Abstract
Purpose
Non-functioning pituitary neuroendocrine tumors are challengingly diagnosed tumors in the clinic. Transsphenoidal surgery remains the first-line treatment. Despite the development of state-of-the-art techniques, no drug therapy is currently approved for the treatment. There are also no randomized controlled trials comparing therapeutic strategies or drug therapy for the management after surgery. Therefore, novel therapeutic interventions for the therapeutically challenging NF-PitNETs are urgently needed.
Methods
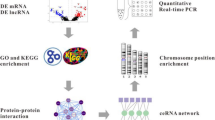
We integrated epigenome and transcriptome data (both coding and non-coding) that elucidate disease-specific signatures, in addition to biological and pharmacological data, to utilize rational pathway and drug prioritization in NF-PitNETs. We constructed an epigenome- and transcriptome-based PPI network and proposed hub genes. The signature-based drug repositioning based on the integration of multi-omics data was performed.
Results
The construction of a disease-specific network based on three different biological levels revealed DCC, DLG5, ETS2, FOXO1, HBP1, HMGA2, PCGF3, PSME4, RBPMS, RREB1, SMAD1, SOCS1, SOX2, YAP1, ZFHX3 as hub proteins. Signature-based drug repositioning using hub proteins yielded repositioned drug candidates that were confirmed in silico via molecular docking. As a result of molecular docking simulations, palbociclib, linifanib, trametinib, eplerenone, niguldipine, and zuclopenthixol showed higher binding affinities with hub genes compared to their inhibitors and were proposed as potential repositioned therapeutics for the management of NF-PitNETs.
Conclusion
The proposed systems’ biomedicine-oriented multi-omics data integration for drug repurposing to provide promising results for the construction of effective clinical therapeutics. To the best of our knowledge, this is the first study reporting epigenome- and transcriptome-based drug repositioning for NF-PitNETs using in silico confirmations.







Similar content being viewed by others
Data availability
The datasets that were used in this study are publicly available at Gene Expression Omnibus (GEO Database) with the following links: GSE115783– https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE115783. GSE77517– https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE77517. GSE63357– https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE63357.
References
Asa SL, Casar-Borota O, Chanson P, Delgrange E, Earls P, Ezzat S, Grossman A, Ikeda H, Inoshita N, Karavitaki N (2017) From pituitary adenoma to pituitary neuroendocrine tumor (PitNET): an international pituitary pathology club proposal. Endocr Relat Cancer 24:C5–C8
Trouillas J, Jaffrain-Rea M-L, Vasiljevic A, Raverot G, Roncaroli F, Villa C (2020) How to classify pituitary neuroendocrine tumors (PitNET) s in 2020. Cancers (Basel) 12:514
Ostrom QT, Gittleman H, Truitt G, Boscia A, Kruchko C, Barnholtz-Sloan JS (2018) CBTRUS statistical report: primary brain and other central nervous system tumors diagnosed in the United States in 2011–2015. Neuro Oncol. https://doi.org/10.1093/neuonc/noy131
Daly AF, Beckers A (2020) The Epidemiology of pituitary adenomas. Endocrinol Metab Clin North Am 49:347–355
