Abstract
This study aimed to estimate human age and gender from panoramic radiographs using various deep learning techniques while using explainability to have a novel hybrid unsupervised model explain the decision-making process. The classification task involved training neural networks and vision transformers on 706 panoramic radiographs using different loss functions and backbone architectures namely ArcFace, a triplet network named TriplePENViT, and the subsequently developed model called PENViT. Pseudo labeling techniques were applied to train the models using unlabeled data. FullGrad Explainable AI was used to gain insights into the decision-making process of the developed PENViT model. The ViT Large 32 model achieved a validation accuracy of 68.21% without ArcFace, demonstrating its effectiveness in the classification task. The PENViT model outperformed other backbones, achieving the same validation accuracy without ArcFace and an improved accuracy of 70.54% with ArcFace. The TriplePENViT model achieved a validation accuracy of 67.44% using hard triplet mining techniques. Pseudo labeling techniques yielded poor performance, with a validation accuracy of 64.34%. Validation accuracy without ArcFace was established at 67.44% for Age and 84.49% for gender. The unsupervised model considered developing tooth buds, tooth proximity and mandibular shape for estimating age within deciduous and mixed dentitions. For ages 20–29, it factored permanent dentition, alveolar bone density, root apices, and third molars. Above 30, it notes occlusal deformity resulting from missing dentition and the temporomandibular joint complex as predictors for age estimation from panoramic radiographs.
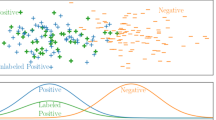
Graphical abstract

Article highlights
-
Development of a novel hybrid model to estimate human age and sex from labeled and unlabeled orthopantomograms.
-
Evaluation of regression task, pseudo-labelling, and Triplet networks for age and sex estimation.
-
Successful application of explainable AI to identify the anatomy responsible for shaping estimation accuracy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Non-invasive age and gender estimation from radiographs has notable roles in dental diagnostics and forensic investigations [1]. Applications outside of the dental practice range from estimate the age of the corpse or to help in determining the identities of the deceased who succumb to calamities such as explosions in law enforcement and judicial trials evaluating the truthfulness of an individual’s age at specified times of interest or in the cases of undocumented children [2, 3]. To provide scientifically backed evidence on age and biological sex, forensic dentistry determines the age of an individual through estimating the stage of development of a tooth and maxillofacial arches. The development of a tooth occurs through several stages, starting from the formation of the tooth bud in the embryonic stage to the eruption and maturation of the tooth in the oral cavity. Panoramic radiographs, also known as orthopantomogram (OPG) or panoramic imaging, is a specialized, easily accessible, and cost-effective dental imaging technique that captures a wide-angle view of the entire oral and maxillofacial region in a single image. It provides a comprehensive 2-dimensional overview of the dentition, maxillofacial and mandibular bone anatomy, temporomandibular joints (TMJs), sinuses, and other surrounding structures and have been used previously to report on root-canal treatment progression [4].
While advanced 3-dimensional techniques such as computed tomography (CT) [5, 6] and cone-beam computed tomography (CBCT) [7, 8] have gained recent popularity, panoramic radiographs still remain the most commonly used technique in both dental diagnostics and deep learning applications related to dentistry [9,10,11,12]. The innovation of high-resolution biosensors and subsequent imaging processes in recent times have produced large quantities of data that can be examined with the help of computer programs. OPGs are considered to contain most two-dimensional landmark information used to reach a preliminary diagnosis and are usually the first step in determining whether three-dimensional computed tomography is required. Traditional automations of dental age estimation involve phases, such as image-preprocessing, segmentation, feature extraction, and classification (categorical) or regression (numerical). In the case of classification, these processes aim to identify age categories for people, whereas the regression phase aims to identify their exact ages.
Deep Learning (DL) methods have been applied in recent years to automate activities utilizing OPG images. DL techniques, most notably convolutional neural networks (CNNs), have shown promise in various applications involving digital images of panoramic radiographs that are able to extract and segment features within the maxilla and mandible to isolate each tooth from other objects in the image such as jaws[13]; the detection and classification of individual teeth involve the identification and labeling of each tooth within a dental image [14,15,16]; the detection of previous treatment, e.g., endodontics [17]; the reconstruction of OPG images where a patient was badly positioned [18]; the diagnosis of osteoporosis [19] and jaw tumors [20].
While there has been extensive exploration of supervised CNN-based age estimation in previous literature, the integration of unsupervised learning and explainability is a relatively nascent area in terms of both design and approach [21]. The application of unsupervised learning to radiographic assessment to reduce operator-related variability is of particular interest. In this context, our research introduces a novel unsupervised deep learning approach, termed PENViT, which combines EfficientNet and Vision Transformer (ViT) models with Addictive Angular Margin Loss (ArcFace). This amalgamation of different deep learning models and loss functions aims to elevate the accuracy and resilience of dental age estimation. The primary objective of the present study was to explore existing methods and devise novel strategies for advancing automated age prediction using weak and minimal supervision. To this end, the study posed the following research questions:
-
a.
Which model architecture can correctly estimate age and biological gender using regression-based neural network?
-
b.
Does margin losses (ArcFace, TripletMarginLoss) increase performance of OPG-based age classification as compared to pure cross entropy loss?
-
c.
Does Hard Triplet Mining task improve validation accuracy of Triplet network?
-
d.
Can the novel PENViT model backbone perform on par in both general form and triplet-like network (TriplePENViT) as compared with any other backbone?
-
e.
Does a two-step semi-supervised pseudo labelling workflow improve validation accuracy of age estimation?
-
f.
Can AI interpretation produce medically sound regions of explainability on radiographs for predicting age?
2 Related literature
The anatomical form of the maxilla and mandible along with the alveolar bone region development has most correlation with the individual’s chronological age [22]. When classifying jaw development, striking age-related features include the development of deciduous dentition, followed by each permanent tooth and finally the root completion of third molars [22]. The current study aimed to implement a series of methodologies from previous literature and generate a hybrid model that can be used to identify biological gender and estimate age.
