Abstract
The use of inoculants carrying diazotrophic and other plant growth–promoting bacteria plays an essential role in the Brazilian agriculture, with a growing use of microorganism-based bioproducts. However, in the last few years, some farmers have multiplied microorganisms in the farm, known as “on farm” production, including inoculants of Bradyrhizobium spp. for soybean (Glycine max L. Merrill.) and Azospirillum brasilense for corn (Zea mays L.) or co-inoculation in soybean. The objective was to assess the microbiological quality of such inoculants concerning the target microorganisms and contaminants. In the laboratory, 18 samples taken in five states were serial diluted and spread on culture media for obtaining pure and morphologically distinct colonies of bacteria, totaling 85 isolates. Molecular analysis based on partial sequencing of the 16S rRNA gene revealed 25 genera of which 44% harbor species potentially pathogenic to humans; only one of the isolates was identified as Azospirillum brasilense, whereas no isolate was identified as Bradyrhizobium. Among 34 isolates belonging to genera harboring species potentially pathogenic to humans, 12 had no resistance to antibiotics, six presented intrinsic resistance, and 18 presented non-intrinsic resistance to at least one antibiotic. One of the samples analyzed with a shotgun-based metagenomics approach to check for the microbial diversity showed several genera of microorganisms, mainly Acetobacter (~ 32% of sequences) but not the target microorganism. The samples of inoculants produced on farm were highly contaminated with non-target microorganisms, some of them carrying multiple resistances to antibiotics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Soybean (Glycine max L. Merr.) and corn (Zea mays L.) are the main Brazilian grain crops [1], with a production ~ 125 million tons in ~ 37 million hectares of soybean, and ~ 102.5 million tons in ~ 18.5 million hectares of corn [2]. The symbiosis between soybean and elite Bradyrhizobium strains can supply the most part of the required N via biological nitrogen fixation (BNF) [3] and grain yield increases by 8% due to inoculation [4]. In corn, yield increase due to inoculation with Azospirillum brasilense has been attributed to bacterial phytohormones [5, 6]. Co-inoculation of soybean with Bradyrhizobium spp. and A. brasilense has doubled the benefits compared with single inoculation [7, 8].
Brazil has a long tradition in research with inoculants containing rhizobia and Azospirillum, and legislation for quality control of inoculants. According to the standards established by the Ministry of Agriculture, Livestock and Food Supply (MAPA), commercial inoculants must have the minimal concentration of 109 viable cells of Bradyrhizobium and 108 cells of Azospirillum per gram or milliliter of inoculant, no contaminants at the 10−5 dilution, and must carry only elite strains with recognized agronomic efficiency [9, 10].
The industrial production of inoculants is a complex process, but improvements in the last two decades have resulted in high-quality products in terms of cell concentrations, no contaminants, and very low cost, probably the cheapest inoculant in the world [11]. However, in the last five years, some farmers have tried to produce their own bioproducts, including inoculants in the farm, using simplified biofactories, known as “on farm” production. In most cases, the production system is rudimentary and varies in terms of installations, equipment, microbiological standards, and technical capacity. Very often the bioproducts are produced in fermenters, open tanks, or even water tanks, without appropriate control of contaminations, which may result in highly contaminated, non-effective products [12, 13].
The objective of this study was to assess the microbiological quality of inoculants based on Bradyrhizobium spp. and A. brasilense produced on farm in Brazil, concerning the intended microorganisms, presence, and characterization of probable contaminants.
Materials and methods
Sampling
Sampling and transportation kits containing Styrofoam box, sterile 50-mL Falcon-type conical tubes, sterile 30-mL disposable syringes, disposable gloves, Parafilm M® for sealing the tubes, and cooling packs were sent to farmers interested to know the microbiological quality of their inoculants produced on farm. The kit included a protocol for sampling, emphasizing aseptic procedures and an identification form. Immediately after sampling, two aliquots per tank or fermenter were packed with cooling packs in the Styrofoam box and sent back by express postal service or personally delivered in the Laboratory for Soil Biotechnology at Embrapa Soja. A total of 18 samples were obtained during 2019/20 cropping season, six aiming Bradyrhizobium and 12 aiming Azospirillum as target microorganisms (Table 1). These samples were obtained from five states: São Paulo (six), Bahia (two), Paraná (five), Rio Grande do Sul (three), and Mato Grosso (two). For comparative purposes, commercial inoculants containing A. brasilense strains Ab-V5 and Ab-V6 (C1, lot 1,108,718), B. diazoefficiens strain SEMIA 5080 and B. japonicum strain SEMIA 5079 (C2, lot 0,135,218), and Bradyrhizobium elkanii strains SEMIA 587 and SEMIA 5019 (C3, lot 19,014,223) were included. It is worth mentioning that, although not mandatory, commercial inoculants in Brazil usually contain two bacterial strains.
Physical–chemical and organoleptic properties
The samples and the commercial inoculants were evaluated for pH using a pH-meter model FiveEasy Plus pH-meter FP20 (METTLER TOLEDO, Ohio, USA) and electrical conductivity in a digital conductivity-meter Tec-4MP (TECNAL, Piracicaba, Brazil). A sensorial analysis was based on the “odor wheel” described by McGinley and McGinley [14], which highlights eight categories of odors.
Isolation of morphotypes
Under aseptic conditions, serial dilutions were made in sterile 0.85% NaCl saline and 100-μL aliquots of the 10−5, 10−6, and 10−7 dilutions were spread on five different culture media: modified YMA (Yeast Mannitol Agar) for Bradyrhizobium [15]; RC (Rojo Congo) [16] for Azospirillum; LB (Luria Bertani) [17]; NA (Nutrient Agar) [18]; and Sabouraud [19]. The different culture media aimed to check for occurrence of typical colonies of the target microorganisms, and increase the chance of obtaining as many as possible contaminating isolates able to grow in these culture media.
After spreading on each medium, plates were incubated at 28 ± 1 °C in the inverted position in a growth room and were daily observed for 7 days. The morphologically distinct colonies in each culture medium were streaked again on the same culture medium to select single colonies. To avoid morphologically distinct isolates due to the growth medium, all isolates were streaked on NA to standardize the morphology of colonies. Finally, morphologically distinct isolates in NA medium were cryopreserved in NA broth with 30% glycerol at − 80 °C for further analysis.
