Abstract
Solanum nigrum Linn is a medicinal herb widely used in traditional Chinese medicine to treat ailments such as fever, inflammation and cancer. Although quite a few compounds have been isolated and characterized, its anticancer mechanism remains elusive. Thus, in this study, we used network pharmacology and molecular docking strategies to identify the major active ingredients in S. nigrum and reveal its putative mechanism against human breast cancer. Six compounds, quercetin, cholesterol, 3-epi-beta-sitosterol, diosgenin, medioresinol and solanocapsine, were identified to be the major active ingredients. Target identification and analysis showed that they regulate 80 breast cancer-related targets. Furthermore, network analysis showed that the six active ingredients regulate multiple pathways including ErbB signaling pathway and estrogen signaling pathway and genes AKT1(AKT serine/threonine kinase 1), ESR1(estrogen receptor 1), EGFR(epidermal growth factor receptor), SRC(proto-oncogene tyrosine-protein kinase Src), AR(androgen receptor) and MMP9(matrix metalloproteinase 9) are crucial genes involved in the regulations. Molecular docking implied that quercetin could form good binding with AKT1, EGFR, SRC and MMP9. Our current study suggests that the anticancer function of S. nigrum is likely via synergistic/additive effects of multiple active ingredients’ regulations of different signaling pathways. Further studies are warranted to establish the standard for S. nuigrum to be used as a CAM (complementary and alternative medicine) in breast cancer treatment and dissect its potential interactions with chemotherapy drugs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Breast cancer is one of the leading causes of cancer death in women. The latest epidemiological report shows that female breast cancer has surpassed lung cancer as the most commonly diagnosed cancer with 2.3 million new cases (11.7% of total cases) worldwide in 2020, closely followed by lung cancer (11.4%) and colorectal cancer (10.0%) [1]. The incidence rate of breast cancer has been decreased slightly in Canada over the past 30 years [2] but increased at 0.5% per year in the United States in the past years [3]. In addition, it has been noticed that the affected population is getting younger [4], which is highly concerning in the society. However, the mortality rate of breast cancer has been decreased steadily in both Canada and the United States since 1990s [3, 5], which is mainly attributed to advances in breast cancer diagnosis and treatment, such as mammography, targeted therapy and immunotherapy.
Although the 5-year survival rate for breast cancer is around 90%, prognosis for advanced stage and recurrent breast cancer remains poor [6, 7]. For example, the 5-year survival rate is around 100% for stage I patients but 22% for stage IV patients in Canada [6]. Chemotherapy of breast cancer usually faces drug resistance and severe adverse drug reactions, which, in turn, significantly affects the quality of life in breast cancer patients [8, 9]. Furthermore, the high cost of oncology drugs may limit their clinical application in certain countries [10]. Thus, many cancer patients, not just limited to advanced stage patients, seek complementary and alternative medicine (CAM) treatments in an attempt to improve therapeutic efficacy and/or reduce adverse drug reactions. A study of cancer patients in Northern Ontario, Canada showed that 51.8% of the patients used CAM products after diagnosis [11]. For patients’ safety, it is critical to enhance researches to understand the mechanism of action (MOA) and side effects of the CAM products.
Medicinal plants are a valuable resource for developing anticancer therapeutics and widely used in CAM. For example, paclitaxel, a chemotherapy drug used to treat breast cancer and ovarian cancer, was discovered from the Pacific Yew tree. Solanum nigrum L., commonly known as black nightshade, is a folklore herb used in traditional Chinese medicine (TCM). It is usually used to treat ailments such as fever, pain and inflammation [12,13,14]. However, its anticancer function has attracted people’s attention in recent years [15,16,17,18]. The water extract of S. nigrum and various active ingredients such as polyphenols have shown potent in vitro anticancer activities [19,20,21,22]. Specifically for breast cancer, it has been reported that S. nigrum can induce apoptosis and autophagy [15] and suppress mitochondrial function and epithelial-mesenchymal transition [19] in breast cancer cells. In addition, our previous studies showed that S. nigrum water extract inhibited cell migration and suppressed aerobic glycolysis towards human breast cancer MCF7 cells [23].
The main active components of S. nigrum and their corresponding targets in human cells have not yet been thoroughly identified, though. The underlying MOA for the entire plant rather than just an active constituent should be clarified because S. nigrum is typically administered as a whole plant in TCM preparations. In the present study, active components of S. nigrum as well as their potential mechanism behind its anticancer action were investigated by using network pharmacology and molecular docking approaches.The findings of this investigation may be able to offer a fresh method for thoroughly comprehending its antineoplastic properties..
2 Materials and methods
2.1 Screening of active ingredients in S. nigrum and corresponding targets in human cells
Information of bioactive compounds in S. nigrum were retrieved from the traditional Chinese medicine systems pharmacology database and analysis platform (TCMSP) (https://tcmspw.com/tcmsp.php) using “Solanum nigrum Linn” as a keyword. Oral bioavailability (OB ≥ 30%) and drug-likeness (DL ≥ 0.18) were set as the thresholds in identifying candidate active ingredients [24, 25]. For each candidate, 3D structure and structural graphic were obtained from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/) and saved in sdf and mol2 format. Subsequently, the sdf file was uploaded to Swiss Target Prediction database (http://swisstargetprediction.ch/index.php) to identify its potential interacting protein targets. Target protein names filtered with “Homo sapiens” and “Probability > 0” were downloaded.