2.1 Regression tasks
Age estimation from orthopantomograms or panoramic radiographs constitutes an application that leverages regression models [2, 3, 23]. The primary objective revolves around gauging an individual’s age, drawing from diverse variables encompassing mandible development, tooth germs, and areas of missing space within the dental arch [3, 23]. Previous inquiries have adopted Mean Absolute Error (MAE) metrics to delineate the efficacy of the regression model [10, 21]. In a manner akin to the present exposition (with results expounded in a subsequent segment), Fan et al. [21] similarly identified a diminished MAE vis-à-vis alternative CNN-only architectures, all pertaining to regression tasks. Demonstrating an automated methodology, Atas et al. modified the InceptionV3 framework to yield an innovative neural network model [10].
A more accurate and relatively faster dental age estimation stemmed from curtailing the array of attributes inherent in the devised model structure. He et al. introduced profound relation learning for regression, aiming to unearth diverse correlations within pairs of input images [24]. In parallel, Fan et al. aspired to formulate a hybrid deep neural network, termed DASE-net, amalgamating Transformer and CNN components. This novel architecture aimed at age prediction via dental x-rays, juxtaposing its performance against CNNs and manual techniques executed by forensic dentistry experts [21]. A contemporaneous study also evaluated gender using dental x-rays, employing DenseNet Architecture alongside comparative models [25]. The authors experimented with four distinct deep learning network structures: VGG, ResNet, EfficientNet, and DenseNet. Out of these, the proposed DenseNet121 model, endowed with fewer parameters, manifested superior outcomes compared to its more parameter-laden counterparts.
An independent exploration introduced the lightweight SFCN model, capable of accurate age prediction through a solitary fully connected layer, thus minimizing parameter count, in contradistinction to multi-layer counterparts [26]. After contrasting SFCN’s performance against that of ResNet18, ResNet50, ResNet101, and ResNet152, it was deduced that deeper models did not inherently outperform shallower counterparts in predicting brain age. Among the gamut of tested architectures, SFCN emerged as the pinnacle performer. Alternatively, prior literature also documented Bayesian convolutional neural networks as a possible approach to estimate age uncertainty [27].
2.2 Classification tasks
Age estimation can alternatively be tackled as a classification task, wherein the objective is to categorize individuals into predetermined age groups or classes. The classes employed for classification in this research are indicated in Table 1, and they were modified based on prevalent patterns in dentistry but in a simplified manner to ensure the limited dataset can generate meaningful and reliable data [28, 29].
An automated approach for determining individuals’ age groups was presented by a group of researchers, employing transfer learning techniques on two convolutional deep neural networks: AlexNet and ResNet-101 [2, 3, 11, 23]. The classification process involved utilizing decision tree (DT), k-nearest neighbor (K-NN), linear discriminant (LD), and support vector machine (SVM) methods. Another study by Vila-Blanco et al. introduced two fully automatic methods for estimating chronological age [12]. The first approach, named DANet, employed a sequential Convolutional Neural Network (CNN) for age estimation. The second approach, known as DASNet, extended this by incorporating a second CNN path to predict gender and leveraging gender-specific features to enhance age estimation performance. Comparative results indicated the superior performance of DASNet over DANet.
In a different context, Almalki et al. delved into object detection using the YOLOv3 deep learning model, creating an automated tool to diagnose and classify dental abnormalities from panoramic dental radiographs. Meanwhile, Farhadian et al. employed the pulp-to-tooth ratio for age estimation [30]. Recent literature also introduced saliency map-enhanced age estimation techniques, capable of automatically estimating age based on lateral cephalometric images [31]. To identify the most suitable convolutional neural network model for automated age estimation, Milosevic et al. employed pre-trained parameters from general-purpose vision models [32]. Through ablation experiments, the authors identified the key anatomical areas within the dental system that significantly contributed to the age estimation process.
2.3 Pseudo labeling
Pseudo-labeling is a semi-supervised learning (SSL) technique that involves using the estimations of a trained model on unlabeled data to generate pseudo-labels, which are then used to augment the labeled dataset and train the model further. Pseudo-labeling can be a useful approach when labeled data is limited but unlabeled data is abundant. Fengbei Liu et al. in 2022 proposed a new and effective semi-supervised learning (SSL) algorithm in medical image analysis (MIA), called anti-curriculum pseudo-labelling (ACPL), which introduced novel selection and balancing techniques of unlabelled samples, that in turn facilitated the model to work with both multi-label and multi-class problems while allowing for the estimation of pseudo labels using ensemble classifiers [33].
More recently, in 2023, the Bayesian Pseudo Labels were used by the Xu et al. to illustrate the entire generalization of pseudo labels under the Bayes principle [34]. Then, by learning a threshold to choose high-quality pseudo labels, they offer a variational technique to learning to approximate Bayesian pseudo labels. A connection was built between pseudo labeling and the Expectation Maximization algorithm which partially explains its empirical successes. Rhee and Cho, through their research, offered a new confidence-based weighting technique for obtaining pseudo-labels with varied contributions based on the confidence in addition to an adaptive threshold adjustment strategy to supply enough and precise pseudo-labels throughout the training [35]. The ambiguity of pseudo-labels for perplexing samples in SSL was then drastically reduced by the unique pseudo-labeling schemes suggested later by Ham et al.[36] The investigators used the Easy-to-Forget (ETF) Sample Finder to execute our approach, Pruning for Pseudo-Label (P-PseudoLabel), comparing outputs of the model versus the trimmed model to find samples that are perplexing. Then, utilizing the perplexing samples, they execute negative learning to reduce the likelihood of giving inaccurate information and to enhance performance.