Mercado M, Melgar V, Salame L, Cuenca D (2017) Clinically non-functioning pituitary adenomas: pathogenic, diagnostic and therapeutic aspects. Endocrinol diabetes y Nutr 64:384–395
Llyod RV, Osamura RY, Klöppel GRJ (2017) WHO classification of tumours of endocrine organs. IARC, Lyon
Torregrosa-Quesada ME, García-Martínez A, Sánchez-Barbie A, Silva-Ortega S, Cámara R, Fajardo C, Pico A (2021) The silent variants of pituitary tumors: demographic, radiological and molecular characteristics. J Endocrinol Invest 44:1637–1648
Vargas G, Gonzalez B, Ramirez C, Ferreira A, Espinosa E, Mendoza V, Guinto G, Lopez-Felix B, Zepeda E, Mercado M (2015) Clinical characteristics and treatment outcome of 485 patients with nonfunctioning pituitary macroadenomas. Int J Endocrinol. https://doi.org/10.1155/2015/756069
Chanson P, Raverot G, Castinetti F, Cortet-Rudelli C, Galland F, Salenave S (2015) Management of clinically non-functioning pituitary adenoma. Annales d’endocrinologie. Elsevier, pp 239–247
Penn DL, Burke WT, Laws ER (2018) Management of non-functioning pituitary adenomas: surgery. Pituitary 21:145–153
Lucas JW, Bodach ME, Tumialan LM, Oyesiku NM, Patil CG, Litvack Z, Aghi MK, Zada G (2016) Congress of neurological surgeons systematic review and evidence-based guideline on primary management of patients with nonfunctioning pituitary adenomas. Neurosurgery 79:E533–E535
Greenman Y, Stern N (2015) Optimal management of non-functioning pituitary adenomas. Endocrine 50:51–55
Even-Zohar N, Greenman Y (2018) Management of NFAs: medical treatment. Pituitary 21:168–175. https://doi.org/10.1007/s11102-018-0865-7
Brochier S, Galland F, Kujas M, Parker F, Gaillard S, Raftopoulos C, Young J, Alexopoulou O, Maiter D, Chanson P (2010) Factors predicting relapse of nonfunctioning pituitary macroadenomas after neurosurgery: a study of 142 patients. Eur J Endocrinol 163:193
Ilie MD, Raverot G (2020) Treatment options for gonadotroph tumors: current state and perspectives. J Clin Endocrinol Metab 105:e3507–e3518
Peverelli E, Giardino E, Treppiedi D, Meregalli M, Belicchi M, Vaira V, Corbetta S, Verdelli C, Verrua E, Serban AL (2017) Dopamine receptor type 2 (DRD2) and somatostatin receptor type 2 (SSTR2) agonists are effective in inhibiting proliferation of progenitor/stem-like cells isolated from nonfunctioning pituitary tumors. Int J cancer 140:1870–1880
Wang H, Chen K, Yang Z, Li W, Wang C, Zhang G, Zhu L, Liu P, Yang Y (2019) Diagnosis of invasive nonfunctional pituitary adenomas by serum extracellular vesicles. Anal Chem 91:9580–9589
Aydin B, Arga KY (2019) Co-expression network analysis elucidated a core module in association with prognosis of non-functioning non-invasive human pituitary adenoma. Front Endocrinol (Lausanne) 10:361
Gossing W, Frohme M, Radke L (2020) Biomarkers for liquid biopsies of pituitary neuroendocrine tumors. Biomedicines. https://doi.org/10.3390/biomedicines8060148
Hernández-Ramírez LC, Morgan RML, Barry S, D’Acquisto F, Prodromou C, Korbonits M (2018) Multi-chaperone function modulation and association with cytoskeletal proteins are key features of the function of AIP in the pituitary gland. Oncotarget. https://doi.org/10.18632/oncotarget.24183
Gadelha MR, Kasuki L, Dénes J, Trivellin G, Korbonits M (2013) MicroRNAs: Suggested role in pituitary adenoma pathogenesis. J Endocrinol Invest 36:889–895
