Abstract
The incidence of infections caused by resistant Gram-negative pathogens has become a critical factor in public health due to the limitation of therapeutic options for the control of infections caused, especially, by Enterobacteriaceae (Escherichia coli and Klebsiella pneumoniae), Pseudomonas spp., and Acinetobacter spp. Thus, given the increase in resistant pathogens and the reduction of therapeutic options, polymyxins were reintroduced into the clinic. As the last treatment option, polymyxins were regarded as the therapeutic key, since they were one of the few classes of antimicrobials that had activity against multidrug-resistant Gram-negative bacilli. Nonetheless, over the years, the frequent use of this antimicrobial has led to reports of resistance cases. In 2015, mcr (mobile colistin resistance), a colistin resistance gene, was described in China. Due to its location on carrier plasmids, this gene is characterized by rapid spread through conjugation. It has thus been classified as a rising threat to public health worldwide. In conclusion, based on several reports that show the emergence of mcr in different regional and climatic contexts and species of isolates, this work aims to review the literature on the incidence of the mcr gene in Brazil in different regions, types of samples identified, species of isolates, and type of carrier plasmid.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Recently, antimicrobial resistance (AMR) has become a critical factor in public health due to the reduction in therapeutic options associated with increased morbidity and mortality rates [1, 2]. First, with the development and commercialization of antimicrobials, along with the increase in the prevalence rates of bacterial resistance, the scientific community has focused on Gram-positive pathogens [3,4,5]. Thus, the change in the susceptibility pattern of pathogens and the multidrug resistance (MDR) presented by Gram-negative bacilli has become a threat to the medical and scientific community worldwide [5]. Indeed, between 2000 and 2010, approximately 70 billion doses of antibiotics were administered globally and 50% of doses are used incorrectly or unnecessarily, thus facilitating the development of AMR [1, 6].
AMR is divided into two types: (i) innate resistance and (ii) acquired resistance. Intrinsic or innate resistance is characterized by the particularities of each microorganism and depends on its biological characteristics. Furthermore, acquired resistance can result from the acquisition of resistance genes, mutation of chromosomal DNA, or both mechanisms [7]. Due to the emergence of resistant pathogens over the years, new classes of drugs have been identified. In the last 20 years, two new classes of antibiotics, lipopeptides and oxazolidinones, have been classified and approved for use. However, the rapid ability of bacteria to acquire resistance to these new classes made them ineffective against some bacterial strains [8].
According to the World Health Organization (WHO), the lack of treatment options for multidrug-resistant Gram-negative bacilli (MDR-GNB), including carbapenem-resistant, such as Escherichia coli and Klebsiella pneumoniae, Pseudomonas spp., and Acinetobacter spp., results in serious infections that can lead to public health problems [9]. The irrational use of carbapenems was due to the emergence of cephalosporinases, which in turn resulted in carbapenem resistance [3, 10]. Therefore, with the increase in carbapenem-resistant pathogens, polymyxins have been reintroduced into the clinic as one of the last options for the treatment of MDR-GNB [5, 10, 11].
Polymyxins were first discovered in 1947, but in the 1970s, they were found to have high nephrotoxic potential after their incorporation into clinical practice [12]. After the arising of frequent cases of patients with renal dysfunction and the need for safer drugs, the use of polymyxins is now reduced, requiring the use of smaller doses for this antibiotic as well as preferential use of PMB over it, or its co-administration with antioxidants [13]. However, in an era of MDR bacteria, old antimicrobials have re-emerged as treatment options, being, in some cases, the only ones with activity against MDR-GNB [13, 14].
Ho et al. [15] in 2010 described polymyxins as a key therapeutic for infections caused by pathogens resistant to carbapenems. However, with the rise in the use of these drugs, an increase in cases of resistance to these lipopeptides has been noted [16]. In 2015, in China, a transferable plasmid-mediated colistin resistance gene, mcr-1, was identified, whose gene product, the MCR-1 protein, modifies the lipid A component of LPS, inhibiting the binding of polymyxins to target [17, 18].
It has been shown that the production of MCR-1 in pathogens such as E. coli leads to an increase in the minimum inhibitory concentrations (MICs) of polymyxins by up to eight times; therefore, even without additional resistance mechanisms, the production of this enzyme is sufficient to develop resistance to polymyxins [16]. In addition, the location of the mcr gene in plasmids makes it favorable for rapid propagation by conjugation, especially in E. coli isolates [17, 19].
Initial reports of the presence of mcr-1 in Brazil were based on 16 samples of E. coli isolated from poultry and swine. Subsequently, the first case of clinical isolates and environmental samples carrying mcr-1 has been reported in 2016 [20, 21]. Nowadays, there are several reports of isolates that produce MCR-1 related to nosocomial infections across the country [22]. Thus, the present study aims to understand, through a narrative literature review, the variants present in reports of the incidence of Gram-negative bacilli carrying the mcr gene in Brazil.
Characteristics of Gram-negative bacilli
Through the action of the virulence factors, bacteria use genes to carry out their infection and resist host defenses [23]. These factors are classified according to their mechanism and/or function, for example, (a) membrane proteins, which play a fundamental role in host adhesion, colonization, and invasion; (b) toxin-secreting proteins capable of modifying the host cell environment and are responsible for some host cell-bacteria interactions; (c) components of the outer membrane, such as lipopolysaccharide (LPS) that can protect against lysis mediated by the complement system; and (d) ability to form biofilms, which are bacterial aggregates contained in extracellular matrices of polysaccharides, proteins, enzymes, and nucleic acids. This matrix confers some resistance to bacteria, restricting the action of antibiotics, reducing the growth rate, and even neutralizing the immune system. Therefore, this last resistance factor associated with highlighted bacterial resistance mechanisms synergistically contributes to the wide dissemination of MDR-GNB strains [24, 25].
Gram-negative pathogens have genetic mutations and acquire mobile genetic elements that confer resistance mechanisms. Due to this, they are among the most common pathogens in community and hospital infections since they have a high transmission rate [24,25,26]. Furthermore, resistance genes have become a serious public health issue worldwide by compromising the effects of antimicrobials, especially β-lactams [15, 22].
As highlighted by Mota et al. [27], Enterobacterales and Gram-negative non-fermenting bacilli (GNNFB) are the most common pathogens associated with nosocomial infections. The author reported that, considering the isolated microorganisms, K. pneumoniae, followed by E. coli, A. baumannii, and P. aeruginosa, showed the highest prevalence in an intensive care unit (ICU) located in the Midwest region of Brazil. Additionally, the profile of bacterial resistance of these isolates to antimicrobials was analyzed and a significant resistance to β-lactams, such as penicillins (56.2%), second-generation cephalosporins (57.9%), third-generation cephalosporins (51.9%) and carbapenems (46.1%), and the class of antimicrobials quinolones (88.8%), was observed. In relation to GNNFB, it was realized that for A. baumannii the resistance could be between 80 and 100% for the β-lactams cefepime and meropenem, as well as for the fluoroquinolone ciprofloxacin, and from 50 to 70% for β-lactams the imipenem, piperacillin associated with beta-lactamase inhibitor tazobactam and ceftazidime, for the cefamicin, cefoxitin, and for the aminoglycoside gentamicin. P. aeruginosa isolates showed a lower rate of resistance in general when compared to other bacterial species, ranging from 30 to 40% for β-lactams cefoxitin, cefuroxime, imipenem, and meropenem [27].
Resistance mechanism in Gram-negative bacilli
The majority of antimicrobials are naturally produced, and co-resident bacteria develop a natural resistance mechanism known as intrinsic resistance [28]. After establishing itself in the genome of a microorganism’s genus or species, innate resistance will spread through subsequent generations. There are several causes for this characteristic, such as lack of affinity of the drug for the target, inaccessibility to the bacterial cell interior, or extrusion of the drug by active exporters and efflux pumps, yet the innate production of enzymes that inactivate the drug [29, 30].
Acquired resistance is defined as a process in which the bacteria that originally presented susceptibility to the drug become resistant. The development of this resistance is the result of gene mutations transferred from the parent cell to the daughter cell or through gene exchange mechanisms [28]. Clinically, acquired resistance is more significant due to the high spread of resistance genes [29]. This transfer can be performed by plasmids, which can carry resistance genes to multiple classes of antibiotics and integrate into the bacterial genome [31].
The mechanism of transfer of genetic material includes conjugation, transduction, and transformation [32]. The gene transfer through conjugation occurs through the contact between the surfaces of the donor and the receptor cells with the transfer of the plasmid carrying the resistance gene. This process has been widely reported in Gram-negative pathogens [7].
The production of antimicrobial-modifying enzymes is another resistance mechanism that can be mediated by plasmids. Resistance of Pseudomonas aeruginosa isolates to fosfomycin is one of the examples of this type of resistance reported in the clinic [7]. Chromosomal mutations that occur in incorrect DNA repair or chromosomal replication have also been a widely reported bacterial mean of acquiring resistance, as well as the expression of efflux systems, which is responsible for the resistance to fluoroquinolones present in several Gram-negative isolates, for example [7, 32, 47].
As a result of multiple resistance mechanisms coexisting within a pathogen, a MDR phenotype has become common among isolates associated with nosocomial infections [22]. In 2011, terminologies were created to standardize and describe the profiles of broadly resistant microorganisms, such as MDR (multidrug-resistant)—resistant to 3 or more classes of antimicrobials; XDR (extensively drug-resistant)—microorganisms that remain susceptible to at least one or two antibiotics belonging to two different classes; PDR (pandrug-resistant)—proven resistance in all categories of antimicrobials tested. In this way, the increase in the detection rate of pathogens with widespread resistance MDR, XDR, and PDR is attributed to a combination of microbial features, uncontrolled use of antibiotics, and dissemination of resistance genes [30, 33].
The alarming lack of effective antimicrobial agents to combat pathogens with a wide spectrum of resistance in recent years has led to the reintroduction of polymyxins B and E, two antibiotics from the 1960s that had promising in vitro activity against resistant pathogens, but their use was limited due to their high incidence of nephrotoxicity and neurotoxicity [34, 35]. The intense use of polymyxins for the absence of equally effective antibiotics consequently led to the identification of some strains resistant to these antibiotics not long after their return, even though this resistance was chromosomal, and therefore there was no risk of rapid distribution and dissemination [36]. However, the whole scenario of the use of polymyxins is incisively threatened with the first report of plasmid resistance, attributed to the new gene mcr-1 [37].
Polymyxins and the plasmid resistance gene
It is essential for antibiotics to penetrate bacterial amphiphilic barriers, allowing the active ingredient to enter the cell. Cyclic antibiotics have the ability to interact with bacterial membranes. In this context, the constituent compounds of this class have been used in the front line against infections caused by MDR Gram-negative isolates [38, 39].
Polymyxin was obtained from the microorganism Paenibacillus polymyxa [40]. Its chemical structure is composed of a decapeptide core that contains an intramolecular cycle of heptapeptides linked by amide-type bonds between the amino group of the side chain of the diaminobutyric acid (Dab) residue at position 4 and the carboxyl group on the C-threonine residue terminal. Another important structural feature of the polymyxin chain is the presence of five non-proteinogenic Dab residues, which are positively charged at physiological pH, hydrophobic residues, and an N-terminal acyl group. This class of antimicrobials is formed by a group of five compounds (A, B, C, D, and E), which differ in terms of amino acid sequences and fatty acid side chains [41]. The chemical structure of polymyxins B and E (colistin) differs by the presence of an amino acid D-Leucine in the colistin molecule, and D-phenylalanine in the polymyxin B molecule [42].
Polymyxins act by binding to lipopolysaccharide (LPS) and phospholipid molecules found in the outer membrane of gram-negative bacteria. This process results in a change in the cell wall of the microorganism, leading to extravasation of intracellular content and, consequently, to the death of the bacteria [43]. Due to the electrostatic interaction between antimicrobial molecules with the phosphate group of lipid A of the bacterial membrane, polymyxins have a greater affinity for bacterial LPS when compared to magnesium and calcium cations, which in turn stabilize LPS [17]. This interaction causes a shift in calcium and magnesium ions, thus allowing the binding of the active principle to the membrane of the microorganism, resulting in a derangement in the membrane and, later, in an increase in the absorption of the antibiotic, which leads to osmotic imbalance, causing cell death [44].
Among the other mechanisms of action of polymyxins are the inhibition of enzymes essential for cellular respiration in bacteria, such as NADH-quinone oxidoreductase type II [45]. However, it has been reported that polymyxin susceptibility has decreased as well as an increase in infections caused by polymyxin-resistant Gram-negative pathogens. Polymyxin resistance was mainly observed in isolates of K. pneumoniae and E. coli. Thus, the clinical efficacy of this class of antimicrobials has become limited when used as monotherapy [22, 46].
The mechanism by which Gram-negative pathogens become resistant to polymyxins involves a chemical modification of the bacterial outer membrane [47]. These modifications are mediated by specific chromosomal mutations in the genes that control the two components system PhoP/Q and PmrA/B, such as the recently discovered CrrAB, which positively regulates the PmrAB system. CrrAB is responsible for the genes crrB that encodes a sensor kinase and crrA that encodes a regulatory protein and the modulator gene crrC regulators of the pbgP operon. Moreover, mgrB gene inhibits the PhoPQ system. This gene encodes a small transmembrane lipoprotein that is responsible for the negative regulation of the PhoPQ system. Since the two systems regulate resistance by altering the cationic charge and the ionic pattern of the outer membrane, changes in their function can lead to a decrease in the anionic charge and, consequently, prevent the electrostatic interaction of the bacterial cell with drug molecules [48,49,50].
Gene regulation results in the modification of LPS that leads to resistance. Gene regulation is mediated by two-component enzyme systems (TCS) that usually regulate resistance and virulence factors [49]. The inactivation of TCS regulator MgrB by non-sensing mutation, premature spot-codon, or gene interruption by insertion sequences, as well as change of the histidine kinase sensor CrrB resulting in PmrA/B overexpression, have been associated with polymyxin resistance in Klebsiella. Additionally, although the function of CrrAB has not yet been fully understood, it has been described that, when dysfunctional, CrrAB may also modify lipid A through the activation of a glycosyltransferase-like protein [49, 50]. Furthermore, the action of efflux pump systems, changes in protein concentrations of the outer membrane (porins), and thickening of the polysaccharide capsule are also extremely important for the development of pathogen resistance to polymyxins among the constituents of the order Enterobacterales [38].
In addition to resistance mediated by mutations involving TCSs, the mechanism encoded by a plasmid localization gene, mcr-1, described in China in 2015, has become the leading cause of polymyxin resistance [17]. There are two types of chemical changes in the outer membrane of Gram-Negative bacilli that lead to resistance to polymyxins, one of the main ones is the addition of 4-amino-4-deoxy-L-arabinose (L-Ara4N) in the lipid A component of LPS, reducing membrane antimicrobial activity. Another way is the addition of phosphoethanolamine (PEA) to the 1-(4′)-phosphate group of lipid A, promoting intrinsic resistance to polymyxins [50].
MCR-1 is described as lipase A-40-PEA-transferase. This family of alkaline phosphatases catalyzes the binding of phosphatidylethanolamine (PEA) to LPS lipid A and consequently leads to colistin resistance [51, 52]. Thus, transferable polymyxin resistance is related to the addition of PEA to LPS lipid A by MCR-1 [48,49,50,51,52]. Furthermore, the location of the mcr gene on plasmids favors rapid propagation by conjugation, especially in E. coli isolates (Fig. 1). The presence of this gene in a variety of plasmids highlights its ability to spread (Table 1) [17, 19, 52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70,71,72,73,74,75,76,77,78,79,80,81,82,83,84,85,86,87,88,89].
Jeannot et al. [62] highlighted the interspecies transfer of mobile genes as a factor responsible for the progressive spread of resistance genes. Additionally, the intrinsic mechanism and rates of polymyxin resistance in Enterobacteriaceae and isolates of K. pneumoniae, A. baumannii, and P. aeruginosa was observed until 2017. In their study, Jeannot et al. [635] also confirmed that mcr genes are located in the transferable plasmid, which, in turn, represents a high propagation factor by conjugation.
According to the studies present on the NCBI platform, ten variants of the mcr gene have been described, so far, in several species of microorganisms [17, 19, 54,55,56,57,58,59,60,61]. The mcr-1 gene has 13 variants, most of which have been reported in E. coli and K. pneumoniae isolates [63, 64]. The variants already described can be found in different classes of plasmids, characterizing a diversity of gene reservoirs. IncI2, IncHI2, and IncX4 are described as the main groups of plasmids responsible for carrying resistance genes. There are many studies describing the spread of colistin resistance related to gene variants, sites, the types of samples analyzed, the species of isolates, and the type of carrier plasmids (Table 2) [17, 21, 30, 34, 55, 60].
Incidence of the mcr gene in samples of clinical isolates in Brazil
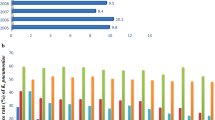
In Brazil, the first Gram-negative bacteria with polymyxin resistance were identified in E. coli isolates [65, 66]. Cases of colistin resistance in clinical isolates in humans began to be reported in 2016 (Fig. 2). Carried by the IncX4 plasmid, the mcr-1 gene was detected in E. coli strains (ST-101) recovered from a diabetic patient with a foot infection [20]. The case occurred in the Northeast Brazilian region and a complete genome analysis was performed [20].
In different studies, IncX4 has been identified as the main carrier of the colistin resistance gene in Enterobacteriaceae; these plasmids have been found in a variety of species, belonging to different “sequence types” (STs) and in a variety of samples, such as food, animals, and clinical isolates [20]. As a result, the presence of this carrier on different continents indicates that it is one of the most important factors behind the propagation of the mcr-1 gene. Correspondingly, Zamparette et al. [21] reported the presence of IncX4 in two Escherichia coli isolates (ST-206 and ST-354) as a carrier plasmid for the mcr-1.1 gene.
Analogously, Girardello et al. [64] isolated 490 colistin-resistant Gram-negative rods from human infections in a hospital setting in the Northeastern Brazilian region, of which eight were mcr-1.1-positive Escherichia coli. The genomes of these bacteria were sequenced, showing that mcr-1.1 was located in a contig that is presumed to be part of an IncX4 plasmid in seven of eight isolates. In the last one, mcr-1.1 was located in a contig that is presumed to be part of an IncHI2A plasmid.
Interestingly, isolates obtained from patients at an outpatient hospital in Santa Catarina (RS) who were not directly exposed to polymyxin showed an affinity of approximately 99.9% to the IncX4 plasmids reported in China and in Brazil for the first time. The IncX4 plasmid was found to be one of the main vectors of mcr spread in both analyses [21, 65]. Moreover, Vasconcelos et al. [67] coded the presence of other genes for MDR, such as blaCTX-M-2.
The coexistence of resistance genes has become a major concern due to the lack of therapeutic options [67]. Several studies have found the existence of more than one resistance gene in isolates of enterobacteria, especially in extended-spectrum β-lactamases (ESBL) producers [66]. Conceição-Neto et al. [68] and Dalmolin et al. [66] described the coexistence of colistin and carbapenem resistance genes. Dalmolin et al. [66] reported a case of K. pneumoniae harboring mcr-1 and blaKPC-2. The analysis was carried out through the screening of 3468 isolates recovered in several hospitals in the Southern region of Brazil, between 2013 and 2016. Another case of K. pneumoniae carrying the mcr-1 and blaKPC genes was described in a patient diagnosed with thrombosis, in a hospital in Vitória (ES) [69]. In another study, a single isolate containing the two resistance genes was reported in Porto Alegre (RS). An analysis of the data revealed that the plasmids carrying mcr-1 and blaKPC belonged to the IncX4 and IncFIB groups, respectively [66].
Nitz et al. [70] assessed the molecular profile of virulence and resistance genes in 99 isolates of P. aeruginosa collected from various clinical specimens in São Luís (MA), one of which had the mcr-1 gene. Except for carbapenems, imipenem, and meropenem, which it was sensitive to, this isolate showed resistance to at least one antibiotic from the following classes: ß-lactams, aminoglycosides, quinolones, and polymyxins, different from those presented in strains isolated from animal meat (Table 2).