Prior to cryopreservation, all isolates were observed at 400 × magnification under an optical microscope (AxioLab A1, Zeiss) coupled to an AxioCam ERc 5 s digital video camera system (Zeiss) for recognition of typical yeast traits such as nucleus, vacuole, and cell dimensions. Isolates identified as yeasts were not submitted to further analysis.
Molecular identification of isolates
Total DNA of morphologically distinct isolates was extracted with the DNeasy Blood and Tissue Kit (Qiagen), according to the manufacturer’s instructions. After extraction, the integrity of DNA was verified by electrophoresis in 1% agarose gel. The 16S rRNA gene was amplified as described [20] with universal primers fD1 (5′-AGAGTTTGATCCTGGCTCAG-3′) and rD1(5′-AAGGAGGTGATCCAGCC-3′) for phylogenetic studies of bacteria, flanking nearly the entire region of the 16S rRNA gene (~ 1,500 bp) [21]. The PCR products were purified with the PureLink™ Quick PCR Purification Kit (Invitrogen), according to the manufacturer’s instructions. Sequencing was performed in an ABI3500xL analyzer (Applied Biosystems) as described [22]. Fragment sequences ranging from 484 to 1139 bp were analyzed using the software Bionumerics version 7.6 and identification was based on comparison with the NCBI GenBank database using the BLAST tool for nucleotides (https://blast.ncbi.nlm.nih.gov/Blast.cgi).
Metagenome analysis
To have a broader view of the diversity of microorganisms that might not have grown on the culture media, or occurring at low concentrations in the sample, metagenomic analysis was performed in sample 10, from Palotina, PR. We used the shotgun approach, sequencing all DNA fragments extracted from the sample, without previous amplification of any specific region, as described before [23]. The shotgun approach detects higher diversity in a sample as well as microorganisms in all domains of life and, if required, can also be used for functional analysis. For the metagenomics analysis, total DNA was extracted with the DNeasy blood and tissue kit (Qiagen) and used to build the library with the Nextera XT kit, according to the manufacturer’s procedure. The library was processed on the MiSeq platform (Illumina) at Embrapa Soja, and the sequences were assembled with the A5-miseq pipeline (de novo) version 20,140,604. The sequenced fragments were uploaded to the MG-RAST v.4.0.4 (RAST—http://metagenomics.anl.gov) and submitted to automatic annotation in the server based on the NCBI BLAST and SEED databases [24].
Susceptibility to antimicrobials
After molecular identification, isolates belonging to potentially pathogenic genera were subjected to evaluation of susceptibility to antimicrobials by the Disk-Diffusion Test [25]. Cells grown for 24–48 h on NA medium were suspended in sterile saline (0.85% NaCl) until a turbidity compatible with the McFarland scale 0.5 (~ 1.5 × 108 CFU mL−1). The suspension was then inoculated on the Müeller-Hinton [26] agar plate using a sterile swab. Then, paper disks impregnated with antimicrobials were added, as indicated in the annual updates of the Clinical and Laboratory Standards Institute (CLSI) [27].
The antimicrobials and their concentrations per disk were as follows: amikacin 30 μg, amoxicillin + clavulanate 20/10 μg, ampicillin 10 μg, ampicillin + sulbactam 10/10 μg, aztreonam 30 μg, cefazolin 30 μg, cefepime 30 μg, cefotaxime 30 μg, cefoxitin 30 μg, ceftazidime 30 μg, ceftriaxone 30 μg, ciprofloxacin 5 μg, clindamycin 2 μg, chlorampheniol 30 μg, erythromycin 15 μg, ertapenem 10 μg, gentamicin 10 μg (120 μg for Enterococcus faecalis), imipenem 10 μg, linezolid 30 μg, levofloxacin 5 μg, meropenem 10 μg, penicillin 10 μg, piperacillin + tazobactam 100/10 μg, streptomycin 10 μg (300 μg for E. faecalis), sulbactam 10 μg, sulfamethoxazole + trimethoprim 1.25/23.75 μg, tetracycline 30 μg, and vancomycin 30 μg. The plates were incubated at 36 °C and the patterns of inhibition halos around each disk were evaluated after 18–24 h, as indicated by CLSI [27].
Results
Physical–chemical and organoleptic properties
The physical–chemical and organoleptic properties, type of equipment used for multiplication (open tanks or fermenters), and growth time (from inoculation up to sampling) of the 18 samples are shown in Table 1. The pH ranged from 3.6 (sample 6) to 7.2 (sample 7), the latter was the only with slightly alkaline pH, whereas the others were acidic, below pH 6.0. The electrical conductivity ranged from 800 (sample 5) to 8390 μS cm−1 (sample 10). Among the commercial inoculants, pH was slightly alkaline and the one containing A. brasilense presented the highest electrical conductivity. The cell concentration in the commercial inoculant C1 (A. brasilense Ab-V5 and Ab-V6) was 1.01 × 109 CFU mL−1; in C2 (Bradyrhizobium spp. SEMIA 5079 and SEMIA 5080) was 6.30 × 109 CFU mL−1; and in C3 (B. elkanii SEMIA 587 and SEMIA 5019) was 8.47 × 109 CFU mL−1. No contaminants were found in the commercial inoculants.
In the sensorial analysis [14], only two samples were classified as “yeast” (samples 13 and 14), whereas the others presented odors classified as “offensive,” which might be attributed to putrefaction processes. The commercial inoculants, however, presented odors classified as “vinegar” and “yeast” for Azospirillum and Bradyrhizobium, respectively (Table 1). Among 18 samples, three were declared as multiplied in fermenters, 13 in open tanks, and two were not informed. The growth time ranged from 4 h (sample 5) to 10 days (sample 3).
Bacterial isolation and molecular identification
The plating for isolation in culture media indicated a variety of colony morphotypes, as exemplified in Fig. 1A, suggesting occurrence of contaminants, as they differed from typical colonies of Bradyrhizobium (Fig. 1B) and Azospirillum (Fig. 1C).