Beylerli O, Beeraka NM, Gareev I, Pavlov V, Yang G, Liang Y, Aliev G (2020) MiRNAs as noninvasive biomarkers and therapeutic agents of pituitary adenomas. Int J Mol Sci 21:7287
Butz H, Likó I, Czirják S, Igaz P, Korbonits M, Rácz K, Patócs A (2011) MicroRNA profile indicates downregulation of the TGFβ pathway in sporadic non-functioning pituitary adenomas. Pituitary 14:112–124
Cheunsuchon P, Zhou Y, Zhang X, Lee H, Chen W, Nakayama Y, Rice KA, Hedley-Whyte ET, Swearingen B, Klibanski A (2011) Silencing of the imprinted DLK1-MEG3 locus in human clinically nonfunctioning pituitary adenomas. Am J Pathol 179:2120–2130
Li Z, Li C, Liu C, Yu S, Zhang Y (2015) Expression of the long non-coding RNAs MEG3, HOTAIR, and MALAT-1 in non-functioning pituitary adenomas and their relationship to tumor behavior. Pituitary 18:42–47. https://doi.org/10.1007/s11102-014-0554-0
Xing W, Qi Z, Huang C, Zhang N, Zhang W, Li Y, Qiu M, Fang Q, Hui G (2019) Genome-wide identification of lncRNAs and mRNAs differentially expressed in non-functioning pituitary adenoma and construction of an lncRNA-mRNA co-expression network. Biol Open. https://doi.org/10.1242/bio.037127
Klutstein M, Nejman D, Greenfield R, Cedar H (2016) DNA methylation in cancer and aging. Cancer Res 76:3446–3450
Srirangam Nadhamuni V, Korbonits M (2020) Novel insights into pituitary tumorigenesis: genetic and epigenetic mechanisms. Endocr Rev 41:821–846
Cheng S, Li C, Xie W, Miao Y, Guo J, Wang J, Zhang Y (2020) Integrated analysis of DNA methylation and mRNA expression profiles to identify key genes involved in the regrowth of clinically non-functioning pituitary adenoma. Aging (Albany NY) 12:2408
Vicchio TM, Aliquò F, Ruggeri RM, Ragonese M, Giuffrida G, Cotta OR, Ferraù F (2020) MicroRNAs expression in pituitary tumors: differences related to functional status, pathological features, and clinical behavior. J Endocrinol Investig 43:947–958
Kober P, Boresowicz J, Rusetska N, Maksymowicz M, Goryca K, Kunicki J, Bonicki W, Siedlecki JA, Bujko M (2018) DNA methylation profiling in nonfunctioning pituitary adenomas. Mol Cell Endocrinol 473:194–204. https://doi.org/10.1016/j.mce.2018.01.020
Edgar R, Domrachev M, Lash AE (2002) Gene Expression omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 30:207–210. https://doi.org/10.1093/nar/30.1.207
Smyth GK, Ritchie M, Thorne N, Wettenhall J (2005) LIMMA: linear models for microarray data. In Bioinformatics and Computational Biology Solutions Using R and Bioconductor, Statistics for Biology and Health
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, Ellis B, Gautier L, Ge Y, Gentry J (2004) Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 5:1–16
Aryee MJ, Jaffe AE, Corrada-Bravo H, Ladd-Acosta C, Feinberg AP, Hansen KD, Irizarry RA (2014) Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 30:1363–1369
Tian Y, Morris TJ, Webster AP, Yang Z, Beck S, Feber A, Teschendorff AE (2017) ChAMP: updated methylation analysis pipeline for Illumina BeadChips. Bioinformatics 33:3982–3984
Peters TJ, Buckley MJ, Statham AL, Pidsley R, Samaras K, Lord RV, Clark SJ, Molloy PL (2015) De novo identification of differentially methylated regions in the human genome. Epigenetics Chromatin 8:1–16
