Abstract
Morphogenesis of the silica based cell walls of diatoms, a large group of microalgae, is a paradigm for the self-assembly of complex 3D nano- and microscale patterned inorganic materials. In recent years, loss-of-function studies using genetic manipulation were successfully applied for the identification of genes that guide silica morphogenesis in diatoms. These studies revealed that the loss of one gene can affect multiple morphological parameters, and the morphological changes can be rather subtle being blurred by natural variations in morphology even within the same clone. Both factors severely hamper the identification of morphological mutants using subjective by-eye inspection of electron micrographs. Here we have developed automated image analysis for objectively quantifying the morphology of ridge networks and pore densities from numerous electron micrographs of diatom biosilica. This study demonstrated differences in ridge network morphology and pore density in diatoms growing on ammonium rather than nitrate as the sole nitrogen source. Furthermore, it revealed shortcomings in previous by-eye evaluation of the biosilica phenotype of the silicanin-1 knockout mutant. We anticipate that the computational methods established in the present work will be invaluable for unraveling genotype–phenotype correlations in diatom biosilica morphogenesis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Diatoms represent a large group of unicellular, eukaryotic microalgae that biosynthesize silica based cell walls. The 3D nano- to microscale architecture of the biosilica is a species specific trait indicating that its morphology is genetically controlled [1]. The intricate and highly regular, hierarchical porous patterns of diatom biosilica are regarded as paradigms for the bottom-up synthesis of mineral based materials under environmentally benign conditions [2,3,4]. Furthermore, diatom biosilica microparticles have interesting materials properties that can be exploited for a wide range of applications in photonics, chemical sensing, catalysis and drug delivery [5,6,7,8]. The diatom cell wall consists of two halves of identical structures that are arranged in a petri-dish like arrangement. The slightly larger half (epitheca) overlaps the other half (hypotheca). Each theca is composed of a plate- or dome-shaped valve, exhibiting a species specific silica pattern, and several ring-shaped girdle bands. The biosilica is perforated with numerous, usually regularly arranged pores enabling a light-weight yet mechanically robust architecture and the exchange of ions and molecules with the environment [9, 10]. The formation of valves and girdle bands takes place inside diatom cells in specialized compartments called silica deposition vesicles (SDVs) [11]. Apart from amorphous SiO2 as the main component, biosilica contains proteins, long-chain polyamines (LCPA), and polysaccharides [12,13,14].
In recent years, genetic manipulation studies have identified genes involved in biosilica morphogenesis in the radial centric model diatom Thalassiosira pseudonana. Individual knockdowns of each of the genes sap1, sap2 and sap3 resulted in various aberrations in biosilica morphology [15]. Knockdown of the gene Thaps3_21880 caused a reduction of the pore density while increasing the mean area of the same [16]. Knockout of the silicanin-1 gene (sin1) had affected valve morphology and caused substantially reduced strength and stiffness of the cell wall [17]. Besides such targeted genetic modifications, the effect of many external parameters on biosilica morphology was investigated. These include, drug-based inhibition of cytoskeleton function [18,19,20,21,22] and of a V-type H+-ATPase [23], as well as controlled changes in (i) salinity and pH of the growth medium [24, 25], (ii) growth temperature [26], and (iii) light wavelength as well as intensity [27, 28] (iv) the insertion of metals such as titanium into the biosilica [29, 30]. In most of these studies a multitude of structural changes was observed upon alteration of only one gene or one environmental parameter. Furthermore, some of the morphological changes were rather subtle and somewhat heterogeneous among individual cells even from the same clone. As a consequence, biosilica morphology changes were prone to biased interpretation or might even have been overlooked. To eliminate such problems, unbiased, automated, and quantitative computational analysis tools are required that enable thorough and impartial evaluation of the effects on biosilica structure by gene disruptions and external perturbations.
Previous work applied supervised learning for the analysis of scanning electron microscopy (SEM) images of the biosilica ridge pattern of the valves of T. pseudonana [26]. This led to the identification of a phenotype switch from a mesh-like to a tree-like ridge pattern in cells that were depleted of silicic acid, the precursor for silica [26]. Artificial neural networks (NNs) were used to identify in SEM images slight structural differences between the valves from T. pseudonana wild type and the Thaps3_21880 gene knockdown mutant [16]. However, although NNs can be used to determine if any morphological differences exist between multiple groups of images, they do not reveal what the specific morphological differences are. As a consequence, NNs offer little aid in reaching a mechanistic understanding of biosilica morphogenesis. Therefore, it is desirable to develop computational methods that not only enable rapid and unbiased in-depth analysis of diatom biosilica phenotypes, but also point to an understanding of the mechanisms that create them.