A Overview of Petri’s dishes containing different culture media inoculated with samples of inoculants produced on farm in the 2019/2020 growth season aiming the multiplication of Bradyrhizobium spp. or Azospirillum brasilense. Petri’s dishes containing pure colonies of Bradyrhizobium (B) and Azospirillum (C), grown on YMA (Yeast Mannitol Agar) and RC (Rojo Congo) culture media, respectively
A total of 84 morphologically distinct isolates were obtained from the 18 samples (Table 2). Sequencing of 16S rRNA gene resulted in sequences that ranged from 484 to 1140 bp, most of them above 1000 bp. Comparisons of sequences in the GenBank showed 44 isolates with similarity ≥ 99% and 28 between 99 and 97.2% with deposited sequences, and coverage between 95 and 100%. Finally, 12 isolates were identified as yeasts based on the cell morphology (size, presence of nucleus, and budding) and were not sequenced.
Among the 84 bacterial isolates, 41 had similarity with species or genera containing at least one species reported as potentially pathogenic to humans (49%): Enterococcus (10), Acinetobacter (seven), Citrobacter (six), Klebsiella (three), Stenotrophomonas (three), Enterobacter (three), Burkholderia (two), Atlantibacter (one), Bacillus (one), Escherichia (one), Kocuria (one), Paenibacillus (one), Pseudomonas (one), and Staphylococcus (one) (Table 2).
Metagenome analysis
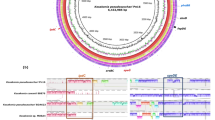
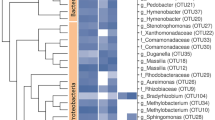
The shotgun approach of the sample no. 10 revealed a total of 2,467,209 sequences. After removal of the low-quality sequences and artificial duplicate reads, a total of 679,917,634 bp with average length of 276 bp was obtained. The rarefaction curve indicated that the number of sequences submitted was capable of detecting the existing diversity in the sample (not shown). Among the good-quality sequences, 1% contained ribosomal RNA genes, 90.68% encoded for proteins with known functions, and 8.14% proteins with unknown functions. Considering the automatic annotation in the MG-RAST v.4.0.4 server, the taxonomic classification of all shotgun sequences indicated that 99.23% belonged to the domain Bacteria, 0.2% to Eukaryota, 0.01% to Archaea, and 0.56% to Viruses (not shown). Among the 14 predominating genera identified in the sample, Acetobacter and Leuconostoc represented more than 50% of the sequences in the microbiome, whereas Azospirillum, the target microorganism in that sample, was not found (Fig. 2).
Susceptibility to antimicrobials
The test of susceptibility to antimicrobials was carried out according to [73,74,75] only with 36 isolates considered of clinical relevance. Considering the CLSI protocol, 12 isolates presented no resistance to at least one antibiotic; six presented intrinsic resistance to at least one antibiotic; and 18 isolates presented single or multiple resistance (Table 3). Noteworthy, some isolates showed multiresistance to antibiotics, e.g., isolates 1.5 and 2.4, which showed high 16S rRNA gene homology with Enterococcus faecalis, and showed resistance to all and to five tested antibiotics, respectively.
Discussion
Among 84 isolates, 25 genera were identified, 44% of which are known to harbor potential human pathogens, whereas only one isolate (5.2) showed 16S rRNA gene homology with the target microorganism A. brasilense. That was a case in which the sample was taken only 4 h after the tank had been inoculated with a commercial inoculant. Thus, the isolate probably originated from the commercial inoculant used as inoculum, not from the multiplication, since the short time between the addition of inoculum and the sampling may still have allowed the microorganism to survive. No other sample provided colonies identified as Azospirillum, showing that the target microorganism is eliminated or suppressed as the growth media become dominated by contaminating microorganisms. In addition, among the six samples aiming to multiply Bradyrhizobium, no isolate corresponded to the target bacteria.
Multiplication of microorganisms must assure several minimal microbiological procedures to guarantee that the target microorganism prevails in the culture medium. In the case of Azospirillum and mainly Bradyrhizobium, a slow-growing bacterium [15], several other microbial contaminants dominate the culture medium as they have shorter generation times, i.e., higher growth rates than the target bacteria. In many cases, the carbon source in the culture medium used for on farm production is not appropriate. For example, the use of sucrose provided as molasses for growth of Bradyrhizobium is not appropriate, as the preferred carbon sources are glycerol or mannitol [15]. Besides competition with contaminating microorganisms, the physical–chemical characteristics in the culture medium are also inappropriate for growth of the target microorganisms. For example, the adequate range of pH for Bradyrhizobium and Azospirillum is between 6.8 and 7.0 [15, 16, 76]; however, 94.4% of the samples had pH ranging from 3.6 to 5.9. The low pH can also favor the growth of contaminating microorganisms adapted to low pH and thus contributing to suppress the target microorganisms.
The lack of standardization in the incubation time is another problem in the samples taken from on farm production in this study. The average growth time of the recommended Bradyrhizobium strains to reach the ideal concentration (at least 1 × 109 cells mL−1) in the inoculant is approximately 7 days [76,77,78,79]. Similarly, A. brasilense has a growth time of about 5 days to reach at least 1 × 108 cells mL−1 [80]. In contrast, many contaminants have much shorter generation times, and dominate the culture medium in less than 24 h. Contaminating microorganisms compete for resources in the growth medium that becomes nutritionally poor and can also release inhibiting byproducts [81]. Therefore, it is reasonable to conclude that the high multiplication rates of the contaminating microorganisms, in addition to the low growth rates of the target microorganisms, result in the rapid depletion of the culture medium and enrichment with metabolites that inhibit the development of slow-growing microorganisms, like Bradyrhizobium and Azospirillum.