Teschendorff AE, Marabita F, Lechner M, Bartlett T, Tegner J, Gomez-Cabrero D, Beck S (2013) A beta-mixture quantile normalization method for correcting probe design bias in Illumina Infinium 450 k DNA methylation data. Bioinformatics 29:189–196. https://doi.org/10.1093/bioinformatics/bts680
Chen Y, Lemire M, Choufani S, Butcher DT, Grafodatskaya D, Zanke BW, Gallinger S, Hudson TJ, Weksberg R (2013) Discovery of cross-reactive probes and polymorphic CpGs in the Illumina Infinium HumanMethylation450 microarray. Epigenetics 8:203–209. https://doi.org/10.4161/epi.23470
Kamburov A, Stelzl U, Lehrach H, Herwig R (2013) The ConsensusPathDB interaction database: 2013 update. Nucleic Acids Res 41:D793–D800. https://doi.org/10.1093/nar/gks1055
Zhou Y, Zhou B, Pache L, Chang M, Khodabakhshi AH, Tanaseichuk O, Benner C, Chanda SK (2019) Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun 10:1523. https://doi.org/10.1038/s41467-019-09234-6
Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, Itoh M, Katayama T, Kawashima S, Okuda S, Tokimatsu T, Yamanishi Y (2008) KEGG for linking genomes to life and the environment. Nucleic Acids Res 36:D480–D484. https://doi.org/10.1093/nar/gkm882
The Gene Ontology Consortium (2019) The gene ontology resource: 20 years and still GOing strong. Nucleic Acids Res 47:D330–D338. https://doi.org/10.1093/nar/gky1055
Fabregat A, Sidiropoulos K, Garapati P, Gillespie M, Hausmann K, Haw R, Jassal B, Jupe S, Korninger F, McKay S, Matthews L, May B, Milacic M, Rothfels K, Shamovsky V, Webber M, Weiser J, Williams M, Wu G, Stein L, Hermjakob H, D’Eustachio P (2016) The reactome pathway knowledgebase. Nucleic Acids Res 44:D481–D487. https://doi.org/10.1093/nar/gkv1351
Cheng L, Wang P, Tian R, Wang S, Guo Q, Luo M, Zhou W, Liu G, Jiang H, Jiang Q (2019) LncRNA2Target v2.0: a comprehensive database for target genes of lncRNAs in human and mouse. Nucleic Acids Res 47:D140–D144. https://doi.org/10.1093/nar/gky1051
Liu C-J, Gao C, Ma Z, Cong R, Zhang Q, Guo A-Y (2017) lncRInter: a database of experimentally validated long non-coding RNA interaction. J Genet Genomics 44:265–268
Gov E, Arga KY (2016) Interactive cooperation and hierarchical operation of microRNA and transcription factor crosstalk in human transcriptional regulatory network. IET Syst Biol 10:219–228. https://doi.org/10.1049/iet-syb.2016.0001
Karagoz K, Sevimoglu T, Arga KY (2016) Integration of multiple biological features yields high confidence human protein interactome. J Theor Biol 403:85–96
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. https://doi.org/10.1101/gr.1239303
Chin C-H, Chen S-H, Wu H-H, Ho C-W, Ko M-T, Lin C-Y (2014) cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol 8(Suppl 4):S11. https://doi.org/10.1186/1752-0509-8-S4-S11
Campillos M, Kuhn M, Gavin A-C, Jensen LJ, Bork P (2008) Drug target identification using side-effect similarity. Science. https://doi.org/10.1126/science.1158140
Duan Q, Reid SP, Clark NR, Wang Z, Fernandez NF, Rouillard AD, Readhead B, Tritsch SR, Hodos R, Hafner M (2016) L1000CDS 2: LINCS L1000 characteristic direction signatures search engine. NPJ Syst Biol Appl 2:1–12
Kim S, Chen J, Cheng T, Gindulyte A, He J, He S, Li Q, Shoemaker BA, Thiessen PA, Yu B, Zaslavsky L, Zhang J, Bolton EE (2019) PubChem 2019 update: improved access to chemical data. Nucleic Acids Res 47:D1102–D1109. https://doi.org/10.1093/nar/gky1033
