Abstract
Medicinal plants have a long tradition of being cultivated and harvested in India. The Indian Forest is the principal repository for many useful medicinal herbs. As a result of their critical role in maintaining people's life, medicinal plants have traditionally been the subject of intensive research and consideration. Yet, correctly identifying plants used in medicine is a laborious process that takes a lot of time and expertise. Because of this, a vision-based approach may aid scientists and regular people in the rapid and precise identification of herb plants. Therefore, this research suggests a vision-based smart method to recognize herb plants by creating a deep learning (DL) model. Although there is a wide variety of useful plants, we limit ourselves to just six from the Kaggle database: betel, curry, tulsi, mint, neem, and Indian beech. For each medicinal plant, we collected 500 images. The data undergo a process of resizing and augmentation to increase the sample size. For the fully automatic identification of medicinal leaves, the MobileNet DL model is selected. To determine the model's effectiveness, it must first be trained, then validated, and ultimately tested. The DL model is evaluated using measures including accuracy, precision, and recall. For this reason, the DL model was able to correctly identify medicinal leaves at an accuracy rate of 98.3%. After being thoroughly investigated, the DL model is uploaded to the cloud, and a mobile app is created for the real-time identification of medicinal leaves. To recognize leaf images, the built mobile app accesses the DL model on the cloud. The automated recognition of plants represents an extremely promising option for filling the taxonomic gap and gaining a lot of interest from the fields of botany and machine vision.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Every life on Earth depends on the oxygen that plants produce. By providing oxygen and water, plants of diverse sizes and forms play a crucial role in maintaining the diversity of life on Earth [1]. Herbs, or medicinal plants, are used to heal many disorders and illnesses in humans [2]. These plants have several medicinal benefits, from the roots to the leaves. Plants have been used by humans for a variety of purposes including medicine, food preparation, and the cosmetics industry. Herbalists must utilize taxonomy to make sense of the plethora of medicinal plant species available, some of which are challenging to distinguish from one another. For the most part, experts in some countries still use antiquated methods of herb identification. Both human and animal health can benefit from the employment of therapeutic herbs. In traditional Chinese medicine, they were known as common plants; today, we refer to them as herbal remedies. Although the plant is rarely used, at least one of its parts can be used to manufacture herbal treatments. Various sections of the same plant may be used for different reasons. Aside from their use in medicine, many plants are also valuable for their use in cooking and the production of safe drinks. People have always looked to nature for answers to their health problems. In humans, herbal medicine is used to treat a wide variety of conditions. Plants were used to treat and prevent ailments before the discovery of iatrochemistry in the sixteenth century [3]. Yet, due to the growing number of negative side effects associated with the use of synthetic pharmaceuticals and their declining efficiency, the usage of natural therapeutic herbs has come back into the spotlight. Plants are the primary source of nourishment and shelter for the great majority of animal species. Furthermore, many people today use fuels such as coal and conventional gas that were formed from plants that have been present for a long time, but humans have seriously degraded the herbal environment in recent years, leading several harvests to fall short. The environmental calamity that followed, on the other hand, had a wide range of severe impacts, including land degradation, climatic abnormalities, disasters, and so on, all of which jeopardized human life and development. The proliferation of low-quality chemicals in the herbal medicine market poses a threat to human health and the industry's potential to expand internationally. As a result, research into new ways to categorize herbal medications has exploded in recent years. It is now widely accepted that the plant's leaf has characteristics that facilitate harvesting and research. Thus, it is naturally used as the primary method of identifying all medicinal plants. Because of recent breakthroughs in image processing, automatic computer image identification is now widely used in this field.