A key goal of diatom based materials synthesis is the development of cell lines that produce biosilica with structures and properties tailored for specific applications. Recently, a genetic engineering based method called live diatom silica incorporation (LiDSI) was established that allows for generating diatom strains, which contain desired fluorescent proteins, receptor proteins, or enzymes in the biosilica [31]. While biosilica functionalization using LiDSI is well developed and even enables regioselective incorporation of proteins [32], the genetic engineering of biosilica structure is still in its infancy. Only quite recently, the first genetically induced morphological mutants were generated in diatoms through knockdown and knockout of specific genes [15, 17]. One of the mutants (Sin1KO), which was generated by knocking out the sin1 gene in T. pseudonana, exhibited a pleiotropic phenotype. Tedious manual inspection of transmission electron microscopy (TEM) and atomic force microscopy images revealed differences in the height and number of cross-connections between the ridges in the valves [17]. The analysis of other structural parameters was not pursued.
Here we describe the development of two complementary computational approaches for impartial and quantitative in-depth image analysis of valve biosilica morphology. These tools were then applied to quantify and compare the influence of growth conditions and sin1 gene knockout on biosilica morphology of T. pseudonana.
2 Results
As test cases for developing computational tools for the analysis biosilica phenotypes we chose wild type cells and three previously described sin1 knockout mutants of T. pseudonana (KO1, KO2, KO3), which exhibited subtle changes in biosilica morphology [17]. The four strains were cultivated under two growth conditions, in which nitrate (nit) or ammonium (am) served as the sole nitrogen source. The reason for varying the nitrogen source is the fact that the nitrate reductase promoter/terminator cassette, Pnr/Tnr, is the only well-established inducible promoter in diatoms [33]. When an introduced gene is placed under control of Pnr/Tnr, it is repressed when the cells are grown in medium containing ammonium as the sole nitrogen source [34]. Expression of the transgene is induced when the cells are transferred into nitrate containing medium that lacks ammonium [34]. The type of nitrogen source exerts large scale effects on metabolism [35], yet it is unknown whether this has an effect on silica morphogenesis. We envision that use of the inducible Pnr/Tnr cassette may become useful for evaluating the effect of candidate genes suspected to be involved in silica morphogenesis. Therefore, it needs to be clarified whether or not biosilica morphology is influenced by the metabolic changes that are elicited solely through changing the nitrogen source in the growth medium.
2.1 Imaging of valves
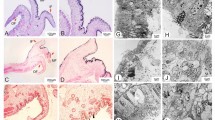
Morphological analyses in the present work were entirely focused on the valve part of the biosilica, which is the structurally most elaborate part of a diatom cell wall. To prepare valves of wild type (WT) cells and sin1 knockout strains for transmission electron microscopy (TEM), cell walls were isolated, valves were severed from the cell walls by mild ultrasound treatment, and the samples air dried on a TEM grid. The valves remained largely intact during the preparation procedure but frequently contained small cracks. T. pseudonana valves exhibit five main structural features: fultoportulae, costae, cross-connections, areola pores, and cribrum pores. A fultoportula is a tubular structure surrounded by three or four satellite pores. Fultoportulae are regularly spaced near the valve rim (Fig. 1a, yellow circles), and in about ~ 50% of the valves one or two fultoportulae are present slightly off the valve center (Fig. 1a, yellow circle). The valve surface contains numerous silica ridges that radiate from the valve center and branch multiple times forming a highly interconnected meshwork (Fig. 1). A radially oriented ridge is called a costa (Fig. 1a, green) and an azimuthally oriented one is called a cross-connection (Fig. 1a, orange). A region bounded by adjacent cross-connections and costa segments is called an areola pore, which typically exposes a trapezoid shape (Fig. 1a, cyan). Inside each areola pore are several circular openings termed cribrum pores (Fig. 1a). Visual inspection of transmission electron microscopy (TEM) images revealed that the sin1 knockout mutants exhibited a reduced number of cross-connections compared to the valve of wild type cells, as expected (Fig. 1 and Additional file 1: Fig. S1). The reduction in cross-connections was apparent for valves from both nitrate and ammonium grown cells (Fig. 1c, b and Additional file 1: Fig. S1). When comparing WT valves from cells grown on different nitrogen sources, the cross-connections seem to be less pronounced in the valves from ammonium growing cells (Fig. 1a, b). In sin1 knockout strains, the cribrum pore densities seemed to be lower in valves from ammonium grown cells compared to valves from nitrate grown cells (Fig. 1c, d and Additional file 1: Fig. S1). However, the differences in the expression of cross-connections and in cribrum pore densities are not equally obvious from each individual valve. As these features are difficult to assess manually, we aimed to develop computational methods for their automated quantitative analysis. The number of valves from each strain included in the analysis is shown in Additional file 1: Table S1.