2.4 Siamese or triplet networks
An age estimation method was proposed by Zhang and Kurita from periods of age using Triplet Network [37]. The proposed model extrapolates the age values from that age period based on similarities across age periods. Triplet Network was utilized to record the age connection between the facial photos in order to achieve this functionality. Next, linear regression is used to estimate each image’s age. Hajamohideen et al. more recently proposed a Siamese Convolutional Neural Network (SCNN) architecture employing the triplet-loss function to represent MRI image inputs as k-dimensional embeddings [38]. To convert images into the embedding space, they employed CNNs that had been trained and those that had not. Afterwards, the 4-way classification of Alzheimer’s disease utilized similar embedding techniques. In Jeong et al.’s work, the investigators trained a convolutional neural network (CNN) model using the deep metric learning method based on a binary classifier Siamese network for class clustering operations [39].
2.5 Explainable AI
Selvaraju et al. presented a method for explaining decision to visual outputs made by a wide range of Convolutional Neural Network (CNN)-based models, which improved their transparency. This method, called Gradient-weighted Class Activation Mapping (Grad-CAM), identifies the most important areas in an image that are relevant to the estimation of a target concept by analyzing the gradients of the concept that flow into the final convolutional layer. Chattopadhay et al.[40] built upon the work of Selvaraju et al. by introducing Grad-CAM++, a refined version of Grad-CAM that provides visual explanations of CNN model estimates object localization and attempts to explain the presence of multiple instances of a class in a single image. Later, Omeiza et al. improved upon this by combining SMOOTH GRAD and Grad-CAM++ to present Smooth Grad-CAM+ +; The result was a model that could explain visual sharpness, object localization, and was adept at explaining multiple occurrences of objects in a single image [41]. Recently, gradient-based visualization techniques have been subject to criticism in academia, and there is ongoing debate about their effectiveness. Ramaswamy proposed a new methodology for generating visual explanations for deep Convolutional Neural Networks (CNN) using Ablation-based Class Activation Mapping (Ablation CAM). This approach applies ablation analysis to determine the importance of individual feature map units with respect to a particular class [42]. The authors later used ablation analysis to visualize the major components of learned representations from convolutional layers, and their innovative Eigen-CAM technique was used to improve explanations of CNN estimates without the need for accurate model classification [42].
Wang et al. [43] propose developing Bayesian deep learning techniques that are both explicable and implementable to quantify uncertainties precisely and identify the causes and potential solutions for reducing their impact. While FullGrad has recently gained attention for its model interpretability capabilities [44], if the highlighted red areas in the FullGrad analysis are consistent with medical theories, then the current study’s model can be deemed interpretable according to explainable AI and medical theory concepts.
3 Methodology
This section outlines the methodology employed in this study, including the dataset description, preprocessing techniques, the workflow, the PENViT and TriplePENViT model architecture, and triplet networks hard triplet mining, as well as the pseudo labelling task for semi-supervised training of neural networks.
3.1 Creation of the dataset
The current study utilized a two-part dataset comprising "Dataset A" and "Dataset B," both of which contain grayscale panoramic radiographs of the maxillofacial region.
Dataset A served as the primary labeled and deidentified dataset, with each image annotated with the patient’s age and gender at the time of data input as obtained from the history sheet. The dataset was expanded with additional radiographs during revision to provide greater support to the models when estimating age and gender across several variables. The age and gender distributions for Dataset A are reported in Table 2. The use of Dataset A was approved by the related organizations.
Dataset B was a publicly available dataset from Tufts University (http://tdd.ece.tufts.edu/Tufts_Dental_Database/Radiographs.zip), which lacks annotated labels for age and gender, similar to those seen in multicenter deep learning implementations [45]. It comprised of a collection of 1000 unannotated radiographs [46]. The said Dataset B was utilized as the source for unlabeled data in the semi-supervised task undertaken in the current study, and for the rest of the task, Dataset A was used. While Dataset B lacked information on chronological age, it added geographic variation to the datasets used to investigate pseudo-labelling technique (5).
The word “age” in context of the current study has two meaning: chronological age (labels) and estimated dental age (estimated by the neural networks). In the context of dataset interpretation, the former is used, however, when discussing neural networks estimation the authors of the current study use the latter.
Dataset A initially consisted of total 525 radiographic images, from where 101 samples either had labelling issues or exhibited distorted features and were therefore discarded leaving 425 images. The images were subsequently split into training and validation datasets. To explore more versatile approaches, Dataset A was later expanded during revision stages to 706. All images in Dataset A consisted of sizes of a minimum dimension of 2000 pixels in width and 1000 pixels in height. To ensure compatibility with the pretrained model, which has fixed input image size, each file was resized depending on the model to either 384 × 384 or 224 × 224 pixels during both training and validation stages.
Stratified resampling was then applied based on age labels to divide the initial dataset of 424 files into a training-validation split of 296 files for training and 129 files for validation, maintaining a ratio of 70:30 [47]. Once the age classifier was ready, the dataset was expanded further to train the gender classifier.
At this stage, the age distribution of the training samples was not balanced, resulting in class imbalance. To address this, medical data augmentation techniques were applied only to the training set, while the validation set remained fixed throughout the study [48]. The augmentation techniques employed included Horizontal Flip, Geometric transformations (Rotate, scale), and Intensity operations (gamma contrast and linear contrast) [48]. By augmenting the training data, the sample size increased to n = 924, and the class imbalance issue was completely mitigated, with each class having 154 images in the training set.
3.2 Model architecture
In this subsection, the proposed PENViT and TriplePENViT model architecture is demonstrated with information about the proper flow of data through the neural network, input–output dimensions, network’s layers details, and the architecture figure.
3.2.1 PENViT model architecture
In the current research, the PENViT model architecture (Fig. 1) incorporates two pretrained models, EfficientNet and Vision Transformer were used in parallel. Each model takes \(3\times 24\times 224\) images as input and produces intermediary vectors \({E}_{1}\in {\mathbb{R}}^{1000}\) and \({E}_{2}\in {\mathbb{R}}^{1000}\), respectively.