There are millions of plant species on Earth; some are poisonous to humans, others are used in medicine, and yet others are on the verge of extinction. Plants are vital not just to human existence, but also to the entire stability of the food chain. The main uses for medicinal plants are herbal, Ayurvedic, and traditional medicine. Herbal plants are plants that can be used to treat illnesses naturally. Almost 80% of people worldwide still practice traditional medicine. Herbal plants are those whose roots, stems, or leaves can be used as components in the production of pharmaceuticals. Most of the time, you may locate this kind of medicinal plant in the woods. By recognizing herbs, they can gain a lot about their characteristics, and one strategy is to examine the leaves. To preserve plant species, accurate plant research and classification are essential [4].
Ayurvedic doctors used to gather plants and make treatments for their patients. This method is still used by a small number of people today. Ayurvedic medicine manufacture and distribution today produces more than Rs. 4000 crores in annual revenue. There are around 8500 Ayurvedic drug manufacturers in India. The quality of the raw ingredients used to make Ayurveda treatments has come under fire as the Ayurvedic industry has become more commercialized. Women and children, who lack the specialized expertise necessary to identify the appropriate therapeutic herbs, now gather the plants from the wild. Incorrect or substitute medicinal plants are frequently delivered to manufacturing facilities. Most of these facilities lack proper quality control procedures to inspect these plants. Additionally, there is a considerable lot of ambiguity due to regional differences in names. Some plants arrive dry, which makes manual identification substantially more difficult. Ayurveda therapy is useless when medicinal herbs are used improperly. Unanticipated consequences are also a possibility. In this context, strong quality control procedures must be imposed on Ayurveda pharmaceuticals and raw materials utilized by the industry to safeguard the industry's current growth while maintaining the efficacy and credibility of medications.
Nonetheless, because expert opinions are available, manually identifying medicinal plants is equally difficult and time-consuming as identifying any other form of the plant [5]. Motivated by these hurdles, researchers constructed various autonomous plants or leaf recognition systems, where most of them applied Machine learning (ML) methodologies.
Literature Survey
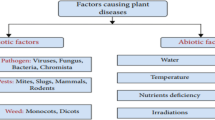
It has become common practice in recent years to use ML and DL techniques to classify plants based on images of their leaves; this article [1] compares the accuracy and prediction abilities of these methods. A few classifiers used to identify leaves and extract important leaf attributes are described in this study's image-processing methodologies. Early detection of plant diseases is crucial because they might stunt the development of some plant types. Current advancements in DL, and ML, seems to offer a tremendous opportunity to enhance accuracy when it comes to identifying and categorizing the symptoms of plant diseases. This article provides a comprehensive review of the ML and DL algorithms used to classify plant leaves.
The research [6] proposes a method for the automated, real-time recognition of plant species, specifically for medicinal herbs native to Borneo. An improved EfficientNet-B1 model was trained and tested on private and open accessible plant species databases to solve the recognition challenge. In both datasets, the proposed model achieved accuracy of over 10% compared to the conventional model on test data. This study stands out because of its innovative utilization of public feedback and geo-mapping of plants within the Borneo area, both of which were made possible using a mobile application. Yet, preliminary results from the suggested method indicated that it had promise as a real-time approach to recognizing medicinal species.
The purpose of the work [7] was to investigate the possibility of using automated recognition to correctly identify medicinal plants. In this study, they present a new database of medicinal herbs that includes images of 10 categories of medicinal and 1 category of non-medicinal species. Then, proposed a model for an affordable, reliable, and effective classification of medicinal plants based on the MobileNetV3 architecture. The proposed model using transfer learning improves accuracy on the tough challenge. Overall, the results showed that a robust and effective classifier for therapeutic plants is possible.
The study [8] analyzes the practicality of utilizing convolutional neural network (CNN)-based approaches to discern between several species of Indian leaves. In recent years, several DL frameworks have been applied to the task of recognizing and classifying plants. The primary goal of this research was to create a database of therapeutic plants that do well in remote locations. A pre-trained CNN architecture called MobileNet was chosen using the transfer learning strategy. Models were evaluated using their pre-trained weights on a dataset consisting of thirty classes of medicinal plants and three thousand images. The trained model achieved 98.05% accuracy on a hidden test set, validating the approach.