Multiplication of microorganisms without strict quality control can be risky to humans, animals, crops, and environment. Many contaminants are potentially pathogenic to humans and may cause various diseases, posing risks to the health of individuals who handle these products, or even final consumers if applied to products consumed in natura. Although potentially pathogenic microorganisms are found in the environment, they usually do not cause risk due to the low potential of inoculum in the environment. However, the multiplication of this microbial population in contaminated culture media could also magnify risks of infections or contaminations. For example, microorganisms from genera like Enterococcus, for which similar sequences were found in 61.1% of the samples, are frequently related to bacteremia, septicemia, urinary tract infections, abscesses, meningitis, and endocarditis [32, 82,83,84]. Some isolates also presented high genetic similarity with Citrobacter freundii [85], Enterobacter cloacae [86, 87], and Paenibacillus polymyxa [88], which are also potentially pathogenic to plants [86,87,88].
The possibility to carry genes of resistance to antimicrobials is a further concern in magnifying the population of potentially pathogenic contaminants in the on farm production. The spread of such genes in the environment may restrict the resources to fight infections. Some opportunist pathogens like Stenotrophomonas maltophilia are intrinsically resistant to several antimicrobials and collaborate to spread genes of resistance in the environment [70]. In this study, 12 isolates presented non-intrinsic resistance to antimicrobials, and 10 isolates presented resistance to two or more antimicrobials (1.1, 1.2, 1.3, 1.5, 2.2, 2.4, 3.4, 4.4, 7.1, and 18.4), what is an additional concerning issue.
Isolates identified microscopically as yeasts were not sequenced for genetic comparisons with sequences deposited in ribosomal databanks. However, some genera of yeasts can also cause injuries to humans and animals. Although yeasts are used in the manufacture of breads and beer, without any risk to humans and animals, like Saccharomyces cerevisiae, the genus Candida is the main pathogenic yeast and comprises approximately 200 species [89].
The approach based on metagenome for sample no. 10 showed that only contaminating microorganisms prevailed in the on farm sample. Although four morphologically distinct colonies were isolated from that sample based on the culture medium approach, the metagenome approach revealed more than 10 genera, including the ones isolated based on the cultivation method. This indicates that the amount of contaminating microorganisms in the on-farm multiplications can be far higher than revealed by the culture-based method. In addition, even using a more sensitive method, the target microorganism was not found in that sample.
Studies on inoculants produced on farm and their impacts on production systems and potential risks to public health are scarce. However, our findings corroborate previous studies on bioinsecticides produced on farm, which revealed low concentration or absence of the target microorganisms Bacillus thuringiensis [12], and absence of Chromobacterium subtsugae and Saccharopolyspora spinosa [13]. However, there was high prevalence of contaminants in the samples, some of them potentially pathogenic to humans [12, 13].
The negative effect of low-quality bioproducts produced on farm goes beyond the risk to Brazilian quality of agricultural products, crops, and environment, because the benefits to the crops cannot be reached with its use. The lack of effect for not containing the target microorganism might put in doubt consolidated technologies that are important to the sustainability of cropping systems like the BNF in soybean by inoculation with Bradyrhizobium [3, 4], and more recently inoculation of grasses and co-inoculation of soybean with Azospirillum [7, 8, 11].
In conclusion, the samples of inoculants produced on farm assessed in this study were highly contaminated with several non-target microorganisms, whereas the target microorganisms Azospirillum and Bradyrhizobium were not detected in the great majority of the samples. In addition, the occurrence of contaminants presenting high genetic similarity with potentially pathogenic microorganisms, some of them carrying non-intrinsic resistance or multiresistance to antimicrobials, may indicate risk to human health.
References
Cattelan AJ, Dall’Agnol A, (2018) The rapid soybean growth in Brazil. OCL 25:D102. https://doi.org/10.1051/ocl/2017058
National Supply Company – CONAB (2020) Monitoring of the Brazilian grain harvest 2019/2020. Twelfth survey, 12. https://www.conab.gov.br/info-agro/safras/graos/boletim-da-safra-de-graos
Hungria M, Franchini JC, Campo RJ, Crispino CC, Moraes JZ, Sibaldelli RNR, Mendes IC, Arihara J (2006) Nitrogen nutrition of soybean in Brazil: contributions of biological N2 fixation and N fertilizer to grain yield. Can J Plant Sci 86:927–939. https://doi.org/10.4141/P05-098
Hungria M, Mendes IC (2015) Nitrogen fixation with soybean: the perfect symbiosis? In: de Bruijn FJ (ed) Biological Nitrogen Fixation. John Wiley & Sons Inc, New Jersey, pp 1005–1019
Fukami J, Ollero FJ, Megías M, Hungria M (2017) Phytohormones and induction of plant-stress tolerance and defense genes by seed and foliar inoculation with Azospirillum brasilense cells and metabolites promote maize growth. AMB Express 7:153. https://doi.org/10.1186/s13568-017-0453-7
Masciarelli O, Urbani L, Reinoso H, Luna V (2013) Alternative mechanism for the evaluation of indole-3-acetic acid (IAA) production by Azospirillum brasilense strains and its effects on the germination and growth of maize seedlings. J Microbiol 51:590–597. https://doi.org/10.1007/s12275-013-3136-3
Hungria M, Nogueira MA, Araujo RS (2013) Co-inoculation of soybeans and common beans with rhizobia and Azospirilla: strategies to improve sustainability. Biol Fert Soils 49:791–801. https://doi.org/10.1007/s00374-012-0771-5
Hungria M, Nogueira MA, Araujo RS (2015) Soybean seed co-inoculation with Bradyrhizobium spp. and Azospirillum brasilense: a new biotechnological tool to improve yield and sustainability. Am J Plant Sci 6:811–817. https://doi.org/10.4236/ajps.2015.66087
Brasil. Ministério da Agricultura, Pecuária e Abastecimento (2011) Instrução Normativa nº. 13, de 24 de março de 2011. Available at: <http://www.agricultura.gov.br/assuntos/insumos-agropecuarios/insumos-agricolas/fertilizantes/legislacao/in-sda-13-de-24-03-2011-inoculantes.pdf>. 2011. Access on August 16, 2017.
Brasil. Ministério da Agricultura, Pecuária e Abastecimento (2010) Instrução Normativa nº. 30, de 12 de novembro de 2010. Available at: <http://www.agricultura.gov.br/assuntos/insumos-agropecuarios/insumos-agricolas/fertilizantes/legislacao/in-30-2010-dou-17-11-10-metodo-inoculantes.pdf>. 2010. Access on August 16, 2017.
