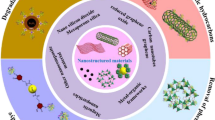
Abstract
Purpose of Review
In the presented review, we have summarized and highlighted recent developments in the use of lignin peroxidase (LiP) to remove a variety of pollutants from water matrices. The high redox potential of LiP is underlined by its excellent catalytic functionalities in the elimination of pharmaceuticals, phenolics, dyes, polycyclic aromatic hydrocarbons (PAHs), endocrine-disrupting chemicals (EDCs), and other miscellaneous pollutants. LiP-based computational frameworks for theoretical bioremediation of multiple pollutants have also been discussed, which have prompted a rise in scientific interest.
Recent Findings
According to current studies, both free and immobilized LiPs are biocatalysts capable of efficient pollutant degradation and LMW transformation. Some immobilized LiP preparations demonstrated excellent recyclability, enabling its reusability in multiple catalytic cycles. Additionally, computational degradability makes it easier to comprehend the mechanisms underlying the degradation of recalcitrant pollutants.
Summary
The capacity of LiP to cleave C–C and C–O–C bonds has led to its widespread application as a biocatalyst. Its outstanding potential to catalyze oxidative cleavage has been effectively used in the remediation of pollutants without needing mediators. Nevertheless, we brought attention to the current LiP system in pollutants remediation and computational framework, which has generated a significant rise in scientific interest.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oxidoreductases have shown a robust catalytic capacity for the removal of a wide array of pollutants as well as deployment in the valorization process [1,2,3,4]. Lignin peroxidase (LiP) is a heme-containing oxidoreductase enzyme that utilizes H2O2 to complete its catalytic reaction. LiP has strong catalytic potential in the cleavage of ether bonds (C–O–C) and carbon–carbon bonds (Cα–Cβ) inside lignin polymers [5,6,7,8]. The discovery of LiP as an extracellular enzyme from white-rot fungi (WRF) was originally demonstrated in ligninolytic cultures of the wood-decomposing basidiomycete Phanerochaete chrysosporium Burds in the 1980s [9,10,11]. The initial step in the LiP catalytic process involves using H2O2 and various substrates as sources of electrons, facilitating the effective oxidation of the substrate to complete the catalytic cycle. The catalytic mechanism of LiP corresponding to substrate binding is illustrated in Fig. 1. The investigation of lignin degradation, especially from the 1980s to the 2020s, has received considerable attention after the discovery of LiP [12]. However, lignin has long been a major impediment to many industrial processes, especially in paper industries. Later, it was observed that numerous bacterial species have been documented to produce LiP and act on lignin as substrate under a controlled environment [8, 13, 14]. During the action on lignin polymer, veratryl alcohol, a secondary metabolite of the WRF, serves as a cofactor for LiP in its interactions with lignin polymer. Owing to its strong oxidative potential (~1450 mV), LiP has been deployed for efficient lignin depolymerization by cleaving the β–O–4 linkage, which seems to be the most prevalent linkage type, accounting for more than 50% of all other ether linkages in this material [7, 15]. Although for three decades it was thought that only fungal LiP could degrade lignin due to its recalcitrant nature, it was later discovered some bacterial LiP was also able to break down lignin under controlled ex-situ conditions for obtaining renewable chemicals [16,17,18]. Fungal systems are more potent than bacterial systems in breaking down lignin, although bacterial systems may also change lignin and produce smaller aromatic molecules that may pass through the wall of membrane cells and be metabolized via aromatic catabolism [19]. Lignin can only be completely degraded by Basidiomycota, which belongs to the aerobic white rot group in nature [13]. The inability to synthesize needed enzymes and the absence of enzymatic machinery make anaerobic fungi unable to mineralize or catabolize lignin [13]. The enzymatic cleavage of the aromatic ring necessitates the presence of oxygen or its partly reduced forms, known as reactive oxygen species (ROS). Consequently, this process is unable to occur in an anaerobic environment [13]. Typical peroxidases used by fungi to degrade lignin are not present in bacteria. These relatively complicated proteins, which are usually glycosylated, contain many disulfide linkages and integrate multiple calcium ions and a heme cofactor. Thus, it may be intrinsically challenging for some organisms to produce them [17]. The mechanisms used by bacterial to produce proteins may not be compatible with the specific conditions needed for the folding and processing of these enzymes [17]. Despite extensive research on white rot basidiomycetes as a source of potent ligninolytic extracellular oxidative enzymes, the ligninolytic potential of fungi remains limited [20]. The use of bacteria as ligninolytic agents is drawing more attention in scientific research. It is brought about by less ethical considerations concerning the usage of bacteria, environmental adaptability, spontaneous development, and the simplicity with which bacteria may be genetically modified [20]. Thus, the investigation of microorganisms as possible ligninolytic agents is gaining interest. Actinomycetes, α-Proteobacteria, and γ-Proteobacteria are the three categories of bacteria that have been found in the screening process for those capable for producing lignin-modifying enzymes (LMEs) [13]. The high redox potential and capacity of LiP to oxidize resistant to degradation materials are major drivers for their sustainable usage in environment protection in biopulping and biobleaching, textile dye decolorization, and other waste effluent treatment [21,22,23]. These strong chemical bond–cleaving characteristics have been used to remove a broad range of pollutants, including pharmaceuticals, phenolics, chlorinated phenols, endocrine-disrupting chemicals (EDCs), polycyclic aromatic hydrocarbons (PAHs), and chlorinated phenols. In some instances, it has been observed that immobilized LiP has improved catalytic outcomes than free-form LiP, also enabling enzyme reusability, which entails the direct deployment of producing microorganisms in different environments with acceptable results [24, 25]. Despite the wide variety of potential industrial and biotechnological applications of LiP, the enzyme has not yet been commercially exploited, owing to a scarcity of commercial LiP preparations and the high cost. Consequently, the discovery and development of efficient ligninolytic peroxidases produced at relatively large amounts and concentrations will remain crucial for a wide array of environmental remediation applications. One of the main matters in using LiP is the necessity of use as co-substrate hydrogen peroxide. This reagent is highly oxidant and able to inactivate enzymes for different reasons [26]. While using the enzyme to produce some valuable product on a reactor, the supply of this reagent may not be a big problem (stepwise addition or in situ production may be a solution); in bioremediation, the supply of this reagent must be carefully considered [26]. Numerous efforts have been made in the last several years to increase the oxidative stability and catalytic attributes of LiP [23, 27]. Besides its potential uses in environmental remediation, LiP has often been used in a computational framework to comprehend the molecular-level degradation mechanism of enzymes and target pollutants [28]. A computational framework including molecular docking (MD) and molecular dynamics simulation (MDS) has been identified in recent studies on binding affinity for understanding real-time binding conformational state, which prompted the catalytic ability of targeted enzymes corresponding to pollutants [4, 28, 29]. To enhance the current limit in enzyme applications for the environment, machine learning (ML) and artificial intelligence (AI) have been incorporated into computational frameworks [30,31,32,33]. The current review highlights the successful progressive journey of LiP over the previous five decades. The most up-to-date updated information on the use of LiP in pollutant remediation in combination with immobilization, lignin depolymerization, biobleaching, and nanocatalysts have been outlined. Furthermore, recent advances of LiP in the computational framework for pre-screening-based enzyme-pollutant catalytic attributes to explore its potential future use in oxidative applications have also been spotlighted.