Rocha et al. [70] showed the presence of mcr-1 gene variants in isolates of Enterobacteriaceae recovered from clinical isolates in a hospital in Recife (PE). Multiple studies have indicated the presence of different variants in Brazil over the years in this context. Kieffer et al. [71], Martins-Sorenson et al. [72], and Dos Santos et al. [74] reported the emergence of mcr-3.12, mcr-4.3, and mcr-3 and 7 alleles, respectively, carried by the IncY plasmid, in different types of samples and different isolates in South and Southeast regions of Brazil. These studies also showed a difference in the plasmids carrying the resistance genes. Kieffer et al. [71] described a plasmid IncA/C2, and Martins-Sorenson [72] described a previously unknown plasmid, pAb-MCR4,3, which carried mcr-4,3 in A. baumannii isolates. The mcr-9.1 allele in Salmonella typhimurium isolates was first identified from a clinical isolate in 2019 collected in the Southern Brazilian region; however, due to rapid spread, this allele has been reported globally among Enterobacteriaceae [75].
In addition, the emergence of the COVID-19 pandemic, caused by the SARS-CoV-2 virus, expanded the number of antimicrobial interactions in hospitals, therefore causing an increase in the occurrence of secondary infections and the spread of microorganisms in this environment. The COVID-19 pandemic has also enhanced the frequent prescription of antibiotics by doctors, since most patients develop mild/moderate symptoms that resemble a bacterial infection at early stages, as well as the search for these drugs in pharmaceutical shops, contributing to the current challenge of antimicrobial resistance [76]. The study conducted by Polly et al. [77] in an acute care hospital in Brazil, for example, already describes an increase in the incidence of carbapenem-resistant Acinetobacter baumannii (CRAB) and methicillin-resistant Staphylococcus aureus (MRSA) infections both in ICUs and non-ICUs units after the pandemic. Thus, these factors may further favor the spread of strains carrying the mcr gene in human clinical isolates [78].
Incidence of the mcr gene in animal and animal meat
The extensive use of polymyxins in production animals was recognized as a major cause of the emergence and rapid spread of the resistance gene. In this context, the first reports of these isolates were obtained from pork and chicken retail fair trade [20, 65]. Vasconcelos et al. [67] identified IncX4 as a carrier of the mcr-1 gene in strains of E. coli (ST-359) obtained from chicken carcasses from a public market in the Northeast Brazilian region.
Brazilian cattle without previous contact with polymyxin showed several resistance genes, according to Palmeira et al. [79]. In the study, the presence of mcr-1 and blaCTX-M-2 was observed in an E. coli isolate (ST-443) obtained in northeastern Brazil. Genes for colistin resistance and extended-spectrum β-lactamases (ESBL) were in separate plasmids, IncX4 and IncF, respectively. Fuentes-Castillo et al. [80], in turn, reported the coexistence of resistance genes in Enterobacter kobei (E11R) strains isolated in Angra dos Reis (RJ). The isolates were recovered from a culture of the respiratory exudate of a franciscana dolphin, an endangered species. The analyses reported the presence of mcr-9.1 and blaCTX-M-15, in addition to aminoglycoside, tetracycline, fluoroquinolone, fosfomycin, and sulfonamide resistance genes.
In 2020, mcr-9-mediated colistin resistance was described by Leite et al. [75] on Salmonella typhimurium (ST-19) isolates found in swine raised in the Northeastern Brazilian region. Resistance was determined by the minimum inhibitory concentration, and from genome sequencing, the presence of plasmids IncHI2 and IncHI2A was reported. Salmonella enterica serovar Typhimurium carrying the mcr-1 was described in 2018, in samples taken from pork reservoirs in Southern Brazil [76]. In a study carried out by Rau et al. [81], Salmonella enterica carrying the colistin resistance gene was detected and characterized in foods produced in Brazil; 90% of the detected isolates belonged to the serovar Typhimurium and harbored the mcr-1 gene in plasmid IncX4.
Studies carried out by Moreno et al. [82] detected another type of Salmonella carrying the colistin resistance gene. In the study, samples of chicken meat from São Paulo (SP) were selected for testing for the presence of the mcr gene. Two isolates that contained the gene were identified and genome analysis determined the presence of the serovar Schwarzengrund (ST-96) harbored in plasmid IncX4. The Schwarzengrund serovar had already been isolated from birds and in poultry meat; however, few studies indicated the presence of bacterial resistance. Furthermore, the presence of mcr in fair trade samples and clinical isolates of enterobacteria has been frequently described. Saidenberg et al. [83] reported the presence of mcr-bearing strains of E. coli MDR (ST131-H22) in domestic poultry from São Paulo (SP). The isolates showed the mcr-5.1 and mcr-9 variants, suggesting the role of poultry farming as one of the main reservoirs of colistin resistance in pathogenic enterobacteria.
Correspondingly, Barbieri et al. [84] tested the presence of mcr genes in 107 E. coli isolates obtained from production poultry in Rio de Janeiro (RJ). The mcr-1 gene was found in 62 (57.9%) isolates, while the mcr-5 gene was discovered in 3 (2.8%) isolates; all of these isolates were phenotypically resistant to colistin (MIC > 8 µg/mL). IncI2, FIB, and B/O were the most prevalent replicon types discovered, accounting for 38, 36, and 34% of mcr-positive isolates, respectively.
A study conducted by Kobs et al. [85], demonstrated the presence of the polymyxin resistance gene in pets from the Brazilian state of Santa Catarina. In his analysis, this study identified the first E. coli, Klebsiella spp., and Enterobacter spp. isolates carrying mcr-1 recovered from dogs and cats. A total of 64 bacterial isolates with phenotypic resistance to polymyxin B were identified, of which 62 isolates were from dogs and two from cats. Furthermore, all over the mcr-1-positive isolates were MDR. As a result, veterinary medicine should pay more attention to antimicrobial resistance.
Additionally, Ramos et al. [86] characterized 216 E. coli strains isolated from dogs fed either a raw meat-based (RMDB) diet or a conventional dry feed in Minas Gerais, southeastern Brazil. Isolates from RMBD-fed dogs were frequently positive for MDR E. coli isolates, and among them, seven ESBL producers were identified and subjected to whole-genome sequencing. The colistin-resistant gene mcr-1 was found to be positive in two strains, ST57 and ST410. Moreover, a BLAST search of the nodes containing the ESBL and mcr-1 genes revealed that the majority of them were found in mobile genetic components of varied replicon types, including the IncFII plasmid. Therefore, this study revealed that due to the risk of spreading these bacteria both within the household and in the community, ESBL and mcr-1 E. coli strains pose a potential risk not only to the host dogs but also to other animals and humans in the vicinity (Table 3).