TEM images of valve biosilica from T. pseudonana wild type (WT) and a representative sin1 knockout mutant (KO1) grown in media containing different nitrogen sources. Representative images of individual valves are shown for the indicated strains (see top right corner). The indices nit (for nitrate) or am (for ammonium) refer to the sole nitrogen source in the growth medium. a WT grown in nitrate. Characteristic structural features of the biosilica are highlighted: green = costa, orange = cross-connection, cyan = areola pore, yellow = fultoportula. b WT grown in ammonium, c KO1 grown in nitrate, d KO1 grown in ammonium. Scale bars: 1 µm
2.2 Automated analysis of the ridge morphology of valves
To quantify valve morphology, we focused on the pattern of silica ridges (costae and cross-connections), which are thickenings of the cell wall that appear as a network of dark lines in TEM images (Fig. 1). We aimed to extract the ridge network as a pixel-thin skeleton image representation. However, extraction of the skeleton is challenging as the contrast of the ridge network varies widely between images, indicating differences in the thickness of the ridges in the valves from different strains (Fig. 1). Indeed, it was previously demonstrated that Sin1KO strains had reduced silica content and less elevated costae [17]. To deal with the challenging contrast issue, we developed two different approaches to extract ridge networks. In the first approach, we implemented an adaptive thresholding method, which uses local contrast differences to extract a binary image from the TEM data. This method mainly detects ridges with high local contrast, which is likely indicative of high silicification. In the following, we refer to the skeletons generated with this method as “type 1 skeletons” (Fig. 2b). Type 1 skeletons extracted from low-contrast images are typically fragmented. However, it is known that during valve formation, the ridges develop as a continuous branched network, with cross-connections between branches appearing at later stages of development [36, 37]. To recover this continuous branching structure, we developed a second approach, designed to extract a connected (i.e. non-fragmented) skeleton even from low-contrast images (Fig. 2c). This approach relies on a so-called “ridge detection image filter”, which can identify dark lines of a chosen thickness by estimating the “main principal curvature” at each point in the image. If the pixel values (gray levels) represented elevation in the landscape, the ridge detection filter would measure the largest vertical curvature at each point, and could thus identify geographical ridges of the chosen broadness, regardless of gradual changes in elevation along the ridgelines. In contrast, thresholding would identify places of high elevation. We refer to the skeletons it generates as “type 2 skeletons”. The two skeleton extraction approaches produce similar results from high-contrast images, but differ when used for low-contrast images (Fig. 3).