2.2 Prediction of breast cancer-related targets for S. nigrum Active Ingredients
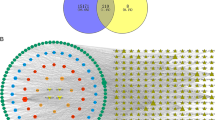
All breast cancer-related target genes were retrieved from GeneCards database (https://www.genecards.org/) using "breast cancer" as a keyword. These targets were subsequently standardized in UniProt (http://www.uniprot.org/) with organism selected as “Homo sapiens” to obtain a list of human breast cancer related targets. Finally, common targets between human breast cancer-related targets and the predicted targets for S. nigrum active ingredients (Sect. 2.1) were extracted and presented in a Venn diagram.
2.3 Construction of protein–protein interaction (PPI) network
The aforementioned common targets (Sect. 2.2) were subjected to protein-protein interaction networks and functional enrichment analysis using STRINGdb (https://string-db.org/). The protein interaction data was then imported into Cytoscape 3.7.2 [26] to construct PPI, analyze key regulatory proteins in the PPI, and identify core proteins with maximal degrees in topological analysis.
2.4 GO and KEGG analysis of key protein targets
Key targets identified above in Sect. 2.2 were uploaded to DAVID (database for annotation, visualization and integrated discovery, https://david.ncifcrf.gov/) for Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis (http://www.genome.jp/kegg/) with the species defined as "Homo Sapiens” and the threshold of significant difference set at p < 0.05. GO function enrichment analysis includes biological process (BP), cellular component (CC) and molecular function (MF). The results were then visualized and plotted using an online bioinformatics tool for data analysis and visualization (http://www.bioinformatics.com.cn).
2.5 Construction of component-target-pathway network
To figure out MOA of S. nigrum against breast cancer, we constructed the component-target-pathway network using the aforementioned information of active ingredients (component), key protein targets (target) and major biological pathways (pathway). The network was subsequently visualized using Cytoscape 3.7.2. In the network, the nodes represent components, targets and associated pathways (shown in pink, blue and green, respectively); while the edges represent component-target and target-pathway interactions.
2.6 Molecular docking of major active ingredients to key protein targets
Molecular docking was used to evaluate the capability and affinity of S. nigrum active ingredients to the selected key protein targets. 3D-structures of the protein targets were retrieved from the RCSB Protein Data Bank (PDB, http://www1.rcsb.org/) and 3D-structures of the active ingredients were prepared using Chimera 1.15 [27]. Molecular docking was performed using AutoDock Vina [28], with binding energy of −5.0 kcal/mol as the selection criteria [29].
3 Results
3.1 Identification of S. nigrum active ingredients and their respective targets in human cells
Seven major active ingredients (OB ≥ 30% and DL ≥ 0.18) were retrieved for S. nigrum from the TCMSP database. They are cholesterol, solanocapsine, diosgenin, medioresinol, beta-carotene, quercetin and 3-epi-beta-sitosterol (Table 1). Then, 190 potential targets were identified for the 7 bioactive compounds from the Swiss Target Prediction database.
3.2 Identification of bioactive compounds targets in S. nigrum for breast cancer
We extracted 14,360 breast cancer-related gene targets from GeneCards and selected 1837 genes for further crossover analysis after two cycles of median operation (score ≥ 9.67). As shown in Fig. 1, 80 gene targets were revealed to be shared between the bioactive compound targets and breast cancer targets. They are related to 6 active ingredients of S. nigrum except beta-carotene. Thus, beta-carotene was removed from the active ingredient list in the following analyses.
3.3 PPI Network Construction
PPI network was constructed for the 80 identified targets using the STRING database with the minimum interaction score set at 0.4 (Fig. 2). After screening, 79 targets were identified to interact with other proteins and 634 edges were observed in the network. This indicates that multiple interactions are present for the targets. The average degree of each node was 15.8. Sucrase-isomaltase (encoded by gene SI) was removed from the PPI network as it did not meet the selection criteria. Furthermore, we noticed that six proteins with highest degrees in the PPI network could be major hubs responsible for the anticancer function of S. nigrum. The six hub proteins are AKT1 (AKT serine/threonine kinase 1), ESR1 (estrogen receptor 1), EGFR (epidermal growth factor receptor), SRC (proto-oncogene tyrosine-protein kinase Src), AR (androgen receptor) and MMP9 (matrix metalloproteinase 9).
3.4 GO and KEGG enrichment analysis
DAVID database was employed for GO enrichment analysis to reveal the biological functions associated with the 80 common targets (p < 0.05). In total, we obtained 211 enrichment items for BP, 27 enrichment items for CC and 71 enrichment items for MF. In Fig. 3, we presented the top 10 enriched terms in each GO category in the order of p-value from low to high. These top 10 enriched terms are protein autophosphorylation, negative regulation of apoptotic process, peptidyl-tyrosine phosphorylation, response to drug, positive regulation of cell proliferation, positive regulation of gene expression, positive regulation of transcription from RNA polymerase II receptor, transmembrane receptor protein tyrosine kinase signaling pathway, oxidation–reduction process and phosphatidylinositol-mediated signaling in BP; cytosol, nucleus, nucleoplasm, plasma membrane, protein complex, extracellular space, extracellular exosome, mitochondrion, phosphatidylinositol 3-kinase complex and cytoplasm in CC; and ATP binding, transmembrane receptor protein tyrosine kinase activity, RNA polymerase II transcription factor activity, enzyme binding, protein tyrosine kinase activity, protein kinase activity, kinase activity, steroid binding, steroid hormone receptor activity and protein binding in MF.
Top 10 enriched items (p-value from low to high) for the 80 common targets (identified in Sect. 3.2) in the three GO categories of biological process (BP, shown in green), cellular component (CC, shown in orange) and molecular function (MF, shown in blue)
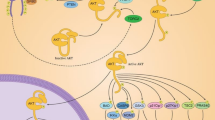
Signaling pathway enrichment for the 80 common targets was performed using the KEGG pathway database in order to reveal the key signaling pathways in breast cancer that are regulated by the active ingredients of S. nigrum. We identified 83 primarily enriched signaling pathways. The top 20 most significantly enriched pathways (p < 0.05) are pathways in cancer, prolactin signaling pathway, proteoglycans in cancer, FoxO signaling pathway, focal adhesion, central carbon metabolism in cancer, progesterone-mediated oocyte maturation, prostate cancer, PI3K-AKT signaling pathway, melanoma, insulin resistance, ErbB signaling pathway, glioma, ovarian steroidogenesis, estrogen signaling pathway, Rap1 signaling pathway, Ras signaling pathway, VEGF signaling pathway, pancreatic cancer and hepatitis C (Fig. 4).
3.5 Construction of component-target-pathway network
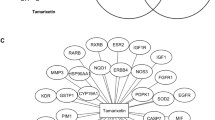
The component-target-pathway network (Fig. 5) was constructed using Cytoscape 3.7.2 by incorporating the 6 bioactive ingredients of S. nigrum, the 80 common targets shared between the bioactive compound targets and breast cancer targets, and the top 20 signaling pathways revealed by KEGG analysis. The network degree value was calculated for the 6 bioactive ingredients (Table 2) with 4 compounds having the value higher than 10. The 4 compounds are quercetin, cholesterol, 3-epi-beta-sitosterol and diosgenin. The network degree was relatively low for medioresinol and solanocapsine. This network analysis suggests that quercetin may play a more important role in contributing to the anticancer function of S. nigrum than the other active ingredients. It is also noteworthy that each active ingredient acts on multiple targets and is involved in various signaling pathways, implying that synergistic, additive and even antagonistic effects might be present among the active ingredients. Further studies are warranted to confirm these effects.
The component-target-pathway network constructed for S. nigrum. The component (active ingredients of S. nigrum), target (80 common targets identified in Sect. 3.2) and pathway (top 20 signaling pathways identified from KEGG) were shown in pink, blue and green, respectively
3.6 Docking the active ingredients of S. nigrum to the six hub proteins revealed from PPI
From the above PPI analysis, we identified 6 key protein targets (AKT1, ESR1, EGFR, SRC, AR and MMP9), which could be the major biological hubs responsible for the anti-breast-cancer function of S. nigrum. To explore whether the active ingredients could bind directly to these hub proteins, we carried out the docking studies between the 6 bioactive compounds and the 6 hub targets using binding energy of -5.0 kcal/mol as a selection cutoff. A binding energy value of less than 0 suggests indicates that the ligand molecules can spontaneously bind to the receptor protein.The lower the binding energy value,the stronger the affinity between the two. A value of less than − 5.0 kal/mol suggests that the ligand molecules have a desirable binding affinity.The results of the molecular docking revealed that 8 ingredient-target pairs were lower than −5 kcal/mol, indicating that 3 active ingredients, namely quercetin, cholesterol and 3-epi-beta-sitosterol, had good binding affinity to each of the six hub protein targets, namely AKT1,ESR1,EGFR,SRC,AR and MMP9 (Table 3). The docking patterns of the core ingredients and hub proteins were shown in Fig. 6. This docking study shows that quercetin could potentially bind to four hub targets, including AKT1, EGFR, SRC, and MMP9, which may trigger synergistic effects among different signaling pathways, suggests the use of quercetin in the treatment of breast cancer and provides data support for further experimental design.
Conformations of the 8 strong bindings identified from molecular docking between the 6 active ingredients of S. nigrum and the 6 hub protein targets, a AKT1 and quercetin, b ESR1 and cholesterol, c ESR1 and 3-epi-beta-sitosterol, d EGFR and quercetin, e SRC and quercetin, f AR and cholesterol, g AR and 3-epi-beta-sitosterol, and h MMP9 and quercetin
4 Discussion
S. nigrum is a medicinal herb widely used to treat fever, pain and inflammation in TCM [12,13,14]. With the discovery of its anticancer activities [15,16,17,18], S. nigrum has been included in integrative therapy of TCM to treat different types of cancer. Our previous studies have shown that the water extract of S. nigrum inhibited proliferation and migration of breast cancer MCF7 cells and suppressed the cells’ aerobic glycolysis [23]. However, the anticancer mechanism of S. nigrum is still far from understanding.
In the present study, we undertook an approach including network pharmacology and molecular docking to probe MOA of S. nigrum against human breast cancer. Six major bioactive compounds, cholesterol, solanocapsine, diosgenin, medioresinol, quercetin and 3-epi-beta-sitosterol, were identified. Out of these bioactive compounds, quercetin, which is probably one of the most-studied nutraceuticals, has been shown to possess strong antioxidant and anti-inflammatory activities and induce apoptosis of breast cancer cells [53, 54]. It can also inhibit cell adhesion and migration of breast cancer cells via downregulating HuR protein [54]. β-sitosterol, which is the most abundant type of phytosterol in various dietary plants such as vegetables and nuts, can inhibit the growth of breast cancer cells [55, 56]. Diosgenin, a steroidal saponin in plants, is also capable of inhibiting the growth and migration of human breast cancer cells [57, 58]. However, the functions of cholesterol, solanocapsine and medioresinol towards breast cancer cells are unknown.
Our subsequent analyses showed that the 6 active ingredients of S. nigrum interact with 80 breast-cancer-related targets and regulate a wide range of biological pathways. From the PPI network, 6 key protein targets (AKT1, ESR1, EGFR, SRC, AR and MMP9) were identified as major biological hubs. They play important roles in normal breast functions, and upregulation of their expressions promote breast cancer proliferation and metastasis. Therefore, we examined the expressions of the genes encoding these 6 hub proteins in breast cancer patients using online software GEPIA (http://gepia.cancer-pku.cn/). As shown in Fig. 7, genes ESR1, AR and MMP9 are upregulated and gene EGFR is downregulated, respectively, in breast cancer patients (p < 0.05 and labeled with *). However, none of them is a prognostic marker for breast cancer.
Boxplots showing the expressions of genes AKT1, ESR1, EGFR, SRC, AR and MMP9 in breast cancer patients (shown in red) versus normal controls (shown in grey) in the patient dataset BRCA deposited in The Cancer Genome Atlas (TCGA). This figure was generated using online software GEPIA (http://gepia.cancer-pku.cn/)
For the four differentially expressed genes in breast cancer patients, gene ESR1 encodes ESR1, which is a major hormone receptor on the surface of breast cancer cells. Its upregulation stimulates breast cancer proliferation, growth, and metastasis [59, 60]. ESR1 introns SNP + 2464 C/T (rs3020314) and SNP -4576 A/C (rs1514348) are correlated with breast cancer susceptibility and the expression status of progesterone receptor [61]. EGFR enhances proliferation and invasion of tumor cells and promotes breast cancer progression via JAK/STAT3 signaling [62]. EGFR is a major target for ΔNp63 regulation that influences cancer cell adhesion in basal-like triple-negative breast cancer (TNBC) [63]. Although EGFR is downregulated in breast cancer patients (without subcategorization), it is overexpressed in up to 76% in TNBC patients and EGFR-based targeted therapy has shown promising effects in TNBC patients [64, 65]. AR is expressed in > 70% breast cancer and across three major breast cancer subtypes [66, 67]. However, it seems that the function of AR is subtype-specific [67]. AR signaling can also crosstalk with other signaling pathways to regulate breast cancer pathology and progression [67]. It can either elicit or diminish oncogenic effects in breast cancer depending on estrogen bioavailability [68]. Thus, both AR agonists and antagonists could be potentially applied as therapeutic agents for breast cancer. A phase II clinical trial has shown that enzalutamide, an AR antagonist approved for treating prostate cancer, improved treatment outcome in AR-positive TNBC patients [69]. MMP9 is a matrix metalloproteinase responsible for breaking down the extracellular matrix (ECM). Its expression is low in normal breast tissues. However, MMP9 is overexpressed in breast cancer cells and promotes breast cancer cell migration and invasion via cleaving E-cadherin and degrading ECM [70, 71]. However, it has been shown that overexpression of MMP9 can trigger the generation and release of endostatin and tumstatin, both of which are capable of suppressing breast cancer progression and metastasis [72, 73]. The expression of MMP9 has been reported to be associated with the presence of AR in epithelial ovarian cancer cells [74]. However, this association is not observed in breast cancer (Fig. 8).
Pearson correlation between the expressions of MMP9 and AR in the patient dataset BRCA deposited in TCGA. This figure was generated using online software GEPIA (http://gepia.cancer-pku.cn/)
For the two genes that are not differentially expressed in breast cancer patients, gene AKT1 is a key player of the PI3K-AKT-mTOR signaling pathway in breast cancer although its expression is not statistically significantly increased in breast cancer patients. Activation of the PI3K-AKT-mTOR signaling pathway promotes cell proliferation, growth and survival and induces drug resistance in breast cancer [74, 75]. This pathway has emerged to be one of the major targets in developing therapeutics to treat solid tumors including breast cancer [76]. SRC (encoding SRC or c-SRC) is a proto-oncogene that is frequently activated in solid tumors including breast cancer [77]. SRC can activate multiple signaling pathways to promote cancer cell proliferation, growth, survival, migration and invasion [78,79,80]. Both epidermal growth factor (EGF) and SRC can bind to EGFR and enhance each respective cancer-promoting functions [80,81,82]. Furthermore, SRC was observed to crosstalk with AR during prostate cancer progression [83].
From the above analyses, it is clear that S. nigrum can regulate multiple signaling pathways via the six critical hub proteins and quercetin, cholesterol and 3-epi-beta-sitosterol are likely to be the major active ingredients responsible for those regulatory actions. Molecular docking suggests that the three active ingredients could directly bind to the hub proteins to exert the anticancer activity of S. nigrum. Therefore, we may conclude that administration of S. nigrum is beneficial for breast cancer patients and the anticancer function of S. nigrum is via a network including multiple bioactive compounds and multiple protein targets. Synergistic effects could be achieved from regulating multiple pathways. However, caution should be taken in co-administration of chemotherapy drugs and medicinal herbs such as S. nigrum since the drug-herb interactions are basically unknown. These drug-herb interactions may reduce the therapeutic efficacy of chemotherapy drugs and could even be detrimental to cancer patients. We highly recommend avoiding co-administration of chemotherapy drugs and medicinal herbs unless the co-administration is clinically proved to be safe, or the drug-herb interaction is clearly defined.
5 Conclusion
In this study, we used network pharmacology and molecular docking approaches to illustrate that S. nigrum may exert its anticancer function via interactions between the 6 active ingredients and 80 protein targets. Out of these interactions, the interactions between quercetin, cholesterol and 3-epi-beta-sitosterol and 6 hub proteins (AKT1, ESR1, EGFR, SRC1, AR and MMP9) are more important. Synergistic effects are likely to be achieved from the multiple signaling pathways. Briefly, our current research indicates that the anticancer activity of S. nigrum is probably due to synergistic effects achieved from multiple bioactive compound-target interactions.
In addition, we would like to point out several limitations of the current study. First, because of the vast amounts of data from the literature, discrepancies may exist in various databases. Secondly, during the decocting process, active compounds with low concentrations may be dismissed and/or unidentified gradients may be generated. Finally, as genes are dynamically regulated by a variety of factors, gene expression information extracted from the databases may not be fully representative for the diseases.Despite several limitations, the current study sets the groundwork for further experimental and clinical confirmation of S. nigrum's molecular mechanism for fighting breast cancer.
Our present approach of determining the MOAs for S. nigrum including multi-component, multi-target, and multi-pathway may be also applicable for identifying the MOAs for other medicinal herbs.These medicinal herbs unquestionably merit further pharmacological and clinical research to validate theoretical predictions and standardize their applications in cancer treatment.
Data availability
The authors do not have permission to share data. The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- S. nigrum :
-
Solanum nigrum Linn
- CAM:
-
Complementary and alternative medicine
- TCM:
-
Traditional Chinese medicine
- MOA:
-
Mechanism of action
- TCMSP:
-
Traditional Chinese medicine systems pharmacology database and analysis platform
- OB:
-
Oral bioavailability
- DL:
-
Drug-likeness
- PPI:
-
Protein–protein interaction
- GO:
-
Gene Ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- BP:
-
Biological process
- CC:
-
Cellular component
- MF:
-
Molecular function
- AKT1:
-
AKT serine/threonine kinase 1
- EGF:
-
Epidermal growth factor
- EGFR:
-
Epidermal growth factor receptor
- SRC:
-
Proto-oncogene tyrosine-protein kinase Src
- MMP9:
-
Matrix metalloproteinase 9
- ESR1:
-
Estrogen receptor 1
- AR:
-
Androgen receptor
- TNBC:
-
Triple-negative breast cancer
- ECM:
-
Extracellular matrix
- TCGA:
-
The Cancer Genome Atlas
References
Sung H, Ferlay J, Siegel RL et al (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71(3):209–249
Breast Cancer (2022) https://www.canada.ca/en/public-health/services/chronic-diseases/cancer/breast-cancer.html. Accessed 17/11/2022
Key Statistics for Breast Cancer (2022) https://www.cancer.org/cancer/breast-cancer/about/how-common-is-breast-cancer.html. Accessed 17/11/2022
He ZY, Chen Z, Tan MD et al (2020) A review on methods for diagnosis of breast cancer cells and tissues. Cell Prolif 53(7):12822
Brenner DR, Poirier A, Woods RR et al (2022) Projected estimates of cancer in Canada in 2022. Can Med Assoc J 194(17):E601–E607
Treating Breast Cancer (2022) https://cancer.ca/en/cancer-information/cancer-types/breast/prognosis-and-survival/survival-statistics. Accessed 17/11/2022
How is cancer treated (2022) https://www.canada.ca/en/public-health/services/chronic-diseases/cancer/cancer-treated.html. Accessed 17/11/2022
Aldossary S (2019) Review on pharmacology of cisplatin: clinical use, toxicity and mechanism of resistance of cisplatin. Biomed Pharmacol J 11:07–15
Waks AG, Winer EP (2019) Breast cancer treatment: a review. JAMA 321(3):288–300
Malik JA, Ahmed S, Jan B et al (2022) Drugs repurposed: an advanced step towards the treatment of breast cancer and associated challenges. Biomed Pharmacother 145:112375
Buckner CA, Lafrenie RM, Denommee JA et al (2018) Complementary and alternative medicine use in patients before and after a cancer diagnosis. Curr Oncol 25(4):E275–E281
Zakaria ZA, Gopalan HK, Zainal H et al (2006) Antinociceptive, anti-inflammatory and antipyretic effects of Solanum nigrum chloroform extract in animal models. Yakugaku Zasshi-J Pharm Soc Jpn 126(11):1171–1178
Pokhrel B, Acharya E (2007) Ethno-medicinal plants used by Bantar of Bhaudaha, Morang, Nepal. Our Nat 4:96–103
Kudesia R, Rani M, Pal A (2009) Reclaiming degraded land in India through the cultivation of medicinal plants. Bot Res Int 2:174–181
Huang HC, Syu KY, Lin JK (2010) Chemical composition of Solanum nigrum linn extract and induction of autophagy by leaf water extract and its major flavonoids in AU565 breast cancer cells. J Agric Food Chem 58(15):8699–8708
Wang CW, Chen CL, Wang CK et al (2015) Cisplatin-, doxorubicin-, and docetaxel-induced cell death promoted by the aqueous extract of Solanum nigrum in human ovarian carcinoma cells. Integr Cancer Ther 14(6):546–555
Swetha M, Keerthana CK, Rayginia TP et al (2022) Augmented efficacy of uttroside b over sorafenib in a murine model of human hepatocellular carcinoma. Pharmaceuticals 15(5):636
Tai CJ, Wang CK, Chang YJ et al (2012) Aqueous extract of Solanum nigrum leaf activates autophagic cell death and enhances docetaxel-induced cytotoxicity in human endometrial carcinoma cells. Evid Based Complement Altern Med 2012:859185
Lai YJ, Tai CJ, Wang CW et al (2016) Anti-cancer activity of Solanum nigrum (AESN) through suppression of mitochondrial function and epithelial-mesenchymal transition (EMT) in breast cancer cells. Molecules 21(5):553
Fu RJ, Wang XH, Hu Y et al (2019) Solamargine inhibits gastric cancer progression by regulating the expression of lncNEAT1_2 via the MAPK signaling pathway. Int J Oncol 54(5):1545–1554
Xie XD, Zhang XM, Chen J et al (2019) Fe3O4-solamargine induces apoptosis and inhibits metastasis of pancreatic cancer cells. Int J Oncol 54(3):905–915
Chen XF, Dai XF, Liu YH et al (2022) Solanum nigrum Linn: an insight into current research on traditional uses, phytochemistry, and pharmacology. Front Pharmacol 13:918071
Ling BB, Xiao SJ, Yang JH et al (2019) Probing the antitumor mechanism of Solanum nigrum L aqueous extract against human breast cancer MCF7 cells. Bioeng Basel 6(4):112
Hsin KY, Matsuoka Y, Asai Y et al (2016) systemsDock: a web server for network pharmacology-based prediction and analysis. Nucleic Acids Res 44(W1):W507–W513
Huang L, Zhang Y, Zhang X et al (2019) Therapeutic potential of Pien-Tze-Huang: a review on its chemical composition, pharmacology, and clinical application. Molecules 24(18):3274
Cytoscape 3.7.2 User Manual. http://manual.cytoscape.org/en/3.7.2/#cytoscape-version-user-manual (accessed 17/11/2022.
Chimera User's Guide. http://www.cgl.ucsf.edu/chimera/current/docs/UsersGuide/index.html (accessed 17/11/2022.
AutoDock User Guide. https://autodock.scripps.edu/documentation/documentation/ (accessed 17/11/2022.
Alam MS, Rahaman MM, Sultana A et al (2022) Statistics and network-based approaches to identify molecular mechanisms that drive the progression of breast cancer. Comput Biol Med 145:105508
Ding SS, Wang WH, Song XJ et al (2021) Based on network pharmacology and molecular docking to explore the underlying mechanism of Huangqi Gegen decoction for treating diabetic nephropathy. Evid-Based Complement Altern Med 2021:1–14
Li N, Liu KX, Yu M et al (2022) Systematic elucidation of the traditional Chinese medicine prescription Danxiong particles via network pharmacology and molecular docking. Trop J Pharm Res 21(5):981–987
Ma XQ, Hao CX, Yu MX et al (2022) Investigating the molecular mechanism of Quercetin protecting against podocyte injury to attenuate diabetic nephropathy through network pharmacology, microarraydata analysis, and molecular docking. Evid-Based Complement Altern Med 2022:7291434
Wu LT, Chen Y, Chen MJ et al (2021) Application of network pharmacology and molecular docking to elucidate the potential mechanism of Astragalus-Scorpion against prostate cancer. Andrologia 53(9):e14165
Xia QD, Xun Y, Lu JL et al (2020) Network pharmacology and molecular docking analyses on Lianhua Qingwen capsule indicate Akt1 is a potential target to treat and prevent COVID-19. Cell Prolif 53(12):e12949
Lin HC, Zhang XY, Li JY et al (2022) Unraveling the molecular mechanism of Xuebijing injection in the treatment of chronic obstructive pulmonary disease by combining network pharmacology and Affymetrix array. Nat Prod Commun 17(4):1934578
Ou L, Kang WQ, Liang ZY et al (2021) Investigation of anti-osteoporosis mechanisms of Rehmanniae Radix Preparata based on network pharmacology and experimental verification. J Orthop Surg Res 16(1):1–12
Fan YH, Liu W, Jin Y et al (2021) Integrated molecular docking with network pharmacology to reveal the molecular mechanism of simiao powder in the treatment of acute gouty arthritis. Evid-Based Complement Altern Med 2021:1–15
Lu DD, Shang JW, Guo XP et al (2022) Assessing the mechanism of action of “Fructus ligustri Lucidi-Cuscutae Semen” in prostate cancer treatment using network pharmacology and molecular docking[J]. Comput Math Methods Med 2022:7543619
Wu CX, Yu QQ, Shou WZ et al (2022) Identification of molecular mechanism of the anti-lung cancer effect of Jin Ning Fang using network pharmacology and its experimental verification. All Life 15(1):745–759
Ko MS, Kim Y, Kim HH et al (2022) Network pharmacology and molecular docking approaches to elucidate the potential compounds and targets of Saeng-Ji-Hwang-Ko for treatment of type 2 diabetes mellitus. Comput Biol Med 149:106041
Wang Q, Du LJ, Hong JN et al (2021) Molecular mechanism underlying the hypolipidemic effect of Shanmei Capsule based on network pharmacology and molecular docking. Technol Health Care 29:S239–S256
Agustina DW, Wahyuningsih MD, Widyarti S et al (2021) Molecular docking study to reveal Morinda citrifolia fruits as a novel EGFR inhibitor for anticancer therapy. IOP Conf Series: Earth Environ Sci 743(1):012082
Rizvi SMD, Hussain T, Mehmood K et al (2021) Molecular docking and dynamic simulation study to explore quercetin as a multi-potent candidate against gliomas. Trop J Pharm Res 20(4):815–823
Tan X, Xian W, Li XR et al (2022) Mechanisms of Quercetin against atrial fibrillation explored by network pharmacology combined with molecular docking and experimental validation. Sci Rep 12(1):9777
Yang L, Hu Z, Zhu J, Liang Q et al (2020) Systematic elucidation of the mechanism of Quercetin against gastric cancer via network pharmacology approach. BioMed Res Int 2020:3860213
Huang J, Teh BM, Xu Z et al (2022) The possible mechanism of Hippophae fructus oil applied in tympanic membrane repair identified based on network pharmacology and molecular docking. J Clin Lab Anal 36(1):e24157
He Y, Wang Y, Xu J et al (2022) In Tilte 6–8:2022
Zhu R, Du BZ, Yuan JY et al (2022) Potential mechanisms of Biejiajian Pill in the treatment of diabetic atherosclerosis based on network pharmacology, molecular docking, and molecular dynamics simulation. Evid-Based Complement Altern Med 2022:3296279
Zeng Y, Xiao S, Yang L et al (2021) Systematic analysis of the mechanism of Xiaochaihu decoction in hepatitis B treatment via network pharmacology and molecular docking. Comput Biol Med 138:104894
Chang Z, He LJ, Tian DF et al (2021) Therapeutic targets and mechanism of Xingpi Jieyu decoction in depression: a network pharmacology study. Evid-Based Complement Altern Med 2021:5516525
Belal A, Elanany MA, Raafat M et al (2022) Calendula officinalis phytochemicals for the treatment of wounds through matrix metalloproteinases-8 and 9 (MMP-8 and MMP-9): in silico approach. Nat Prod Commun 17(5):1934578X221098848
Tan X, Xian W, Chen Y et al (2021) Exploring the therapeutic mechanism of quercetin for heart failure based on network pharmacology and molecular docking. J South Med Univ 41(8):1198–1206
Gol RM, Kheirouri S (2022) The effects of quercetin on the apoptosis of human breast cancer cell lines MCF-7 and MDA-MB-231: a systematic review. Nutr Cancer Int J 74(2):405–422
Umar SM, Patra S, Kashyap A et al (2022) Quercetin impairs HuR-driven progression and migration of triple negative breast cancer (TNBC) cells. Nutr Cancer Int J 74(4):1497–1510
Awad AB, Chinnam M, Fink CS et al (2007) beta-sitosterol activates Fas signaling in human breast cancer cells. Phytomedicine 14(11):747–754
Afzal O, Akhter MH, Ahmad I et al (2022) A beta-sitosterol encapsulated biocompatible alginate/chitosan polymer nanocomposite for the treatment of breast cancer. Pharmaceutics 14(8):1711
Liu YL, Zhou ZJ, Yan JZ et al (2020) Diosgenin exerts antitumor activity via downregulation of Skp2 in breast cancer cells. Biomed Res Int 2020:8072639
He ZM, Chen HY, Li GF et al (2014) Diosgenin inhibits the migration of human breast cancer MDA-MB-231 cells by suppressing Vav2 activity[J]. Phytomedicine 21(6):871–876
Brett JO, Spring LM, Bardia A, Wander SA (2021) ESR1 mutation as an emerging clinical biomarker in metastatic hormone receptor-positive breast cancer. Breast Cancer Res 23:1–15
Robinson DR, Wu YM, Vats P et al (2013) Activating ESR1 mutations in hormone-resistant metastatic breast cancer. Nat Genet 45(12):1446-U1197
Lipphardt MF, Deryal M, Ong MF et al (2013) ESR1 single nucleotide polymorphisms predict breast cancer susceptibility in the central European Caucasian population. Int J Clin Exp Med 6(4):282–288
Song X, Liu ZY, Yu ZY (2020) EGFR promotes the development of triple negative breast cancer through JAK/STAT3 signaling. Cancer Manag Res 12:703–717
Holcakova J, Nekulova M, Orzol P et al (2017) Delta Np63 activates EGFR signaling to induce loss of adhesion in triple-negative basal-like breast cancer cells. Breast Cancer Res Treat 163(3):475–484
Zakaria Z, Zulkifle MF, Hasan WANW et al (2019) Epidermal growth factor receptor (EGFR) gene alteration and protein overexpression in Malaysian triple-negative breast cancer (TNBC) cohort. Onco Targets Ther 12:7749–7756
Matsuda N, Lim B, Wang XP et al (2017) Early clinical development of epidermal growth factor receptor targeted therapy in breast cancer. Expert Opin Investig Drugs 26(4):463–479
Chia K, O’Brien M, Brown M et al (2015) Targeting the androgen receptor in breast cancer. Curr Oncol Rep 17(2):1–6
Anestis A, Zoi I, Papavassiliou AG et al (2020) Androgen receptor in breast cancer-clinical and preclinical research insights. Molecules 25(2):358
Need EF, Selth LA, Harris TJ et al (2012) Research resource: interplay between the genomic and transcriptional networks of androgen receptor and estrogen receptor alpha in luminal breast cancer cells. Mol Endocrinol 26(11):1941–1952
Traina TA, Miller K, Yardley DA et al (2018) Enzalutamide for the treatment of androgen receptor-expressing triple-negative breast cancer. J Clin Oncol 36(9):884–890
Augoff K, Hryniewicz-Jankowska A, Tabola R, Stach K (2022) MMP9: a tough target for targeted therapy for cancer. Cancers 14(7):1847
Quintero-Fabian S, Arreola R, Becerril-Villanueva E et al (2019) Role of matrix metalloproteinases in angiogenesis and cancer. Front Oncol 9:1370
Okamoto T, Akaike T, Sawa T et al (2001) Activation of matrix metalloproteinases by peroxynitrite-induced protein S-glutathiolation via disulfide S-oxide formation. J Biol Chem 276(31):29596–29602
Ali MAM, Garcia-Vilas JA, Cromwell CR et al (2021) Matrix metalloproteinase-2 mediates ribosomal RNA transcription by cleaving nucleolar histones. FEBS J 288(23):6736–6751
Miricescu D, Totan A, Stanescu-Spinu II et al (2021) PI3K/AKT/mTOR signaling pathway in breast cancer: from molecular landscape to clinical aspects. Int J Mol Sci 22(1):1–15
Dong C, Wu J, Chen Y et al (2012) Activation of PI3K/AKT/mTOR pathway causes drug resistance in breast cancer. Front Pharmacol 12:628690
LoRusso PM (2016) Inhibition of the PI3K/AKT/mTOR pathway in solid tumors. J Clin Oncol 34(31):3803–3815
Dehm SM, Bonham K (2004) SRC gene expression in human cancer: the role of transcriptional activation. Biochem Cell Biol 82(2):263–274
Ma HH, Zhang J, Zhou L et al (2020) c-SRC promotes tumorigenesis and tumor progression by activating PFKFB3. Cell Rep 30(12):4235
Finn RS (2008) Targeting SRC in breast cancer. Ann Oncol 19(8):1379–1386
Belli S, Esposito D, Servetto A et al (2020) c-SRC and EGFR inhibition in molecular cancer therapy: What else can we improve ? Cancers 12(6):1489
Wasilenko WJ, Payne DM, Fitzgerald DL et al (1991) Phosphorylation and activation of epidermal growth factor receptors in cells transformed by the src oncogene. Mol Cell Biol 11(1):309–321
Biscardi JS, Maa M-C, Tice DA et al (1999) c-SRC-mediated phosphorylation of the epidermal growth factor receptor on Tyr845 and Tyr1101 is associated with modulation of receptor function. J Biol Chem 274(12):8335–8343
Gao L, Ouyang Y, Li R et al (2022) Icaritin inhibits migration and invasion of human ovarian cancer cells via the Akt/mTOR signaling pathway. Front Oncol 12:843489
Funding
This research was funded by a Jiangsu Government Scholarship for Overseas Studies (Grant No.: JS-2020–046), The Open Project of Nanjing Research Center for Infectious Diseases of Integrated Traditional Chinese and Western Medicine (Grant No.: NCMIC-2022–06) and the Research on Undergraduate Education and Teaching Reform of Nanjing University of Chinese Medicine (Grant No.: NZYJG2022067).
Author information
Authors and Affiliations
Contributions
Conceptualization, YS and JY; methodology, YS; investigation, YS and JY; resources, MKS; data curation, YS; writing—original draft preparation, YS; writing—review and editing, YS and JY; visualization, YS; supervision, JY; funding acquisition, YS. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Conflicts of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Song, Y., Sakharkar, M.K. & Yang, J. Probing the mechanism of action (MOA) of Solanum nigrum Linn against breast cancer using network pharmacology and molecular docking. SN Appl. Sci. 5, 133 (2023). https://doi.org/10.1007/s42452-023-05356-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42452-023-05356-1