Santos MS, Nogueira MA, Hungria M (2019) Microbial inoculants: reviewing the past, discussing the present and previewing an outstanding future for the use of beneficial bacteria in agriculture. AMB Expr 9:205. https://doi.org/10.1186/s13568-019-0932-0
Lana UGP, Tavares ANG, Aguiar FM, Gomes EA, Valicente FH (2019) Avaliação da qualidade de biopesticidas à base de Bacillus thuringiensis produzidos em sistema “on farm”. Boletim de Pesquisa e Desenvolvimento, Embrapa Milho e Sorgo, Sete Lagoas. ISSN: 1679–0154
Santos AFJ, Dinnas SSE, Feitoza AFA (2020) Microbiological quality of bioproducts multiplied on farm in the São Francisco valley: preliminary data. Enc Biotr 17:429–443. https://doi.org/10.18677/EnciBio_2020D33
McGinley C, McGinley M (2002) Odor testing biosolids for decision making. Water Environment Federation Specialty Conference, Austin, pp. 3–6.
Hungria M, O'Hara G, Zilli J, Araujo RS, Deaker, R, Howieson J (2016) Isolation and growth of rhizobia. In: Howieson JG, Dilworth MJ (eds.). Working with Rhizobia. Canberra: Australian Centre for International Agriculture Research (ACIAR), pp. 39–60.
Cassán F, Penna C, Creus C, Radovancich D, Monteleone E, Salamone IG, Salvo LD, Mentel I, Garcia J, Pasarello MCM, Lett L, Puente M, Correa O, Punschke Valerio K, Massa R, Rossi A, Diaz M, Catafesta M, Righes S, Carletti S, Cáceres ER (2010) Protocolo para el control de calidad de inoculantes que contienen Azospirillum sp. Documento de Procedimientos de la REDCAI número 2. Associación Argentina de Microbiología, Buenos Aires, 13 p. CD-ROM. ISBN: 978–987–98475–9–6.
Sambrook J, Fritch EF, Maniatis T (1989) Molecular cloning - A Laboratory Manual, 2nd edn. Cold Spring Harbor Laboratory Press, New York, p 1626
Association APH, - APHA, (2017) Standard methods of water analysis, 23rd edn. American Public Health Association, New York
Sabouraud R (1892). Contribution à l'Etude de la Trichophytie humaine. Etude clinique, microscopique et bactériologique sur la pluralité des trichophytons de l'homme. Ann Dermatol Syphilig, 3rd ed, pp. 1061–1087.
Menna P, Hungria M, Barcellos FG, Bangel EV, Hess PN, Martínez-Romero E (2006) Molecular phylogeny based on the 16S rRNA gene of elite rhizobial strains used in Brazilian commercial inoculants. Syst Appl Microbiol 29:315–332. https://doi.org/10.1016/j.syapm.2005.12.002
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703. https://doi.org/10.1128/jb.173.2.697-703
Delamuta JRM, Ribeiro RA, Menna P, Hungria M (2017) Phylogenies of symbiotic genes of Bradyrhizobium symbionts of legumes of economic and environmental importance in Brazil support the definition of the new symbiovars pachyrhizi and sojae. Syst Appl Microbiol 40:254–265. https://doi.org/10.1016/j.syapm.2017.04.005
Souza RC, Mendes IC, Reis-Junior FB, Carvalho FM, Vasconcelos ATR, Vicente VA (2016) Hungria M (2016) Shifts in taxonomic and functional microbial diversity with agriculture: how fragile is the Brazilian Cerrado? BMC Microbiol 16:42. https://doi.org/10.1186/s12866-016-0657-z
Meyer F, Paarmann D, D’Souza M, Olson R, Glass EM, Kubal M, Paczian T, Rodriguez A, Stevens R, Wilke A, Wilkening J, Edwards RA (2008) The metagenomics RAST server – a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinformatics 9:386. https://doi.org/10.1186/1471-2105-9-386
Bauer AW, Kirby WMM, Sherris JC, Turck M (1966) Antibiotic susceptibility testing by a standardized single disk method. Am J Clin Pathol 36:493–496. https://doi.org/10.1093/ajcp/45.4_ts.493
Mueller JH, Hinton J (1941) A protein-free medium for primary isolation of the Gonococcus and Meningococcus. Exp Biol Med 48:330–333. https://doi.org/10.3181/00379727-48-13311
Clinical and Laboratory Standards Institute – CLSI (2018) Performance standards for antimicrobial susceptibility testing. 28th ed. CLSI supplement M100, Clinical and Laboratory Standards Institute, Wayne, PA, 257p. ISBN: 1–56238–839–8
Hirai J, Uechi K, Hagihara M, Sakanashi D, Kinjo T, Haranaga S, Fujita J (2016) Bacteremia due to Citrobacter braakii: a case report and literature review. J Infect Chemother 12:819–821. https://doi.org/10.1016/j.jiac.2016.07.003
Pati NB, Doijad SP, Schultze T, Mannala GK, Yao Y, Jaiswal S, Ryan D, Suar M, Gwozdzinski K, Bunk B, Mrahei MA, Marahiel MA, Hegemann JD, Spröer C, Goesmann A, Falgenhauer L, Hain T, Imirzalioglu C, Mshana SE, Overmann O, Chakraborty T (2018) Enterobacter bugandensis: a novel enterobacterial species associated with severe clinical infection. Sci Rep 8:5392. https://doi.org/10.1038/s41598-018-23069-z