Based on the provided leaf sample, the author [9] presents a technique that can determine the plant species. With the help of ExG-ExR, an enhanced vegetation index, additional data may be gleaned from images about vegetation. Since that it comes with an inherent threshold of zero, setting a threshold value of OTSU is unnecessary. Although ExG employing the Otsu’s technique produces more flora and fauna information, their ExG-ExR index functions brilliantly irrespective of the illuminating backdrop. This means that the ExG-ExR index can be used to pinpoint the location of a study focused on binary species. The mask was utilized to split the leaves to separate images. Every leaf's colour and texture data are retrieved, and then a Logistic Regression model is utilized to correctly recognize the plant species.
Based on the findings [10], a deep CNN framework may be developed, and its parameters adjusted to improve the recognition rate. This research highlights the critical role of the multi-layer strategy's impacts on low-number samples in achieving satisfactory outcomes. In addition, data augmentation has larger positive effects on productivity. Simple transformations such as resizing, flipping, and rotating can greatly improve accuracy provided invariance is incorporated and the model is prevented from acquiring unnecessary information. The experiment results improved as a new leaf database from several Malaysian medicinal plants was developed.
In the study [11], the categorization of 6 distinct medicinal species is examined using a range of morphological, colour, and textural characteristics. On the outskirts of Assam, India, they gathered 90 leaf images representing 6 different medicinal plant species. For the classification job, each attribute was used separately before being combined for more precision. The back propagation neural network (BPNN) is employed for efficient leaf recognition. Extensive testing on ninety images of leaves from six distinct types demonstrates that the proposed method improves precision. The outcomes show that the suggested strategy successfully differentiates between leaves with varying colour, morphological, and textural characteristics.
Research [12] introduces an ensemble learning approach to the DL subject in artificial intelligence, which enables quick identification of plant species from images of their leaves. Thirty distinct types of medicinal leaves are included in the dataset. A transfer learning technique was used to build and pre-train the neural network model. The input image features were retrieved using these models, which were learned with the relevant database and a classifier made of a SoftMax connected to a Dense Layer. The 3 and fivefold cross-validation was employed to confirm the accuracy of the results. They used a classification called Ensemble Deep Learning-Automated Medicinal Leaf Identification (EDL-AMLI) that averaged the results from several different models. Better results were achieved with the EDL-AMLI compared to the most advanced pre-trained algorithms.
The proposed method [13] highlights incomplete issues in the datasets to increase the detection accuracy for herb recognition. The inclusion of dimension variables in the datasets improves the image segmentation method. Deep knowledge-based identification is the process of validating a result using an ML classifier and an Exclusive-OR gate operation. The two-stage authentication (TSA) method improves the recognition accuracy required for detecting herbal leaves. By combining image segmentation with ML, they can create an architecture that is both efficient and resilient. Using the most appropriate image segmentation methods for extracting leaves from images also aids in the enhancement of the accuracy rate. The outcomes demonstrate that the proposed method improves upon traditional performance metrics, such as accuracy.
Methodology
The research focuses on identifying the medicinal herb in real-time using the mobile app. The research is composed of three stages training testing and deployment.
-
Training: in this stage, first the data are collected from the Kaggle website. Next, the data are split into training, validation, and testing. The gathered data undergo processing techniques like resizing and augmentation. The training and validating data are used to train the designed DL model. For evaluation, the precision, recall, accuracy, and loss metrics are chosen.
-
Testing: after training and validating the DL model, the testing was done on test data. This phase is used to finalize the DL model if the model gives a satisfactory response.
-
Deployment: in many cases, the research gives good results, but it is not deployed in real-time. Normal people cannot taste the use of the DL model. To break this barrier, the mobile app is designed. And the designed DL model is deployed in the cloud. The user can use the mobile app to capture the leaf images, then the app collects the images and sent them to the DL model present in the cloud. The DL model helps to identify the medicinal species. All the above-mentioned details are represented in Fig. 1 using the flow chart.
Data and its Processing
Many medicinal plants are available on earth. Many drugs are derived from medicinal plants and have been used to treat human illness for many centuries. Classifying medicinal plant species is essential for the development and preservation of pharmaceuticals. The general community lacks a sufficient understanding of the properties and applications of their medicinal herbs. To solve this DL model is employed. The DL model requires many leaf image samples to train them. In this research, the data were collected from Kaggle [14]. There are many medicinal species available on earth, we focus on six species Betel, Curry, Tulsi, Mint, Neem, and Indian Beech. The sample images of each species are given in Fig. 2. The Kaggle database holds 500 images of each species. We took 350 images of each species for training, 100 for validation, and 50 for testing. The data distribution of this research is given in Table 1 and Fig. 3. 70% of the data was taken for training, 20% for validation, and 10% for testing.
After image collection, the pre-processing is done on the images. The pre-processing stages involve resizing and augmentation. The various CNN networks have different image size requirements [15]. Hence, to fit the MobileNet model, the images were downsized to 224 × 224. To get better accuracy, the trained data must be more. But we have only 350 images of each species. Data augmentation comes into the picture to increase the training sample.
Data augmentation techniques are employed to artificially increase the size of the dataset [16]. Most of the work with tiny datasets uses this method. It helps to give more images for training, which helps avoid overfitting, and it also helps to supply more images for testing purposes. The images are rotated, flipped, and had their colour altered to build a larger dataset using data augmentation. The training images were rotated both clockwise and anticlockwise by an angle ranging from 5 to 90 degrees. The flipping of images is horizontal and vertical out of an image. The process of altering an image's colour is known as image manipulation. Each medicinal species before and after augmentation such as rotation, flip, and colour manipulation are given in Table 2.
Deep Learning Model Construction
The DL model is employed to classify the medicinal species. We have chosen the MobileNet model for this purpose. The MobileNet [17] model, which is built on a CNN, is commonly used to categorize images. Since the MobileNet model uses fewer computing resources than the standard CNN approach, it can be used on low-powered smartphones and desktop computers [18]. The MobileNet framework is a basic model that can be employed to partition data based on two controllable criteria that effectively switch between accuracy and latency. When it comes to saving space on a network, the MobileNet method comes out on top.
With fewer features, the MobileNet architecture still works well [19]. MobileNet is organized in a hierarchy. The core of the architecture is made up of multiple abstraction layers, which look to be the quantized setup that accurately evaluates the complexity of a typical task. Abstraction layers that pass through a rectified linear unit (ReLU) [20] are used to generate depth platforms. To decrease the dimensions in both the source images and every layer's inner depiction, the quality multiplier variable ω is introduced.
The dimension of the feature vector map is \({F}_{m}*\) \({F}_{m}\) and the dimension of the filter is \({F}_{s}*\) \({F}_{s}\). The input can be denoted by \(p\), and the resulting outcome by \(q\). The variable ce represents the entire amount of computing labour for the architecture's core abstract layers, and it may be calculated using the equation below (1).
Context determines the appropriate multiplier value ω; for this study on leaf disease classification techniques, we will assume a multiplier value of between 1 and \(n\). It is assumed that the resolution multiplier α, indicated by 1. Using the below-mentioned Eq. (2), the computation efforts can be determined via variable coste.
The suggested approach includes depth and point-wise convolutions, both of which are constrained by a depletion element, \(d\), which is represented by the following Eq. (3):
When it comes to making an accurate estimation, the width and resolution multiplying hyper-features are invaluable tools. Images having dimensions of 224 by 224 by 3 pixels are supported by the proposed model. The image's dimensions are indicated by the first 2 digits (224*224). These digits must always be at least 32. Based on its third digit, we can deduce that it collects information from three sources. The suggested architecture consists of 32 filters, each dimension is 3*3*3.