To obtain the combined embedding vector \({E}_{c}\in {\mathbb{R}}^{2000}\), the following operation was performed:
The combined intermediary vector \({E}_{c}\) was then fed into a fully connected block consisting of a 60% dropout layer, followed by a Dense layer with 512 units, ReLU activation, another 60% dropout layer, and finally a Dense layer with 256 units. This fully connected block outputs the final embedding vector \(E\in {\mathbb{R}}^{256}\) for PENViT.
The authors in the current study performed \({l}_{2}\) normalization of the embedding vector \(E\). They then used the dot product of the weight matrix \(W \in {\mathbb{R}}^{256\times 6}\), where the weight matrix is in its \({l}_{2}\) normalized form. The dot product is equivalent to \(\mathrm{cos}({\theta }_{{y}_{i}})\) in the context of ArcFace Loss [49]. To obtain the logit projection, the authors applied the following transformations:
Here, \(s\) represents the scale factor, and m is the additive margin [49]. By applying the ArcFace loss, the final cross-entropy loss (also known as SoftMax loss) is computed as:
For performance evaluation, the ArcFace component is omitted, and the logits are calculated as follows:
In the current study, Pretrained frozen EfficientNet B0 and Pretrained Frozen Vision Transformer Large Patch Size 32 models were used for the proposed PENViT architecture. Both models were pretrained on ImageNet and subsequently fine-tuned on ImageNet1k.
3.2.2 TriplePENViT model architecture
In the current study, for the Triplet Network experiments (Fig. 2), a partial PENViT backbone was utilized that was reused up until the combined intermediary vector \({E}_{c}\in {\mathbb{R}}^{2000}\). \({E}_{c}\) then underwent a 60% dropout followed by a Dense layer with 256 units, resulting in the backbone’s embedding vector \(E\in {\mathbb{R}}^{256}\).
In the Triplet Network, the backbone is denoted as M with parameterization θ. The same parameterization θ was used in three of the models within the triplet, denoted as \({M}_{\theta }\). For each triplet, consisting of an anchor, positive, and negative sample, three embeddings were produced: \({E}_{a}, {E}_{p},\) and \({E}_{n}\) each having a dimension of \({\mathbb{R}}^{256}\). The Triplet Margin Loss, denoted as \({L}_{2}({E}_{a}, {E}_{p}, {E}_{n})\), was employed to train the triplet network:
Here, the distance function used was the Euclidean distance, and \(m\) represents the margin value for the triplet loss.
For the evaluation of TriplePENViT, two methods were used. The first involved training a classification block, which consisted of a single dense layer, with the triplet network using only the anchor’s embedding. The second method involved calculating the distance between all pair embeddings and predicting the label of the current image as the label of the closest embedding. Both methods were reported in the results section.
Additionally, when training a classifier with the anchor’s embedding alongside training the triplet network, the authors proposed a customized task-specific loss function denoted as \({L}_{3}\):
The equation \({L}_{c}\) represents the classifier’s classification loss, which is the cross-entropy loss, and \({L}_{2}\) is the triplet margin loss. The scales, \(scal{e}_{1}\) and \(scal{e}_{2}\), are multiplied with each loss to ensure that none overpowers the others. However, during the initial warmup periods, \(scal{e}_{1}\) was forcefully assigned a value of zero (0). This was done to allow the triplet network to focus on learning to discriminate among the embeddings in the latent space before attempting to train the classifier. This customized loss function allowed for a balanced optimization of the classifier and the triplet network, incorporating both the classification, and embedding objectives in a joint training process.
Hard Triplet mining was used to choose the three samples (positive, negative, and anchor) as the input of TriplePENViT network. (Fig. 3).
3.3 Details of the training process
Initially, the regression experiments were conducted with various backbone architectures. For regression tasks, the mean squared error (MSE) loss function was employed during training, while for validation, we reported the mean absolute error (MAE) loss in line with the metrics used by existing literatures [9, 21].
For classification task, continuous age values from the labels of Dataset A were converted into six class for the neural networks to classify. Different loss functions were used depending on the specific task and model type. The loss functions employed were: Cross Entropy (also known as SoftMax Loss), ArcFace loss, and Triplet Margin Loss [49]. However, all Triplet networks, including our proposed TriplePENViT, employs Triplet Margin Loss. All ArcFace experiments were conducted with an additive margin value of 34.3 degrees, and all Triplet Margin Loss experiments margin was 1.0 (unless mentioned otherwise).
The classification tasks were further challenged by conducting pseudo-labelling semi-supervised experiments (Fig. 4) using Dataset B that had unlabeled radiographs, i.e., no chronological age information was made available. A PENViT model was trained with Dataset A and trained up until 68.99% validation accuracy, and later used in the workflow of (Fig. 4) to complete the experiments of Table 6.
In all experiments, an initial learning rate of \({10}^{-2}\) was utilized and employed the “Reduced Learning Rate on Plateau” training scheduler, with a patience level of 5 and a gamma factor of 0.9, to adjust the learning rate based on the validation loss. All experiments were conducted for a minimum of 400 epochs to evaluate the performance of various backbones. A weight decay value of 0.9 was utilized for the purpose.
For classification task, after getting the best performing backbone from above training process, the best backbone (PENViT as denoted in results section’s Table 4) was subjected to a three-day experiment consisting of 3000 epochs using 1 × Nvidia Tesla M60 on the Microsoft Azure ML Compute platform. However, it is worth noting that the model converged in less than 600 epochs. All other experiments were conducted on 1 × Nvidia Tesla T4. A minimum of 70 + experiments were conducted during this study, with only the most significant ones being presented in the results section.
In the case of the triplet network with a classifier (\({L}_{3}\) Loss), the authors experimented with initial warmup periods ranging from 20 to 100 epochs. During this warmup phase, the \(scal{e}_{1}\) value of \({L}_{3}\) was forcefully set to zero (0), allowing the triplet network to initially focus on learning to discriminate among the embeddings in the latent space, rather than attempting to train the classifier. Following the warmup period, the scale values of \({L}_{3}\) were set to be equal, with \(scal{e}_{1}\) and \(scal{e}_{2}\) both set to 1.0.