H2O2 dependent catalytic scheme of lignin peroxidase. A general schematic of the LiP chemical cycle. B Catalytic action of LiP on dimer lignin model compound, yield monomer, and LMW transformed compound. C The crystal structure of LiP from Phanerochaete chrysosporium (PDB: 1LGA). The left panel portrayed the crystal structure (teal color) containing heme (Fe) as a native ligand (Khaki/yellow color) while the right panel portrayed the active site of LiP consisting of HIS 47, ASP 165, GLU 168, TRP 171, HIS 176, ASP 238, PHE 265, and PHE 267 residues (coded in red color). HIS 176 forms hydrogen bonds with ASP 238, which facilitates the stabilization of Fe (IV)−O intermediate in compound I
LiP Immobilization
Enzyme immobilization started as a solution to facilitate enzyme recovery and reuse of these relatively expensive biocatalysts [34]. However, nowadays, enzyme immobilization has become a powerful tool for the final refinement of enzyme features [35]. To take full advantage of enzyme immobilization, the whole immobilization protocol should be performed considering the objectives, the application, the final reactor, etc. [36]. The immobilization protocol should consider all the components, starting with the selection of support compatible with the reactor and the biocatalyst application, the active group in its surface that is intended to interact with the enzyme, the immobilization conditions that determine the enzyme orientation on the support surface, and the immobilized enzyme incubation conditions to favor the enzyme-support reactivity (e.g., when pursuing multipoint covalent immobilization) [36, 37]. As a general rule, the researcher should select supports that, after the enzyme immobilization, can become chemically and physically inert (e.g., glyoxyl agarose beads after reduction) [38]. To prevent undesired enzyme-support interactions that can stabilize incorrect enzyme conformation during operation [39]. This is not possible using physical immobilization (except in some instances affinity immobilization), as the supports must remain physically active throughout time [40]. In other instances, the researcher did not realize that the support matrix is hydrophobic or has an ionic character. The research in this area is still relevant, as the interactions between immobilized enzymes and supports are not fully understood and less controlled [36]. In any case, it has been shown that proper immobilization can improve enzyme stability for diverse reasons [37]. For example, if there are several enzyme-support strong bonds, the so-called multipoint covalent, the relative positions of all involved points must be maintained throughout the length of the spacer arm under any conformational change of the enzyme, increasing the overall rigidity of the enzyme and, that way, its stability/activity under distorting conditions [37]. Moreover, if it is a multimeric enzyme, the multi-subunit immobilization may prevent enzyme dissociation, a first cause of inactivation in many of these enzymes [41]. The support can also promote some partition of reagents, and if some exert a negative effect on enzyme stability and are partitioned way, the enzyme stability may be increased [26]. However, immobilization cannot prevent the release of cations or cofactors from the enzyme; that way, it is convenient to know the causes of enzyme inactivation before trying to solve the enzyme stability limitations via immobilization. Immobilization may be coupled to enzyme purification when first using immobilization event specific for the target enzyme (e.g., a genetically added tag, the protein size, the enzyme catalytic mechanism) [42]. This can reduce the final cost of the biocatalyst and enable it to fully cover the support surface with the target enzyme. In this sense, the use of heterobifunctional supports reveals itself as an exciting possibility [43]. When an enzyme is immobilized, it suffers some conformational changes due to the reaction with the support or the later interactions with other moieties with the support, which are not usually fully inert. These conformational changes can lead to significant alterations in enzyme specificity, selectivity, activity, and even inhibition [44, 45]. Enzyme immobilization has become a very important step in preparing an industrial enzymatic biocatalyst.
In the context of LiP, there are some specific considerations. If it is going to be utilized in the degradation of small pollutants in a clear solution, no special precautions are necessary. However, if the enzyme is intended to degrade its natural substrate, insoluble lignin, it should be considered that the substrate is very large or even a solid and that, in many instances, the final product remains a suspension. This is very important when selecting a support. A standard porous support, easy to handle in clear solutions, may not be recommendable if the substrate/product is a suspension, and the biocatalyst recovery by standard filtration will not be possible due to the existence of solid material that can block the filters and clog the pores of the biocatalyst during filtration. A possible solution is the use of magnetic materials, such as a magnet, to recover the biocatalyst. If the substrate itself is solid or extremely large, there is no other option than the use of non-porous materials for enzyme immobilization, as enzymes inside the particle cannot access the substrate. One possibility is to immobilize the enzyme directly on the surface of the reactor wall [46,47,48]. One more general solution is the use of nanoparticles as the enzyme loading capacity of these materials depends on the size; tiny ones are required, and to facilitate their handling, magnetic nanoparticles are preferred. In both cases, some general protection caused by the enzyme immobilization on porous supports is lost: the enzyme can interact with interfaces and suffer surface interaction in the presence of a fluid flow. Using nanoparticles, there is a possibility of enzyme autolysis and aggregation. Therefore, the selection of an immobilization matrix must be chosen carefully, depending on the intended final use. Another point to consider is the possibility of producing hydrogen peroxide “in situ” using an oxidase [49]. This can reduce the risks of adding exogenous hydrogen peroxide. In these instances, co-immobilized oxidase/LiP may offer some advantages, as the concentration of the hydrogen peroxide may be higher inside the particles than when using two independently immobilized enzymes. However, enzyme co-immobilization presents some problems, like the complexity of finding good enough protocols valid for both enzymes or the possibility of one enzyme being much less stable than the other, making it necessary to withdraw the whole biocatalysts even when one enzyme remains fully active [50]. There are some solutions to these matters, but these strategies complicate the design of the biocatalysts and should only be employed when the advantages compensate for the drawbacks [51,52,53].
Microbial Species Involved in LiP Production
Bacterial species cannot often synthesize peroxidases, but fungi possess distinct enzymatic machinery dedicated to the degradation and utilization of lignin [17]. This might be owing to inherent challenges in producing such highly complex proteins, which are normally glycosylated, include numerous disulfide linkages, and incorporate many calcium ions and a heme cofactor [17]. Folding and processing may necessitate conditions incompatible with bacterial protein synthesis machinery [17]. Oxidizing mediators produce small oxidizing agents, which can penetrate the branching lignin polymer and cause depolymerization via radical chemistry [17]. Microbial lignin degradation in species other than fungi has not been extensively investigated; however, there have been reports of bacteria that can break down lignin. Three kinds of lignin-degrading bacteria are predominant: α-Proteobacteria, Actinomycetes, and γ-Proteobacteria [13, 54, 55]. This ubiquitous collection of microorganisms is present in terrestrial and aquatic environments and is widely dispersed throughout global natural ecosystems [13, 56]. Streptomyces viridosporus–derived LiP was first identified in 1988. Unfortunately, despite several publications on the cloning of the relevant gene, neither the sequence of this protein nor the corresponding gene has ever been deposited in the respective database [17, 57, 58]. LiPs were originally identified in the well-known WRF Phanerochaete chrysosporium, Trametes versicolor, Bjerkandera sp., and Phlebia tremellosa [9, 13, 59,60,61]. LiP can effectively oxidize a wide range of substrates, including phenolic and non-phenolic chemicals, along with several other organic compounds. The enzyme LiP has two glycosylation sites, two Ca2+ binding sites, and four disulfide bridges, which contribute to the stabilization of its three-dimensional structure. The molecular weight (MW), isoelectric point (pI), and other metrics exhibit variations depending on the characteristics of the producing species. A few of the most recent LiP-producing microbial species are listed in Table 1.
Molecular and Computational Aspects of LiP
In certain instances, structural and molecular aspects are critical for understanding the functional aspects of enzymes, especially in substrate binding, catalysis, and inhibition. Structural and molecular attributes of LiP are crucial in diverse substrate binding and catalysis. The binding site is catalytic residues dependent and may be specific to each substrate; however, similar structural substrates may have similar active site residues. Only a few critical amino acids are typically involved in catalytic activity during the catalysis process, whether by experimental or computational methods. For instance, the protein structure of the fungal LiP from Trametes cervina (PDB: 3Q3U) comprised 338 AA residues in a single chain [68, 69]. Although there are differences in the structure and residue content across members of the peroxidase family, it is important to emphasize that these enzymes could function in an identical way. Many fungal LiP structures are readily accessible for study in the protein data bank (PDB). Nevertheless, no accurate bacterial crystal structure has been submitted to date [17]. However, it is worth noting that several peroxidase or LiP variants exhibit differences in terms of their amino acid composition and protein structure, as shown by their existence in the database. For example, DyP (PDB: 5VJ0) from Enterobacter lignolyticus comprised 318 AA residues distributed in four chains (A, B, C, and D) [70, 71]. Likewise, another variant of DyP (PDB:4HOV) from Rhodococcus jostii RHA1 contains 353 AA residues distributed in three chains(A, B, and C) only [72, 73]. Modest alterations induce changes in protein functionality. As a result, structural information through protein engineering may be altered to increase catalytic efficiency under variable environmental conditions. However, it should be noted that the active sites of LiP variations exhibit substrate specificity. Therefore, in order to identify these active sites, a binding assay with the corresponding substrates needs to be employed. From a secondary structure perspective, the LiP from Trametes cervina (PDB: 3Q3U) has the following composition: 35.80% of its structure is comprised of helices, 7.69% is in the form of extended strands, 4.14% is present as beta turns, and 52.37% is in the conformation of random coils. A schematic representation of LiP structure and its comparison with known variants has portrayed in Fig. 2.
Molecularly rendered depiction of lignin peroxidase variants in schematic form. Based on the oxidizing activity of lignin, three variants of LiP (PDB: 5VJ0, 4HOV, 3Q3U) have been compared for their structural architecture and structural annotation. The basic secondary structural element in α-helix, β-sheet, and loop annotation has been depicted at the bottom of each panel. The structural architecture and SSE annotation can be noticed for non-similar peculiarities
Catalytic Action of LiP in Lignin Valorization and Depolymerization
Lignin exhibited tremendous potential as a renewable feedstock for converting HMW to LMW in a range of high-value chemicals and products for medicinal, energy, chemical, and biotechnological applications [74,75,76]. Lignin and valorization might be modified by various microbial enzymes [15, 77, 78]. Approximately 6.2 × 107 and 5 × 107 tons of lignin, comprising kraft lignin, lignosulfonate, and soda lignin, are produced annually by the biomass refinery and pulp and paper sectors, respectively [15, 79]. Currently, most lignin is either disposed of as scrap or utilized as an energy source. Due to its large carbon-to-oxygen ratio and abundant aromatic structure, lignin is a viable feedstock for producing biofuels and biochemicals. The majority of processes to modify lignin require high temperatures, catalysts, and complex reactors, among other elements that enhance lignin cleavage into low molecular weights compounds [80]. On the other hand, these techniques frequently produce waste compounds that are frequently unusable, such as byproducts of the catalysts or solvent [80]. Despite certain advancements in the field of lignin valorization, the full potential of biological systems to be utilized for lignin valorization will require significant effort in research and development. One of the main issues to be taken into account throughout the lignin valorization process is the choice of both the lignin source and the depolymerization capacity of the employed technique. The following are some potential strategic solutions to improve the microbial biotransformation of lignin: (1) selecting a lignin source and the microorganism’s ability to depolymerize, (2) selection of ideal microorganisms, (3) enzyme-microbe and microbe-microbe synergy, and (4) optimization of catabolic pathways and yield of targeted product. Understanding the degradation process and creating effective metabolic pathways for conversion are urgently needed to take advantage of lignin valorization. The first step to lignin valorization is degradation through peroxidase activity or any other enzyme with auxiliary activities. Through the sequential activity of many enzymes, including the possible oxidative production of H2O2, auxiliary enzymes facilitate the degradation process of lignin. These enzymes are pyranose 2-oxidase (POX; EC 1.1.3.10), cellobiose dehydrogenase (CDH; EC 1.1.99.18), glucose oxidase (EC 1.1.3.4), aryl alcohol oxidases (AAO; EC 1.1.3.7), and glyoxal oxidase (GLOX; EC 1.2.3.5). These enzymes have frequently been discovered in WRF secretomes [13]. As described extensively in recent literatures, significant effort has been undertaken in each step, from lignin characterization through lignocellulosic biomass pretreatment, depolymerized components separation, and catalytic or thermochemical transformation to allow lignin valorization efficiently [81,82,83,84,85,86]. Although several different LMEs have been identified in different bacterial species, the activity of these enzymes is still required for the extracellular degradation of HMW to LMW lignin. LiP has limited substrate specificity and interacts with a broad range of lignins as well as other chemicals. It varies from the ability to oxidize aromatic methoxylated rings without a free phenol group, generating cation radicals that may then react through different mechanisms, including ring-opening, demethylation, and phenol dimerization. In contrast to laccase, LiP does not require any mediators to break down high-redox compounds; nonetheless, H2O2 is required as substrate. Consolidated bioprocessing has been developed and proved to depolymerize lignin simultaneously with chemical processing. Nonetheless, this bioprocessing of lignin is industrially feasible; however, there are several concerns to be answered, the two primary targets: (1) efficient lignin depolymerization to supply considerable LMW lignin-related carbon substrate, and (2) a significant increase in yield of value-added product from aromatic compounds [6, 77, 87]. The activity of lignin breakdown enzymes may be enhanced by protein engineering [88]. Nevertheless, other heme peroxidases, including chemical methods, have shown the excellent capability of lignin depolymerization by cleaving C–C and C–O–C bonds to yield monomer chemicals in recent years [14, 88,89,90,91,92,93,94]. Many strategies for depolymerizing lignin structures have been established, including thermochemical and biological methods. The particular cleavage of lignin linkages, i.e., β–O–4 has been shown to provide many benefits from a biological standpoint, and this process is environmentally friendly [80]. Nevertheless, there are several disadvantages, such as the bacterial high sensitivity to changes in the reaction system (e.g., variations in pH, temperature, and oxygen concentrations), and obstacles in genetically modifying the microbes [80]. A schematic illustration of the LiP-assisted depolymerization process is portrayed in Fig. 3.
Role of lignin peroxidase (LiP) in valorization/depolymerization of lignin/lignocellulosic feedstock. A The upper panel depicts the lignin polymer and its distinct motif/linkages, and the oxidative cleavage of the C–C and C–O–O bonds has been explained. B The bottom panel depicts the value-added monomeric compounds and Dimeric lignin model compounds by the oxidative catalytic action of LiP
Deployment of LiP in Environmental Remediation
Textile and Pulp and Paper Industries
The textile industry extensively employs a wide range of dyes to fulfill the demand for high-quality and finished products. However, this production process generates a significant quantity of colored wastewater that poses potential hazards to the environment [95, 96]. Textile wastewater often comprises hazardous chemicals such as phenolics, dyes, aromatic and benzene ring nitro compounds, phthalates, and organochlorines. The aforementioned compounds have been shown to have detrimental environmental effects, including carcinogenicity, mutagenicity, and ecotoxicity. The aforementioned impacts have been observed via experimental studies conducted inside controlled laboratory settings as well as in natural aquatic environments [95,96,97]. In addition to textiles, several other industries employ the dyeing process, including food, cosmetics, paper, photography, and plastics, which are the leading causes of pollution in the effluent water ecosystem and have a detrimental influence on the native flora and fauna [97, 98]. The use of microorganisms and their dye-degrading enzymes for decolorizing and metabolizing dye components in a controlled laboratory environment has been well-established and has shown to be the most effective bioremediation process. Giap et al. reported that the WRF Lentinus squarrosulus MPN12 biodegraded lignin and synthetic dyes, including Acid Blue 62, Porocion Brilliant blue HGR, Acid Blue 281, Acid Blue 113, Acid red 266, and Acid red 299, adequately in lab conditions [99]. The pulp and paper production process is a complex method that involves the simultaneous generation of enormous amounts of environmental contaminants from pulping and bleaching steps that disrupt the environmental balance through environmental irregularities if drained without treating wastewater into water bodies [100, 101]. The effluent from the pulp and paper industries often contained lignin and hazardous waste by-products, i.e., black liquor, lignin derivatives, cellulose, phenolics, resins, fatty acids, and mixed tannins, which constitute major environmental hazards [102, 103]. The complex and resistant lignin with chlorinated lignin derivatives contained in the paper pulp mill effluent is the main contaminant that causes mutations in genes in exposed organisms [104]. Because it prevents sunlight from penetrating the water, lignin conveys the water a dark brown color, making the natural process of photosynthesis far more difficult to carry out [104]. Additionally, elevated parameters including pH, biological chemical demand (BOD), chemical oxygen demand (COD), color, and lignin content of the alkaline effluent render this a potential hazard to the receiving aquatic and soil environments. As a result, it is critical to treat pulp and paper wastes and wastewater appropriately before releasing them into water bodies. However, the biological methods and their potential ligninolytic enzyme-mediated deployment seem more economical and environmentally friendly for the bioremediation of such wastewater. However, small extrinsic factors such as pH, temperature, nutritional supply, and so on might influence and fail to produce LiP by LiP-producing microbes. Even though the enzyme is expensive, and yields are limited, efforts are being made to address these limitations and drive progress in the future. However, bioleaching is still in its early stages when applied to using enzymes in the pulp industry. This could lead to eco-friendly biobleaching methods for large amounts of hardwood and soft pulp [105, 106]. A basic illustrated representation portrayal of LiP deployment in paper industries is depicted in Fig. 4.
