.00358
Moffat CS, Ingle RA, Wathugala DL, Saunders NJ, Knight H, Knight MR (2012) ERF5 and ERF6 play redundant roles as positive regulators of JA/Et-mediated defense against Botrytis cinerea in Arabidopsis. PLoS ONE 7:e35995. https://doi.org/10.1371/journal.pone.0035995
Muroi A, Ishihara A, Tanaka C, Ishizuka A, Takabayashi J, Miyoshi H, Nishioka T (2009) Accumulation of hydroxycinnamic acid amides induced by pathogen infection and identification of agmatine coumaroyltransferase in Arabidopsis thaliana. Planta 230:517–527. https://doi.org/10.1007/s00425-009-0960-0
Nakano T, Fujisawa M, Shima Y, Ito Y (2014) The AP2/ERF transcription factor SlERF52 functions in flower pedicel abscission in tomato. J Exp Bot 65:3111–3119. https://doi.org/10.1093/jxb/eru154
Nakano T, Suzuki K, Fujimura T, Shinshi H (2006) Genome-wide analysis of the ERF gene family in Arabidopsis and rice. Plant Physiol 140:411–432. https://doi.org/10.1104/pp.105.073783
Nakayasu M, Shioya N, Shikata M, Thagun C, Abdelkareem A, Okabe Y, Ariizumi T, Arimura GI, Mizutani M, Ezura H, Hashimoto T, Shoji T (2018) JRE4 is a master transcriptional regulator of defense-related steroidal glycoalkaloids in tomato. Plant J 94:975–990. https://doi.org/10.1111/tpj.13911
Ngou BPM, Jones JDG, Ding P (2022) Plant immune networks. Trends Plant Sci 27:255–273. https://doi.org/10.1016/j.tplants.2021.08.012
Ohme-Takagi M, Shinshi H (1995) Ethylene-inducible DNA binding proteins that interact with an ethylene-responsive element. Plant Cell 7:173–182. https://doi.org/10.1105/tpc.7.2.173
Okamuro JK, Caster B, Villarroel R, Van Montagu M, Jofuku KD (1997) The AP2 domain of APETALA2 defines a large new family of DNA binding proteins in Arabidopsis. Proc Natl Acad Sci U S A 94:7076–7081. https://doi.org/10.1073/pnas.94.13.7076
Paul P, Singh SK, Patra B, Liu X, Pattanaik S, Yuan L (2020) Mutually regulated AP2/ERF gene clusters modulate biosynthesis of specialized metabolites in plants. Plant Physiol 182:840–856. https://doi.org/10.1104/pp.19.00772
Paul P, Singh SK, Patra B, Sui X, Pattanaik S, Yuan L (2017) A differentially regulated AP2/ERF transcription factor gene cluster acts downstream of a MAP kinase cascade to modulate terpenoid indole alkaloid biosynthesis in Catharanthus roseus. New Phytol 213:1107–1123. https://doi.org/10.1111/nph.14252
Peng Y, Yang J, Li X, Zhang Y (2021) Salicylic acid: biosynthesis and signaling. Annu Rev Plant Biol 72:761–791. https://doi.org/10.1146/annurev-arplant-081320-092855
Pireyre M, Burow M (2015) Regulation of MYB and bHLH transcription factors: a glance at the protein level. Mol Plant 8:378–388. https://doi.org/10.1016/j.molp.2014.11.022
Pirrello J, Prasad BC, Zhang W, Chen K, Mila I, Zouine M, Latché A, Pech JC, Ohme-Takagi M, Regad F, Bouzayen M (2012) Functional analysis and binding affinity of tomato ethylene response factors provide insight on the molecular bases of plant differential responses to ethylene. BMC Plant Biol 12:190. https://doi.org/10.1186/1471-2229-12-190
Qi H, Liang K, Ke Y, Wang J, Yang P, Yu F, Qiu F (2023) Advances of Apetala2/Ethylene response factors in regulating development and stress response in Maize. Int J Mol Sci 24:5416. https://doi.org/10.3390/ijms24065416