Throughout all the experiments, the batch size varied between 296 and 500, depending on the available GPU memory during runtime. Additionally, data augmentation techniques were applied during training to enhance the robustness and generalization of the models.
3.4 Evaluation methods
For the current study, the validation set remained the same for all the task and experiments. Therefore, validation accuracy was used as the sole performance metrics in the current study’s classification task, whereas for regression reliability or performance metrics, MAE was used.
4 Results
This section illustrates the results of the regression task, classification task, comparison between cross entropy and ArcFace performance, hard triplet minding task results, PENViT backbones performance, pseudo labelling workflow’s performance, and FullGrad images for model interpretability. MAE outcomes of the regression task have been highlighted in Table 3. The comparison between using SoftMax Loss and ArcFace loss was later reported in Table 4. The outputs have been described in Tables 3,4,5,6.
4.1 Regression: estimation of age using different model architecture
Table 3 presents the results of regression tasks using multiple layer CNN backbone, ViT, autoencoder, fully connected layers, and RESNET-like backbone. Among these popular backbones, pretrained ViT demonstrated superior performance in regression tasks. Compared to pure ViTs, the pretrained ViT achieved a lower Mean Absolute Error (MAE) of 2.83 years in the regression task.
4.2 Pure cross entropy versus ArcFace margin loss: rigorous experiments
Initially, cross entropy loss was utilized during the experimentation phase. Both ViT L32 and the novel PENViT model emerged as top performers, achieving validation accuracies of 68.11%.
To further enhance performance, the top-five models were selected and investigated the application of ArcFace Loss. Notably, the PENViT model demonstrated superior performance, reaching a validation accuracy of 70.54%. It was also observed that combining ArcFace Loss with a ResNet backbone led to increased validation accuracy in certain cases. However, it is important to highlight that in three out of the five cases evaluated, the application of ArcFace Loss resulted in a decline in overall performance.
These findings highlight the effectiveness of PENViT in conjunction with ArcFace Loss, consistently outperforming other models. The synergy between the ResNet backbone and ArcFace Loss was found to be beneficial in specific scenarios. The gender classifier alongside the age classifier resulted in partial degradation of validation accuracy from 68.21% to 67.44%. Nevertheless, also affirming that the PENViT architecture performed better for panoramic radiographs when combining age and gender classifiers.
4.3 Evaluating hard triplet mining task
In the Siamese or Triplet network family, the authors introduced the TriplePENViT model architecture, which outperformed other models by incorporating hard triplet mining and a classification block that utilizes only the anchor’s embedding. The TriplePENViT model, with its specific loss function, achieved an accuracy of 67.44%.
Interestingly, increasing the margin value of the loss function L3 did not lead to an improvement in the validation accuracy of the TriplePENViT model. This suggests that the chosen margin value was already optimal for the given task, and further adjustments did not yield significant performance gains.
4.4 PENViT backbone’s effectiveness against other backbones
In Tables 4 and 5, it is evident that the PENViT model and its variation, TriplePENViT, consistently outperformed other models, whether they were triplet networks or other types of neural networks. The PENViT model achieved a validation accuracy of 70.54%, while others achieved a maximum of 68.21%, resulting in a 2.33% increase in accuracy.
Similarly, in another instance with TriplePENViT utilizing the triplet network and its classification block, it surpassed other models by achieving a validation accuracy of 67.44%, compared to a maximum of 65.11% achieved by others. This again resulted in a 2.33% increase in validation accuracy.
Therefore, in both cases, when using either the PENViT backbone or its triplet network variation, TriplePENViT, there was a consistent improvement of 2.33% in validation accuracy compared to other models (Table 6).
4.5 Evaluating pseudo-labelling technique
The application of pseudo-labelling technique to train medical image data, which typically has a limited number of available samples, did not result in an increase in validation accuracy beyond the performance of the best model. Despite attempting pseudo labelling, the validation accuracy remained consistent and did not surpass the accuracy achieved by the best model.
4.6 PENViT’s model interpretability of classification
The model’s Explainable AI results (FullGrad) are provided in Fig. 5:
For Deciduous dentition (Fig. 5A) group and Mixed Dentition (Fig. 5B) groups, model took account of the developing tooth buds of permanent teeth and the relative proximity to the overlying deciduous dentition with some prioritization on the mandibular shape. For ages 20 to 29 (Fig. 5C), the model accounted for the permanent dentition and the alveolar bone density surrounding the formed root apices and the root formation and eruption status of the third molars. For ages above 30 years (Fig. 5D), the model additionally notes occlusal deformity resulting from missing permanent dentition and the condylar regions highlighting temporomandibular joints to be an important predictor for automated age estimation from 2D panoramic radiographs.
5 Discussion
To attain the objectives outlined, regression tasks were first adopted as the means to ascertain the deep learning models that yield the minimal Mean Absolute Error (MAE) when trained on a designated training set and validated on the same data sets. Notably, Vision Transformers utilizing self-attention mechanisms outperformed their counterparts, signifying the superiority of Vision Transformer-based backbones, particularly within Orthopantomograms (OPGs), for subsequent computer vision tasks such as classification. Consequently, the subsequent stage involved a transition from regression to classification, wherein the continuous age labels were discretized into six distinct classes. This also served as another test for the efficacy of the Vision Transformer hypothesis, reaffirming the dominance of Vision Transformers over Convolutional Neural Network (CNN) based backbones in the present study.
With the said pattern in consideration, the authors proposed the PENViT architecture as a hybrid solution, synergizing both CNN-based and self-attention-based backbones, which resulted in a notable performance boost. In further endeavors to enhance performance, a comparative analysis of two loss functions for multi-class classification was conducted, namely, SoftMax/CrossEntropy and the ArcFace Margin Loss function. Impressively, ArcFace exhibited even greater performance improvements. Although ArcFace is conventionally employed in Face Recognition models, its geometric interpretation elucidates its role in creating a margin in the latent vector space for class differentiation, as depicted in Fig. 6.