Analysis pipelines for the characterization of the ridge pattern of a valve. a The region of interest is identified as a circle that is interior to the circular pattern of the fultoportulae at the rim. The ridge pattern is extracted as b type 1 skeleton (maintaining high-contrast features) and c type 2 skeleton (maintaining connectedness). d Cross-connections are identified based on their orientation and length in both skeletons
Once the ridge network is extracted, several of its features can be measured. In earlier work, valve morphology was quantified by manually counting the number of cross-connections per valve [17]. Therefore, the first feature we analyzed in the extracted skeletons was the number of “geometric cross-connections”, defined as valve ridges oriented in an azimuthal direction (Fig. 2d). We developed an algorithm to classify parts of the network as belonging to costae or cross-connections. Each segment of the ridge network was classified based on a geometric criterion: the angle it forms with a line connecting it to the valve’s centerpoint. Segments oriented at an angle smaller than a threshold (here: 45°) were considered to be parts of costae, while those forming an angle greater than the threshold were classified as cross-connections. Additionally, azimuthal segments shorter than 70 nm were discarded. This length was chosen by comparing the azimuthal segments for a range of lengths and choosing a value around which the cross-connection count varied the least. Geometric cross-connections per unit area (i.e. cross-connections densities) were determined in both, type 1 skeletons (representing highly silicified features) and type 2 skeletons (representing the fully connected ridge network) (Fig. 4). Using densities instead of total counts is justified because the cross-connection density of individual valves does not correlate with their area (Additional file 1: Fig. S3). The cross-connection densities in type 1 skeletons of the valves from wild type cells (WT) grown on the two different nitrogen sources were very similar (Fig. 4a). However, it was higher in the valves from nitrate grown sin1 knockout strains (KO1-3) compared to the valves from the same strains grown in ammonium (Fig. 4a). These data suggested that in the absence of the sin1 gene, the cross-connection density is highly dependent on the nitrogen source. To further investigate this unexpected result, the geometric cross-connection density was analyzed in the type 2 skeletons. Here, the densities of cross-connections between the valves from cells grown in nitrate and ammonium were very similar for all strains irrespective of the nitrogen source (Fig. 4b).
The discrepancy between the cross-connection densities in the two types of skeletons was caused by the fact that the two methods for skeletonization have different capabilities for identifying cross-connections. The method for extracting type 2 skeletons is geared towards identifying costae and cross-connections also in low contrast regions, where the extraction method for type 1 skeletons yielded fragmented networks and thus a lower number of cross-connections. The ridge networks in valves from ammonium grown cells exhibited consistently lower contrast of the ridge network in TEM images compared to valves from nitrate gown cells (Additional file 1: Fig. S2). Therefore, the discrepancy between the cross-connection counts by the two skeletonization methods is more drastic in valves from ammonium grown cells (Fig. 4).
2.3 Automated analysis of the density of cribrum pores
A characteristic feature of T. pseudonana are the ~ 20 nm sized cribrum pores, which are seemingly randomly distributed in the areas between the costae. Each areola usually contains two or more cribrum pores (Fig. 5a). The center of each pore was located in the TEM images by first accentuating pores using the “difference of Gaussians” method followed by identification for tall local maxima. For all strains, the cribrum pore density was consistently higher in valves from nitrate grown cells than in valve from cells grown in ammonia (Fig. 5b). Furthermore, in nitrate medium the sin1 knockout mutant strains exhibited higher cribrum pore densities than the wild type (Fig. 5b). The same trend applied for the ammonium grown strains, except for the KO3 strain, which had the same cribrum pore density as the wild type (Fig. 5b).
3 Discussion
In the present work we have developed automated image analysis for evaluating the ridge patterns and pore densities from large numbers of TEM images of diatom biosilica. The new methods for morphological analysis replaced previously used manual methods [17], enabling significantly faster evaluation of much larger data sets. To analyze valve morphologies, we established two different approaches for extracting the characteristic networks of ridges (costae and cross-connections), which we termed type 1 and type 2 skeletons. Both approaches are suitable for determining the number of cross-connections from images in which the gray-level contrast between the ridge network and the surrounding areas is high. However, when the contrast is low only the method extracting type 2 skeletons is able to identify the connected ridge network, because it is based on a “ridge detection image filter” which identifies continuous lines comparable to geographic ridgelines.
It is reasonable to assume that gray-level contrast in TEM images of biosilica reflects differences in silica content. Diatom biosilica is composed of ~ 90% hydrated silica and ~ 10% organic material, and thus electron contrast in TEM images of biosilica is dominated by silicon, which is the element with by far the highest electron density in this material. Consistent with this assumption is the observation that the TEM contrast between the ridges and the surrounding areas is lower in valves of sin1 knockout mutants (see Fig. 1a, c), which have a ~ 40% reduced z-height of costae compared to wild type [17]. For reasons described above, skeletonization method 1 is more susceptible than method 2 to differences in TEM contrast. Therefore, strongly reduced silicification of a cross-connection ridge is often interpreted as a missing cross-connection in type 1 skeletons (see Fig. 3). This is consistent with the interpretation gained from by-eye inspection of TEM micrographs (see Fig. 1 and Ref. [17]). In contrast, skeletonization method 2 is capable of detecting weakly silicified cross-connections (i.e. low TEM contrast), and thus revealed that the ridge network of T. pseudonana wild type is still fully intact in the sin1 knockout mutants (see Fig. 4b). The previous interpretation of missing cross-connection in the sin1 knockout mutants [17], was thus a misinterpretation of a phenotype that exhibits weakly silicified cross-connections. This result demonstrated that by applying both skeletonization methods, a more realistic description of the valve phenotypes could be obtained.