Turanli B, Gulfidan G, Arga KY (2017) Transcriptomic-Guided drug repositioning supported by a new bioinformatics search tool: Genexpharma. Omi A J Integr Biol. https://doi.org/10.1089/omi.2017.0127
Davis AP, Grondin CJ, Johnson RJ, Sciaky D, Wiegers J, Wiegers TC, Mattingly CJ (2021) Comparative Toxicogenomics Database (CTD): update 2021. Nucleic Acids Res 49:D1138–D1143. https://doi.org/10.1093/nar/gkaa891
Berman H, Henrick K, Nakamura H (2003) Announcing the worldwide protein data bank. Nat Struct Mol Biol 10:980
UniProt Consortium (2021) UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res. 49(D1):D480–D489. https://doi.org/10.1093/nar/gkaa1100
Velankar S, Best C, Beuth B, Boutselakis CH, Cobley N, Sousa Da Silva AW, Dimitropoulos D, Golovin A, Hirshberg M, John M (2010) PDBe: protein data bank in Europe. Nucleic Acids Res 38:D308–D317
Wass MN, Kelley LA, Sternberg MJE (2010) 3DLigandSite: predicting ligand-binding sites using similar structures. Nucleic Acids Res 38:W469–W473
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31:455–461
Barabási A-L, Gulbahce N, Loscalzo J (2011) Network medicine: a network-based approach to human disease. Nat Rev Genet 12:56–68. https://doi.org/10.1038/nrg2918
Beermann J, Piccoli M-T, Viereck J, Thum T (2016) Non-coding RNAs in development and disease: background, mechanisms, and therapeutic approaches. Physiol Rev 96:1297–1325
Mormando M, Puliani G, Barnabei A, Lauretta R, Bianchini M, Chiefari A, Russillo M, Cognetti F, Romano L, Appetecchia M (2020) A rare case of pituitary melanoma metastasis: a dramatic and prolonged response to dabrafenib-trametinib therapy. Front Endocrinol (Lausanne) 11:471
Anderson E, Heller RS, Lechan RM, Heilman CB (2018) Regression of a nonfunctioning pituitary macroadenoma on the CDK4/6 inhibitor palbociclib: case report. Neurosurg Focus 44:E9. https://doi.org/10.3171/2018.2.FOCUS17660
Chen Y, Li Z, Fang Q, Wang H, Li C, Gao H, Zhang Y (2020) CDKN2A (p16INK4A) affects the anti-tumor effect of CDK inhibitor in somatotroph adenomas. Int J Mol Med 47:500–510. https://doi.org/10.3892/ijmm.2020.4807
Campana C, Nista F, Castelletti L, Caputo M, Lavezzi E, Marzullo P, Gatto F (2022) Clinical and radiological presentation of parasellar ectopic pituitary adenomas: case series and systematic review of the literature. J Endocrinol Investig. https://doi.org/10.1007/s40618-022-01758-x
Fang Y, Fullwood MJ (2016) Roles, functions, and mechanisms of long non-coding RNAS in cancer. Genom Proteom Bioinformat 14:42–54. https://doi.org/10.1016/j.gpb.2015.09.006
Slaby O, Laga R, Sedlacek O (2017) Therapeutic targeting of non-coding RNAs in cancer. Biochem J 474:4219–4251. https://doi.org/10.1042/BCJ20170079
Du Q, Hu B, Feng Y, Wang Z, Wang X, Zhu D, Zhu Y, Jiang X, Wang H (2019) circOMA1-Mediated miR-145-5p suppresses tumor growth of nonfunctioning pituitary adenomas by targeting TPT1. J Clin Endocrinol Metab 104:2419–2434. https://doi.org/10.1210/jc.2018-01851
Cui M, Zhang M, Liu H-F, Wang J-P (2017) Effects of microRNA-21 targeting PITX2 on proliferation and apoptosis of pituitary tumor cells. Eur Rev Med Pharmacol Sci 21:2995–3004
Trivellin G, Butz H, Delhove J, Igreja S, Chahal HS, Zivkovic V, McKay T, Patócs A, Grossman AB, Korbonits M (2012) MicroRNA miR-107 is overexpressed in pituitary adenomas and inhibits the expression of aryl hydrocarbon receptor-interacting protein in vitro. Am J Physiol Endocrinol Metab 303:E708–E719. https://doi.org/10.1152/ajpendo.00546.2011