Environmental incidence of the mcr gene
The existence of the mcr gene in pathogens that produce β-lactamases and carbapenemases, as shown in Table 4, is of concern for the scientific community [88]. Sacramento et al. [88] described the presence of the mcr-1 and blaCTX-M-8 genes in E. coli isolates recovered from a mangrove ecosystem in the Northeast region of Brazil. In this study, the presence of plasmids Incl1, IncX1, IncFII, IncFIB, and IncX4 was described. The colistin resistance gene was carried by the plasmid IncX4, corroborating the concept of this plasmid as one of the main agents of gene dissemination in enterobacteria [62]. Furthermore, the identification of pathogens with more than one resistance gene, especially those that present the mcr-9.1 variant, has represented a sign of greater propagation of the mcr gene [80].
Cordeiro-Moura et al. [89] detected the presence of mcr-1 in an E. coli strain isolated from a river sample in João Pessoa, Paraíba, as part of a project to analyze bacterial resistance in coastal waters. The strain isolated with the gene had an MIC of 4 µg/mL. In addition, plasmids IncX4, IncN, IncI1/Iγ, IncF, and IncFIB were identified in the isolate, in which the gene was identified as belonging to the IncX4 incompatibility group. Similarly, Fernandes et al. [65] detected the presence of the mcr-1 gene mediated by plasmid IncX4 and blaCTX-M in E. coli strains isolated from coastal waters of the city of São Paulo (Table 4).
Final considerations
Due to its wide dissemination, bacterial resistance has become a challenge for the scientific community. Resistant Gram-negative bacilli are present mainly in clinical cases. Unfortunately, the development of effective drugs does not keep up with the prevalence rates of MDR strains, even antimicrobials that have high activity against Gram-negative strains, such as polymyxins, have become ineffective in the treatment of infections. This fact is aggravated by the lack of Brazilian laws that prohibit the use of antibiotics of this class as growth promoters in animals for consumption, considered one of the main sources of AMR dissemination. In this context, the present study highlights the importance of the One Health concept in controlling the spread of AMR, emphasizing the influence of animal health and the environment on the prevalence, and spread of bacterial resistance genes. The trajectory of the incidence of the mcr gene in Gram-negative bacteria can be used as an example of the need for studies that highlight and use the One Heath approach, in addition to the importance of molecular epidemiology in the control of bacterial resistance. Furthermore, the pandemic scenario that began in 2019 will possibly be a major accelerator of the spread of resistant bacteria in Brazil.
Data availability
All data generated or analysed during this study are included in this published article.
References
Cruz-López F, Villarreal-Treviño L, Camacho-Ortiz A et al (2020) Acquired genetic elements that contribute to antimicrobial resistance in frequent Gram-negative causative agents of healthcare-associated infections. Am J Med Sci 360:631–640. https://doi.org/10.1016/j.amjms.2020.06.028
Bedos JP, Daikos G, Dodgson AR et al (2021) Early identification and optimal management of carbapenem-resistant Gram-negative infection. J Hosp Infect 108:158–167. https://doi.org/10.1016/j.jhin.2020.12.001
Durand GA, Raoult D, Dubourg G (2019) Antibiotic discovery: history, methods and perspectives. Int J Antimicrob Agents 53:371–382. https://doi.org/10.1016/j.ijantimicag.2018.11.010
Neto LVP, Oliveira MS, Orsi TD et al (2020) Alternative drugs against multiresistant Gram-negative bacteria. J Glob Antimicrob Resist 23:33–37. https://doi.org/10.1016/j.jgar.2020.07.025
Diekema DJ, Hsueh P-R, Mendes RE et al (2019) The microbiology of bloodstream infection: 20-year trends from the SENTRY Antimicrobial Surveillance Program. Antimicrob. Agents Chemother. 63:e00355-19. https://doi.org/10.1128/aac.00355-19
Van Boeckel TP, Gandra S, Ashok A et al (2014) Global antibiotic consumption 2000 to 2010: an analysis of national pharmaceutical sales data. Lancet Infect Dis 14:742–750. https://doi.org/10.1016/s1473-3099(14)70780-7
Khameneh B, Diab R, Ghazvini K, Fazly Bazzaz BS (2016) Breakthroughs in bacterial resistance mechanisms and the potential ways to combat them. Microb Pathog 95:32–42. https://doi.org/10.1016/j.micpath.2016.02.009
Brown ED, Wright GD (2016) Antibacterial drug discovery in the resistance era. Nature 529:336–343. https://doi.org/10.1038/nature17042
Brown P, Abdulle O, Boakes S et al (2020) Direct modifications of the cyclic peptide Polymyxin B leading to analogues with enhanced in vitro antibacterial activity. Bioorganic Med Chem Lett 30:127. https://doi.org/10.1016/j.bmcl.2020.127163
Harris PNA, Tambyah PA, Lye DC et al (2018) Effect of piperacillin-tazobactam vs meropenem on 30-day mortality for patients with E.coli or Klebsiella pneumoniae bloodstream infection and ceftriaxone resistance. JAMA 320:984. https://doi.org/10.1001/jama.2018.12163
Jayol A, Kieffer N, Poirel L et al (2018) Evaluation of the Rapid Polymyxin NP test and its industrial version for the detection of polymyxin-resistant Enterobacteriaceae. Diagn Microbiol Infect Dis 92:90–94. https://doi.org/10.1016/j.diagmicrobio.2018.05.006
Roberts KD, Azad MAK, Wang J et al (2015) Antimicrobial activity and toxicity of the major lipopeptide components of polymyxin B and colistin: last-line antibiotics against multidrug-resistant Gram-negative bacteria. ACS Infect Dis 1:568–575. https://doi.org/10.1021/acsinfecdis.5b00085
Pogue JM, Ortwine JK, Kaye KS (2016) Are there any ways around the exposure-limiting nephrotoxicity of the polymyxins? Int J Antimicrob Agents 48:622–626. https://doi.org/10.1016/j.ijantimicag.2016.11.001
Falagas ME, Skalidis T, Vardakas KZ, Legakis NJ (2017) Activity of cefiderocol (S-649266) against carbapenem-resistant Gram-negative bacteria collected from inpatients in Greek hospitals. J Antimicrob Chemother 72:1704–1708. https://doi.org/10.1093/jac/dkx049
Ho J, Tambyah PA, Paterson DL (2010) Multiresistant Gram-negative infections: a global perspective. Curr Opin Infect Dis 23:546–553. https://doi.org/10.1097/qco.0b013e32833f0d3e
Poirel L, Jayol A, Nordmann P (2017) Polymyxins: antibacterial activity, susceptibility testing, and resistance mechanisms encoded by plasmids or chromosomes. Clin Microbiol Rev 30:557–596. https://doi.org/10.1128/cmr.00064-16
Liu Y-Y, Wang Y, Walsh TR et al (2016) Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect Dis 16:161–168. https://doi.org/10.1016/S1473-3099(15)00424-7
Xu Y, Wei W, Lei S et al (2018) An evolutionarily conserved mechanism for intrinsic and transferable polymyxin resistance. mBio 9:e02317-17. https://doi.org/10.1128/mbio.02317-17
Xavier BB, Lammens C, Ruhal R et al (2016) Identification of a novel plasmid-mediated colistin-resistance gene, mcr-2, in Escherichia coli, Belgium, June 2016. Eurosurveillance 21:30280. https://doi.org/10.2807/1560-7917.es.2016.21.27.30280
Fernandes MR, McCulloch JA, Vianello MA et al (2016) First report of the globally disseminated IncX4 plasmid carrying the mcr-1 gene in a colistin-resistant Escherichia coli sequence type 101 isolate from a human infection in Brazil. Antimicrob Agents Chemother 60:6415–6417. https://doi.org/10.1128/aac.01325-16
Zamparette CP, Schorner M, Campos E et al (2020) IncX4 plasmid-mediated mcr-1.1 in polymyxin-resistant Escherichia coli from outpatients in Santa Catarina. Southern Brazil Microb Drug Resist 26:1326–1333. https://doi.org/10.1089/mdr.2019.0203
Rossi F, Girardello R, Morais C et al (2017) Plasmid-mediated mcr-1 in carbapenem-susceptible Escherichia coli ST156 causing a blood infection: an unnoticeable spread of colistin resistance in Brazil? Clinics 72:642–644. https://doi.org/10.6061/clinics/2017(10)09
Keyser P, Elofsson M, Rosell S, Wolf-Watz H (2008) Virulence blockers as alternatives to antibiotics: type III secretion inhibitors against Gram-negative bacteria. J Intern Med 264:17–29. https://doi.org/10.1111/j.1365-2796.2008.01941.x
Wu H-J, Wang AH-J, Jennings MP (2008) Discovery of virulence factors of pathogenic bacteria. Curr Opin Chem Biol 12:93–101. https://doi.org/10.1016/j.cbpa.2008.01.023
Dumaru R, Baral R, Shrestha LB (2019) Study of biofilm formation and antibiotic resistance pattern of gram-negative Bacilli among the clinical isolates at BPKIHS. Dharan. BMC Res. Notes 12:1–6. https://doi.org/10.1186/s13104-019-4084-8
Nordmann P, Naas T, Poirel L (2011) Global spread of carbapenemase-producing Enterobacteriaceae. Emerg Infect Dis 17:1791–1798. https://doi.org/10.3201/eid1710.110655
Mota FS, Oliveira HA, Souto RCF (2018) Perfil e prevalência de resistência aos antimicrobianos de bactérias Gram-negativas isoladas de pacientes de uma unidade de terapia intensiva - Revista RBAC. RBAC 50:270–277. https://doi.org/10.21877/2448-3877.201800740
Munita JM, Arias CA (2016) Mechanisms of antibiotic resistance. Microbiol Spectr 4:10. https://doi.org/10.1128/microbiolspec.vmbf-0016-2015
Tenover FC (2006) Mechanisms of antimicrobial resistance in bacteria. Am J Infect Control 34:S3–S10. https://doi.org/10.1016/j.ajic.2006.05.219
Blair JMA, Webber MA, Baylay AJ et al (2014) Molecular mechanisms of antibiotic resistance. Nat Rev Microbiol 13:42–51. https://doi.org/10.1038/nrmicro3380
Carattoli A (2009) Resistance plasmid families in Enterobacteriaceae. Antimicrob Agents Chemother 53:2227–2238. https://doi.org/10.1128/aac.01707-08
Leavitt A, Chmelnitsky I, Ofek I et al (2009) Plasmid pKpQIL encoding KPC-3 and TEM-1 confers carbapenem resistance in an extremely drug-resistant epidemic Klebsiella pneumoniae strain. J Antimicrob Chemother 65:243–248. https://doi.org/10.1093/jac/dkp417
Magiorakos A-P, Srinivasan A, Carey RB et al (2012) Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect 18:268–281. https://doi.org/10.1111/j.1469-0691.2011.03570.x
Nang SC, Li J, Velkov T (2019) The rise and spread of mcr plasmid-mediated polymyxin resistance. Crit Rev Microbiol 45:131–161. https://doi.org/10.1080/1040841x.2018.1492902
Giamarellou H (2016) Epidemiology of infections caused by polymyxin-resistant pathogens. Int J Antimicrob Agents 48:614–621. https://doi.org/10.1016/j.ijantimicag.2016.09.025
Lima WG, Alves MC, Cruz WS, Paiva MC (2018) Chromosomally encoded and plasmid-mediated polymyxins resistance in Acinetobacter baumannii: a huge public health threat. Eur J Clin Microbiol Infect Dis 37:1009–1019. https://doi.org/10.1007/s10096-018-3223-9
Khondker A, Rheinstädter MC (2020) How do bacterial membranes resist polymyxin antibiotics? Commun Biol 3:1–4. https://doi.org/10.1038/s42003-020-0803-x
Zavascki AP, Goldani LZ, Li J, Nation RL (2007) Polymyxin B for the treatment of multidrug-resistant pathogens: a critical review. J Antimicrob Chemother 60:1206–1215. https://doi.org/10.1093/jac/dkm357
Velkov T, Thompson PE, Nation RL, Li J (2010) Structure−activity relationships of polymyxin antibiotics. J Med Chem 53:1898–1916. https://doi.org/10.1021/jm900999h
Ortwine JK, Kaye KS, Li J, Pogue JM (2014) Colistin: understanding and applying recent pharmacokinetic advances. Pharmacotherapy: J Human Pharmacol Drug Ther 35:11–16. https://doi.org/10.1002/phar.1484
Martin NI, Hu H, Moake MM et al (2003) Isolation, structural characterization, and properties of mattacin (polymyxin M), a cyclic peptide antibiotic produced by Paenibacillus kobensis M. J Biol Chem 278:13124–13132. https://doi.org/10.1074/jbc.m212364200
Falagas ME, Rafailidis PI, Matthaiou DK (2010) Resistance to polymyxins: mechanisms, frequency and treatment options. Drug Resist Updat 13:132–138. https://doi.org/10.1016/j.drup.2010.05.002
Nikoo HR, Ardebili A, Mardaneh J (2017) Systematic review of antimicrobial resistance of clinical Acinetobacter baumannii isolates in Iran: an update. Microb Drug Resist 23:744–756. https://doi.org/10.1089/mdr.2016.0118
Velkov T, Roberts KD, Nation RL et al (2013) Pharmacology of polymyxins: new insights into an “old” class of antibiotics. Future Microbiol 8:711–724. https://doi.org/10.2217/fmb.13.39
Deris ZZ, Akter J, Sivanesan S et al (2013) A secondary mode of action of polymyxins against Gram-negative bacteria involves the inhibition of NADH-quinone oxidoreductase activity. J Antibiot 67:147–151. https://doi.org/10.1038/ja.2013.111
Maalej S, Meziou M, Mahjoubi F, Hammami A (2012) Epidemiological study of Enterobacteriaceae resistance to colistin in Sfax (Tunisia). Med Mal Infect 42:256–263. https://doi.org/10.1016/j.medmal.2012.04.008
Kaye KS, Pogue JM, Tran TB et al (2016) Agents of last resort: polymyxin resistance. Infect Dis Clin North Am 30:391–414. https://doi.org/10.1016/j.idc.2016.02.005
Rocha IV, dos Santos Silva N, das Neves Andrade CA et al (2020) Diverse and emerging molecular mechanisms award polymyxins resistance to Enterobacteriaceae clinical isolates from a tertiary hospital of Recife, Brazil. Infect Genet Evol 85:104. https://doi.org/10.1016/j.meegid.2020.104584
Mmatli M, Mbelle NM, Maningi NE, Osei Sekyere J (2020) Emerging transcriptional and genomic mechanisms mediating carbapenem and polymyxin resistance in Enterobacteriaceae: a systematic review of current reports. mSystems 5:e00783-20. https://doi.org/10.1128/mSystems.00783-20
Trimble MJ, Mlynárčik P, Kolář M, Hancock REW (2016) Polymyxin: alternative mechanisms of action and resistance. Cold Spring Harb Perspect Med 6:a025288. https://doi.org/10.1101/cshperspect.a025288
Girardello R, Gales AC (2012) Resistência às Polimixinas: velhos antibióticos, últimas opções terapêuticas. Revista de Epidemiologia e Controle de Infecção 2:66. https://doi.org/10.17058/reci.v2i2.2504
Miller AK, Brannon MK, Stevens L et al (2011) PhoQ mutations promote lipid A modification and polymyxin resistance of Pseudomonas aeruginosa found in colistin-treated cystic fibrosis patients. Antimicrob Agents Chemother 55:5761–5769. https://doi.org/10.1128/aac.05391-11
Anandan A, Evans GL, Condic-Jurkic K et al (2017) Structure of a lipid A phosphoethanolamine transferase suggests how conformational changes govern substrate binding. Proc Natl Acad Sci USA 114:2218–2223. https://doi.org/10.1073/pnas.1612927114
Yin W et al (2017) Novel plasmid-mediated colistin resistance gene mcr-3 in Escherichia coli. MBio 8(3):e00543-e617. https://doi.org/10.1128/mBio.00543-17
Carretto E et al (2018) Detection of mcr-4 positive Salmonella enterica serovar Typhimurium in clinical isolates of human origin, Italy, October to November 2016. Eurosurveillance 23(2):17–00821. https://doi.org/10.2807/1560-7917.ES.2018.23.2.17-00821
Borowiak M et al (2017) Identifcation of a novel transposon-associated phosphoethanolaminetransferase gene, mcr-5, conferring colistin resistance in d-tartrate fermenting Salmonella enterica subsp. enterica serovar Paratyphi B. J Antimicrob Chemother 72(12):3317–3324. https://doi.org/10.1093/jac/dkx327
AbuOun M et al (2017) mcr-1 and mcr-2 variant genes identifed in Moraxella species isolated from pigs in Great Britain from 2014 to 2015. J Antimicrob Chemother 72(10):2745–2749. https://doi.org/10.1093/jac/dky272
Yang Y-Q et al (2018) Novel plasmid-mediated colistin resistance gene mcr-7.1 in Klebsiella pneumoniae. J AntimicrobChemother 73(7):1791–1795. https://doi.org/10.1093/jac/dky111
Wang X et al (2018) Emergence of a novel mobile colistin resistance gene, mcr-8 NDM-producing Klebsiella pneumoniae. Emerg Microbes Infect 7(1):1–9. https://doi.org/10.1038/s41426-018-0124-z
Carroll LM et al (2019) (2019) Identifcation of novel mobilized colistin resistance gene mcr-9 in a multidrug-resistant, colistin susceptible Salmonella enterica serotype Typhimurium isolate. mBio 10(3):e00853-19. https://doi.org/10.1128/mBio.00853-19
Wang C et al (2020) Identifcation of novel mobile colistin resistance gene mcr-10. Emerg Microbes Infect 9(1):508–516. https://doi.org/10.1080/22221751.2020.1732231
Jeannot K, Bolard A, Plésiat P (2017) Resistance to polymyxins in Gram-negative organisms. Int J Antimicrob Agents 49:526–535. https://doi.org/10.1016/j.ijantimicag.2016.11.029
Di Pilato V, Arena F, Tascini C et al (2016) mcr-1.2, a new mcr variant carried on a transferable plasmid from a colistin-resistant KPC carbapenemase-producing Klebsiella pneumoniae strain of sequence type 512. Antimicrob Agents Chemother 60:5612–5615. https://doi.org/10.1128/aac.01075-16
Girardello R, Piroupo CM, Martins J et al (2021) Genomic characterization of mcr-1.1-producing Escherichia coli recovered from human infections in São Paulo Brazil. Front Microbiol 12:663. https://doi.org/10.3389/fmicb.2021.663414
Fernandes MR, Moura Q, Sartori L et al (2016) Silent dissemination of colistin-resistant Escherichia coli in South America could contribute to the global spread of the mcr-1 gene. Eurosurveillance 21:30214. https://doi.org/10.2807/1560-7917.es.2016.21.17.30214
Dalmolin TV, Martins AF, Zavascki AP et al (2018) Acquisition of the mcr-1 gene by a high-risk clone of KPC-2-producing Klebsiella pneumoniae ST437/CC258. Brazil Diagn Microbiol Infect Dis 90:132–133. https://doi.org/10.1016/j.diagmicrobio.2017.09.016
Vasconcelos PC, Leite EL, Araújo WJ et al (2020) Draft genome sequence of mcr-1-mediated colistin-resistant Escherichia coli ST359 from chicken carcasses in Northeastern Brazil. J Glob Antimicrob Resist 23:135–136. https://doi.org/10.1016/j.jgar.2020.08.016
Conceição-Neto OC, Aires CAM, Pereira NF et al (2017) Detection of the plasmid-mediated mcr-1 gene in clinical KPC-2-producing Escherichia coli isolates in Brazil. Int J Antimicrob Agents 50:282–284. https://doi.org/10.1016/j.ijantimicag.2017.05.003
Aires CAM, da Conceição-Neto OC, Tavares e Oliveira TR et al (2017) Emergence of the plasmid-mediated mcr-1 gene in clinical KPC-2-producing Klebsiella pneumoniae sequence type 392 in Brazil. Antimicrob Agents Chemother 61:e00317-17. https://doi.org/10.1128/aac.00317-17
Nitz F, de Melo BO, da Silva LCN et al (2021) Molecular detection of drug-resistance genes of blaOXA-23-blaOXA-51 and mcr-1 in clinical isolates of Pseudomonas aeruginosa. Microorganisms 9:786. https://doi.org/10.3390/microorganisms9040786
Kieffer N, Nordmann P, Moreno AM et al (2018) Genetic and functional characterization of an MCR-3-like enzyme-producing Escherichia coli isolate recovered from swine in Brazil. Antimicrob Agents Chemother 62:e00278-18. https://doi.org/10.1128/aac.00278-18
Martins-Sorenson N, Snesrud E, Xavier DE et al (2019) A novel plasmid-encoded mcr-4.3 gene in a colistin-resistant Acinetobacter baumannii clinical strain. J Antimicrob Chemother 75:60–64. https://doi.org/10.1093/jac/dkz413
Silveira LS (2019) Investigação do gene mcr-1 em isolados resistentes à colistina na região de Santa Maria-RS. Disciplinarum Scientia | Saúde 20:341–352
dos Santos LDR, Furlan JPR, Ramos MS et al (2020) Co-occurrence of mcr-1, mcr-3, mcr-7 and clinically relevant antimicrobial resistance genes in environmental and fecal samples. Arch Microbiol 202:1795–1800. https://doi.org/10.1007/s00203-020-01890-3
Leite EL, Araújo WJ, Vieira TR et al (2020) First genome of a mcr-9-mediated colistin-resistant Salmonella typhimurium from Brazilian livestock. J Glob Antimicrob Resist. 23:394–397. https://doi.org/10.1016/j.jgar.2020.09.012
Rossato L, Negrão FJ, Simionatto S (2020) Could the COVID-19 pandemic aggravate antimicrobial resistance? Am J Infect Control 48(9):1129–1130
Polly M, de Almeida BL, Lennon RP, Cortês MF, Costa SF, Guimarães T (2022) Impact of the COVID-19 pandemic on the incidence of multidrug-resistant bacterial infections in an acute care hospital in Brazil. Am J Infect Control 50(1):32–38
Grinbaum RS, Kiffer CRV (2021) Bacterial infections in COVID-19 patients: a review. Rev Assoc Med Bras 67(12):1863–1868. https://doi.org/10.1590/1806-9282.20210812
Palmeira JD, Ferreira H, Madec J-Y, Haenni M (2018) Draft genome of a ST443 mcr-1 - and bla CTX-M-2 -carrying Escherichia coli from cattle in Brazil. J Glob Antimicrob Resist 13:269–270. https://doi.org/10.1016/j.jgar.2018.05.010
Fuentes-Castillo D, Sellera FP, Goldberg DW et al (2020) Colistin-resistant Enterobacter kobei carrying mcr-9.1 and bla CTX-M-15 infecting a critically endangered franciscana dolphin (Pontoporia blainvillei ), Brazil. Transbound Emerg Dis 68:3048–3054. https://doi.org/10.1111/tbed.13980
Rau RB, de Lima-Morales D, Wink PL et al (2020) Salmonella enterica mcr-1 positive from food in Brazil: detection and characterization. Foodborne Pathog Dis 17:202–208. https://doi.org/10.1089/fpd.2019.2700
Moreno LZ, Gomes VTM, Moreira J et al (2019) First report of mcr-1-harboring Salmonella enterica serovar Schwarzengrund isolated from poultry meat in Brazil. Diagn Microbiol Infect Dis 93:376–379. https://doi.org/10.1016/j.diagmicrobio.2018.10.016
Saidenberg ABS, Stegger M, Price LB et al (2020) mcr-positive Escherichia coli ST131-H22 from poultry in Brazil. Emerg Infect Dis 26:1951–1954. https://doi.org/10.3201/eid2608.191724
Barbieri NL, Pimenta RL, de Melo DA et al (2021) mcr-1 Identified in fecal Escherichia coli and avian pathogenic E. coli (APEC) from Brazil. Front Microbiol 12:659. https://doi.org/10.3389/fmicb.2021.659613
Kobs VC, Valdez RE, de Medeiros F et al (2020) mcr-1-carrying Enterobacteriaceae isolated from companion animals in Brazil. Pesqui Vet Bras 40:690–695. https://doi.org/10.1590/1678-5150-pvb-6635
Ramos CP, Kamei CYI, Viegas FM et al (2022) Fecal shedding of multidrug resistant Escherichia coli isolates in dogs fed with raw meat-based diets in Brazil. Antibiotics 11:534. https://doi.org/10.3390/antibiotics11040534
Pontes LDS, Pimenta R, Silveira MC et al (2020) Escherichia fergusonii harboring IncHI2 plasmid containing mcr-1 gene: a novel reservoir for colistin resistance in Brazil. Microb Drug Resist 27(5):721–725. https://doi.org/10.1089/mdr.2020.0041
Sacramento AG, Fernandes MR, Sellera FP et al (2018) Genomic analysis of MCR-1 and CTX-M-8 co-producing Escherichia coli ST58 isolated from a polluted mangrove ecosystem in Brazil. J Glob Antimicrob Resist 15:288–289. https://doi.org/10.1016/j.jgar.2018.10.024
Cordeiro-Moura JR, Kraychete GB, de Araújo Longo LG et al (2022) Description and comparative genomic analysis of a mcr-1-carrying Escherichia coli ST683/CC155 recovered from touristic coastal water in Northeastern Brazil. Infect Genet Evol 97:105. https://doi.org/10.1016/j.meegid.2021.105196
Funding
The current study was supported by the Brazilian National Council for Scientific and Technological Development (CNPq) [n. 474777/2013–8] and Pro-Rectory of Research and Innovation/Federal University of Pernambuco (Propesqi/UFPE) (n. 09/2021).
Author information
Authors and Affiliations
Contributions
SDCJ contributed to writing the initial draft. SDCJ, YLAF, MAAA, and AEFA contributed to drawing the charts. IMFC and MBMO contributed to reviewing and revising the paper, and the financial support for this work.
Corresponding author
Ethics declarations
Consent to participate
All authors approved the manuscript.
Consent for publication
Written informed consent for publication was obtained from all participants.
Conflict of interest
The authors declare no competing interests.
Additional information
Responsible Editor: Roxane M Piazza
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Costa-Júnior, S.D., Ferreira, Y.L.A., Agreles, M.A.A. et al. Gram-negative bacilli carrying mcr gene in Brazil: a pathogen on the rise. Braz J Microbiol 54, 1009–1020 (2023). https://doi.org/10.1007/s42770-023-00948-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-023-00948-w