Another unexpected result of this study was the observation that the type of nitrogen source had an effect on the density of cribrum pores and the cross-connection density of the ridge network in type 1 skeletons. The effect on cross-connection density was very mild in wild type cells but much more pronounced in the sin1 knockout mutants, with valves from cells grown on ammonium exhibiting lower cross-connection densities than valves from nitrate grown cells (see Fig. 4a). Analysis of the type 2 skeletons demonstrated for each strain that the cross-connection densities were independent of the nitrogen source (see Fig. 4a), thus indicating that reduced silicification is responsible for the diminished cross-connection densities in the type 1 skeletons from ammonium grown cells. This finding raises a new question: How can the nitrogen source influence silica morphogenesis in diatoms? We believe that part of the answer lies in the observation that the nitrogen source effect on silicification is stronger in the sin1 knockout mutants than in wild type. There is circumstantial evidence that the Sin1 protein enhances the silica forming activity of long-chain polyamines (LCPA) [38], which are general components of the biosilica forming machinery in diatoms [12]. We hypothesize that LCPA biosynthesis might be downregulated in diatoms grown on ammonium medium compared to nitrate grown diatoms, which would be expected to lead to reduced silica formation. In sin1 knockout mutants, the silica formation capability may be further reduced due to the lack of the Sin1 protein. Despite these shortcomings, there is still sufficient silica forming activity present to biosynthesize a complete valve including the ride network. However, the thickening of the ridges (called z-expansion), which is the final stage of silica biosynthesis [36] may be hampered by reduced levels of LCPA and the lack of Sin1. Reduced silicification of the ridge network compromises the mechanical performance of the valves [17], which is problematic as the cell wall needs to counteract the turgor pressure exerted by the protoplast. One countermeasure of the cell for increasing the stability of valves with reduced silica content could be to change the pattern of cribrum pores. Whether a reduction of cribrum pore density, as we observed here (see Fig. 5), generally leads to an increase in mechanical performance of the valve, remains to be determined.
The results from the present study clearly demonstrate that automated computational analysis is highly useful for unraveling the influences of external and internal factors on diatom biosilica morphology. We anticipate that the computational geometric methods established here will be instrumental in future work for characterizing the function of biomineralization proteins in silica morphogenesis in T. pseudonana and possibly many other diatoms species.
4 Methods
4.1 Culture conditions
T. pseudonana (clone CCMP 1335) was grown at 18 °C with a 16 h light/8 h dark cycle at 5000–10,000 lx, in artificial seawater (ASW) [37] lacking the nitrogen source. Sterile filtered NH4Cl or KNO3 was added to a final concentration of 550 µM.
4.2 Preparation of biosilica for electron microscopy
For biosilica isolation 100 mL of a cell culture (∼ 106 cells mL−1) were harvested by centrifugation (3220×g, 10 min). The pellet was resuspended in 15 mL extraction buffer (2% SDS, 100 mM EDTA pH 8, and 1 mM PMSF), and continuously shaken at 55 °C for 30 min to solubilize intracellular material. The biosilica was pelleted (3220×g, 2 min) and washed twice in 15 mL 10 mM EDTA pH 8, and 1 mM PMSF. Extraction and washing were repeated until the biosilica was colorless. The colorless biosilica was washed with 1 mL acetone, two times with 15 mL H2O, and finally resuspended in 1 mL H2O.
To detach girdle bands from valves, 500 µL of isolated biosilica was transferred in a 1.5-mL tube and sonicated with an MS72 sonotrode tip (Bandelin) applying a total of 0.12 kJ over 5 s. For transmission electron microscopy (TEM) 10 µL of sonicated biosilica was transferred onto Formvar-coated gold-coated finder grids (G200F2-Au from EMS) that were strengthened with evaporated carbon. The majority of the liquid was removed by blotting with a piece of filter paper, washed two times with 50 µL H2O and samples were air-dried over-night before imaging. TEM images were taken using a Jeol JEM-1400 TEM instrument at an acceleration voltage of 80 kV.