Boresowicz J, Kober P, Rusetska N, Maksymowicz M, Paziewska A, Dąbrowska M, Zeber-Lubecka N, Kunicki J, Bonicki W, Ostrowski J (2020) DNA Methylation Influences miRNA expression in gonadotroph pituitary tumors. Life 10:59
Haddick PCG, Tom I, Luis E, Quiñones G, Wranik BJ, Ramani SR, Stephan J-P, Tessier-Lavigne M, Gonzalez LC (2014) Defining the ligand specificity of the deleted in colorectal cancer (DCC) receptor. PLoS One 9:e84823
Nakamura H, Sudo T, Tsuiki H, Miyake H, Morisaki T, Sasaki J, Masuko N, Kochi M, Ushio Y, Saya H (1998) Identification of a novel human homolog of the Drosophila dlg, P‐dlg, specifically expressed in the gland tissues and interacting with p55. FEBS Lett 433:63–67
Dwyer J, Li HE, Xu D, LIU J (2007) Transcriptional regulation of telomerase activity: roles of the the Ets transcription factor family. Ann N Y Acad Sci 1114:36–47
Nakae J, Kitamura T, Kitamura Y, Biggs III WH, Arden KC, Accili D (2003) The forkhead transcription factor Foxo1 regulates adipocyte differentiation. Dev Cell 4:119–129
Sampson EM, Haque ZK, Ku M, Tevosian SG, Albanese C, Pestell RG, Paulson KE, Yee AS (2001) Negative regulation of the Wnt–β‐catenin pathway by the transcriptional repressor HBP1. EMBO J 20:4500–4511
Taherbhoy AM, Huang OW, Cochran AG (2015) BMI1-RING1B is an autoinhibited RING E3 ubiquitin ligase. Nat Commun 6:7621. https://doi.org/10.1038/ncomms8621
Taherbhoy AM, Huang OW, Cochran AG (2015) BMI1-RING1B is an autoinhibited RING E3 ubiquitin ligase. Nat Commun 6:7621. https://doi.org/10.1038/ncomms8621
Qian M-X, Pang Y, Liu CH, Haratake K, Du B-Y, Ji D-Y, Wang G-F, Zhu Q-Q, Song W, Yu Y, Zhang X-X, Huang H-T, Miao S, Chen L-B, Zhang Z-H, Liang Y-N, Liu S, Cha H, Yang D, Zhai Y, Komatsu T, Tsuruta F, Li H, Cao C, Li W, Li G-H, Cheng Y, Chiba T, Wang L, Goldberg AL, Shen Y, Qiu X-B (2013) Acetylation-mediated proteasomal degradation of core histones during DNA repair and spermatogenesis. Cell 153:1012–1024. https://doi.org/10.1016/j.cell.2013.04.032
Sun Y, Ding L, Zhang H, Han J, Yang X, Yan J, Zhu Y, Li J, Song H, Ye Q (2006) Potentiation of Smad-mediated transcriptional activation by the RNA-binding protein RBPMS. Nucleic Acids Res 34:6314–6326 . https://doi.org/10.1093/nar/gkl914
Date S, Nibu Y, Yanai K, Hirata J, Yagami K-I, Fukamizu A (2004) Finb, a multiple zinc finger protein, represses transcription of the human angiotensinogen gene. Int J Mol Med 13:637–642
Lin W, Karin M (2007) A cytokine-mediated link between innate immunity, inflammation, and cancer. Am Soc Clin Investig 117:1175–1183
Kamizono S, Hanada T, Yasukawa H, Minoguchi S, Kato R, Minoguchi M, Hattori K, Hatakeyama S, Yada M, Morita S, Kitamura T, Kato H, Ki N, Yoshimura A (2001) The SOCS box of SOCS-1 accelerates ubiquitin-dependent proteolysis of TEL-JAK2. J Biol Chem 276:12530–12538 . https://doi.org/10.1074/jbc.M010074200
Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, Tomoda K, Yamanaka S (2007) Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131:861–872 . https://doi.org/10.1016/j.cell.2007.11.019
Zhao B, Wei X, Li W, Udan RS, Yang Q, Kim J, Xie J, Ikenoue T, Yu J, Li L, Zheng P, Ye K, Chinnaiyan A, Halder G, Lai Z-C, Guan K-L (2007) Inactivation of YAP oncoprotein by the Hippo pathway is involved in cell contact inhibition and tissue growth control. Genes Dev 21:2747–2761. https://doi.org/10.1101/gad.1602907