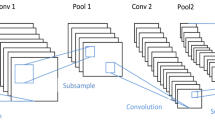
The goal behind MobileNet structures is to replace complex convolutional layers (CL) with simple ones. Every layer consists of a CL with dimension 3*3 that delays the inputs, followed by a CL with dimension 1*1 pointwise that combines these filtering variables to produce a new element. The purpose of the abovementioned technique is to compress the model and improve its effectiveness over a normal convolutional network. The architecture of MobileNet is given in Fig. 4.
Deep Learning Model Deployment
Google is a major competitor in the market for cloud computing and has its cloud service called Google Cloud Platform (GCP). Data in the tens of thousands of terabytes range is created, saved, and analysed daily by individuals around the globe. Google has spent a lot of money over the years building out its processing and storage facilities to handle this massive amount of data [21]. These massive computational facilities are available to the public via GCP and may be used to save ML and DL models. Securing, saving, distributing, and analysing the data are some of the functions offered by GCP. Such cloud services provide a protected cloud perimeter around data, letting it undergo a wide range of processing and conversion without exiting the cloud environment. Google Cloud Storage provides extensible, real-time access to both present and past data inside the confines of the cloud. With using cloud storage, you can access the files from anywhere in the globe, at any time. When you factor in the magnitude and monetary importance of the data being saved, the price tag for this kind of storage power is surprisingly low. The expense is also justified when you consider the convenience, safety, and consistency that cloud storage offers. Data mining, exploration, streaming, batch analysis, cloud-based Hadoop ecosystems, and communications systems are just some of the solutions offered by GCP. A wide range of methods for mining and producing real-time insight from big data is provided by such services.
On computed platforms with sufficient resources, there are numerous well-established DL frameworks [22]. They have been utilized extensively and perform well in many DL tasks. These substantial structures, however, are inappropriate for mobile systems with limited resources. First off, despite being comprehensive, most of these frameworks require additional components, making them too large and resource-intensive to be used effectively on low-powered mobile devices. Second, the inference phase of the mobile DL system must be prioritized in contrast to a more traditionally designed framework. Furthermore, unlike a non-mobile DL framework, portable DL optimization focuses on model simplification and hardware-oriented computation efficiency. Even though mobile versions of many models exist, these models match perfectly with the powerful GPU processors that aid in training but provide little in the way of utility during inferences. To train and execute DL models on low-powered mobiles, there has been a recent focus on developing professional software packages. To this end, we will be integrating existing frameworks and accelerating the inference process of trained models to substantially reduce the number of resources required to execute a mobile app.
Results and Discussion
The purpose of this study is to identify medicinal plants in real-time. The necessary medicinal leaf images were obtained from Kaggle to complete this challenge. Because the collected images are of various sizes, they have been scaled to a specific dimension. Geometrical augmentation was performed on the leaf images because the collected data is insufficient to train the DL model. The augmented data is fed into the DL model, which is trained with an epoch of 15.
Training and Validation Result
Metrics like loss, recall, accuracy, and precision are used to assess the performance of a DL model during the training and validation stages. Table 3 displays the obtained metrics values. At the train and validate stages, the DL model's loss value is 1.07 and 1.64 at the 0th epoch. The loss value gradually lowers and reaches values of 0.08 and 0.18 as the epoch count increases. The second metric considered is recall performance. During training, the score begins at 0.53 on the 0th epoch and steadily rises to 0.97; during validation, the recall values at the initial and final epochs are 0.67 and 0.94. The accuracy of the DL model is assessed in the third step. The training and validation accuracy scores of the DL model are 0.64 and 0.68 at the 0th epoch. The accuracy scores for both phases are 0.97 and 0.95 at the final, 14th epoch. The precision of the DL model throughout the training and validation phases is then evaluated. Precision at the initial epoch yields a value of 0.80 and 0.70, which reaches 0.97 and 0.95 at the final epoch.