4.3 Segmenting the high-contrast valve ridges (type 1 skeleton)
Each grayscale TEM image was converted into a binary image by applying adaptive thresholding to differentiate the network structure from the background. From the network structure, a single-pixel thick skeleton was extracted using the Ridge Detection Plugin of ImageJ [39]. Examples of skeletons are shown in Figs. 2b and 3.
4.4 Segmenting the connected valve ridge network (type 2 skeleton)
First, we corrected any uneven brightness in the electron micrograph: The local brightness was estimated by removing (inpainting) the pores, removing the valve ridges using a closing filter, and finally applying Gaussian blur. The brightness field obtained this way was subtracted from the image. Then the location of valve ridges (costae and cross-connections) was identified with a ridge detection image filter that computes the main principal curvature at each point using Gaussian derivates [40]. The scale parameter of the ridge detection filter was set at \(\sigma = 29\;{\text{nm}}\), in accordance with the typical costa width, and converted to pixels according to each image’s resolution. Finally, the result was thresholded (choosing the threshold using Otsu’s cluster variance maximization method [41]) and skeletonized to create a binary one-pixel thin representation of the ridge network (see Fig. 2c). The extraction of the ridge network requires care, so that we may obtain a network with a large connected component while making sure that the parts separating neighboring pores can be reliably distinguished from true ridges, even in low-contrast images. We manually fine-tuned the ridge detector’s scale parameter to obtain good results across all image datasets. Examples of final skeletons are shown in Figs. 2c and 3. These examples have one level of open branches pruned away to aid comparison with the original image. We note that such pruning has no effect on the topological ridge network analysis.
4.5 Classifying cross-connections using geometric criteria
We designed an algorithm to identify geometric cross-connections from the electron micrograph of a diatom valve based on their orientation (Additional file 1: Fig. S4). Since T. pseudonana is a centric diatom with radial symmetry for the valves, we manually marked the centerpoint of each valve. Cross-connections would appear nearly perpendicular to radial lines (i.e. azimuthally oriented), while costae are approximately parallel to radial lines. In a first step, each pixel of the skeleton network was classified based on the angle the line segment it is part of forms with the radial line of the pixel: A pixel is considered to be part of a cross-connection if it has fewer than five neighboring pixels (within a seven-pixel radius) forming an angle less than a threshold (which we choose as 45°); and as part of a costa otherwise. Connected components of pixels filtered in this fashion were considered candidates for cross-connections. In a second step, such candidate segments shorter than a minimum length of 70 nm were discarded. The resulting pattern (see Fig. 2d) consists of only valid cross-connections, which we count by connected component labeling [42].
4.6 Pore detection
The pores were located using the difference of Gaussians method. This method will accentuate features at a certain chosen length scale in the image. The center of pores was detected by choosing the pore size as the length scale and looking for maxima of sufficient height in the image. In order to find an appropriate length scale for each image, an initial pore detection pass was run with an approximate, assumed pore size. The mean size of the initially detected pores was then measured, and used in subsequent passes to refine the result. This method usually converges in at most 2-3 iterations.
Data availability
The data are available from the corresponding authors upon request.
Code availability
The code used for image analysis is available from the corresponding authors upon request.
References
Round FE, Crawford RM, Mann DG. The diatoms. Cambridge: Cambridge University Press; 1990.
Jeffryes C, Agathos SN, Rorrer G. Biogenic nanomaterials from photosynthetic microorganisms. Curr Opin Biotechnol. 2015;33:23–31.
Kröger N, Sandhage KH. From diatom biomolecules to bioinspired syntheses of silica- and titania-based materials. MRS Bull. 2010;35:122–6.
Coradin T, Lopez PJ. Biogenic silica patterning: simple chemistry or subtle biology? ChemBioChem. 2003;4:251–9.
Rea I, De Stefano L. Recent advances on diatom-based biosensors. Sensors. 2019;19:5208.
Ragni R, Cicco SR, Vona D, Farinola GM. Multiple routes to smart nanostructured materials from diatom microalgae: a chemical perspective. Adv Mater. 2018;30:e1704289.
De Tommasi E. Light manipulation by single cells: the case of diatoms. J Spectrosc. 2016;2016:2490128.
Kröger N, Brunner E. Complex-shaped microbial biominerals for nanotechnology. WIREs Nanomed Nanobiotechnol. 2014;6:615–27.
Hamm CE, Merkel R, Springer O, Jurkojc P, Maier C, Prechtel K, Smetacek V. Architecture and material properties of diatom shells provide effective mechanical protection. Nature. 2003;421:841–3.
Aitken ZH, Luo S, Reynolds SN, Thaulow C, Greer JR. Microstructure provides insights into evolutionary design and resilience of Coscinodiscus sp frustule. Proc. Natl Acad. Sci. USA. 2016;113:2017–22.
Drum RW, Pankratz HS. Post mitotic fine structure of Gomphonema parvulum. J Ultrastruct Res. 1964;10:217–23.
Kröger N, Poulsen N. Diatoms-from cell wall biogenesis to nanotechnology. Annu Rev Genet. 2008;42:83–107.
Sumper M, Brunner E. Silica biomineralisation in diatoms: the model organism Thalassiosira pseudonana. ChemBioChem. 2008;9(8):1187–94.
Hildebrand M, Lerch SJL, Shrestha RP. Understanding diatom cell wall silicification - moving forward. Front Mar Sci. 2018;5:1–19.
Tesson B, Lerch SJL, Hildebrand M. Characterization of a new protein family associated with the silica deposition vesicle membrane enables genetic manipulation of diatom silica. Sci Rep. 2017;7:1–13.
Trofimov AA, Pawlicki AA, Borodinov N, Mandal S, Mathews TJ, Hildebrand M, et al. Deep data analytics for genetic engineering of diatoms linking genotype to phenotype via machine learning. npj Comput Mater. 2019;5:67.
Görlich S, Pawolski D, Zlotnikov I, Kröger N. Control of biosilica morphology and mechanical performance by the conserved diatom gene Silicanin-1. Commun Biol. 2019;2:245.
Schmid AMM. Valve morphogenesis in diatoms: a pattern-related filamentous system in pennates and the effect of APM, colchicine and osmotic pressure. Nova Hedw. 1980;33:811–47.
Kharitonenko KV, Bedoshvili YD, Likhoshway YV. Changes in the micro-and nanostructure of siliceous valves in the diatom Synedra acus under the effect of colchicine treatment at different stages of the cell cycle. J Struct Biol. 2015;190:73–80.
Blank GS, Sullivan CW. Diatom mineralization of silicic acid. VII. Influence of microtubule drugs on symmetry and pattern formation in valves of Navicula saprophila during morphogenesis. J Phycol. 1983;19:294–301.
Van de Meene AM, Pickett-Heaps JD. Valve morphogenesis in the centric diatom Rhizosolenia setigera (Bacillariophyceae, Centrales) and its taxonomic implications. Eur J of Phycol. 2004;39(1):93–104.
Tesson B, Hildebrand M. Dynamics of silica cell wall morphogenesis in the diatom Cyclotella cryptica: substructure formation and the role of microfilaments. J Struct Biol. 2010;169:62–74.
Yee DP, Hildebrand M, Tresguerres M. Dynamic subcellular translocation of V-type H+-ATPase is essential for biomineralization of the diatom silica cell wall. New Phytologist. 2020;225:2411–22.
Vrieling EG, Sun Q, Tian M, Kooyman PJ, Gieskes WWC, van Santen RA, Sommerdijk NAJM. Salinity-dependent diatom biosilicification implies an important role of external ionic strength. Proc Natl Acad Sci USA. 2007;104:10441–6.
Hervé V, Derr J, Douady S, Quinet M, Moisan L, Lopez PJ. Multiparametric analyses reveal the pH-dependence of silicon biomineralization in diatoms. PLoS ONE. 2012;7:e46722.
Javaheri N, Dries R, Burson A, Stal LJ, Sloot PMA, Kaandorp JA. Temperature affects the silicate morphology in a diatom. Sci Rep. 2015;5:11652.
Su Y, Lundholm N, Friis SM, Ellegaard M. Implications for photonic applications of diatom growth and frustule nanostructure changes in response to different light wavelengths. Nano Res. 2015;8:2363–72.