Depiction of LiP deployment in wastewater treatment and paper industries. A The employment of LiP in the paper industry for wastewater treatment. B The exploitation of LiP in biobleaching in the pulp industry. C The widespread use of LiP in bioremediation, which has been performed using different LiP producers. Associated remediation was assessed using the toxicity assessment, which was shown to be minimized after the treatment
Biodegradation of Pharmaceutical Compounds
The enzymatic activity of LiP has been demonstrated to oxidize lignin derivatives and phenolic chemicals in addition to lignin. In addition to its vital role in lignin breakdown/depolymerization, LiP has been investigated for its ability to remove a wide range of pharmaceutical compounds from wastewater [107, 108]. Nonsteroidal anti-inflammatory drugs (NSAIDs) have been detected in aquatic ecosystems, and the existing techniques for eliminating these compounds exhibit a range of limitations [109]. The identification of NSAIDs as emerging contaminants has raised significant concern within the scientific community due to their potentially hazardous impacts on the environment and public health [110,111,112,113]. NSAIDs constitute a substantial contribution to the overall quantity of chemicals discharged into the wastewater treatment system annually [114, 115]. Numerous physicochemical techniques have been developed to eliminate pharmaceutical compounds from wastewater. Among such physiochemical methods, photochemical UV/TiO2 oxidation, adsorption of organic compounds on activated carbon and polymer resins, chemical coagulation/flocculation of lignin using synthetic or natural coagulants, and catalytic wet air oxidation (WAO) are some of the methods that have been developed and experimentally tested over the course of many years for the goal of treating wastewater and removing organic pollutants [116]. Nevertheless, these processes can be quite costly, have a negative impact on the environment, and frequently lack efficiency [116]. In addition, during these processes, lignin is not degraded but rather converted from a state suspended in water to a solid or absorbed state, essentially just shifting the issue elsewhere [116]. Biological treatments, including LiP, offer a viable solution for removing organic pollutants from pulp and paper wastewater [99, 117]. EDCs have been removed from water using adsorption, filtration, chlorination, coagulation/flocculation, Fenton/photo-Fenton degradation, sonochemical degradation, photochemical/photocatalytic oxidation, ozonation, and hybrid physical–thermal processes [118,119,120,121,122,123]. The aforementioned methods are costly and sometimes produce toxic secondary contaminants. Alternatively, WRF treatment of polluted water is advantageous and sustainable. However, WRF requires an additional source of carbon since the degradation of EDC mentioned earlier occurs as part of its secondary metabolism. Unlike bacteria, WRF has the ability to break down EDC even when present in low concentrations. Currently, organic contaminants are primarily eliminated through microbial degradation due to their cost-effectiveness and simple operational requirements. Microorganisms have the ability to eliminate organic contaminants, either on their own or in a collective effort, through metabolic consumption. In addition, they eliminate the pollutants by breaking them down using secreted enzymes [124]. Nevertheless, only a limited number of these methods are practically viable due to their high costs [125, 126]. Guo et al. investigated the efficacy of utilizing immobilized LiP on Fe3O4@SiO2@polydopamine nanoparticles for the degradation of several organic pollutants, including tetracycline, dibutyl phthalate, 5-chlorophenol, phenol, phenanthrene, fluoranthene, and benzo(a)pyrene [63]. Subsequently, the immobilized LiP exhibited a reaction efficacy of 100% for tetracycline, dibutyl phthalate, 5-chlorophenol, and phenol. Additionally, it demonstrated reaction efficacies of 79%, 73%, and 65% for phenanthrene, fluoranthene, and benzo(a)pyrene, respectively [63]. Conversely, the inactivated immobilized LiP only adsorbed less than 25% of phenanthrene and fluoranthene [63]. Ding et al. highlighted the in-vitro degradation of propranolol using LiP [127]. Throughout the investigation, the findings indicated that significant degradation of 94.2% of propranolol occurred when exposed to a LiP activity level of 30 U L−1 [127]. Furthermore, a multipath degrading mechanism was conferred onto LiP a noteworthy capability to efficiently remove a wide range of recalcitrant pollutants in the context of environmental remediation [127]. The efficacy of fungal enzymatic activity, including LiP, in removing carbamazepine, diclofenac, and ibuprofen from aquatic environments was reported by Kasonga et al. [128].
Biodegradation of Endocrine-Disrupting Chemicals and Polycyclic Aromatic Hydrocarbons
EDCs are a broad category of substances known to interfere with the function of the endocrine system. In light of their structural similarities to natural steroid hormones, they can bind to hormone receptors, which results in negative health impacts once exposed. Bisphenol A (BPA), bisphenol S (BPS), and nonylphenol (NP) stand out as three of the EDCs that seem to be particularly hazardous to humans [129]. As a result of the substantial growth of the industry, there has been a significant increase in the production of BPA, BPS, and NP, resulting in notable impacts on the environment. WRFs are a cost-effective, environmentally friendly, and potentially acceptable method of removing EDCs [129]. Ligninolytic enzymes, including LiPs, are secreted extracellularly by WRF. These enzymes can degrade a wide variety of EDCs because of their low substrate specificity (Fig. 5). LiP has a wide substrate specificity, which allows it to eliminate a variety of xenobiotics, including EDCs. Due to their hydrophobic characteristics, EDC compounds are thought to only biodegrade slowly under aerobic conditions. EDCs can be effectively degraded by purified peroxidase and ligninolytic fungi. A typical instance of how NPs can transform is through carbon–carbon (C–C) and carbon–oxygen (C–O) coupling and oxidation at the terminal alkyl chain carbon. This metabolite is likely formed through the action of LMEs on phenolic hydroxyl groups, leading to the creation of phenoxyl radicals. These radicals then undergo various spontaneous reactions, including oxidative coupling. Consequently, it can be concluded that the use of WRF has significant potential in accomplishing the aforementioned challenge of EDCs during wastewater treatment operations [129]. PAHs are persistent organic pollutants with carcinogenic, teratogenic, and mutagenic properties. In PAH-contaminated soils, it has been shown that HMW-PAHs with four or more rings exhibit greater prevalence than their LMW-PAH counterparts. PAHs exhibit high resistance to degradation processes. Fortunately, new studies have been conducted to determine the efficiency of LiP in the degradation of PAHs. Zhang et al. investigated the degradation of HMW-PAHs by LiP activity using the fungal species Fusarium sp. ZH-H2 [130]. This study was shown to have used different parameters during the treatment process [130]. Afterward, results showed that the DF treatment combined with starch and humic acid as carbon sources yielded the highest soil LiP activity (27.42 U L−1) [130]. In another recent work, Chai et al. reported immobilized LiP on chitosan-modified halloysite nanotubes for PAH degradation [131]. Their study achieved the optimal enzyme activity and immobilization efficiency with an enzyme dosage of 8 mg and an immobilization time of 8 h [131]. The immobilized LiP possessed greater pH resistance, thermal stability, and storage stability than the free LiP, and after eight cycles, it retained more than 40% of its initial enzyme activity [131]. Immobilized LiP had the highest removal efficiency of phenanthrene (55.7%) and fluoranthene (41.2%) in spiked soil after 4 days [131]. The immobilized LiP removed 21.1% of 16 PAHs from aged soil, with the removal efficacy of LMW PAHs being higher than that of medium- and HMW PAHs [131].
LiP-Assisted Dye Degradation
WRFs are the most common source of LiP. However, not all WRFs produce LiP. LiP has a greater redox potential compared to laccases and MnP. This characteristic enables LiP to oxidize both phenolic and non-phenolic components of lignin, even in the absence of mediators. Consequently, LiP demonstrates superior catalytic efficiency in comparison to laccase. LiP activity relies on the presence of H2O2; nevertheless, excessive quantities of H2O2 may have detrimental effects on the functionality of LiP. Applying this noted oxidative ability has shown potential for the degradation of several dyes [24, 67]. Bilal et al. documented the immobilization of LiP by covalent binding onto Ca-alginate beads, employing glutaraldehyde as a cross-linking agent [25]. Subsequently, the immobilized biocatalytic system demonstrated the ability to efficiently decolorize dye (Remazol Brilliant Blue R) in five consecutive batch operations, maintaining a dye-removal efficiency of over 80% even after the fifth cycle [25]. In another study, Pham et al. described a novel LiP-producing Bacillus sp. React3 for methylene blue dye biodegradation [67]. Subsequently, After 48 h of incubation, it was observed that the removal rate of methylene blue achieved 99.5% [67]. The most favorable parameters for the degradation of methylene blue were seen at a pH of 7 and a temperature of 35 °C, under static conditions, with an inoculum concentration of 4% and a concentration of 1000 mg/L of methylene blue [67]. Tryptone was used as the carbon source, while yeast extract was the nitrogen source [67].
Enzyme Engineering for Enhancing Catalytic Potential
Chemical modifications have been made to the enzyme to improve its catalytic efficiency [31, 132, 133]. Enhancing the performance of native or indigenous enzymes for industrial applications may be achieved by boosting factors such as enzyme selectivity and catalytic efficiency. Nevertheless, the achievement of such optimization may only be realized by the use of a protein engineering approach [134]. Increasing the activity of an enzyme via amino acid sequence modifications and making it appropriate for its use in industrial processes is the primary goal of enzyme engineering. Determining the composition and structure of the enzyme that has to be changed is the main goal of various enzyme engineering approaches. The underlying theory is that information on the molecular complexity of an enzyme structure may be used to change the enzyme’s functional properties. The creation of the enzyme-substrate complex during biocatalysis is a critical step in determining the kinetic efficiency of the enzyme. Since the amino acid residues found in the catalytic domain of the enzyme are often involved in the interactions between the enzyme and substrate, it is therefore usually viewed as crucial to modify alterations in those residues when designing engineered enzymes. Nevertheless, by creating a general change in the enzyme conformation and functional activity, changes to amino acids other than those at the active site may also aid in controlling the interactions between the enzyme and the substrate. In recent years, rational design and directed molecular evolution approaches have facilitated the implementation of oxidoreductase engineering for specific industrial applications [135]. Recombinant LiP production in native and heterogeneous hosts has proven more efficient than in native variants. Nevertheless, it is usually noted that certain extrinsic factors, such as increased temperature, the presence of a solvent, and susceptibility to inactivation by H2O2, often impede the desired outcomes. Therefore, it is essential to thoroughly examine different protein engineering approaches to enhance the production of LiP and improve its enzymatic activity. Đurđić et al. reported improved degradation of azo dyes by mutagenesis in LiP [23]. Their study used LiP from Phanerochaete chrysosporium for protein engineering [23]. Following that, mutations exhibited up to tenfold higher affinity for three different azo dyes (Evans blue, amido black 10B, and Guinea green) and up to 13-fold higher catalytic activity [23]. In another study, the WRF (Phanerodontia chrysosporium) LiP gene was cloned and heterologously produced in the food-grade yeast (Cyberlindnera jadinii) [136]. At 96 h, its LiP activity reached its highest level, 68.52 U/L [136].
LiP in Theoretical Degradation of Pollutants Deploying Computational Technologies
A variety of computational tools have been developed to investigate and comprehend the fundamental principles that enable enzymes to degrade pollutants or biotransformed substrates into smaller fragments employing non-testing methods [28, 29, 32, 137, 138]. As a result, an abundance of computational toolsets is now accessible, and they may play an important role in comprehending the complex steps occurring at the molecular level of the degradation mechanism involving the target enzyme-pollutant interaction [139,140,141]. The conventional degradation processes often exhibit imperfections in molecular interaction, complete transformed compounds, and binding affinity between enzyme-pollutants. These limitations may be overcome by employing theoretical degradation approaches. The aforementioned methods use intelligent, non-testing approaches and are reliant on computational algorithms to enhance the reliability of pre-screening-based pollutant degradation [29, 139, 142,143,144]. In recent years, a few pieces of research have been conducted using LiP, and its variants to investigate the degradation mechanisms of various pollutants at a molecular level [28, 145, 146]. In our previous study, we conducted an in silico investigation of the enzyme LiP to gain insights into the degradation process of 12 lignin derivatives [28]. Such a study deploys glide docking and high-performance Desmond MDS approaches in order to elucidate the underlying theoretical degradation mechanism [28]. The trimer model compound exhibited the lowest XP Gscore of −8.136 kcal/mol among all docked compounds. This was attributed to the presence of Pi-Pi stacking and H-bond (side chain) type bonding interactions corresponding to the LiP-trimer compound. Another similar study was carried out using LiP and multimeric lignin model compounds, and the significant degradation potential of LiP was estimated using docking, and Desmond simulation [29]. Table 2 explains the deployment of LiP in computational frameworks for the theoretical exploration of various pollutants at the molecular level. Integrating computational tools, which are non-testing approaches, along with conventional methods, may enhance the current extent of degradation in the context of green degradation design for regulatory consideration. A schematic representation of computational methodology with undertaking LiP for pollutants degradability prediction is shown in Fig. 6.
The foremost advance in environmental research by employing lignin peroxidase (LiP) from the domain of computer-aided research. Artificial intelligence (AI) and machine learning (ML)-based methodologies in certain computational tools, such as docking and molecular dynamics simulation, have been deployed in computational remediation or theoretical degradation. Three key computational strategies have been demonstrated, which have been described to be used along with LiP in smart remediation technologies. A The existing flaws in conventional remediation technologies. B The computational techniques that could overcome the existing flaws. C The molecular interplay for unraveling the degradation mechanism through LiP
Future Outlook and Concluding Remarks
Considering its low cost, rapid remediation process, and bypassing of secondary pollution, microorganism-based biological remediation is one of the most often investigated approaches. Therefore, microbial remediation technology is one of the most effective and environmentally friendly solutions for managing hazardous pollutants. The removal of a wide range of pollutants, including pharmaceuticals, EDCs, PAHs, dyes, chlorinated phenols, and other emerging contaminants of concern can be mitigated by deploying LiP. Extrinsic factors, including pH, temperature, solvent, and others, have the potential to influence thermal stability and broaden the scope of catalysis. LiP exhibits strong catalytic capabilities, as well as a high redox potential in comparison to other oxidoreductases. It was first used extensively in lignin degradation and oxidation of identical molecular weight compounds. It was thought it could only oxidize lignin through the degradation assays under controlled environmental conditions. However, throughout the years, its catalytic ability has been unlocked in a variety of applications, most notably bioremediation and lignin depolymerization. During the decade of the 2000s–2010s, the protein crystal structures of LiP derived from fungi were solved and deposited into a protein data repository. Later, a heme peroxidase known as DyP was identified, which has a catalytic action similar to breakdown/depolymerization on lignin. Further crystal structure refinement and molecular investigation revealed the evolutionary relationship among the closest LiP-producing species. No bacterial LiP crystal structure is available in the protein databank, but plenty of structures from fungi are known to date. Despite its high redox potential and oxidizing capabilities, LiP has been remarkably deployed in the pulp and paper industry for wastewater biobleaching, decolorization, and lignin degradation. Furthermore, it has been used in nano-biocatalysts, and immobilization in recent decades. However, the molecular interaction during pollutant degradation and its magnitude remained unresolved until the last decade (the 2010s). Later, remarkable studies were carried out using, for example, docking and molecular dynamics simulation techniques to determine the molecular behavior of LiP during the catalytic action on lignin, lignin derivatives, and comparable pollutants. With the implementation of a computational framework, LiP has attracted considerable scientific attention compared to other oxidoreductases. It has been witnessed that several strategies must be investigated to attain a high yield of LiP. Despite significant progress in producing LiP with improved biofunctional properties, which might also pave the way for prospective future applications of the enzyme. Even though major improvements and progress in molecular biology techniques, engineered bacteria and fungi for recombinant enzyme production, and engineered systems for yielding high levels of LiP, the availability of LiP in widely varying industrial and biotechnological applications remains a challenge. Therefore, such attributes could be enhanced a bit more to maximize the catalytic functionalities of LiP under harsh conditions. Furthermore, LiP may be turned into a “green design” for environmental pollutant biodegradation, combining in-silico and conventional biodegradation assay. For future views on expanding our comprehension and addressing the issues we have identified, there are plenty of gaps to be filled as follows:
-
1.
No bacterial crystal structure of LiP is deposited into protein databases; such can provide the protein engineering framework for enhancing the catalytic role
-
2.
Enhancing the catalytic potential of LiP with conventional and in-silico efforts
-
3.
Enhancing the yield from indigenous and transgenic species
-
4.
Implementation of theoretical degradation into a real-time assay for improving the degradation efficiency
-
5.
LiP-assisted green design for biodegradation of diverse phenolic compounds
-
6.
Exploration of future prospective oxidative application for miscellaneous use
LiP has demonstrated its strong redox potential in catalysis, depolymerization, and H2O2-dependent cleavage of lignin C–C and C–O–C bonds. A considerable deployment of LiP as a viable biocatalyst for acting on lignin has been noted for lignin breakdown. Likewise, lignin polymer is a crucial, plentiful source for generating aromatic high-value compounds, which might be accomplished with the oxidative activity of LiP. With excellent redox potential activity, LiP has drawn enormous attention for enhancing the yield as well as the catalytic activity through engineering strategies. However, advances and reports evidence that increasing the yield for demanding industrial and biotechnological utilization remains a great challenge. Nonetheless, attempts to harness its potential for future use are underway.
References
Kołodziejczak-Radzimska, A., et al., Exploring the functionality of an active ZrF-laccase biocatalyst towards tartrazine decolorization. Environmental Technology & Innovation, 2023. 31: p. 103201.
Rybarczyk A, et al. 3D printed polylactide scaffolding for laccase immobilization to improve enzyme stability and estrogen removal from wastewater. Biores Technol. 2023;381:129144.
Vu HP, et al. A comprehensive review on the framework to valorise lignocellulosic biomass as biorefinery feedstocks. Sci Total Environ. 2020;743:140630.
Ahmad I, et al. In silico insights into the arsenic binding mechanism deploying application of computational biology-based toolsets. ACS Omega. 2024;9(7):7529–44.
Singh AK, et al. Lignin peroxidase in focus for catalytic elimination of contaminants — a critical review on recent progress and perspectives. Int J Biol Macromol. 2021;177:58–82.
Pham LTM, et al. Experimental and theoretical insights into the effects of pH on catalysis of bond-cleavage by the lignin peroxidase isozyme H8 from Phanerochaete chrysosporium. Biotechnol Biofuels. 2021;14(1):108.
Marinović M, et al. Selective cleavage of lignin β-O-4 aryl ether bond by β-etherase of the white-rot fungus Dichomitus squalens. ACS Sustain Chem Eng. 2018;6(3):2878–82.
Rashid GMM, Bugg TDH. Enhanced biocatalytic degradation of lignin using combinations of lignin-degrading enzymes and accessory enzymes. Catal Sci Technol. 2021;11(10):3568–77.