McConnell MJ, Actis L, Pachón J (2013) Acinetobacter baumannii: human infections, factors contributing to pathogenesis and animal models. FEMS Microbiol Rev 37:130–155. https://doi.org/10.1111/j.1574-6976.2012.00344.x
Her J, Kim J (2013) Rummeliibacillus suwonensis sp. nov., isolated from soil collected in a mountain area of South Korea. J Microbiol 51:268–272. https://doi.org/10.1007/s12275-013-3126-5
Poulsen LL, Bisgaard M, Son NT, Trung NV, An HM, Dalsgaard A (2012) Enterococcus faecalis clones in poultry and in humans with urinary tract infections. Vietnam Emerg Infect Dis 18:1096–1100. https://doi.org/10.3201/eid1807.111754
Faccin DJL, Rech R, Secchi AR, Cardozo NSM, Ayub MAZ (2013) Influence of oxygen transfer rate on the accumulation of poly (3-hydroxybutyrate) by Bacillus megaterium. Process Biochem 48:420–425. https://doi.org/10.1016/j.procbio.2013.02.004
Brenner DJ, O’hara CM, Grimont PD, Janda JM, Falsen E, Aldova E, Ageron E, Schindler J, Abbott SL, Steigerwalt AG (1999) Biochemical identification of Citrobacter species defined by DNA hybridization and description of Citrobacter gillenii sp. nov. (formerly Citrobacter genomospecies 10) and Citrobacter murliniae sp. nov. (formerly Citrobacter genomospecies 11). J Clin Microbiol 37:2619–2624. https://doi.org/10.1128/JCM.37.8.2619-2624.1999
Hasan S, Sultana M, Hossain MA (2019) Complete genome arrangement revealed the emergence of a poultry origin superbug Citrobacter portucalensis strain NR-12. J Glob Antimicrob Resist 18:126–129. https://doi.org/10.1016/j.jgar.2019.05.031
Forson AO, Tsidi WB, Nana-Adjei D, Quarchie MN, Obeng-Nkrumah N (2018) Escherichia coli bacteriuria in pregnant women in Ghana: antibiotic resistance patterns and virulence factors. BMC Res Notes 11:901. https://doi.org/10.1186/s13104-018-3989-y
Guerra PV (2018) Evaluation of the immunomodulatory potential of Hsp65-producing Lactococcus lactis in Cutaneous Leishmaniasis caused by Leishmania braziliensis. 101 f. Thesis (PhD in Pathology) - Gonçalo Moniz Institute, Oswaldo Cruz Foundation, Federal University of Bahia.
Dworkin M, Falkow S, Rosenberg E, Schleifer KH, Stackebrandt E (2006) The Prokaryotes: Bacteria: Firmicutes, Cyanobacteria. Springer, New York.
Aghazadeh Z, Pouralibaba F, Yari Khosroushahi A (2017) The prophylactic effect of Acetobacter syzygii probiotic species against squamous cell carcinoma. J Dent Res Dent Clin Dent Prospects 11:208–214. https://doi.org/10.15171/joddd.2017.037
Endo A, Okada S (2007) Lactobacillus farraginis sp. nov. and Lactobacillus parafarraginis sp. nov., heterofermentative lactobacilli isolated from a compost of distilled shochu residue. Int J Syst Evol Microbiol 57:708–712. https://doi.org/10.1099/ijs.0.64618-0
Jung YO, Jeong H, Cho Y, Lee EO, Jang HW, Kim J, Nam KT, Lim KM (2019) Lysates of a probiotic, Lactobacillus rhamnosus, can improve skin barrier function in a reconstructed human epidermis model. Int J Mol Sci 20:4289. https://doi.org/10.3390/ijms20174289
Camargo CH, Bruder-Nascimento A, In Lee SH, Fernandes Júnior A, Kaneno R, Rall VLM (2014) Prevalence and phenotypic characterization of Enterococcus spp. isolated from food in Brazil. Braz J Microbiol 45:111–115. https://doi.org/10.1590/S1517-83822014000100016
Knight DB, Rudin SD, Bonomo RA, Rather PN (2018) Acinetobacter nosocomialis: defining the role of efflux pumps in resistance to antimicrobial therapy, surface motility, and biofilm formation. Front Microbiol 9:1902. https://doi.org/10.3389/fmicb.2018.01902
Moyad MA (2018) Brewer’s/baker’s yeast (Saccharomyces cerevisiae) and preventive medicine: Part II. Urol Nurs 28:73–75 (PMID: 18335702)
Hafed L, Farag H, El-Rouby D, Shaker O, Shabaan H-A (2019) Candida albicans alcohol dehydrogenase 1 gene in oral dysplasia and oral squamous cell carcinoma. Pol J Pathol 70:210–216. https://doi.org/10.5114/pjp.2019.90398
Kus JV, Burrows LL (2016) Infections due to Citrobacter and Enterobacter. In: Enna SJ, Bylund DB (eds) xPharm: The Comprehensive Pharmacology Reference. Elsevier. https://doi.org/10.1016/B978-008055232-3.60868-2
Ramírez-Quintero JD, Chavarriaga-Restrepo A (2017) Bacteriemia por Raoultella planticola de origen gastrointestinal. Iatreia 30:67–71. https://doi.org/10.17533/udea.iatreia.v30n1a06
Kasper DL, Fauci AS, Hauser SL, Longo DL, Jameson L, Loscalzo J (2017) Manual de Medicina de Harrison. AMGH, Porto Alegre. ISBN: 978–85–8055–582–0
Chagas TPG (2015) Characterization of Acinetobacter spp. multiresistant producers of carbapenemases, types OXA and NDM, isolated from different regions of Brazil. Thesis (PhD in Sciences) Oswaldo Cruz Institute, Rio de Janeiro.