Su Y, Lundholm N, Ellegaard M. Effects of abiotic factors on the nanostructure of diatom frustules - ranges and variability. App Microbiol Biotechnol. 2018;102:5889–99.
Qin T, Gutu T, Jiao J, Chang C, Rorrer GL. Biological fabrication of photoluminescent nanocomb structures by metabolic incorporation of germanium into the biosilica of the diatom Nitzschia frustulum. ACS Nano. 2008;2:1296–304.
Jeffryes C, Gutu T, Jiao J, Rorrer GL. Metabolic insertion of nanostructured TiO2 into the patterned biosilica of the diatom Pinnularia sp by a two-stage bioreactor cultivation process. ACS Nano. 2008;2:2103–12.
Kröger N, Dubey NC, Kumari E. Immobilization of proteins on diatom biosilica. In: Losic D, editor. Diatom nanotechnology. London: Royal Society of Chemistry; 2017. p. 126–49.
Kumari E, Görlich S, Poulsen N, Kröger N. Genetically programmed regioselective immobilization of enzymes in biosilica microparticles. Adv Funct Mater. 2020;30:2000442.
Kroth PG, Bones AM, Daboussi F, Ferrante MI, Jaubert M, Kolot M, Nymark M, Río Bártulos C, Ritter A, Russo MT, Serif M, Winge P, Falciatore A. Genome editing in diatoms: achievements and goals. Plant Cell Rep. 2018;37:1401–8.
Poulsen N, Chesley PM, Kröger N. Molecular genetic manipulation of the diatom Thalassiosira pseudonana (Bacillariophyceae). J Phycol. 2006;42:1059–65.
Smith SR, Dupont CL, McCarthy JK, Broddrick JT, Oborník M, Horák A, et al. Evolution and regulation of nitrogen flux through compartmentalized metabolic networks in a marine diatom. Nat Commun. 2019;10:4552.
Hildebrand M, York E, Kelz JI, Davis AK, Frigeri LG, Allison DP. Nanoscale control of silica morphology and three-dimensional structure during diatom cell wall formation. J Mater Res. 2006;21:2689–98.
Heintze C, Formanek P, Pohl D, Hauptstein J, Rellinghaus B, Kröger N. An intimate view into the silica deposition vesicles of diatoms. BMC Mat. 2020;2:11.
Kotzsch A, Gröger P, Pawolski D, Bomans PHH, Sommerdijk NAJM, Schlierf M, Kröger N. Silicanin-1 is a conserved diatom membrane protein involved in silica biomineralization. BMC Biol. 2017;15:9–11.
Steger C. An unbiased detector of curvilinear structures. IEEE Trans Pattern Anal Mach Intell. 1998;20(113–125):389.
Lindeberg T. Scale-space theory: a basic tool for analyzing structures at different scales. J Appl Stat. 1994;21:225–70.
Otsu N. A threshold selection method from gray-level histograms. IEEE Trans. Sys. Man. Cyber. 1979;9:62–6.
Marchand-Maillet S, Yazid MS. Binary digital image processing: a discrete approach. London: Academic Press; 2000.
Acknowledgements
This work was funded by the Deutsche Forschungsgemeinschaft (DFG) through Research Unit 2038 “NANOMEE” (DFG Grants KR 1853/8-2 and SCHL 1896/2-2). This work was supported by the Electron Microscopy Facility, a core facility of the CMCB at Technische Universität Dresden.
Funding
Open Access funding enabled and organized by Projekt DEAL. This work was funded by the Deutsche Forschungsgemeinschaft (DFG) through Research Unit 2038 “NANOMEE” (DFG Grant KR 1853/8-2 and SCHL 1896/2-2).
Author information
Authors and Affiliations
Contributions
NK, CDM and SzH conceived the study. All authors planned and interpreted experiments and computational analyses. SG conducted experiments, SzH and AF developed analysis algorithms and performed computational analyses. All authors wrote the paper. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors have no conflict of interest and no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Horvát, S., Fathima, A., Görlich, S. et al. Computational analysis of the effects of nitrogen source and sin1 knockout on biosilica morphology in the model diatom Thalassiosira pseudonana. Discov Mater 1, 8 (2021). https://doi.org/10.1007/s43939-021-00008-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s43939-021-00008-w