Mabuchi M, Kataoka H, Miura Y, Kim T-S, Kawaguchi M, Ebi M, Tanaka M, Mori Y, Kubota E, Mizushima T, Shimura T, Mizoshita T, Tanida S, Kamiya T, Asai K, Joh T (2010) Tumor suppressor, AT motif binding factor 1 (ATBF1), translocates to the nucleus with runt domain transcription factor 3 (RUNX3) in response to TGF-beta signal transduction. Biochem Biophys Res Commun 398:321–325. https://doi.org/10.1016/j.bbrc.2010.06.090
Lv T, Zhang Z, Yu H, Ren S, Wang J, Li S et al (2022) Tamoxifen exerts anticancer effects on pituitary adenoma progression via inducing cell apoptosis and inhibiting cell migration. Int J Mol Sci 23:2664
Raverot G, Burman P, McCormack A, Heaney A, Petersenn S, Popovic V et al (2018) European society of endocrinology clinical practice guidelines for the management of aggressive pituitary tumours and carcinomas. Eur J Endocrinol 178:G1–24
Yavropoulou MP, Tsoli M, Barkas K, Kaltsas G, Grossman A (2020) The natural history and treatment of non-functioning pituitary adenomas (non-functioning PitNETs). Endocr Relat Cancer 27:R375–R390
Toogood PL, Harvey PJ, Repine JT, Sheehan DJ, VanderWel SN, Zhou H, Keller PR, McNamara DJ, Sherry D, Zhu T, Brodfuehrer J, Choi C, Barvian MR, Fry DW (2005) Discovery of a potent and selective inhibitor of cyclin-dependent kinase 4/6. J Med Chem 48:2388–2406. https://doi.org/10.1021/jm049354h
Turner NC, Ro J, André F, Loi S, Verma S, Iwata H, Harbeck N, Loibl S, Huang Bartlett C, Zhang K, Giorgetti C, Randolph S, Koehler M, Cristofanilli M (2015) Palbociclib in hormone-receptor–positive advanced breast cancer. N Engl J Med 373:209–219. https://doi.org/10.1056/NEJMoa1505270 (Epub 2015 Jun 1. PMID: 26030518)
Italiano A, Bianchini L, Keslair F, Bonnafous S, Cardot-Leccia N, Coindre J, Dumollard J, Hofman P, Leroux A, Mainguené C (2008) HMGA2 is the partner of MDM2 in well-differentiated and dedifferentiated liposarcomas whereas CDK4 belongs to a distinct inconsistent amplicon. Int J cancer 122:2233–2241
Fedele M, Visone R, De Martino I, Troncone G, Palmieri D, Battista S, Ciarmiello A, Pallante P, Arra C, Melillo RM (2006) HMGA2 induces pituitary tumorigenesis by enhancing E2F1 activity. Cancer Cell 9:459–471
Aversa C, Leone F, Zucchini G, Serini G, Geuna E, Milani A, Valdembri D, Martinello R, Montemurro F (2015) Linifanib: current status and future potential in cancer therapy. Expert Rev Anticancer Ther 15:677–687
Tan E-H, Goss GD, Salgia R, Besse B, Gandara DR, Hanna NH, Yang JC-H, Thertulien R, Wertheim M, Mazieres J (2011) Phase 2 trial of Linifanib (ABT-869) in patients with advanced non-small cell lung cancer. J Thorac Oncol 6:1418–1425
Cainap C, Qin S, Huang W-T, Chung IJ, Pan H, Cheng Y, Kudo M, Kang Y-K, Chen P-J, Toh H-C (2015) Linifanib versus Sorafenib in patients with advanced hepatocellular carcinoma: results of a randomized phase III trial. J Clin Oncol 33:172
Wang ES, Yee K, Koh LP, Hogge D, Enschede S, Carlson DM, Dudley M, Glaser K, McKeegan E, Albert DH (2012) Phase 1 trial of linifanib (ABT-869) in patients with refractory or relapsed acute myeloid leukemia. Leuk Lymphoma 53:1543–1551
Albert DH, Tapang P, Magoc TJ, Pease LJ, Reuter DR, Wei R-Q, Li J, Guo J, