Abstract
The current pandemic caused by the SARS CoV-2, tracing back its origin possibly to a coronavirus associated with bats, has ignited renewed interest in understanding zoonotic spillovers across the globe. While research is more directed towards solving the problem at hand by finding therapeutic strategies and novel vaccine techniques, it is important to address the environmental drivers of pathogen spillover and the complex biotic and abiotic drivers of zoonoses. The availability of cutting-edge genomic technologies has contributed enormously to preempt viral emergence from wildlife. However, there is still a dearth of studies from species-rich South Asian countries, especially from India. In this review, we outline the importance of studying disease dynamics through environmental sampling from wildlife in India and how ecological parameters of both the virus and the host community may play a role in mediating cross-species spillovers. Non-invasive sampling using feces, urine, shed hair, saliva, shed skin, and feathers has been instrumental in providing genetic information for both the host and their associated pathogens. Here, we discuss the advances made in environmental sampling protocols and strategies to generate genetic data from such samples towards the surveillance and characterization of potentially zoonotic pathogens. We primarily focus on bat-borne or small mammal-borne zoonoses and propose a conceptual framework for non-invasive strategies to tackle the threat of emerging zoonotic infections.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
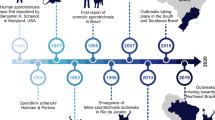
Over the past few decades, the world has encountered several zoonotic spillovers beginning with multiple Influenza outbreaks in the 1900s up to the recent SARS-CoV-2 outbreak in 2019. The World Health Organization (WHO), Food and Agriculture Organization (FAO) of the United Nations, and World Organization for Animal Health (OIE) have identified the increasing incidence of emerging of infectious diseases (EID) capable of transmission from animals to humans (zoonoses) as a major threat to biosecurity and public health1, 2 (Fig. 1). Zoonotic spillovers, which constitute more than 60% of all EIDs3, often involve three primary components, namely, animals, humans and their immediate environment (Fig. 1). The consequences of these spillovers have proven to be devastating for environmental, social, economic, and political systems. The same has been demonstrated through the Ebola virus outbreak of 2014 and the recent SARS-CoV-2 outbreak of 2019, resulting in an economic loss of approximately 28 trillion USD in 2019–20204, 5. However, we are still at a nascent stage when it comes to understanding the diversity of potential zoonotic pathogens and the causes of their spillovers. A more rigorous evaluation of these aspects is critical to preempt future spillover incidents and install appropriate preventive measures. Modern sequencing technologies have contributed immensely to pathogen discovery and are increasingly being used not only towards surveillance for zoonotic pathogens, but also towards a more holistic understanding of disease biology and host–pathogen coevolution6. The purpose of this review is to discuss the role of recent developments in sequencing techniques for pathogen discovery, particularly concentrating on the development of non-invasive environmental sampling and its applications thereof (Fig. 2).
Bats serve as potential reservoirs for a wide variety of viruses across the globe. Along with the plausible cross-species transmission, they form the connecting link between different zoonotic spillovers. A In India and Bangladesh, bat urine and feces contaminate the date palm juice with Nipah virus thereby exposing the palm juice handlers to deadly pathogens. B In Southeast Asia, the same virus passes through an amplification host, i.e., pigs and transmit it to humans, ultimately leading to human–human transmission through aerosols. C In Australia, Hendra virus has been isolated from bats and has caused outbreaks in horses, sometimes further leading to human fatalities. D In Africa, despite being detected in bats and non-human primates, the theories on the origin of Ebola virus remain to be deciphered. The red ticks denote points, where non-invasive sampling can be instrumental in revealing the pathogen load and help inform public health professionals to take necessary interventions.
The mechanistic processes involved in pathogen spillover from zoonotic reservoirs remain a poorly understood phenomenon. Wolfe et al.7 proposed a multistage model of how zoonotic microbes become specialized human pathogens. In the initial stages of this transformation, one may document microbes to be restricted to specific animal groups excluding humans. The consecutive stages of transformation may involve primary transmission between animals and humans, followed by occasional or continuous human to human secondary transmission events7. Infections caused by the West Nile virus or Dengue virus are ideal examples of the presence of a sylvatic cycle (transmission cycle in the wild) involving transmission of microbes between zoonotic reservoirs and humans in the wild followed by repeated secondary transmissions between humans8. The final stage comprises pathogens causing diseases exclusively to humans as seen in the case of measles, mumps, and rubella7. Such a mechanism has evolved either through co-specialization events between microbes and host or through a transformation event from an animal-associated microbe to a specialist human microbe7. Owing to the low fidelity of RNA-dependent RNA polymerase, RNA viruses exhibit a high mutation rate, raising the possibility of altering host tropism and giving room for successful cross-species transmissions to take place further. The transmission of HIV from non-human primates to humans is one of the classical examples that shows how human–wildlife interactions shape disease dynamics9. Similarly, genomic, and serological evidence hints at bats serving as potential reservoirs of Ebola virus. Transmission cycle of Ebola virus between chimpanzees and humans in Africa has been well established. However, involvement of many intermediate hosts during spillover events may further complicate our understanding of the origin of the pathogen10,11,12.
Essentially, reservoirs are the habitats in which the infectious agent lives, grows and multiplies13. Identifying such reservoir species is beneficial to managing emerging infections14. There has been sufficient evidence that a larger population size of the reservoir species is associated with a higher diversity of viruses15, placing small mammals such as bats and rodents at a critical position to serve as reservoirs. The wider range of these species also correlates to a higher chance of cross-species spillovers. Increased contact rate between individual animals ultimately increases the risk of pathogen transmission to humans16. However, the need for rapid and reliable detection along with subsequent characterizations of novel microbes, particularly in the wild, remains one of the major challenges while studying zoonotic spillovers.
2 Approaches to Detect Pathogens from Wildlife
2.1 Classical Approaches
Conventionally, several invasive methods have been employed for the detection of pathogens in wildlife. Serological methods such as ELISA have enabled the detection of antibodies in blood samples, shedding light on the recent or past exposure of the host to a specific infectious agent17. Targeted PCR approaches, classical methods involving isolation of bacteria in culture media or of viruses using embryonated eggs, cell lines and cutting-edge genomics have significantly improved the chances of pathogen discovery18. Detection of genomic fragments in clinical samples from humans or wildlife, using unique pathogen markers can help decipher whether the infection is acute. Serology is useful in estimating the infection history, especially in cases, where the pathogen titre may be significantly low as in the case of Henipa virus infections in bats19, or if the infection is latent as applied to pseudorabies in wild swine20. However, cross reactivity of antibodies to related etiological agents and determination of an optimum threshold to distinguish between positive and negative reactions can result in incorrect estimates of antibody prevalence21. For pathogens circulating in blood or excreted through feces and urine, detection of genomic fragments or antigen is possible but the sensitivity of PCR-based approaches can affect the detection probability. Finally, development of primary bat cell lines for isolation of bat viruses and subsequent host-virus studies is lagging, because bats are enormously diverse, and a given species-specific virus may not infect cells obtained from another species. Controlling optimum temperatures and other physiological parameters resembling that of a natural reservoir as well as allowing efficient cell culture can also be challenging22. While performing blind passage of viruses in cell lines, one may encounter adaptive mutations which attenuates the virus strain as seen during the isolation of Tacaribe virus from Artibeus bats, which acquired point mutations after 20 passages in newborn mice22.
One of the major challenges in formulating novel diagnostic assays for wildlife is attributed to the limitations imposed by smaller sample size and the challenges of test validation. One may encounter poor or degraded nucleic acid in samples, inaccessibility of post-mortem samples, and issues with the capture–recapture of infected individuals in the wild. Long-term monitoring of susceptible individuals in the wild is extremely difficult, further raising two issues: (a) stages of disease progression and pathogen shedding cannot be studied longitudinally (b) in vitro molecular diagnostics developed in the lab may not necessarily reflect observations in the wild, because community interactions amongst species in the wild is unpredictable, making pathogen transmission dynamics even more complex. One must be careful while validating data on physiological stress markers which are potentially associated with any infection that may have a different profile for captive animals in comparison to free ranging ones23, 24. This also holds true for interpreting any stress or immunological profile through serum biomarkers during experimental infection studies.
2.2 The Need for Metagenomics
The gold standard for establishing the true association of an etiological agent with diseases is the isolation and culturing of microbes, followed by validation in accordance with Koch's postulates. There have been exceptions to this rule too; for instance, viruses could not be grown under in vitro conditions for the longest time. Pathogen discovery from wildlife has faced several setbacks, some of the factors contributing to the setback are the specific growth requirement of fastidious microbes, lack of specific cell lines, ecological dependencies on other microbial or non-microbial partners, and lack of pathogen-specific identification markers. The major problem associated with PCR-based surveillance is the lack of prior knowledge of the microbial entity.
Less than 2% of the total microbes can be cultured25. Thus, metagenomics has served as a boon to study the unculturable microbes in nature26. The evolution and diversity of microbial communities in which individual microbial species are strongly dependent on each other’s function in a given habitat, can be studied through the lens of metagenomics. It is a powerful unbiased tool which has helped in a very detailed investigation of the human microbiome27 and similar approaches could be employed to microbiome studies in the wild. Metagenomics has helped to acquire genetic data on the host as well, suggesting better theories on how individual differences can shape the virome diversity. A recent report by Brussel and Holmes shows that 30% of the bat virus discovery has been achieved with the aid of next-generation sequencing tools28, and metagenomics has become an important tool to identify and characterize pathogens of zoonotic potential in the wild29. For example, metagenomics contributed significantly towards the discovery of the SARS coronavirus30. Malayan pangolin coronaviruses which show high similarity in their receptor binding domain to that of SARS-CoV-2 were identified with the aid of metagenomic sequencing31.
2.3 Metagenomics Workflow
The workflow of any metagenomic study starts from sample collection, followed by extraction of nucleic acid, library preparation, sequencing and bioinformatic analyses. One of the major challenges in wildlife viromics is the computational complexity of analyzing vast amounts of data generated through next-generation sequencing platforms. Metagenomics can help to understand the virome composition in reservoirs. However, one should be careful while interpreting which pathogens persist for a longer period in reservoirs against certain microbes which are merely a component of diet. Bats show the presence of insect viruses and bacteriophages which are predominantly acquired through diet32. Whole-genome sequencing of microbes is extremely expensive. Therefore, amplicon sequencing based on the 16S rRNA gene is extensively used to determine taxonomic identities in a microbial community as it is evolutionarily stable. Amplicon sequence variants provide higher resolution data of 16S sequences. As opposed to all these technologies, metagenomics or whole microbiome/virome shotgun sequencing involves the amplification of all the DNA/RNA within a sample, increasing the probability of detecting any and every microbe present in the study system33. However, few challenges of using metagenomics for establishing the etiology of a disease can be posed by the limited access of such technologies in developing or under-developed countries. Depletion of the host DNA with an aim to enrich the microbial genomes, the lack of standardized laboratory protocols of extracting genomic material from varied sources, contamination and working complexity associated during library preparation are some of the major challenges of implementing metagenomics approaches34.
The DNA or RNA sequence can be used to extract complete or partially assembled genomes of pathogens and further employ genome annotation studies to predict the virulence and understand their molecular evolution. Deep sequencing of samples potentially containing diverse bacterial communities is based on consensus gene sequence, i.e., 16S ribosomal RNA. On the other hand, viral communities do not have any such conserved sequence. Hence, discovery of viruses rely majorly on metatranscriptomics for RNA viruses or metagenomics for DNA viruses. However, many studies have identified viruses based on family-specific gene sequences of RNA-dependent RNA polymerase or DNA-dependent DNA polymerase. Isolation of viruses has also been successful from urine or fecal samples of wildlife. Paramyxoviruses were isolated from urine samples of Pteropus species35 and close relatives of the SARS coronavirus from Chinese horseshoe bats36.
There have been issues related to the nature of the buffer to be used for collection of environmental samples. Majority of the studies utilize RNA LATER. for preserving the samples which prevents RNA degradation, but it is ideal to store samples in RNA shield buffer, with the aim to inactivate viruses at its source. Storage of the environmental samples in Viral Transport Media (VTM) gives an opportunity to isolate viruses by maintaining its viability and enables further study of the agent through molecular virological approaches.
2.4 Novel Approaches
To make pathogen quantification reliable, more sensitive, and specific, digital droplet PCR (ddPCR) is an effective tool for samples which have low abundance of nucleic acids. The costs associated with ddPCR is high, but the current need is to make it affordable and accessible to a wide range of labs globally. ddPCR helps in accurate and faster quantification of molecules, having each molecule in one microdroplet. ddPCR involves individual PCR reactions after the initial template has been diluted in individual wells and then enumerates the pathogen load based on PCR positive percentage among all reactions. Using 180 fecal samples, the detection of human-associated Bacteroidales genetic markers, BacHum and B. theta was demonstrated employing dd-PCR37, 38. Detection of HIV, Hepatitis B, Human Herpes virus and malarial parasite is also possible using ddPCR and blood samples; however, this method remains to be explored in the context of environmental sampling of wildlife, as sufficient information about zoonotic pathogens in wild animals is lacking.
Once discovered as adaptive immune systems in bacteria against phages, Microbial Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) and CRISPR-associated (CRISPR-Cas) programmable endonucleases, the genome editing tool have also found its place in diagnosing emerging infectious diseases. SHERLOCK strategy It can be a revolutionary approach in rapid detection of EIDs in the wild as it contributed to the detection of attomolar concentrations of Dengue and Zika virus as well as pathogenic bacteria, such as drug resistant Klebsiella pneumoniae39. It combines the properties of RNA guided RNA detection, promiscuous activity of Cas 13a and isothermal amplification40. Lyophilization of SHERLOCK reagents enabling easy storage, reconstitution on paper-based strip test enabling portability to the field, costs going as low as $0.61/test, applicability to a broad sample range including urine and sera are some of the reasons that make it ideal for environmental sampling especially for the developing countries40. In addition, CRISPR-based methodologies have made it feasible to detect deadly hemorrhagic fevers caused by Ebola and Lassa virus. A modified protocol of the SHERLOCK strategy, called HUDSON, has made it possible to detect BSL-4 viruses without involving any nucleic acid extraction step through the heat inactivation step with TCEP: EDTA directly from non-invasive samples, such as saliva and urine41. The same could be employed for surveillance of bat roosts and rodent samples. CRISPR-based lateral flow assays have also become handy in detecting SARS-CoV-2 from RNA extracted from throat swabs42.
Among others, a rapid and robust approach utilizing high-throughput targeted proteomics assay, towards the clinical diagnostics of viruses such as the novel coronavirus (SARS-CoV-2) is of significant value. Coupling two techniques, turbulent flow chromatography and tandem mass spectroscopy (TFC–MS/MS), such a diagnostic test proves to have specificity towards the detection of SARS-CoV-2, while maintaining peptides from other coronavirus strains, rhinovirus, enterovirus, and influenza viruses undetected. With a higher analyte stability and sensitivity, this strategy proves to be more advantageous than the gold standard real-time RT-PCR. In situations which demand large-scale population testing in a shorter duration and the supply for real-time RT-PCR equipment is insufficient, targeted proteomics provides an alternative strategy to real-time RT-PCR and would be an interesting addition to the viral testing panels43.
The use of biosensors also offers the possibility and potential as a rapid, sensitive, and specific detection platform scalable to global surveillance of pathogens and their control. It does not rely on cold chain storage and can aid in sample detection without additional equipment44, 45.
Ideally, a combination of targeted approaches and metagenomics, complementing the drawbacks of each other should be adapted and brought to the forefront of screening potential pathogens, driving advances in the prediction and control of zoonotic spillovers.
3 Environmental Sampling: Non-invasive Strategies
Environmental sampling refers to the isolation, extraction, and identification of DNA from an environmental sample, such as skin, mucous, saliva, urine, feces, leaves, secretions, water, soil, and rotting bodies. It also includes sampling of abiotic sources, potentially fomites, supplementing an investigation of an outbreak when environmental reservoirs are implicated epidemiologically in disease transmission46 (Fig. 1). During the last few decades, environmental DNA (eDNA) sampling has enhanced our abilities to identify a plethora of pathogenic organisms from the wildlife and predict zoonotic disease and outbreaks, thus aiding in disease preparedness. It has proven to be a potential tool in the detection of common species, along with those which are endangered, invasive, or elusive, and their microflora47. This non-invasive approach is significant in monitoring wildlife diseases and detecting infectious agents as eDNA allows one to sequence information obtained from the environment without encountering the target species and minimizes disruption of already fragile habitats48. It neither utilizes any on-site biosafety equipment which would have been required for dissection or blood sample collection nor is prior monitoring required to localize ideal trapping sites, such as in invasive sampling, thus making feces, urine and saliva an ideal source to understand microbial diversity in the wildlife49. In addition to being less labor-intensive and cost-effective, the process is rapid, allowing multiple microbial species detection. Furthermore, even certain infectious diseases in the wildlife which may cause extinction or loss of biodiversity of species, thereby impacting the ecological balance, can be detected, and managed at an early stage of disease spread. A quantitative analysis of the contents of eDNA aids in estimating the relative prevalence of microbes in natural systems besides its mere presence or absence.
eDNA sampling has been rendered as a vital tool for the early detection of infectious agents at a low density, which improves the success of pathogen eradication and decreases the costs of control and impact on public health. There have been enough examples to prove the use of eDNA as a promising non-invasive and sensitive tool for pathogenic surveillance in wildlife. For instance, non-invasive collection of feces in the wild is one of the most economical, non-hazardous ways of gathering information on the host taxonomy, their population structure as well as understanding the shedding patterns of various pathogens in the environment50 (Fig. 2). Recently, there has been much focus on the surveillance of SARS-CoV-2 in wastewater, relevant for early monitoring and subsequent measures that can be taken by government authorities in treating wastewater51.
Thus, in the coming years, a well-rounded network of eDNA surveillance can be envisioned to understand host–pathogen dynamics, transitioning eDNA from an emerging field to one at the forefront of disease transmission and public health.
There are several biases involved in non-invasive sampling. While implementing a pooled sampling strategy, where individual fecal pellets cannot be differentiated from one another. For instance, collecting bat guano from a cave or social animals performing communal desiccation, it is difficult to identify the infection status at an individual level. One practical solution is to investigate pooled sampling from a collection of individuals. Even while estimating the abundance, considering that eDNA has variable decay rates under different environmental conditions52, 53, it will result in biased estimates and hence be a limitation to this approach. Therefore, one should ideally collect fresh fecal samples and subject them to freezing temperatures immediately to investigate the pathogen diversity.
Giles et al.54 attempted to optimize non-invasive or under roost sampling of bat-borne viruses. Despite many research groups looking at the viral or other microbial diversity using bat guano and urine samples in Asia, Australia, and Africa; this was the first study presenting a theoretical model of what an ideal strategy for sampling under roost would look like and the various sampling biases one may encounter while calculating the actual prevalence of viruses from such field data (Fig. 2). As opposed to the approach, wherein the presence–absence data of diseases is tabulated from pooled samples and then tracking individual level data from the positive pools to estimate the true prevalence; the authors presented a theoretical model of an ideal sampling strategy and showed that a small-sheet design, increased number of sheets spread across the roost area, combined with ‘multi-stage’ pooling of urine at a spatial scale offers a better alternative to estimate virus presence both with high sensitivity and specificity54, 55. Such sample pooling methods prove to be advantageous by minimizing the number of tests, time and costs56. The same has found its applications in recent times as well. While the testing for SARS-CoV-2 using the RT-PCR tests has gained an overwhelming response and has now become a bottleneck, sample pooling strategies are an attractive alternative that increases testing throughput while maintaining high sensitivity57. While pooling of samples, as well as sample libraries for NGS platforms, uneven mixing of pools, may give a low number of sequencing reads58. Furstenau et al. provided a detailed overview of sample pooling to screen pathogens and has concluded that despite the merits, it is important to consider that choosing an optimized pooling approach is often influenced by the epidemiology of the infectious agent, availability of the specimen, and the accuracy and sensitivity of the assay56.
Considering the current advancement in sampling strategies, we have proposed various strategies which can be employed for environmental sample collection (Fig. 2). Plastic sheets can be spread across a larger area for under roost sampling of fresh bat guano samples and urine droplets can be pipetted out in screw cap vials containing RNA/DNA shield buffer and transported to investigation laboratories, maintaining a proper cold chain and three-layered packaging. Alternatively, one could also collect fecal pellets of small mammals such as rodents or birds along with sample collection of nearby abiotic environments to serve as appropriate controls, accounting for any environmental contamination. As far as companion animals and wet markets are concerned, rectal and oral swabs can be taken easily. However, standardization of protocols to collect saliva from potentially infected fruits and investigate the virome or microbiome in the same remains to be explored. Water bodies and ticks in the wild can provide a broader perspective of the different microbes co-circulating in the wildlife and can serve as alternate sources to track the different pathogens that can spread through the feco-oral route.
While developing countries in South Asia and Southeast Asia are known for their high population density, increased wildlife habitat fragmentation and innumerable human–wildlife interactions, they are not yet adequately equipped with facilities to carry out long-term disease surveillance studies, particularly in wildlife. Having 125 reference laboratories certified by World Organization for Animal Health (OIE) to screen one or more pathogen, a disproportionality exists in the distribution of the reference laboratories, with almost 62% of these labs being in Europe and North America and the remaining labs being located mostly in China, while some of them being dispersed across Australia, South America, Africa and Japan59. Moreover, places that serve as potential zoonotic hotspots require the involvement of local communities skilled in lab safety, biosafety and having sufficient knowledge such that they contribute to the efforts of estimating the incidence and prevalence of zoonotic diseases. Non-invasive sampling can play a significant role in developing countries for pathogen bio surveillance studies. Furthermore, training of local communities to collect environmental samples of urine, feces, or oral swabs of dead animals along with individuals equipped with ‘on field’ genomic technologies and subsequently proper communication with regional laboratories can aid in faster epidemiologic analyses and inform outbreak management teams.
4 Small Mammals are Important!
The two most diverse groups of mammals, bats and rodents, account for an overwhelming proportion of pathogens with zoonotic potential and have been implicated in numerous zoonotic outbreaks14. Order Chiroptera within mammals has been associated with deadly, high-risk pathogens, such as Ebola virus and Nipah virus (Fig. 1). A similar trend has also been observed for Order Rodentia, which constitutes the largest percentage of living mammals in the world60. Hantavirus Pulmonary syndrome, Lassa fever, Plague, and Leptospirosis are caused by rodents61. Given the enormous species diversity, it is expected that the associated microbiome and virome for these species would also be diverse. Bats and rodents, also known to be synanthropes, have eventually become accustomed to artificial habitats created by humans62. Rapid globalization, urbanization, anthropogenic activities and increase in wildlife–human interactions have provided ample opportunities for spillovers to take place. There can be direct and indirect ways of transmitting pathogens from wildlife to humans either mediated through (a) consumption of contaminated bushmeat, (b) inhalation of rodents/ bat excreta aerosols, (c) direct contact with pathogens shed on animate household objects or during activities such as farming and camping, (d) intermediate hosts such as pigs and horses, serving as amplifying hosts. Intermediate hosts play a significant role due to their close association with humans, primarily in animal farms or being consumed as a nutrient source63. In addition, many cultural practices can expose humans to pathogens of zoonotic origins. For instance, the presence of filovirus surface protein reactive antibodies in bat harvesters of Mimi village of Nagaland, India, indicated possible exposure to filoviruses, possibly from the bat’s meat obtained during the bat harvests64. Behaviorally, bats are known to chew fruits, leaves, insects, followed by the leftover chewed parts of the food fallen on the ground becoming infected with viruses from the saliva of bats. Other animals such as palm civets (potential intermediate hosts for SARS) and swine consume these contaminated fruits and carry forward the transmission cycle of the virus65. The Nipah virus outbreak in Malaysia, Ebola virus outbreak in Africa and Hendra virus outbreak in Australia involve an intermediate host and are associated with bat reservoirs through a similar transmission route66 (Fig. 1). On the contrary, it has also been observed that the Nipah virus outbreak in Bangladesh involves the direct transmission of viruses from bats to humans through the consumption of contaminated date palm juice67 (Fig. 1). Bats are also long-lived and the only volant mammals with a unique immune system which restricts viruses from showing overt pathologies while persistently evolving within each individual bat and dispersing across wide spatial ranges through their excreta, saliva, and interactions during movement68.
Similar to bats, rodents also play a key role as reservoirs for many medically important pathogens, such as Hantavirus, Arenavirus, Bartonella, Leptospira, Yersinia pestis69, 70. ‘Anthropocene defaunation’—owing to globalization, has reduced the diversity of larger mammals and consequently an increase in rodent population. This further increases the probability of rodent borne spillovers in the future.
5 Environmental Sampling, Metagenomics, and Reservoirs—Foundation Pillars for Disease Surveillance
While we talk about the increasing number of infectious diseases, it is crucial to tap into the microbiome and virome of wildlife (a) to understand the ecological parameters driving the evolution and functional importance of host associated microbes, and (b) to investigate the trends in emergence of novel pathogens or increased shedding rates of previously well-characterized pathogens. However, a major drawback is the lack of baseline data on diversity and prevalence of microbes in wildlife, particularly small mammals, such as bats and rodents. The virome refers to the collection of all viruses present in any organism71. The Global Virome project was launched with an aim to create an atlas of all the viruses that are present in the world, especially in low income or middle-income countries72. The economic losses associated with managing an outbreak outnumbers the amount of investment that would facilitate research in understanding the diversity of viruses which are present naturally in different biological systems. Investments in viral discovery can aid the Global Preparedness Monitoring Board (GPMB) to take necessary measures to mitigate pandemic risks. Such global projects would foster collaborations between countries and initiate exchange of ideas to solve gray areas in the domain of emerging infectious diseases.
Non-invasive sampling of wildlife, especially with a concentrated focus on small mammals such as bats and rodents can mediate important signals prior to any outbreak. Table 1 summarizes the wide range of questions explored using environmental sampling, in the context of reservoirs and infectious agents.
6 The Future
We propose a very detailed investigation approach and strongly recommend coordinated and collaborative efforts comprising epidemiologists, evolutionary biologists, and infectious disease experts to unravel the mystery of zoonotic spillovers for the future (Fig. 3). Currently, in South and Southeast Asian countries, advanced biosafety level facilities are present in restricted numbers. The ecological aspects of any wildlife associated disease have been neglected globally and such studies have almost been negligible in South Asia. While being widely known for its rich biodiversity and population explosion, India can potentially serve as an ideal model system to study cross-species transmission of pathogens (Fig. 4).Through this review, we looked at the wide range of questions one can address simply with the help of non-invasive sampling. We envision environmental sampling to be the ideal solution for informing public health professionals about any unnatural incidence of a disease. Early and rapid monitoring of any emerging infectious agent can help better prepare for the outbreak. Understanding what cues lead to pathogen spillover can help prevent the same. Investigation of genomic data from the field and various mathematical projections can help quantitate the disease burden and predict the pathogenicity of any EIDs.
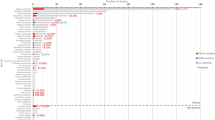
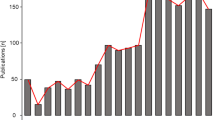
BatVir is a database software for all the bat viruses reported across the world. In this graphical representation, we show that there have been very few studies in India which have investigated viruses from bat reservoirs. No virome study has been done in India, which points out at the urgency of such studies in the country.
The regional laboratories should be well equipped with state-of-the-art genomics facilities and should be able to carry out serosurveys (Fig. 3). However, the entire process of disease investigation must begin with environmental sampling at the field laboratories (Fig. 2). There should be dedicated teams to collect not only non-invasive samples from the wild, but also implement a routine screening of fecal, urine and saliva samples from local animal farms. Farfetched as it is, companion animals should also be screened monthly as they are the ones in closest proximity with humans. Investments need to be made enormously to improve the genomics facilities at the district level. Decentralizing surveillance efforts will make the detection of infectious agents much faster. Regional and District level laboratories should be able to carry out functional genomics studies independently and rely on the apex wildlife disease investigation lab only when the isolation of pathogens is required (Fig. 3). The idea of having one apex laboratory in India should not be encouraged; alternatively, five to six apex laboratories spanning the entire country would be significantly more beneficial. This would again be instrumental in reducing the workload of apex laboratories and prompt outbreak decisions at an early stage.
Discussions, collaborations, research, and outreach are the four foundational pillars that can help in policy making towards preventing zoonotic spillovers. Educating the young generation and engaging local communities can significantly reduce misinformation regarding ecologically important disease reservoirs, such as bats. Adding information about zoonoses in school curricula can generate interest in studying the field and hopefully disseminate the knowledge in their families who may be involved in professions (for instance—date palm sap handlers), where there is a high risk of getting exposed to pathogenic organisms. A major challenge is the time span in which we wish to see the change as age-old beliefs are deep-rooted in many societies. Model predictions and knowledge of bat or rodent diversity in various habitats is necessary to predict the hotspots of spillovers. A well-coordinated disease surveillance effort (Fig. 3) will certainly attract many non-governmental funders, providing more resources. Sharing of information and knowledge regarding any EID is of utmost importance within and outside the research community and calls for a pan-India virome project venture. Finally, pilot environmental sampling studies should be initiated using the current resources available in India which can lay down the proof of concept for any long-term disease monitoring efforts. With such information in hand, it will be possible to tackle the threat of emerging zoonotic spillovers, which in turn will shape the future of public health.
References
Plowright RK, Parrish CR, McCallum H, Hudson PJ, Ko AI, Graham AL, Lloyd-Smith JO (2017) Pathways to zoonotic spillover. Nat Rev Microbiol 15:502–510
World Health Organization (2004) Report of the WHO/FAO/OIE joint consultation on emerging zoonotic diseases / in collaboration with the Health Council of the Netherlands. World Health Organization. https://apps.who.int/iris/handle/10665/68899
Jones KE, Patel NG, Levy MA, Storeygard A, Balk D, Gittleman JL, Daszak P (2008) Global trends in emerging infectious diseases. Nature 451(7181):990–993
Sánchez CA, Venkatachalam-Vaz J, Drake JM (2021) Spillover of zoonotic pathogens: a review of reviews. Zoonoses Public Health 68(6):563–577
International Monetary Fund (IMF), World Economic Outlook (2020) A long and difficult ascent. International Monetary Fund, Washington, DC, p 2020
Näpflin K, O’Connor EA, Becks L et al (2019) Genomics of host-pathogen interactions: challenges and opportunities across ecological and spatiotemporal scales. PeerJ 7:e8013
Wolfe ND, Dunavan CP, Diamond J (2007) Origins of major human infectious diseases. Nature 447(7142):279–283
Young PR (2018) Arboviruses: a family on the move. Adv Exp Med Biol 1062:1–10. https://doi.org/10.1007/978-981-10-8727-1_1
Sharp PM, Hahn BH (2011) Origins of HIV and the AIDS pandemic. Cold Spring Harb Perspect Med 1(1):a006841
Leroy EM, Kumulungui B, Pourrut X et al (2005) Fruit bats as reservoirs of Ebola virus. Nature 438(7068):575–576
Heeney JL, Dalgleish AG, Weiss RA (2006) Origins of HIV and the evolution of resistance to AIDS. Science 313(5786):462–466
Gire SK, Goba A, Andersen KG et al (2014) Genomic surveillance elucidates Ebola virus origin and transmission during the 2014 outbreak. Science 345(6202):1369–1372
Centers for Disease Control and Prevention (2012) Principles of epidemiology. Centers for Disease Control and Prevention. https://www.cdc.gov/csels/dsepd/ss1978/lesson1/section10.html. Accessed 20 January 2022
Luis AD, Hayman DT, O’Shea TJ et al (2013) A comparison of bats and rodents as reservoirs of zoonotic viruses: are bats special? Proc Bol Sci 280(1756):20122753
Patterson JE, Ruckstuhl KE (2013) Parasite infection and host group size: a meta-analytical review. Parasitology 140(7):803–813
Rohr JR, Barrett CB, Civitello DJ et al (2019) Emerging human infectious diseases and the links to global food production. Nat Sustain 2(6):445–456
Aggarwal R, Jameel S (2011) Hepatitis E. Hepatology 54(6):2218–2226
Leland DS, Ginocchio CC (2007) Role of cell culture for virus detection in the age of technology. Clin Microbiol Rev 20(1):49–78
Halpin K, Hyatt AD, Fogarty R et al (2011) Pteropid bats are confirmed as the reservoir hosts of henipaviruses: a comprehensive experimental study of virus transmission. Am J Trop Med Hyg 85(5):946–951
Wittmann G, Rziha HJ (1989) Aujeszky’s disease (pseudorabies) in pigs. In: Wittmann G (ed) Herpesvirus diseases of cattle, horses, and pigs. Springer, Boston, MA, pp 230–325
Gilbert AT, Fooks AR, Hayman DT et al (2013) Deciphering serology to understand the ecology of infectious diseases in wildlife. EcoHealth 10(3):298–313
Banerjee A, Misra V, Schountz T, Baker ML (2018) Tools to study pathogen–host interactions in bats. Virus Res 248:5–12
Michel AL, Lane EP, de Klerk-Lorist LM, Hofmeyr M, van der Heijden EMDL, Botha L, van Helden P, Miller M, Buss P (2017) Experimental Mycobacterium bovis infection in three white rhinoceroses (Ceratotherium simum): susceptibility, clinical and anatomical pathology. PLoS ONE 12(7):e0179943
Dickens MJ, Delehanty DJ, Romero LM (2009) Stress and translocation: alterations in the stress physiology of translocated birds. Proc Biol Sci 276(1664):2051–2056
Wade W (2002) Unculturable bacteria—the uncharacterized organisms that cause oral infections. J R Soc Med 95(2):81–83
Mokili JL, Rohwer F, Dutilh BE (2012) Metagenomics and future perspectives in virus discovery. Curr Opin Virol 2(1):63–77
Handelsman J (2004) Metagenomics: application of genomics to uncultured microorganisms. Microbiol Mol Biol Rev 68(4):669–685
Palmer C, Bik EM, Eisen MB, Eckburg PB, Sana TR, Wolber PK, Relman DA, Brown PA (2006) Rapid quantitative profiling of complex microbial populations. Nucleic Acids Res 34(1):e5
Van Brussel K, Holmes EC (2021) Zoonotic disease and virome diversity in bats. Curr Opin Virol 52:192–202
Rota PA, Oberste MS, Monroe SS et al (2003) Characterization of a novel coronavirus associated with severe acute respiratory syndrome. Science 300(5624):1394–1399
Lam TTY, Jia N, Zhang YW, Shum MHH et al (2020) Identifying SARS-CoV-2-related coronaviruses in Malayan pangolins. Nature 583(7815):282–285
Cobbin JC, Charon J, Harvey E, Holmes EC, Mahar JE (2021) Current challenges to virus discovery by meta-transcriptomics. Curr Opin Virol 51:48–55
New FN, Brito IL (2020) What is metagenomics teaching us, and what is missed? Annu Rev Microbiol 74:117–135
Chiu CY, Miller SA (2019) Clinical metagenomics. Nat Rev Genet 20(6):341–355
Barr J, Smith C, Smith I, de Jong C, Todd S, Melville D, Broos A, Crameri S, Haining J, Marsh G, Crameri G (2015) Isolation of multiple novel paramyxoviruses from pteropid bat urine. J Gen Virol 96(1):24–29
Ge XY, Li JL, Yang XL, Chmura AA, Zhu G, Epstein JH, Mazet JK, Hu B, Zhang W, Peng C, Zhang YJ et al (2013) Isolation and characterization of a bat SARS-like coronavirus that uses the ACE2 receptor. Nature 503(7477):535–538
Nshimyimana JP, Cruz MC, Wuertz S, Thompson JR (2019) Variably improved microbial source tracking with digital droplet PCR. Water Res 159:192–202
Li H, Bai R, Zhao Z et al (2018) Application of droplet digital PCR to detect the pathogens of infectious diseases. Biosci Rep 38(6):BSR20181170
Myhrvold C, Freije CA, Gootenberg JS, Abudayyeh OO et al (2018) Field-deployable viral diagnostics using CRISPR-Cas13. Science 360(6387):444–448
Gootenberg JS, Abudayyeh OO, Lee JW et al (2017) Nucleic acid detection with CRISPR-Cas13a/C2c2. Science 356(6336):438–442
Barnes KG, Lachenauer AE, Nitido A et al (2020) Deployable CRISPR-Cas13a diagnostic tools to detect and report Ebola and Lassa virus cases in real-time. Nat Commun 11(1):1–10
Broughton JP, Deng X, Yu G et al (2020) CRISPR-Cas12-based detection of SARS-CoV-2. Nat Biotechnol 38(7):870–874
Cardozo KHM, Lebkuchen A, Okai GG, Schuch RA, Viana LG, Olive AN, Lazari CDS, Fraga AM, Granato CFH, Pintão MCT, Carvalho VM (2020) Establishing a mass spectrometry-based system for rapid detection of SARS-CoV-2 in large clinical sample cohorts. Nat Commun 11(1):6201
Jiang L, Luo J, Dong W, Wang C, Jin W, Xia Y, Wang H, Ding H, Jiang L, He H (2015) Development and evaluation of a polydiacetylene based biosensor for the detection of H5 influenza virus. J Virol Methods 219:38–45
Ostrov N, Jimenez M, Billerbeck S, Brisbois J, Matragrano J, Ager A, Cornish VW (2017) A modular yeast biosensor for low-cost point-of-care pathogen detection. Sci Adv 3(6):e1603221
Lee SE, Lee DY, Lee WG, Kang B, Jang YS, Ryu B, Lee S, Bahk H, Lee E (2020) Detection of novel coronavirus on the surface of environmental materials contaminated by COVID-19 patients in the Republic of Korea. Osong Public Health Res Perspect 11(3):128–132
Thomsen PF, Willerslev E (2015) Environmental DNA—an emerging tool in conservation for monitoring past and present biodiversity. Biol Conserv 183:4–18
Aylward ML, Sullivan AP, Perry GH, Johnson SE, Louis EE Jr (2018) An environmental DNA sampling method for aye-ayes from their feeding traces. Ecol Evol 8(18):9229–9240
Mengüllüoğlu D, Fickel J, Hofer H, Förster DW (2019) Non-invasive faecal sampling reveals spatial organization and improves measures of genetic diversity for the conservation assessment of territorial species: Caucasian lynx as a case species. PLoS ONE 14(5):e0216549
Dib LV, Palmer JP, de Souza Carvalho Class C, Pinheiro JL, Ramos RC, Dos Santos CR, Fonseca AB, Rodríguez-Castro KG, Gonçalves CF, Galetti PM, Bastos OM (2020) Non-invasive sampling in Itatiaia National Park, Brazil: wild mammal parasite detection. BMC Vet Res 16:295
Kitajima M, Ahmed W, Bibby K, Carducci A, Gerba CP, Hamilton KA, Haramoto E, Rose JB (2020) SARS-CoV-2 in wastewater: state of the knowledge and research needs. Sci Total Environ 739:139076
Foote AD, Thomsen PF, Sveegaard S, Wahlberg M, Kielgast J, Kyhn LA et al (2012) Investigating the potential use of environmental DNA (eDNA) for genetic monitoring of marine mammals. PLoS ONE 7(8):e41781
Bohmann K, Evans A, Gilbert MTP, Carvalho GR, Creer S, Knapp M, Yu DW, de Bruyn M (2014) Environmental DNA for wildlife biology and biodiversity monitoring. Trends Ecol Evol 29(6):358–367
Giles JR, Peel AJ, Wells K, Plowright RK, McCallum H, Restif O (2021) Optimizing noninvasive sampling of a zoonotic bat virus. Ecol Evol 11(18):12307–12321
Martin SW, Shoukri M, Thorburn MA (1992) Evaluating the health status of herds based on tests applied to individuals. Prev Vet Med 14(1–2):33–43
Furstenau TN, Cocking JH, Hepp CM, Fofanov VY (2020) Sample pooling methods for efficient pathogen screening: practical implications. PLoS ONE 15(11):e0236849
Ben-Ami R, Klochendler A, Seidel M, Sido T, Gurel-Gurevich O, Yassour M, Meshorer E, Benedek G, Fogel I, Oiknine-Djian E, Gertler A (2020) Large-scale implementation of pooled RNA extraction and RT-PCR for SARS-CoV-2 detection. Clin Microbiol Infect 26(9):1248–1253
Gu W, Miller S, Chiu CY (2019) Clinical metagenomic next-generation sequencing for pathogen detection. Annu Rev Pathol 14:319–338
Watsa M, Wildlife Disease Surveillance Focus Group (2020) Rigorous wildlife disease surveillance. Science 369(6500):145–147
Rabiee MH, Mahmoudi A, Siahsarvie R, Kryštufek B, Mostafavi E (2018) Rodent-borne diseases and their public health importance in Iran. PLoS Negl Trop Dis 12(4):e0006256
Centers for Disease Control and Prevention (2017) Diseases directly transmitted by rodents. Centers for Disease Control and Prevention. https://www.cdc.gov/rodents/diseases/direct.html. Accessed 21 January 2022
McFarlane R, Sleigh A, McMichael T (2012) Synanthropy of wild mammals as a determinant of emerging infectious diseases in the Asian-Australasian region. EcoHealth 9(1):24–35
Weaver SC, Barrett AD (2004) Transmission cycles, host range, evolution and emergence of arboviral disease. Nat Rev Microbiol 2(10):789–801
Dovih P, Laing ED, Chen Y et al (2019) Filovirus-reactive antibodies in humans and bats in Northeast India imply zoonotic spillover. PLoS Negl Trop Dis 13(10):e0007733
Dobson AP (2005) What links bats to emerging infectious diseases? Science 310(5748):628–629
Letko M, Seifert SN, Olival KJ, Plowright RK, Munster VJ (2020) Bat-borne virus diversity, spillover and emergence. Nat Rev Microbiol 18(8):461–471
Luby SP, Gurley ES, Hossain MJ (2009) Transmission of human infection with Nipah virus. Clin Infect Dis 49(11):1743–1748
Banerjee A, Baker ML, Kulcsar K, Misra V, Plowright R, Mossman K (2020) Novel insights into immune systems of bats. Front Immunol 11:26
Golden JW, Hammerbeck CD, Mucker EM, Brocato RL (2015) Animal models for the study of rodent-borne hemorrhagic fever viruses: arenaviruses and hantaviruses. Biomed Res Int 2015:793257
Mawanda P, Rwego I, Kisakye JJ, Sheil D (2020) Rodents as potential hosts and reservoirs of parasites along the edge of a Central African forest: Bwindi impenetrable national park, South Western Uganda. Afr Health Sci 20(3):1168–1178
Wylie KM, Weinstock GM, Storch GA (2012) Emerging view of the human virome. Transl Res 160(4):283–290
Carroll D, Watson TE et al (2018) Building a global atlas of zoonotic viruses. Bull World Health Organ 96(4):292
Bergner LM, Mollentze N, Orton RJ, Tello C, Broos A, Biek R, Streicker DG (2021) Characterizing and evaluating the zoonotic potential of novel viruses discovered in vampire bats. Viruses 13(2):252
Zhou H, Ji J, Chen X, Bi Y, Li J, Wang Q, Hu T, Song H, Zhao R, Chen Y, Cui M, Zhang Y, Hughes A, Holmes EC, Shi W (2021) Identification of novel bat coronaviruses sheds light on the evolutionary origins of SARS-CoV-2 and related viruses. Cell 184(17):4380-4391.e14
Mortlock M, Geldenhuys M, Dietrich M, Epstein JH, Weyer J, Pawęska JT, Markotter W (2021) Seasonal shedding patterns of diverse henipavirus-related paramyxoviruses in Egyptian rousette bats. Sci Rep 11(1):1–12
Ge X, Li Y, Yang X, Zhang H, Zhou P, Zhang Y, Shi Z (2012) Metagenomic analysis of viruses from bat fecal samples reveals many novel viruses in insectivorous bats in China. J Virol 86(8):4620–4630
Li Y, Ge X, Zhang H, Zhou P, Zhu Y, Zhang Y, Yuan J, Wang LF, Shi Z (2010) Host range, prevalence, and genetic diversity of adenoviruses in bats. J virol 84(8):3889–3897
Banskar S, Bhute SS, Suryavanshi MV, Punekar S, Shouche YS (2016) Microbiome analysis reveals the abundance of bacterial pathogens in Rousettus leschenaultii guano. Sci Rep 6(1):1–13
De Leon M, Montecillo AD, Pinili DS, Siringan MAT, Park DS (2018) Bacterial diversity of bat guano from Cabalyorisa Cave, Mabini, Pangasinan, Philippines: a first report on the metagenome of Philippine bat guano. PLoS ONE 13(7):e0200095
Gerbáčová K, Maliničová L, Kisková J, Maslišová V, Uhrin M, Pristaš P (2020) The faecal microbiome of building-dwelling insectivorous bats (Myotis myotis and Rhinolophus hipposideros) also contains antibiotic-resistant bacterial representatives. Curr Microbiol 77(9):2333–2344
Afonso E, Goydadin AC (2018) Molecular detection of Anaplasma phagocytophilum DNA in the lesser horseshoe bat (Rhinolophus hipposideros) guano. Epidemiol Infect 146(10):1253–1258
Hornok S, Szőke K, Estók P et al (2018) Assessing bat droppings and predatory bird pellets for vector-borne bacteria: molecular evidence of bat-associated Neorickettsia sp. in Europe. Antonie Van Leeuwenhoek 111(9):1707–1717
Field H, de Jong C, Melville D, Smith C, Smith I, Broos A, Kung YH, McLaughlin A, Zeddeman A (2011) Hendra virus infection dynamics in Australian fruit bats. PLoS ONE 6(12):e28678
Wood R, Bangura U, Mariën J, Douno M, Fichet-Calvet E (2021) Detection of Lassa virus in wild rodent feces: Implications for Lassa fever burden within households in the endemic region of Faranah. Guinea One health 13:100317
Seidlova V, Nemcova M, Pikula J et al (2021) Urinary shedding of leptospires in palearctic bats. Transbound Emerg Dis 68(6):3089–3095
Peel AJ, Wells K, Giles J et al (2019) Synchronous shedding of multiple bat paramyxoviruses coincides with peak periods of Hendra virus spillover. Emerg Microbes Infect 8(1):1314–1323
Zhang M, You F, Wu F, He H, Li Q, Chen Q (2021) Epidemiology and genetic characteristics of murine kobuvirus from faecal samples of Rattus losea, Rattus tanezumi and Rattus norvegicus in southern China. J Gen Virol 102(9):001646
Peres MG, Bacchiega TS, Appolinário CM, Vicente AF, Mioni M, Ribeiro B, Fonseca C, Pelícia VC, Ferreira F, Abrahão JS, Megid J (2018) Vaccinia virus in Feces and Urine of Wild Rodents from São Paulo State, Brazil. Viruses 10(2):51
Xiong YQ, Zhou JH, Zhang MY, You FF, Li DL, Chen Q (2018) Presence of rat bocavirus in oropharyngeal and fecal samples from murine rodents in China. Arch Virol 163(11):3099–3103
Monastiri A, Martín-Carrillo N, Foronda P, Izquierdo-Rodríguez E, Feliu C, López-Roig M, Miquel J, Ar Gouilh M, Serra-Cobo J (2021) First coronavirus active survey in rodents from the Canary Islands. Front Vet Sci 8:708079
Lo VT, Yoon SW, Noh JY, Kim Y, Choi YG, Jeong DG, Kim HK (2020) Long-term surveillance of bat coronaviruses in Korea: diversity and distribution pattern. Transbound Emerg Dis 67(6):2839–2848
Dietrich M, Kearney T, Seamark ECJ, Markotter W (2017) Excreted microbiota of bats: Evidence of niche specialisation based on multiple body habitats. FEMS Microbiol Lett 364(1):fnw284
Šimić I, Zorec TM, Lojkić I et al (2020) Viral Metagenomic profiling of Croatian bat population reveals sample and habitat dependent diversity. Viruses 12(8):891
Williams SH, Che X, Garcia JA, Klena JD, Lee B, Muller D, Ulrich W, Corrigan RM, Nichol S, Jain K, Lipkin WI (2018) Viral diversity of house mice in New York City. MBio 9(2):e01354-17
García-Livia K, Martín-Alonso A, Foronda P (2020) Diversity of Cryptosporidium spp. in wild rodents from the Canary Islands, Spain. Parasites Vectors 13(1):445
Zhao W, Zhou H, Yang L, Ma T, Zhou J, Liu H, Lu G, Huang H (2020) Prevalence, genetic diversity and implications for public health of Enterocytozoon bieneusi in various rodents from Hainan Province, China. Parasites Vectors 13(1):438
Kilonzo C, Li X, Vivas EJ, Jay-Russell MT, Fernandez KL, Atwill ER (2013) Fecal shedding of zoonotic food-borne pathogens by wild rodents in a major agricultural region of the central California coast. Appl Environ Microbiol 79(20):6337–6344
Phan TG, Kapusinszky B, Wang C, Rose RK, Lipton HL, Delwart EL (2011) The fecal viral flora of wild rodents. PLoS Pathog 7(9):e1002218
Hansen TA, Mollerup S, Nguyen NP et al (2016) High diversity of picornaviruses in rats from different continents revealed by deep sequencing. Emerg Microbes Infect 5(1):1–8
Walker L, Levine H, Jucker M (2006) Koch’s postulates and infectious proteins. Acta Neuropathol 112(1):1–4
Acknowledgements
B.C. acknowledges the startup funding from Trivedi School of Biosciences, Ashoka University, India and K.M.G. acknowledges the support from the DBT-Ramalingaswami Fellowship (No. BT/HRD/35/02/2006). A.S. was funded by the Trivedi School of Biosciences, Ashoka University, India. We are grateful to the Trivedi School of Biosciences, Ashoka University and National Centre for Biological Sciences (NCBS) for their support. We thank Ansil BR, Pilot Dovih and Som Banerjee for helpful discussions and comments on drafts of figures.
Author information
Authors and Affiliations
Contributions
AS, BC, and UR conceived the ideas. AS and SA prepared the manuscript with critical contribution and inputs from BC, KMG, and UR. AS and SA developed the figures. All authors reviewed and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sanyal, A., Agarwal, S., Ramakrishnan, U. et al. Using Environmental Sampling to Enable Zoonotic Pandemic Preparedness. J Indian Inst Sci 102, 711–730 (2022). https://doi.org/10.1007/s41745-022-00322-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s41745-022-00322-z