Santini JMK, Buzetti S, Teixeira Filho MCM, Galindo FS, Coaguila DN, Boleta EHM (2018) Doses and forms of Azospirillum brasilense inoculation on maize crop. Rev Bras Eng Agríc Ambient 22:373–377. https://doi.org/10.1590/1807-1929/agriambi.v22n6p373-377
Selvakumar G, Kundu S, Joshi P, Nazim S, Gupta AD, Gupta HS (2010) Growth promotion of wheat seedlings by Exiguobacterium acetylicum 1P (MTCC 8707) a cold tolerant bacterial strain from the Uttarakhand Himalayas. Indian J Microbiol 50:50–56. https://doi.org/10.1007/s12088-009-0024-y
Van Dijl JM, Hecker M (2013) Bacillus subtilis: from soil bacterium to super-secreting cell factory. Microb Cell Fact 12:3. https://doi.org/10.1186/1475-2859-12-3
Dai W, Zhu Y, Wang X, Sakenova N, Yang Z, Wang H, Li G, He J, Huang D, Cai Y, Guo W, Wang Q, Feng T, Fan Q, Zheng T, Han A (2016) Draft genome sequence of the bacterium Comamonas aquatica CJG. Genome Announc 4:6. https://doi.org/10.1128/genomeA.01186-16
Boszczowski I, Salomão MC, Moura ML, Freire MP, Guimarães T, Cury AP, Rossi F, Rizek CF, Martins RCR, Costa SF (2019) Multidrug-resistant Klebsiella pneumoniae: genetic diversity, mechanisms of resistance to polymyxins and clinical outcomes in a tertiary teaching hospital in Brazil. Rev Inst Med Trop 61:29. https://doi.org/10.1590/s1678-9946201961029
Ioannou P (2019) Escherichia hermannii infections in humans: a systematic review. Trop Med Infect Dis 4:17. https://doi.org/10.3390/tropicalmed4010017
Delgado S, Leite AMO, Ruas-Madiedo P, Mayo B (2015) Probiotic and technological properties of Lactobacillus spp. strains from the human stomach in the search for potential candidates against gastric microbial dysbiosis. Front Microbiol 5: 766. https://doi.org/10.3389/fmicb.2014.00766
Power RF, Linnane B, Martin R, Power N, Harnett P, Casserly B, O’connell NH, Dunne CP (2016) The first reported case of Burkholderia contaminans in patients with cystic fibrosis in Ireland: from the Sargasso Sea to Irish Children. BMC Pulm Med 16:57. https://doi.org/10.1186/s12890-016-0219-z
Kommanee J, Tanasupawat S, Yukphan P, Thongchul N, Moonmangmee D, Yamada Y (2012) Identification of Acetobacter strains isolated in Thailand based on the phenotypic, chemotaxonomic, and molecular characterization. Sci Asia 38:44–53. https://doi.org/10.2306/scienceasia1513-1874.2012.38.044
Kandi V, Palange P, Vaish R, Bhatti AB, Kale V, Kandi MR, Bhoomagiri MR (2016) Emerging bacterial infection: identification and clinical significance of Kocuria species. Cureus 8:e731. https://doi.org/10.7759/cureus.731
Krishnamurthi S, Chakrabarti TProposal for transfer of Pelagibacillus goriensis Kim, et al (2008) 2007 to the genus Terribacillus as Terribacillus goriensis comb. nov. Int J Syst Evol Microbiol 58:2287–2291. https://doi.org/10.1099/ijs.0.65579-0
Tuazon CU, Murray HW, Levy C, Solny MN, Curtin JA, Sheagren JN (1979) Serious infections from Bacillus sp. JAMA 241:1137–1140. https://doi.org/10.1001/jama.1979.03290370041026
Amin M, Rakhisi Z, Ahmady AZ (2015) Isolation and identification of Bacillus species from soil and evaluation of their antibacterial properties. Avicenna J Clin Microb Infect 2: 23233. https://doi.org/10.17795/ajcmi-23233
Sáez-Nieto JA, Medina-Pascual MJ, Carrasco G, Garrido N, Fernandez-Torres MA, Villalón P, Valdezate S (2017) Paenibacillus spp. isolated from human and environmental samples in Spain: detection of 11 new species. New Microbes New Infect 19:19–27. https://doi.org/10.1016/j.nmni.2017.05.006
Bourafa N, Loucif L, Boutefnouchet N, Rolain JM (2015) Enterococcus hirae, an unusual pathogen in humans causing urinary tract infection in a patient with benign prostatic hyperplasia: first case report in Algeria. New Microbes New Infect 8:7–9. https://doi.org/10.1016/j.nmni.2015.08.003
Ieranò T, Silipo A, Sturiale L, Garozzo D, Bryant C, Lanzetta R, Parrilli M, Aldridge C, Gould FK, Corris PA, Khan CMA, De Soyza A, Molinaro A (2009) First structural characterization of Burkholderia vietnamiensis lipooligosaccharide from cystic fibrosis-associated lung transplantation strains. Glycobiology 19:1214–1223. https://doi.org/10.1093/glycob/cwp112
Cañete-Rodríguez AM, Santos-Dueñas IM, Torija-Martínez MJ, Mas A, Jiménez-Hornero JE, García-García I (2016) An approach for estimating the maximum specific growth rate of Gluconobacter japonicus in strawberry purée without cell concentration data. Biochem Eng J 105:314–320. https://doi.org/10.1016/j.bej.2015.10.005
Libonatti C, Agüeria D, García C, Basualdo M (2018) Encapsulation and its application in the use of fish waste. Rev Argent Microbiol 51:81–83. https://doi.org/10.1016/j.ram.2018.03.001
Nguyen TH, Park MD, Otto M (2017) Host response to Staphylococcus epidermidis colonization and infections. Front Cell Infect Microbiol 7:90. https://doi.org/10.3389/fcimb.2017.00090
Morello E, Pérez-Berezo T, Boisseau C, Baranek T, Guillon A, Bréa D, Lanotte P, Carpena X, Pietrancosta N, Hervé V, Ramphal R, Cenac N, Si-Tahar M (2019) Pseudomonas aeruginosa lipoxygenase LoxA contributes to lung infection by altering the host immune lipid signaling. Front Microbiol 10:1826. https://doi.org/10.3389/fmicb.2019.01826
Almeida MTG, Bertelli ECP, Rossit ARB, Bertollo EMG, Martinez M (2005) Infecções hospitalares por Stenotrophomonas maltophilia: aspectos clínico-epidemiológicos, microbiológicos e de resistência antimicrobiana. Arq Ciênc Saúde 12:141–145
Hungria M, Ribeiro RA, Nogueira MA (2018) Draft genome sequences of Azospirillum brasilense strains Ab-V5 and Ab-V6, commercially used in inoculants for grasses and legumes in Brazil. Genome Announc 6:e00393-e418. https://doi.org/10.1128/genomeA.00393-18
Ghanbarinia F, Kheirbadi M, Mollania N (2015) Comamonas sp. halotolerant bacterium from industrial zone of Jovein of Sabzevar introduced as good candidate to remove industrial pollution. Iran J Microbiol 7:273–280
Moellering RC, Graybill JR, Mcgowan JE, Corey L (2007) Antimicrobial resistance prevention initiative-an update: Proceedings of an expert panel on resistance. Am J Infect Control 35(1):17. https://doi.org/10.1016/j.amjmed.2007.04.001
Rice LB (2008) Editorial commentary. Federal funding for the study of antimicrobial resistance in nosocomial pathogens: No ESKAPE. J Infect Dis 197:1079–1081. https://doi.org/10.1086/533452
Rice LB (2010) Progress and challenges in implementing the research on ESKAPE Pathogens. Infect Control Hosp Epidemiol 31:7–64. https://doi.org/10.1086/655995
Vincent JM (1970) A manual for the pratical study of rooot-nodule bacteria. Oxford: Blackwell Scientific, 164p. (International Biological Programme Handbook, 15).