Reboledo G, Agorio A, Vignale L, Alvarez A, Ponce De León I (2022) The moss-specific transcription factor PpERF24 positively modulates immunity against fungal pathogens in Physcomitrium patens. Front Plant Sci 13:908682. https://doi.org/10.3389/fpls.2022.908682
Riechmann JL, Meyerowitz EM (1998) The AP2/EREBP family of plant transcription factors. Biol Chem 379:633–646. https://doi.org/10.1515/bchm.1998.379.6.633
Sachan N, Falcone DL (2002) Wound-induced gene expression of putrescine N-methyltransferase in leaves of Nicotiana tabacum. Phytochemistry 61:797–805. https://doi.org/10.1016/s0031-9422(02)00427-2
Sakuma Y, Liu Q, Dubouzet JG, Abe H, Shinozaki K, Yamaguchi-Shinozaki K (2002) DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration- and cold-inducible gene expression. Biochem Biophys Res Commun 290:998–1009. https://doi.org/10.1006/bbrc.2001.6299
Sears MT, Zhang H, Rushton PJ, Wu M, Han S, Spano AJ, Timko MP (2014) NtERF32: a non-NIC2 locus AP2/ERF transcription factor required in jasmonate-inducible nicotine biosynthesis in tobacco. Plant Mol Biol 84:49–66. https://doi.org/10.1007/s11103-013-0116-2
Shoji T, Kajikawa M, Hashimoto T (2010) Clustered transcription factor genes regulate nicotine biosynthesis in tobacco. Plant Cell 22:3390–3409. https://doi.org/10.1105/tpc.110.078543
Shoji T, Mishima M, Hashimoto T (2013) Divergent DNA-binding specificities of a group of ethylene response factor transcription factors involved in plant defense. Plant Physiol 162:977–990. https://doi.org/10.1104/pp.113.217455
Sieber P, Schorderet M, Ryser U, Buchala A, Kolattukudy P, Métraux JP, Nawrath C (2000) Transgenic Arabidopsis plants expressing a fungal cutinase show alterations in the structure and properties of the cuticle and postgenital organ fusions. Plant Cell 12:721–738. https://doi.org/10.1105/tpc.12.5.721
Sohn KH, Lee SC, Jung HW, Hong JK, Hwang BK (2006) Expression and functional roles of the pepper pathogen-induced transcription factor RAV1 in bacterial disease resistance, and drought and salt stress tolerance. Plant Mol Biol 61:897–915. https://doi.org/10.1007/s11103-006-0057-0
Tezuka D, Kawamata A, Kato H, Saburi W, Mori H, Imai R (2019) The rice ethylene response factor OsERF83 positively regulates disease resistance to Magnaporthe oryzae. Plant Physiol Biochem 135:263–271. https://doi.org/10.1016/j.plaphy.2018.12.017
Thagun C, Imanishi S, Kudo T, Nakabayashi R, Ohyama K, Mori T, Kawamoto K, Nakamura Y, Katayama M, Nonaka S, Matsukura C, Yano K, Ezura H, Saito K, Hashimoto T, Shoji T (2016) Jasmonate-Responsive ERF transcription factors regulate steroidal glycoalkaloid biosynthesis in tomato. Plant Cell Physiol 57:961–975. https://doi.org/10.1093/pcp/pcw067
Tian Z, He Q, Wang H, Liu Y, Zhang Y, Shao F, Xie C (2015) The potato ERF transcription factor StERF3 negatively regulates resistance to Phytophthora infestans and salt tolerance in potato. Plant Cell Physiol 56:992–1005. https://doi.org/10.1093/pcp/pcv025
Udomsom N, Rai A, Suzuki H, Okuyama J, Imai R, Mori T, Nakabayashi R, Saito K, Yamazaki M (2016) Function of AP2/ERF transcription factors involved in the regulation of specialized metabolism in Ophiorrhiza pumila revealed by transcriptomics and metabolomics. Front Plant Sci 7:1861. https://doi.org/10.3389/fpls.2016.01861
Van der Does D, Leon-Reyes A, Koornneef A, Van Verk MC, Rodenburg N, Pauwels L, Goossens A, Körbes AP, Memelink J, Ritsema T, Van Wees SC, Pieterse CM (2013) Salicylic acid suppresses jasmonic acid signaling downstream of SCFCOI1-JAZ by targeting GCC promoter motifs via transcription factor ORA59. Plant Cell 25:744–761. https://doi.org/10.1105/tpc.112.108548
Van der Hoorn RA, Kamoun S (2008) From Guard to Decoy: a new model for perception of plant pathogen effectors. Plant Cell 20:2009–2017. https://doi.org/10.1105/tpc.108.060194
Vanholme R, De Meester B, Ralph J, Boerjan W (2019) Lignin biosynthesis and its integration into metabolism. Curr Opin Biotechnol 56:230–239. https://doi.org/10.1016/j.copbio.2019.02.018
Vogel MO, Moore M, König K, Pecher P, Alsharafa K, Lee J, Dietz KJ (2014) Fast retrograde signaling in response to high light involves metabolite export, Mitogen-activated protein kinase6, and AP2/ERF transcription factors in Arabidopsis. Plant Cell 26:1151–1165. https://doi.org/10.1105/tpc.113.121061
Wan L, Zhang J, Zhang H, Zhang Z, Quan R, Zhou S, Huang R (2011) Transcriptional activation of OsDERF1 in OsERF3 and OsAP2-39 negatively modulates ethylene synthesis and drought tolerance in rice. PLoS ONE 6:e25216. https://doi.org/10.1371/journal.pone.0025216
Wang JH, Gu KD, Han PL, Yu JQ, Wang CK, Zhang QY, You CX, Hu DG, Hao YJ (2020) Apple ethylene response factor MdERF11 confers resistance to fungal pathogen Botryosphaeria dothidea. Plant Sci 291:110351. https://doi.org/10.1016/j.plantsci.2019.110351
Wang L, Wang Y (2019) Transcription factor VqERF114 regulates stilbene synthesis in Chinese wild Vitis quinquangularis by interacting with VqMYB35. Plant Cell Rep 38:1347–1360. https://doi.org/10.1007/s00299-019-02456-4
Wang P, Du Y, Zhao X, Miao Y, Song CP (2013) The MPK6-ERF6-ROS-responsive cis-acting element7/GCC box complex modulates oxidative gene transcription and the oxidative response in Arabidopsis. Plant Physiol 161:1392–1408. https://doi.org/10.1104/pp.112.210724
Wessler SR (2005) Homing into the origin of the AP2 DNA binding domain. Trends Plant Sci 10:54–56. https://doi.org/10.1016/j.tplants.2004.12.007
Withers J, Dong X (2017) Post-translational regulation of plant immunity. Curr Opin Plant Biol 38:124–132. https://doi.org/10.1016/j.pbi.2017.05.004
Xie W, Ding C, Hu H, Dong G, Zhang G, Qian Q, Ren D (2022) Molecular events of rice AP2/ERF transcription factors. Int J Mol Sci 23:12013. https://doi.org/10.3390/ijms231912013
Xie Z, Nolan TM, Jiang H, Yin Y (2019) AP2/ERF transcription factor regulatory networks in hormone and abiotic stress responses in Arabidopsis. Front Plant Sci 10:228. https://doi.org/10.3389/fpls.2019.00228
Xing L, Di Z, Yang W, Liu J, Li M, Wang X, Cui C, Wang X, Wang X, Zhang R, Xiao J, Cao A (2017) Overexpression of ERF1-V from Haynaldia villosa can enhance the resistance of wheat to powdery mildew and increase the tolerance to salt and drought stresses. Front Plant Sci 8:1948. https://doi.org/10.3389/fpls.2017.01948
Yang C, Liu R, Pang J, Ren B, Zhou H, Wang G, Wang E, Liu J (2021a) Poaceae-specific cell wall-derived oligosaccharides activate plant immunity via OsCERK1 during Magnaporthe oryzae infection in rice. Nat Commun 12:2178. https://doi.org/10.1038/s41467-021-22456-x
Yang H, Sun Y, Wang H, Zhao T, Xu X, Jiang J, Li J (2021b) Genome-wide identification and functional analysis of the ERF2 gene family in response to disease resistance against Stemphylium lycopersici in tomato. BMC Plant Biol 21:72. https://doi.org/10.1186/s12870-021-02848-3
Yin J, Yi H, Chen X, Wang J (2019) Post-translational modifications of proteins have versatile roles in regulating plant immune responses. Int J Mol Sci 20:2807. https://doi.org/10.3390/ijms20112807
Yu ZX, Li JX, Yang CQ, Hu WL, Wang LJ, Chen XY (2012) The jasmonate-responsive AP2/ERF transcription factors AaERF1 and AaERF2 positively regulate artemisinin biosynthesis in Artemisia annua L. Mol Plant 5:353–365. https://doi.org/10.1093/mp/ssr087
Yuan M, Cai B, Xin XF (2023) Plant immune receptor pathways as a united front against pathogens. PLoS Pathog 19:e1011106. https://doi.org/10.1371/journal.ppat.1011106
Zander M, Thurow C, Gatz C (2014) TGA transcription factors activate the salicylic acid-suppressible branch of the ethylene-induced defense program by regulating ORA59 expression. Plant Physiol 165:1671–1683. https://doi.org/10.1104/pp.114.243360
Zang Z, Wang Z, Zhao F, Yang W, Ci J, Ren X, Jiang L, Yang W (2021) Maize ethylene response factor ZmERF061 is required for resistance to Exserohilum turcicum. Front Plant Sci 12:630413. https://doi.org/10.3389/fpls.2021.630413
Zarei A, Körbes AP, Younessi P, Montiel G, Champion A, Memelink J (2011) Two GCC boxes and AP2/ERF-domain transcription factor ORA59 in jasmonate/ethylene-mediated activation of the PDF1.2 promoter in Arabidopsis. Plant Mol Biol 75:321–331. https://doi.org/10.1007/s11103-010-9728-y
Zenkner FF, Margis-Pinheiro M, Cagliari A (2019) Nicotine biosynthesis in nicotiana: a metabolic overview. Tobacco Sci 56:1–9. https://doi.org/10.3381/18-063
Zhai Y, Fan Z, Cui Y, Gu X, Chen S, Ma H (2022) APETALA2/ethylene responsive factor in fruit ripening: Roles, interactions and expression regulation. Front Plant Sci 13:979348. https://doi.org/10.3389/fpls.2022.979348
Zhan X, Chen Z, Chen R, Shen C (2022) Environmental and genetic factors involved in plant protection-associated secondary metabolite biosynthesis pathways. Front Plant Sci 13:877304. https://doi.org/10.3389/fpls.2022.877304
Zhang C, Gao H, Sun Y, Jiang L, He S, Song B, Liu S, Zhao M, Wang L, Liu Y, Wu J, Xu P, Zhang S (2021a) The BTB/POZ domain protein GmBTB/POZ promotes the ubiquitination and degradation of the soybean AP2/ERF-like transcription factor GmAP2 to regulate the defense response to Phytophthora sojae. J Exp Bot 72:7891–7908. https://doi.org/10.1093/jxb/erab363
Zhang G, Chen M, Li L, Xu Z, Chen X, Guo J, Ma Y (2009) Overexpression of the soybean GmERF3 gene, an AP2/ERF type transcription factor for increased tolerances to salt, drought, and diseases in transgenic tobacco. J Exp Bot 60:3781–3796. https://doi.org/10.1093/jxb/erp214
Zhang H, Wang F, Song W, Yang Z, Li L, Ma Q, Tan X, Wei Z, Li Y, Li J, Yan F, Chen J, Sun Z (2023a) Different viral effectors suppress hormone-mediated antiviral immunity of rice coordinated by OsNPR1. Nat Commun 14:3011. https://doi.org/10.1038/s41467-023-38805-x
Zhang H, Pan X, Liu S, Lin W, Li Y, Zhang X (2021b) Genome-wide analysis of AP2/ERF transcription factors in pineapple reveals functional divergence during flowering induction mediated by ethylene and floral organ development. Genomics 113:474–489. https://doi.org/10.1016/j.ygeno.2020.10.040
Zhang M, Su J, Zhang Y, Xu J, Zhang S (2018) Conveying endogenous and exogenous signals: MAPK cascades in plant growth and defense. Curr Opin Plant Biol 45:1–10. https://doi.org/10.1016/j.pbi.2018.04.012
Zhang M, Zhang S (2022) Mitogen-activated protein kinase cascades in plant signaling. J Integr Plant Biol 64:301–341. https://doi.org/10.1111/jipb.13215
Zhang S, Miao W, Liu Y, Jiang J, Chen S, Chen F, Guan Z (2023b) Jasmonate signaling drives defense responses against Alternaria alternata in chrysanthemum. BMC Genomics 24:553. https://doi.org/10.1186/s12864-023-09671-0
Zhang X, Tubergen PJ, Agorsor IDK, Khadka P, Tembe C, Denbow C, Collakova E, Pilot G, Danna CH (2023) Elicitor-induced plant immunity relies on amino acids accumulation to delay the onset of bacterial virulence. Plant Physiol 192:601. https://doi.org/10.1093/plphys/kiad048
Zhang Y, Li X (2019) Salicylic acid: biosynthesis, perception, and contributions to plant immunity. Curr Opin Plant Biol 50:29–36. https://doi.org/10.1016/j.pbi.2019.02.004
Zhang Y, Zhang L, Ma H, Zhang Y, Zhang X, Ji M, van Nocker S, Ahmad B, Zhao Z, Wang X, Gao H (2021) Overexpression of the Apple (Malus x domestica) MdERF100 in Arabidopsis increases resistance to powdery mildew. Int J Mol Sci 22:5713. https://doi.org/10.3390/ijms22115713
Zhao Y, Chang X, Qi D, Dong L, Wang G, Fan S, Jiang L, Cheng Q, Chen X, Han D, Xu P, Zhang S (2017) A novel soybean ERF transcription factor, GmERF113, increases resistance to phytophthora sojae infection in soybean. Front Plant Sci 8:299. https://doi.org/10.3389/fpls.2017.00299
Zheng X, Xing J, Zhang K, Pang X, Zhao Y, Wang G, Zang J, Huang R, Dong J (2019) Ethylene response factor ERF11 activates BT4 transcription to regulate immunity to Pseudomonas syringae. Plant Physiol 180:1132–1151. https://doi.org/10.1104/pp.18.01209
Zhu Y, Zhang X, Zhang Q, Chai S, Yin W, Gao M, Li Z, Wang X (2022) The transcription factors VaERF16 and VaMYB306 interact to enhance resistance of grapevine to Botrytis cinerea infection. Mol Plant Pathol 23:1415–1432. https://doi.org/10.1111/mpp.13223
Zhuang J, Peng R-H, Cheng Z-M, Zhang J, Cai B, Zhang Z, Gao F, Zhu B, Fu X-Y, Jin X-F, Chen J-M, Qiao Y-S, Xiong A-S, Yao Q-H (2009) Genome-wide analysis of the putative AP2/ERF family genes in Vitis vinifera. Sci Hortic 123:73–81. https://doi.org/10.1016/j.scienta.2009.08.002
Acknowledgements
We apologize to all colleagues whose work could not be cited due to space limitations.
Funding
Research of related works were supported by Fruit Industrial Technology System of Shandong Province (SDAIT-06–03).
Author information
Authors and Affiliations
Contributions
NM, PS, ZYL, FJZ, and CLZ collected the references and drafted the manuscript. NM made all the figures. NM and ZZ revised the manuscript. XFW and CXY supervised the projects related to this work.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no conflict of interests.
Additional information
Handling Editor: Dr. Vincent Bus.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ma, N., Sun, P., Li, ZY. et al. Plant disease resistance outputs regulated by AP2/ERF transcription factor family. Stress Biology 4, 2 (2024). https://doi.org/10.1007/s44154-023-00140-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44154-023-00140-y