The backbone acquires the ability to differentiate samples within the latent vector space; however, this differentiation becomes more pronounced with the introduction of ArcFace. The margin imposed by ArcFace enforces greater separation among samples, thereby augmenting the discriminative prowess of the neural network and consequently elevating accuracy.
Future research could contemplate the integration of alternative data modalities alongside OPGs for weakly supervised age estimation. While the present study focused solely on OPGs for this purpose, Schmeling et al.’s [50] suggestion of including hand-wrist radiographs in sequential aging analysis, in conjunction with OPGs[51], could yield further advancements in age estimation performance.
The pursuit of age estimation holds significance in dentistry, forensic science, legal proceedings, court hearings, and related domains. Clinical application of unsupervised age estimation may be present in aiding computer vision-based diagnostics of restorative treatment needs based on predictive age of exfoliation [52, 53]. Within the aforementioned contexts, the utilization of OPGs for the task emerges as a cost-effective and straightforward approach compared to other methodologies [54]. While alternate techniques such as Cone Beam Computed Tomography (CBCT) and Computed Tomography (CT) exist in specialized dental practices and have been documented in literature, it is important to note that authors such as Yuan et al. [55] applied pelvic radiographs and supervised CNNs to train 1498 images, while Othmani et al. [56] applied supervised CNNs on 45,000 facial photographs to attain an MAE of 2.35. In comparison, the current study attained an MAE of 2.83 using unsupervised learning using 706 radiographs.
5.1 Limitation of current study
The study lacked an evaluation of diagnostic accuracy, which could provide valuable clinical insights into the neural network model’s performance on unknown data, potentially affecting its clinical utility given the reported model’s performance of 70.54%. Secondly, the absence of a comparison between the model’s performance and that of a human diagnostician remains a research gap, although the potential for such a comparison through tools like FullGrad, GradCAM, or other EX-AI techniques is acknowledged for future research. Despite recognizing the challenge of immediate execution, this study’s limitation lies in not implementing a comparison with human diagnostic practices, as seen in similar studies [57]. Additionally, the proposed network architectures, PENViT and TripplePENViT, may lean heavily towards computer vision and deep learning techniques rather than a clinician’s perspective, indicating a potential mismatch in orientation. The study also lacks an investigation into class imbalances and which class has higher representation, despite the clear focus on evaluating different backbone performance, proposing concatenation of the best-performing ones, and assessing the impact of margin loss and pseudo-labelling. While prioritizing these aspects over reporting diagnostic accuracy, the study acknowledges the importance of this limitation. Lastly, the dataset’s size and class imbalance (across age and gender) are acknowledged to potentially yield unintended outcomes, despite artificial dataset size increase and balancing efforts. The study recognizes that more optimal results might have been achievable with larger, better-balanced datasets as is crucial for clinical applications.
6 Conclusion
From the current study the following answers can be inferred for the posed research questions:
-
1.
ViT demonstrated superior performance over CNN architectures in regression tasks.
-
2.
ArcFace showed mixed results, with instances of improved performance compared to pure cross entropy loss, but also cases where it deteriorated performance. In contrast, Triplet Margin Loss in a triplet network consistently outperformed other experiments, except for ViT L32 with Cross Entropy Loss, which performed slightly better.
-
3.
The use of hard triplet margin alone resulted in poor performance but combining it with a classifier yielded comparable results to the best-performing approach.
-
4.
The proposed PENViT backbone consistently outperformed other backbones, achieving higher validation accuracy.
-
5.
Training the model with pseudo labelling did not yield satisfactory results compared to using annotated data only.
-
6.
The FullGrad approach for explainability of the model highlighted that the most influential areas for predicting age brackets were deciduous teeth, areas of anodontia, extent of sinus cavities, periodontal regions, third molar regions, medullary regions of the mandible, and the temporomandibular joint complex that are consistent with medical explainability.
Data availability
All codes and scripts used to obtain the outcomes described have been made available on GitHub (https://github.com/ari1337an/OPGAgeClassify) Last accessed on 24 July 2023. All scripts were written adhering to the PEP-8 guidelines.
References
Lu J, Liong VE, Zhou J (2015) Cost-sensitive local binary feature learning for facial age estimation. IEEE Trans Image Process 24:5356–5368
Cunha E, Baccino E, Martrille L, Ramsthaler F, Prieto J, Schuliar Y, Lynnerup N, Cattaneo C (2009) The problem of aging human remains and living individuals: a review. Forensic Sci Int 193:1–13
Schmidt S, Schiborr M, Pfeiffer H, Schmeling A, Schulz R (2013) Sonographic examination of the apophysis of the iliac crest for forensic age estimation in living persons. Sci Justice 53:395–401
Al Hasan H, Saad FH, Ahmed S, Mohammed N, Farook TH, Dudley J (2023) Experimental validation of computer-vision methods for the successful detection of endodontic treatment obturation and progression from noisy radiographs. Oral Radiol. https://doi.org/10.1007/s11282-023-00685-8
Bassed RB, Briggs C, Drummer OH (2011) Age estimation using CT imaging of the third molar tooth, the medial clavicular epiphysis, and the spheno-occipital synchondrosis: a multifactorial approach. Forensic Sci Int 212:273-e1
Aboshi H, Takahashi T, Komuro T (2010) Age estimation using microfocus X-ray computed tomography of lower premolars. Forensic Sci Int 200:35–40
Asif MK, Nambiar P, Mani SA, Ibrahim NB, Khan IM, Sukumaran P (2018) Dental age estimation employing CBCT scans enhanced with Mimics software: comparison of two different approaches using pulp/tooth volumetric analysis. J Forensic Leg Med 54:53–61
Asif MK, Nambiar P, Mani SA, Ibrahim NB, Khan IM, Lokman NB (2019) Dental age estimation in Malaysian adults based on volumetric analysis of pulp/tooth ratio using CBCT data. Leg Med 36:50–58
Čular L, Tomaić M, Subašić M, Šarić T, Sajković V, Vodanović M (2017) Dental age estimation from panoramic X-ray images using statistical models, In: Proceedings of the 10th International Symposium on Image and Signal Processing and Analysis, IEEE, pp. 25–30.
Atas I, Ozdemir C, Atas M, Dogan Y (2022) Forensic dental age estimation using modified deep learning neural network. Comput Vision. https://doi.org/10.48550/arXiv.2208.09799
Mualla N, Houssein EH, Hassan MR (2020) Dental age estimation based on X-ray images. Comput Mater Continua. https://doi.org/10.32604/cmc.2020.08580
Vila-Blanco N, Carreira MJ, Varas-Quintana P, Balsa-Castro C, Tomas I (2020) Deep neural networks for chronological age estimation from OPG images. IEEE Trans Med Imaging 39:2374–2384
Silva G, Oliveira L, Pithon M (2018) Automatic segmenting teeth in X-ray images: trends, a novel data set, benchmarking and future perspectives. Expert Syst Appl 107:15–31
Jader G, Fontineli J, Ruiz M, Abdalla K, Pithon M, Oliveira L (2018) Deep instance segmentation of teeth in panoramic X-ray images, In: 2018 31st SIBGRAPI Conference on Graphics, Patterns and Images (SIBGRAPI), IEEE, pp. 400–407.
A.B. Oktay, Tooth detection with convolutional neural networks. In: 2017 Medical Technologies National Congress (TIPTEKNO), IEEE, 2017: pp. 1–4
Tuzoff DV, Tuzova LN, Bornstein MM, Krasnov AS, Kharchenko MA, Nikolenko SI, Sveshnikov MM, Bednenko GB (2019) Tooth detection and numbering in panoramic radiographs using convolutional neural networks. Dentomaxillofacial Radiol 48:20180051
Kuo Y-F, Lin S-Y, Wu CH, Chen S-L, Lin T-L, Lin N-H, Mai C-H, Villaverde JF (2017) A convolutional neural network approach for dental panoramic radiographs classification. J Med Imaging Health Inform 7:1693–1704
Du X, Chen Y, Zhao J, Xi Y (2018) A convolutional neural network based auto-positioning method for dental arch in rotational panoramic radiography. In: 2018 40th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), IEEE, pp. 2615–2618
Chu P, Bo C, Liang X, Yang J, Megalooikonomou V, Yang F, Huang B, Li X, Ling H (2018) Using octuplet siamese network for osteoporosis analysis on dental panoramic radiographs. In: 2018 40th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), IEEE, pp. 2579–2582
Poedjiastoeti W, Suebnukarn S (2018) Application of convolutional neural network in the diagnosis of jaw tumors. Healthc Inform Res 24:236–241
Fan F, Ke W, Dai X, Shi L, Liu Y, Lin Y, Cheng Z, Zhang Y, Chen H, Deng Z (2023) Semi-supervised automatic dental age and sex estimation using a hybrid transformer model. Int J Legal Med 137:721–731
Thevissen P, Willems G, Van de Voorde W, Solheim T (2013) Dental age estimation in sub-adults: striving for an optimal approach
Sironi E, Taroni F, Baldinotti C, Nardi C, Norelli G-A, Gallidabino M, Pinchi V (2018) Age estimation by assessment of pulp chamber volume: a Bayesian network for the evaluation of dental evidence. Int J Legal Med 132:1125–1138
He S, Feng Y, Grant PE, Ou Y (2022) Deep relation learning for regression and its application to brain age estimation. IEEE Trans Med Imaging 41:2304–2317
Atas I (2022) Human gender prediction based on deep transfer learning from panoramic radiograph images, ArXiv Preprint ArXiv:2205.09850.
Peng H, Gong W, Beckmann CF, Vedaldi A, Smith SM (2021) Accurate brain age prediction with lightweight deep neural networks. Med Image Anal 68:101871
De Back W, Seurig S, Wagner S, Marré B, Roeder I, Scherf N (2019) Forensic age estimation with Bayesian convolutional neural networks based on panoramic dental X-ray imaging
Stahl F, Grabowski R (2003) Orthodontic findings in the deciduous and early mixed dentition–inferences for a preventive strategy. J Orofac Orthop 64:401–416
Hadler-Olsen E, Jönsson B (2021) Oral health and use of dental services in different stages of adulthood in Norway: a cross sectional study. BMC Oral Health 21:257
Farhadian M, Salemi F, Saati S, Nafisi N (2019) Dental age estimation using the pulp-to-tooth ratio in canines by neural networks, Imaging Sci. Dent 49:19–26
Liu N (2021) Chronological age estimation of lateral cephalometric radiographs with deep learning, ArXiv Preprint ArXiv:2101.11805
Milošević D, Vodanović M, Galić I, Subašić M (2022) Automated estimation of chronological age from panoramic dental X-ray images using deep learning. Expert Syst Appl 189:116038
Liu F, Tian Y, Chen Y, Liu Y, Belagiannis V, Carneiro G (2022) ACPL: Anti-curriculum pseudo-labelling for semi-supervised medical image classification. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp. 20697–20706.
Xu M.-C, Zhou Y, Jin C, de Groot M, Alexander D.C, Oxtoby N.P, Hu Y, Jacob J (2023) Expectation maximization pseudo labelling for segmentation with limited annotations, ArXiv Preprint ArXiv:2305.01747
Rhee H, Cho N.I (2019) Efficient and robust pseudo-labeling for unsupervised domain adaptation. In: 2019 Asia-Pacific Signal and Information Processing Association Annual Summit and Conference (APSIPA ASC), IEEE, pp. 980–985
Ham G, Cho Y, Lee J-H, Kim D (2022) P-pseudolabel: enhanced pseudo-labeling framework with network pruning in semi-supervised learning. IEEE Access 10:115652–115662
Zhang G, Kurita T (2021) Age Estimation from the Age Period by Using Triplet Network, In: Frontiers of Computer Vision: 27th International Workshop, IW-FCV 2021, Daegu, February 22–23, 2021, Revised Selected Papers 27, Springer, pp. 81–92.
Hajamohideen F, Shaffi N, Mahmud M, Subramanian K, Al Sariri A, Vimbi V, Abdesselam A (2023) Four-way classification of Alzheimer’s disease using deep Siamese convolutional neural network with triplet-loss function. Brain Inform 10:1–13
Jeong Y, Lee S, Park D, Park KH (2018) Accurate age estimation using multi-task siamese network-based deep metric learning for frontal face images. Symmetry (Basel) 10:385
Chattopadhay A, Sarkar A, Howlader P, Balasubramanian V.N (2018) Grad-cam++: Generalized gradient-based visual explanations for deep convolutional networks. In: 2018 IEEE Winter Conference on Applications of Computer Vision (WACV), IEEE, pp. 839–847
Omeiza D, Speakman S, Cintas C, Weldermariam K (2019) Smooth grad-cam++: An enhanced inference level visualization technique for deep convolutional neural network models, ArXiv Preprint ArXiv:1908.01224
Ramaswamy H.G (2020) Ablation-cam: visual explanations for deep convolutional network via gradient-free localization. In: Proceedings of the IEEE/CVF Winter Conference on Applications of Computer Vision, pp. 983–991.
Wang H, Joshi D, Wang S, Ji Q (2023) Gradient-based uncertainty attribution for explainable bayesian deep learning. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp. 12044–12053.
Srinivas S, Fleuret F (2019) Full-gradient representation for neural network visualization. Adv Neural Inf Process Syst 2019:32
Farook TH, Dudley J (2023) Automation and deep (machine) learning in temporomandibular joint disorder radiomics. A systematic review. J Oral Rehabil. https://doi.org/10.1111/joor.13440
Panetta K, Rajendran R, Ramesh A, Rao SP, Agaian S (2021) Tufts dental database: a multimodal panoramic x-ray dataset for benchmarking diagnostic systems. IEEE J Biomed Health Inform 26:1650–1659
Ferrari A, Lombardi S, Signoroni A (2017) Bacterial colony counting with convolutional neural networks in digital microbiology imaging. Pattern Recognit 61:629–640
Chlap P, Min H, Vandenberg N, Dowling J, Holloway L, Haworth A (2021) A review of medical image data augmentation techniques for deep learning applications. J Med Imaging Radiat Oncol 65:545–563
Deng J, Guo J, Xue N, Zafeiriou S (2019) Arcface: additive angular margin loss for deep face recognition. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp. 4690–4699.
Schmeling A, Grundmann C, Fuhrmann A, Kaatsch H-J, Knell B, Ramsthaler F, Reisinger W, Riepert T, Ritz-Timme S, Rösing FW (2008) Criteria for age estimation in living individuals. Int J Legal Med 122:457–460
Wang Y, Zhang Q, Han J, Jia Y (2018) Application of deep learning in bone age assessment. IOP Publishing, Bristol, p 032012
Farook TH, Ahmed S, Bin Jamayet N, Dudley J (2023) Computer vision with smartphone microphotography for detection of carious lesions. Intell Based Med 8:100105
Tareq A, Faisal MI, Islam MS, Rafa NS, Chowdhury T, Ahmed S, Farook TH, Mohammed N, Dudley J (2023) Visual diagnostics of dental caries through deep learning of non-standardised photographs using a hybrid YOLO ensemble and transfer learning model. Int J Environ Res Public Health 20:5351
Poongodi V, Kanmani R, Anandi MS, Krithika CL, Kannan A, Raghuram PH (2015) Prediction of age and gender using digital radiographic method: a retrospective study. J Pharm Bioallied Sci 7:S504
Li Y, Huang Z, Dong X, Liang W, Xue H, Zhang L, Zhang Y, Deng Z (2019) Forensic age estimation for pelvic X-ray images using deep learning. Eur Radiol 29:2322–2329
Othmani A, Taleb AR, Abdelkawy H, Hadid A (2020) Age estimation from faces using deep learning: a comparative analysis. Comput Vis Image Underst 196:102961
Razzaki S, Baker A, Perov Y, Middleton K, Baxter J, Mullarkey D, Sangar D, Taliercio M, Butt M, Majeed A (2018) A comparative study of artificial intelligence and human doctors for the purpose of triage and diagnosis, ArXiv Preprint ArXiv:1806.10698
Funding
The authors have not disclosed any funding.
Author information
Authors and Affiliations
Contributions
Conceptualization: SA, NM, THF Methodology Design: SHA, TAR, SA, NM Data Collection: THF, JD Data Processing: SHA, TAR, SA Codes and Metadata synthesis: SHA, TAR, SAl Data Interpretation: SHA, TAR, SAl, NM, THF, JD Manuscript writing: SHA, TAR, SAl Manuscript revision: THF, SA, JD, NM Final Approval for Submission: SA, THF
Corresponding author
Ethics declarations
Conflict of interest
The authors have not disclosed any competing interests.
Ethical approval
The study was approved by the University of Adelaide Human Research and Ethics Committee (HREC-2023-073) and Institutional Review Board of North South University (2023/OR-NSU/IRB/0503). All methods were carried out in accordance with relevant guidelines and regulations.
Consent to participate
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Arian, M.S.H., Rakib, M.T.A., Ali, S. et al. Pseudo labelling workflow, margin losses, hard triplet mining, and PENViT backbone for explainable age and biological gender estimation using dental panoramic radiographs. SN Appl. Sci. 5, 279 (2023). https://doi.org/10.1007/s42452-023-05503-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42452-023-05503-8