Jordan DC (1982) Transfer of Rhizobium japonicum Buchanan 1980 to Bradyrhizobium sp. nov., a genus of slow-growing, root nodule bacteria from leguminous plants. Int J Syst Bacteriol 32:136–139. https://doi.org/10.1099/00207713-32-1-136
Kuykendall LD, Saxena B, Devine TE, Udell SE (1992) Genetic diversity in Bradyrhizobium japonicum Jordan 1982 and a proposal for Bradyrhizobium elkanii sp. nov. Can J Microbiol 38:501–505. https://doi.org/10.1139/m92-082
Delamuta JRM, Ribeiro RA, Ormenõ-Orrillo E, Melo IS, Martinéz-Romero E, Hungria M (2013) Polyphasic evidence supporting the reclassification of Bradyrhizobium japonicum group Ia strains as Bradyrhizobium diazoefficiens sp. nov. Int J Syst Evol Microbiol 63:3342–3351. https://doi.org/10.1099/ijs.0.049130-0
Döbereiner J (1991) The genera Azospirillum and Herbaspirillum. In: Ballows A, Trüper HG, Dworkin M, Harder W, Shleifer K. (eds) The Prokaryotes, Springer-Verlag, New York, pp. 2236–2253. https://doi.org/10.1007/978-1-4757-2191-1
Hibbing ME, Fuqua C, Parsek MR, Peterson SB (2010) Bacterial competition: surviving and thriving in the microbial jungle. Nat Rev Microbiol 8:15–25. https://doi.org/10.1038/nrmicro2259
Cauwerts K, Decostere A, De Graef EM, Haesebrouck F, Pasmans F (2007) High prevalence of tetracycline resistance in Enterococcus isolates from broilers carrying the erm (B) gene. Avian Pathol 36:395–399. https://doi.org/10.1080/03079450701589167
Kense MJ, Landman WJM (2011) Enterococcus cecorum infections in broiler breeders and their offspring: molecular epidemiology. Avian Pathol 40:603–612. https://doi.org/10.1080/03079457.2011.619165
Zou LK, Wang HN, Zeng B, Li JN, Li XT, Zhang AY, Zhou YS, Yang X, Xu CW, Xia QQ (2011) Erythromycin resistance and virulence genes in Enterococcus faecalis from swine in China. New Microbiol, 34: 73–80. https://pubmed.ncbi.nlm.nih.gov/21344149
Allahverdi T, Rahimian H, Ravanlou A (2016) First report of bacterial canker in mulberry caused by Citrobacter freundii in Iran. Plant Dis 100:1774. https://doi.org/10.1094/PDIS-01-16-0020-PDN
García-Gonzales T, Sáenz-Hidalgo HK, Silva-Rojas HV, Morales-Nieto C, Vancheva T, Koebnik R, Ávila-Quezada GD (2018) Enterobacter cloacae, an emerging plant-pathogenic bacterium affecting chili pepper seedlings. Plant Pathol J 34:1–10. https://doi.org/10.5423/PPJ.OA.06.2017.0128
Schroeder BK, Du Toit LJ, Schwartz HF (2009) First report of Enterobacter cloacae causing onion bulb rot in the Columbia basin of Washington State. Plant Dis 93:323. https://doi.org/10.1094/PDIS-93-3-0323A
Zhang RY, Zhao SX, Tan ZQ, Zhu CH (2017) First report of bacterial stem rot disease caused by Paenibacillus polymyxa on Hylocereus undulatus in China. Plant Dis 101:1031–1031. https://doi.org/10.1094/PDIS-11-16-1577-PDN
Almirante B, Rodríguez D, Park BJ, Cuenca-Estrella M, Planes AM, Almela M, Mensa J, Sanchez F, Ayats J, Gimenez M, Saballs P, Fridkin SK, Morgan J, Rodriguez-Tudela JL, Warnock DW, Pahissa A (2005) Epidemiology and predictors of mortality in cases of Candida bloodstream infection: results from population-based surveillance, Barcelona, Spain, from 2002 to 2003. J Clin Microbiol 43:1829–1835. https://doi.org/10.1128/JCM.43.4.1829-1835.2005
Acknowledgements
The authors acknowledge the support by the INCT Plant Growth-Promoting Microorganisms for Agricultural Sustainability and Environmental Responsibility (CNPq 465133/2014-2, Fundação Aaucária-STI 043/2019, CAPES) and CNPq 433656/2018-2 (MCTIC/CNPq 28/2018). M.A. Nogueira and M. Hungria are CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico) research fellows. This paper was approved for publication by the Editorial Board of Embrapa Soja as manuscript number 219/2020.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Responsible Editor: Luc F.M. Rouws
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bocatti, C.R., Ferreira, E., Ribeiro, R.A. et al. Microbiological quality analysis of inoculants based on Bradyrhizobium spp. and Azospirillum brasilense produced “on farm” reveals high contamination with non-target microorganisms. Braz J Microbiol 53, 267–280 (2022). https://doi.org/10.1007/s42770-021-00649-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-021-00649-2