Tien M, Kirk TK. Lignin-degrading enzyme from the Hymenomycete Phanerochaete chrysosporium Burds. Science. 1983;221(4611):661–3.
Archibald FS. A new assay for lignin-type peroxidases employing the dye azure B. Appl Environ Microbiol. 1992;58(9):3110–6.
Tien M, Kirk TK. Lignin-degrading enzyme from Phanerochaete chrysosporium: purification, characterization, and catalytic properties of a unique H(2)O(2)-requiring oxygenase. Proc Natl Acad Sci USA. 1984;81(8):2280–4.
Biko ODV, Viljoen-Bloom M, van Zyl WH. Microbial lignin peroxidases: applications, production challenges and future perspectives. Enzyme Microb Technol. 2020;141:109669.
Janusz G, et al. Lignin degradation: microorganisms, enzymes involved, genomes analysis and evolution. FEMS Microbiol Rev. 2017;41(6):941–62.
Riyadi FA, et al. Enzymatic and genetic characterization of lignin depolymerization by Streptomyces sp. S6 isolated from a tropical environment. Sci Reports. 2020;10(1):7813.
Weng C, Peng X, Han Y. Depolymerization and conversion of lignin to value-added bioproducts by microbial and enzymatic catalysis. Biotechnol Biofuels. 2021;14(1):84.
Bugg TDH, Rahmanpour R. Enzymatic conversion of lignin into renewable chemicals. Curr Opin Chem Biol. 2015;29:10–7.
de Gonzalo G, et al. Bacterial enzymes involved in lignin degradation. J Biotechnol. 2016;236:110–9.
Bugg TDH, et al. The emerging role for bacteria in lignin degradation and bio-product formation. Curr Opin Biotechnol. 2011;22(3):394–400.
Grgas D, et al. The bacterial degradation of lignin—a review. Water. 2023. https://doi.org/10.3390/w15071272.
Dube SL, et al. Isolation and characterization of potential lignin peroxidase-producing bacteria from compost samples at Richards Bay (South Africa). Pol J Microbiol. 2023;72(2):117–24.
Wei S, et al. Application of enzyme technology in biopulping and biobleaching. Cellulose. 2021;28(16):10099–116.
Falade AO, et al. Lignin peroxidase functionalities and prospective applications. MicrobiologyOpen. 2017;6(1):e00394.
Ilić Đurđić K, et al. Improved degradation of azo dyes by lignin peroxidase following mutagenesis at two sites near the catalytic pocket and the application of peroxidase-coated yeast cell walls. Front Environ Sci Eng. 2020;15(2):19.
Parveen S, Asgher M, Bilal M. Lignin peroxidase-based cross-linked enzyme aggregates (LiP-CLEAs) as robust biocatalytic materials for mitigation of textile dyes-contaminated aqueous solution. Environ Technol Innov. 2021;21:101226.
Bilal M, Iqbal HMN. Lignin peroxidase immobilization on Ca-alginate beads and its dye degradation performance in a packed bed reactor system. Biocatal Agric Biotechnol. 2019;20:101205.
Hernandez K, et al. Hydrogen peroxide in biocatalysis. A dangerous liaison Current organic chemistry. 2012;16(22):2652–72.
Ilić Đurđić K, et al. Flow cytometry-based system for screening of lignin peroxidase mutants with higher oxidative stability. J Biosci Bioeng. 2020;129(6):664–71.
Singh AK, et al. In silico exploration of lignin peroxidase for unraveling the degradation mechanism employing lignin model compounds. RSC Adv. 2021;11(24):14632–53.
Singh AK, et al. In silico analytical toolset for predictive degradation and toxicity of hazardous pollutants in water sources. Chemosphere. 2022;292:133250.
Mey F, et al. Improving the performance of machine learning models for biotechnology: the quest for deus ex machina. Biotechnol Adv. 2021;53:107858.
Giri P, et al. Chemical modification of enzymes to improve biocatalytic performance. Biotechnol Adv. 2021;53:107868.
Planas-Iglesias J, et al. Computational design of enzymes for biotechnological applications. Biotechnol Adv. 2021;47:107696.
He L, et al. Applications of computational chemistry, artificial intelligence, and machine learning in aquatic chemistry research. Chem Eng J. 2021;426:131810.
Sheldon RA, van Pelt S. Enzyme immobilisation in biocatalysis: why, what and how. Chem Soc Rev. 2013;42(15):6223–35.
Mateo C, et al. Improvement of enzyme activity, stability and selectivity via immobilization techniques. Enzyme Microb Technol. 2007;40(6):1451–63.
Bolivar JM, Woodley JM, Fernandez-Lafuente R. Is enzyme immobilization a mature discipline? Some critical considerations to capitalize on the benefits of immobilization. Chem Soc Rev. 2022;51(15):6251–90.
Rodrigues RC, et al. Stabilization of enzymes via immobilization: multipoint covalent attachment and other stabilization strategies. Biotechnol Adv. 2021;52:107821.
Mateo C, et al. Glyoxyl agarose: a fully inert and hydrophilic support for immobilization and high stabilization of proteins. Enzyme Microb Technol. 2006;39(2):274–80.
Santos JCSD, et al. Importance of the support properties for immobilization or purification of enzymes. ChemCatChem. 2015;7(16):2413–32.
Virgen-Ortíz JJ, et al. Desorption of lipases immobilized on octyl-agarose beads and coated with ionic polymers after thermal inactivation. Stronger adsorption of polymers/unfolded protein composites. Molecules. 2017;22(1).
Fernandez-Lafuente R. Stabilization of multimeric enzymes: strategies to prevent subunit dissociation. Enzyme Microb Technol. 2009;45(6):405–18.
Barbosa O, et al. Strategies for the one-step immobilization-purification of enzymes as industrial biocatalysts. Biotechnol Adv. 2015;33(5):435–56.
Barbosa O, et al. Heterofunctional supports in enzyme immobilization: from traditional immobilization protocols to opportunities in tuning enzyme properties. Biomacromol. 2013;14(8):2433–62.
Rodrigues RC, et al. Modifying enzyme activity and selectivity by immobilization. Chem Soc Rev. 2013;42(15):6290–307.
Garcia-Galan C, et al. Potential of different enzyme immobilization strategies to improve enzyme performance. Adv Synth Catal. 2011;353(16):2885–904.
Yamaguchi H, Miyazaki M. Enzyme-immobilized reactors for rapid and efficient sample preparation in MS-based proteomic studies. Proteomics. 2013;13(3–4):457–66.
Ma J, et al. Immobilized enzyme reactors in proteomics. TrAC, Trends Anal Chem. 2011;30(5):691–702.
Cipolatti EP, et al. Nanomaterials for biocatalyst immobilization – state of the art and future trends. RSC Adv. 2016;6(106):104675–92.
Kumar A, et al. Co-immobilization of cholesterol oxidase and horseradish peroxidase in a sol–gel film. Anal Chim Acta. 2000;414(1):43–50.
Arana-Peña S, et al. Enzyme co-immobilization: always the biocatalyst designers’ choice…or not? Biotechnol Adv. 2021;51:107584.
Morellon-Sterling R, et al. Advantages of supports activated with divinyl sulfone in enzyme coimmobilization: possibility of multipoint covalent immobilization of the most stable enzyme and immobilization via ion exchange of the least stable enzyme. ACS Sustain Chem Eng. 2021;9(22):7508–18.
Arana-Peña S, et al. New applications of glyoxyl-octyl agarose in lipases co-immobilization: Strategies to reuse the most stable lipase. Int J Biol Macromol. 2019;131:989–97.
Arana-Peña S, et al. Multi-combilipases: co-immobilizing lipases with very different stabilities combining immobilization via interfacial activation and ion exchange. The reuse of the most stable co-immobilized enzymes after inactivation of the least stable ones. Catalysts. 2020. https://doi.org/10.3390/catal10101207.
Bugg TDH, et al. Pathways for degradation of lignin in bacteria and fungi. Nat Prod Rep. 2011;28(12):1883–96.
Huang XF, et al. Isolation and characterization of lignin-degrading bacteria from rainforest soils. Biotechnol Bioeng. 2013;110(6):1616–26.
Yang CX, et al. Isolation, identification and characterization of lignin-degrading bacteria from Qinling, China. J Appl Microbiol. 2017;123(6):1447–60.
Wang Z, et al. Cloning and expression of a lignin peroxidase gene from Streptomyces viridosporus in Streptomyces lividans. J Biotechnol. 1990;13(2–3):131–44.
Thomas L, Crawford DL. Cloning of clustered Streptomyces viridosporus T7A lignocellulose catabolism genes encoding peroxidase and endoglucanase and their extracellular expression in Pichia pastoris. Can J Microbiol. 1998;44(4):364–72.
Paszczyński A, Huynh VB, Crawford R. Comparison of ligninase-I and peroxidase-M2 from the white-rot fungus Phanerochaete chrysosporium. Arch Biochem Biophys. 1986;244(2):750–65.
Kirk TK, Farrell RL. Enzymatic “combustion”: the microbial degradation of lignin. Annu Rev Microbiol. 1987;41:465–505.
Johansson T, Nyman PO. Isozymes of lignin peroxidase and manganese(II) peroxidase from the white-rot basidiomycete Trametes versicolor. I. Isolation of enzyme forms and characterization of physical and catalytic properties. Arch Biochem Biophys. 1993;300(1):49–56.
Shaheen R, et al. Immobilized lignin peroxidase from Ganoderma lucidum IBL-05 with improved dye decolorization and cytotoxicity reduction properties. Int J Biol Macromol. 2017;103:57–64.
Guo J, et al. Immobilized lignin peroxidase on Fe3O4@SiO2@polydopamine nanoparticles for degradation of organic pollutants. Int J Biol Macromol. 2019;138:433–40.
Agnestisia R, et al. Lignin-degrading enzymes from a pathogenic canker-rot fungus Inonotus obliquus strain IO-B2. AMB Express. 2023;13(1):59.
Vandana T, et al. Purification, characterization, and biodelignification potential of lignin peroxidase from immobilized Phanerochaete chrysosporium. BioResources. 2019;14(3):5380–99.
Sung HJ, Khan MF, Kim YH. Recombinant lignin peroxidase-catalyzed decolorization of melanin using in-situ generated H2O2 for application in whitening cosmetics. Int J Biol Macromol. 2019;136:20–6.
Pham VH, et al. Biodegradation of methylene blue using a novel lignin peroxidase enzyme producing bacteria, named Bacillus sp. React3, as a promising candidate for dye-contaminated wastewater treatment. Ferment. 2022. https://doi.org/10.3390/fermentation8050190.
PDB. https://www.rcsb.org/structure/3Q3U.(Accessed 9 April 2024).
Miki Y, et al. Crystallographic, kinetic, and spectroscopic study of the first ligninolytic peroxidase presenting a catalytic tyrosine. J Biol Chem. 2011;286(17):15525–34.
PDB. https://www.rcsb.org.(Accessed 9 April 2024).
Shrestha R, et al. Mechanistic insights into dye-decolorizing peroxidase revealed by solvent isotope and viscosity effects. ACS Catal. 2017;7(9):6352–64.
PDB. https://www.rcsb.org/structure/4HOV.(Accessed 9 April 2024).
Singh R, et al. Improved manganese-oxidizing activity of DypB, a peroxidase from a lignolytic bacterium. ACS Chem Biol. 2013;8(4):700–6.
Schneider WDH, Dillon AJP, Camassola M. Lignin nanoparticles enter the scene: a promising versatile green tool for multiple applications. Biotechnol Adv. 2021;47:107685.
Becker J, Wittmann C. A field of dreams: lignin valorization into chemicals, materials, fuels, and health-care products. Biotechnol Adv. 2019;37(6):107360.
Stevens JC, Shi J. Biocatalysis in ionic liquids for lignin valorization: Opportunities and recent developments. Biotechnol Adv. 2019;37(8):107418.
Lee S, et al. Bacterial valorization of lignin: strains, enzymes, conversion pathways, biosensors, and perspectives. Front Bioeng Biotechnol. 2019;7:209.
Xu Z, et al. Recent advances in lignin valorization with bacterial cultures: microorganisms, metabolic pathways, and bio-products. Biotechnol Biofuels. 2019;12(1):32.
Zakzeski J, et al. The catalytic valorization of lignin for the production of renewable chemicals. Chem Rev. 2010;110(6):3552–99.
Lopez Camas K, Ullah A. Depolymerization of lignin into high-value products. Biocatal Agric Biotechnol. 2022;40:102306.
Beckham GT, et al. Opportunities and challenges in biological lignin valorization. Curr Opin Biotechnol. 2016;42:40–53.
Cao L, et al. Lignin valorization for the production of renewable chemicals: state-of-the-art review and future prospects. Bioresour Technol. 2018;269:465–75.
Ponnusamy VK, et al. A review on lignin structure, pretreatments, fermentation reactions and biorefinery potential. Bioresour Technol. 2019;271:462–72.
Wang H, et al. From lignin to valuable products-strategies, challenges, and prospects. Bioresour Technol. 2019;271:449–61.
Wang A, He P, Song H. 6 - Lignin valorization, in recent advances in bioconversion of lignocellulose to biofuels and value-added chemicals within the biorefinery concept. In: Ferreira Filho EX, et al. editors. Elsevier; (2020). p. 133–152.
Bilal M, et al. Exploring the potential of ligninolytic armory for lignin valorization – a way forward for sustainable and cleaner production. J Clean Prod. 2021;326:129420.
Sellami K, et al. Peroxidase enzymes as green catalysts for bioremediation and biotechnological applications: a review. Sci Total Environ. 2022;806:150500.
Brissos V, et al. Engineering a bacterial DyP-type peroxidase for enhanced oxidation of lignin-related phenolics at alkaline pH. ACS Catal. 2017;7(5):3454–65.
Yang C, et al. Biodegradation of lignin by Pseudomonas sp. Q18 and the characterization of a novel bacterial DyP-type peroxidase. J Ind Microbiol Biotechnol. 2018;45(10):913–27.
De Santi A, et al. New Mechanistic insights into the lignin β-O-4 linkage acidolysis with ethylene glycol stabilization aided by multilevel computational chemistry. ACS Sustain Chem Eng. 2021;9(5):2388–99.
Zhu Q, Nocera DG. Catalytic C(β)–O bond cleavage of lignin in a one-step reaction enabled by a spin-center shift. ACS Catal. 2021;11(22):14181–7.
Van den Bosch S, et al. Catalytic strategies towards lignin-derived chemicals. Top Curr Chem (Cham). 2018;376(5):36.
Sun Z, et al. Bright side of lignin depolymerization: toward new platform chemicals. Chem Rev. 2018;118(2):614–78.
Barta K, et al. Depolymerization of organosolv lignin to aromatic compounds over Cu-doped porous metal oxides. Green Chem. 2014;16(1):191–6.
Lellis B, et al. Effects of textile dyes on health and the environment and bioremediation potential of living organisms. Biotechnol Res Innovation. 2019;3(2):275–90.
Berradi M, et al. Textile finishing dyes and their impact on aquatic environs. Heliyon. 2019;5(11):e02711.
Tang AYL, Lo CKY, Kan C-W. Textile dyes and human health: a systematic and citation network analysis review. Color Technol. 2018;134(4):245–57.
Slama HB, et al. Diversity of synthetic dyes from textile industries, discharge impacts and treatment methods. Appl Sci. 2021;11(14).
Dinh Giap V, et al. Lignin peroxidase from the white-rot fungus Lentinus squarrosulus MPN12 and its application in the biodegradation of synthetic dyes and lignin. BioResources. 2022;17(3):4480.
Dixit M, et al. Pulp and paper industry based pollutants, their health hazards and environmental risks. Curr Opin Environ Sci Health. 2019.
Servos MR, Environmental fate and effects of pulp and paper: mill effluents. 2020: CRC Press.
Singh AK, et al. Bioremediation of lignin derivatives and phenolics in wastewater with lignin modifying enzymes: status, opportunities and challenges. Sci Total Environ. 2021;777:145988.
Singh AK, et al. Biotransformation and cytotoxicity evaluation of kraft lignin degraded by ligninolytic Serratia liquefaciens. Front Microbiol. 2019;10.
Mandal DD, et al. Challenges in developing strategies for the valorization of lignin—a major pollutant of the paper mill industry. Environ Sci Pollut Res. 2023;30(5):11119–40.
Kumar A. Biobleaching: an eco-friendly approach to reduce chemical consumption and pollutants generation. Phys Sci Rev. 2021;6(4).
Hamedi J, Vaez Fakhri A, Mahdavi S. Biobleaching of mechanical paper pulp using Streptomyces rutgersensis UTMC 2445 isolated from a lignocellulose-rich soil. J Appl Microbiol. 2020;128(1):161–70.
Falade AO, et al. Ligninolytic enzymes: versatile biocatalysts for the elimination of endocrine-disrupting chemicals in wastewater. MicrobiologyOpen. 2018;7(6):e00722–e00722.
Pylypchuk IV, et al. Removal of diclofenac, paracetamol, and carbamazepine from model aqueous solutions by magnetic sol–gel encapsulated horseradish peroxidase and lignin peroxidase composites. 2020;10(2):282.
Nieto-Juárez JI, et al. Pharmaceuticals and environmental risk assessment in municipal wastewater treatment plants and rivers from Peru. Environ Int. 2021;155:106674.
Apriceno A, et al. A new laccase-mediator system facing the biodegradation challenge: Insight into the NSAIDs removal. Chemosphere. 2019;215:535–42.
Jia Y, et al. Influence of ibuprofen and its biotransformation products on different biological sludge systems and ecosystem. Environ Int. 2021;146:106265.
Madikizela LM, Ncube S. Occurrence and ecotoxicological risk assessment of non-steroidal anti-inflammatory drugs in South African aquatic environment: what is known and the missing information? Chemosphere. 2021;280:130688.
Patel M, et al. Pharmaceuticals of emerging concern in aquatic systems: chemistry, occurrence, effects, and removal methods. Chem Rev. 2019;119(6):3510–673.
Madikizela LM, et al. Uptake, occurrence, and effects of nonsteroidal anti-inflammatory drugs and analgesics in plants and edible crops. J Agric Food Chem. 2022;70(1):34–45.
Thalla AK, Vannarath AS. Occurrence and environmental risks of nonsteroidal anti-inflammatory drugs in urban wastewater in the southwest monsoon region of India. Environ Monit Assess. 2020;192(3):193.
Costa S, et al. Lignin biodegradation in pulp-and-paper mill wastewater by selected white rot fungi. Water. 2017. https://doi.org/10.3390/w9120935.
Khan SI, et al. Degradation of lignin by Bacillus altitudinis SL7 isolated from pulp and paper mill effluent. Water Sci Technol. 2021;85(1):420–32.
Ghosh S, Sahu M. Ultrasound for the degradation of endocrine disrupting compounds in aqueous solution: a review on mechanisms, influence of operating parameters and cost estimation. Chemosphere. 2024;349:140864.
Gamarra-Güere CD, et al. Application of fenton, photo-fenton and electro-fenton processes for the methylparaben degradation: a comparative study. J Environ Chem Eng. 2022;10(1):106992.
Azevedo TDS, et al. Removal of endocrine-disrupting chemical mixtures in water using chlorination and photolysis. Water Supply. 2023;23(3):1284–96.
Abdullah MS, et al. The treatment of endocrine-disruptive chemicals in wastewater through asymmetric reverse osmosis membranes: a review. Sym. 2023. https://doi.org/10.3390/sym15051049.
Azizi D, et al. A comprehensive review on current technologies for removal of endocrine disrupting chemicals from wastewaters. Environ Res. 2022;207:112196.
Cuerda-Correa EM, Alexandre-Franco MF, Fernández-González C. Advanced oxidation processes for the removal of antibiotics from water. An overview. Water. 2020. https://doi.org/10.3390/w12010102.
Loganathan P, et al. Treatment trends and combined methods in removing pharmaceuticals and personal care products from wastewater-a review. Membranes (Basel). 2023;13(2).
Oluwole AO, Omotola EO, Olatunji OS. Pharmaceuticals and personal care products in water and wastewater: a review of treatment processes and use of photocatalyst immobilized on functionalized carbon in AOP degradation. BMC Chem. 2020;14(1):62.
Massima Mouele ES, et al. Removal of pharmaceutical residues from water and wastewater using dielectric barrier discharge methods-a review. Int J Environ Res Public Health. 2021;18(4).
Ding Y, et al. Lignin peroxidase-catalyzed direct oxidation of trace organic pollutants through a long-range electron transfer mechanism: using propranolol as an example. J Hazard Mater. 2022;431:128544.
Kasonga TK, et al. Assessing the fungal simultaneous removal efficiency of carbamazepine, diclofenac and ibuprofen in aquatic environment. Front Microbiol. 2021;12.
Grelska A, Noszczyńska M. White rot fungi can be a promising tool for removal of bisphenol A, bisphenol S, and nonylphenol from wastewater. Environ Sci Pollut Res Int. 2020;27(32):39958–76.
Zhang X, et al. Enhancement of soil high-molecular-weight polycyclic aromatic hydrocarbon degradation by Fusarium sp. ZH-H2 using different carbon sources. Ecotoxicol Environ Saf. 2023;249:114379.
Chai C, et al. Immobilized lignin peroxidase on chitosan-modified halloysite nanotubes for degradation of polycyclic aromatic hydrocarbons in soil. Int J Environ Sci Technol. 2023;20(10):10877–86.
Wu L, et al. Computer-aided understanding and engineering of enzymatic selectivity. Biotechnol Adv. 2022;54:107793.
Li W, et al. Broadening the scope of biocatalysis engineering by tailoring enzyme microenvironment: a review. Catal Lett. 2022.
Wang Y, Chen J, Kang Z. In silico protein design promotes the rapid evolution of industrial enzymes. Biochemistry. 2019;58(11):1451–3.
Martínez AT, et al. Oxidoreductases on their way to industrial biotransformations. Biotechnol Adv. 2017;35(6):815–31.
Gong G, et al. Construction and identification of a food-grade recombinant cyberlindnera jadinii strain expressing lignin peroxidase. Appl Sci. 2023. https://doi.org/10.3390/app13106277.
Singh AK, et al. A predictive toolset for the identification of degradation pattern and toxic hazard estimation of multimeric hazardous compounds persists in water bodies. Sci Total Environ. 2022;824:153979.
Bhatt P, et al. Bioremediation potential of laccase for catalysis of glyphosate, isoproturon, lignin, and parathion: molecular docking, dynamics, and simulation. J Hazard Mater. 2023;443:130319.
Singh AK, et al. Trends in predictive biodegradation for sustainable mitigation of environmental pollutants: recent progress and future outlook. Sci Total Environ. 2021;770:144561.
Liu Z, et al. Application of molecular docking for the degradation of organic pollutants in the environmental remediation: a review. Chemosphere. 2018;203:139–50.
Li X, et al. Combined molecular docking, homology modelling and density functional theory studies to modify dioxygenase to efficiently degrade aromatic hydrocarbons. RSC Adv. 2019;9(20):11465–75.
Singh AK, Chowdhary P, Raj A. 13 - In silico bioremediation strategies for removal of environmental pollutants released from paper mills using bacterial ligninolytic enzymes, in Microorganisms for sustainable environment and health. In: Chowdhary P, et al. editors. Elsevier; 2020. p. 249–285.
Singh AK, et al. Deployment of oxidoreductases for sustainable biocatalytic degradation of selected endocrine-disrupting chemicals. Sustain Chem Pharm. 2023;31:100934.
Singh AK, et al. Assessing chemical hazard and unraveling binding affinity of priority pollutants to lignin modifying enzymes for environmental remediation. Chemosphere. 2023;313:137546.
Chen M, et al. Understanding lignin-degrading reactions of ligninolytic enzymes: binding affinity and interactional profile. PLoS ONE. 2011;6(9):e25647.
Alzate-Morales J, et al. Computational modeling methods for understanding the interaction of lignin and its derivatives with oxidoreductases as biocatalysts. Lignin Trends Appl. 2018;121.
Francesca Gerini M, et al. Molecular dynamics simulations of lignin peroxidase in solution. Biophys J. 2003;84(6):3883–93.
Phillips JC, et al. Scalable molecular dynamics with NAMD. J Comput Chem. 2005;26(16):1781–802.
Ecker J, Fülöp L. Lignin peroxidase ligand access channel dysfunction in the presence of atrazine. Sci Rep. 2018;8(1):5989.
Acknowledgements
AKS thankfully acknowledged the Academy of Scientific & Innovative Research (AcSIR)—an Institute of National Importance, the CSIR-Indian Institute of Toxicology Research, and also thanks to the Council of Scientific and Industrial Research (CSIR), India.
Funding
The NOBELIUM JOINING GDANSK TECH RESEARCH COMMUNITY program is graciously acknowledged for supporting this work under the international collaboration project awarded to Muhammad Bilal (grant number 44/2023/IDUB/I.1).
Author information
Authors and Affiliations
Contributions
Anil Kumar Singh: conceptualization, data curation, writing—original draft, project administration, figures, and final draft. Roberto Fernandez-Lafuente: writing—review and editing. Grzegorz Boczkaj: conceptualization, data curation, validation, and writing—review and editing. Jens Ejbye Schmidt: writing—review and editing, Muhammad Bilal: conceptualization, writing—original draft, review and editing, supervision, project administration, and funding acquisition.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Singh, A.K., Fernandez-Lafuente, R., Schmidt, J.E. et al. Biocatalytic Functionalities of Lignin Peroxidase-Based Systems in Lignin Depolymerization and Pollutants Removal from Environmental Matrices. Curr Pollution Rep (2024). https://doi.org/10.1007/s40726-024-00310-0
Accepted:
Published:
DOI: https://doi.org/10.1007/s40726-024-00310-0