Line plots of the data from Table 3 are shown in Fig. 5. The MobileNet's loss plot is depicted in Fig. 5a, and the recall plot is shown in Fig. 5b. Figures 5c, d display a precision and accuracy graph. The epochs are plotted along the x-axis and the metrics are along the y-axis in all of the graphs. All plots, except for the loss plot, rise as the number of epochs increases.
Testing Result
For DL model testing, test data are used after the model has been trained and validated. Accuracy, precision, and recall are three measures considered to examine the testing outcome. The accuracy, recall, and precision scores of the DL model are 98.33, 97.36, and 99.32%, respectively, on test data. The results show that the developed MobileNet is effective in identifying medicinal plants, with a total average score of above 97% across all measures. Figure 6 and Table 4 shows the metrics score of the DL model applied to test data in the form of a bar graph.
Deployment
During the testing phase, the constructed model produces satisfactory results. So, we have finalized the MobileNet model and deployed it to the cloud named GCP. The next step is to design the mobile app with the target user in mind. Minimal options were included in the final version of the mobile app. The login screen shows automatically when the user launches the app. A user registers for an app by providing basic information about themselves. The homepage is displayed after login. The user can only see one option on the home page, which is to scan or upload the leaf. Now, we have the option of using the phone's camera to perform a direct scan of the plant or uploading has previously taken pictures of the leaves. The prediction option enables after the images have been uploaded. When a user taps the app's prediction button, the uploaded images are forwarded to the cloud-based DL model for analysis. The DL model is useful for determining the correct name of the healing herb. The results of the DL model are again sent to the mobile app from the cloud. The leaf's name and its health advantages are displayed in the app. Scanning the tulsi crop is used to test the developed mobile app. Figure 7 also illustrates how the mobile app works. The name of the plant is identified accurately and the software also displays a dialogue box detailing the advantages of using the plant.
Conclusion
Plants have a crucial role in maintaining human life. Traditional herbal remedies have been used by indigenous communities for thousands of years. Clinicians often identify herbs by years of accumulated familiarity with their smell or taste. Automatic herb identification has been greatly aided by recent developments in analytical technologies. Many people appreciate this, especially newcomers to the world of herb identification. Furthermore, laboratory-based analysis requires proficiency in sample collection and data interpretation, both of which add extra time and effort to already-lengthy procedures. So, it is necessary to have a quick and accurate procedure for detecting herbs. In this study, we use DL to develop an automated system for identifying medicinal plants. Only six herbs are used in this study. Over 97% in accuracy, precision, and recall are achieved by the created DL model. This model will be deployed in the cloud and create a smart mobile app that can instantly identify medicinal plants. For those who lack access to costly measuring instruments, this DL and mobile-based technique will be the method of choice for the rapid detection of medicinal plants. Future studies will focus on enhancing or maintaining the model's classification performance by taking more medicinal plant species.
Data Availability
The sample data set information is included in the article that support the findings of this research.
References
Chanyal H, Yadav RK, Saini DKJ. Classification of medicinal plants leaves using deep learning technique: a review. Int J Intell Syst Appl Eng. 2022;10(4):78–87.
Javid A, Haghirosadat BF. A review of medicinal plants effective in the treatment or apoptosis of cancer cells. Cancer Press J. 2017;3(1):22–6.
Barimah KB, Akotia CS. The promotion of traditional medicine as enactment of community psychology in Ghana. J Community Psychol. 2015;43(1):99–106.
Rao RU, Lahari MS, Sri KP, Srujana KY, Yaswanth D. Identification of medicinal plants using deep learning. Int J Res Appl Sci Eng Technol. 2022;10:306–22.
Singh V, Misra AK. Detection of plant leaf diseases using image segmentation and soft computing techniques. Inf Process Agric. 2017;4(1):41–9.
Malik OA, Ismail N, Hussein BR, Yahya U. Automated real-time identification of medicinal plants species in natural environment using deep learning models—a case study from Borneo Region. Plants. 2022;11(15):1952.
Valdez DB, Aliac CJG, Feliscuzo LS. Medicinal plant classification using convolutional neural network and transfer learning. In: 2022 IEEE International Conference on Artificial Intelligence in Engineering and Technology (IICAIET). IEEE; 2022. p. 1–6.
Abdollahi J. Identification of medicinal plants in ardabil using deep learning: identification of medicinal plants using deep learning. In: 2022 27th International Computer Conference, Computer Society of Iran (CSICC). IEEE; 2022. p. 1–6.
Sivaranjani C, Kalinathan L, Amutha R, Kathavarayan RS, Kumar KJJ. Real-time identification of medicinal plants using machine learning techniques. In: 2019 International Conference on Computational Intelligence in Data Science (ICCIDS). IEEE; 2019. p. 1–4.
Zin IAMd, Ibrahim Z, Isa D, Aliman S, Sabri N, Mangshor NNA. Herbal plant recognition using deep convolutional neural network. Bull Electr Eng Inform. 2020;9(5):2198–205.
Saikia AP, Hmangaihzuala PVL, Datta S, Gope S, Deb S, Singh KR. Medicinal plant species classification using neural network classifier. In: 2021 6th International Conference on Communication and Electronics Systems (ICCES). IEEE; 2021. p. 1805–11.
Sachar S, Kumar A. Deep ensemble learning for automatic medicinal leaf identification. Int J Inf Technol. 2022;14(6):3089–97.
Manoharan JS. Flawless detection of herbal plant leaf by machine learning classifier through two stage authentication procedure. J Artif Intell Capsule Netw. 2021;3(2):125–39.
https://www.kaggle.com/datasets/vishnuoum/medicinal-plant-dataset-augmented?select=data.
Saponara S, Elhanashi A. Impact of image resizing on deep learning detectors for training time and model performance. In: Applications in Electronics Pervading Industry, Environment and Society: APPLEPIES 2021. Cham: Springer International Publishing; 2022. p. 10–17.
Shorten C, Khoshgoftaar TM. A survey on image data augmentation for deep learning. J Big Data. 2019;6(1):1–48.
Sae-Lim W, Wettayaprasit W, Aiyarak P. Convolutional neural networks using MobileNet for skin lesion classification. In: 2019 16th International Joint Conference on Computer Science and Software Engineering (JCSSE). IEEE; 2019. p. 242–7.
Wang W, Hu Y, Zou T, Liu H, Wang J, Wang X. A new image classification approach via improved MobileNet models with local receptive field expansion in shallow layers. Comput Intell Neurosci. 2020.
Rabano SL, Cabatuan MK, Sybingco E, Dadios EP, Calilung EJ. Common garbage classification using MobileNet. In: 2018 IEEE 10th International Conference on Humanoid, Nanotechnology, Information Technology, Communication and Control, Environment and Management (HNICEM). IEEE; 2018. p. 1–4.
Patel R, Chaware A. Quantizing MobileNet models for classification problem. In: 2021 8th International Conference on Computing for Sustainable Global Development (INDIACom). IEEE; 2021. p. 348–51.
Bisong, E. Building machine learning and deep learning models on google cloud platform: a comprehensive guide for beginners, 1st edn; 2019.
Wang Y, Wang J, Zhang W, Zhan Y, Guo S, Zheng Q, Wang X. A survey on deploying mobile deep learning applications: a systemic and technical perspective. Digit Commun Netw. 2022;8(1):1–17.
Funding
Open access funding provided by Manipal Academy of Higher Education, Manipal. There is no funding for this research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors do not have any conflicts of interest with this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the topical collection “Diverse Applications in Computing, Analytics and Networks” guest edited by Archana Mantri and Sagar Juneja.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kavitha, S., Kumar, T.S., Naresh, E. et al. Medicinal Plant Identification in Real-Time Using Deep Learning Model. SN COMPUT. SCI. 5, 73 (2024). https://doi.org/10.1007/s42979-023-02398-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42979-023-02398-5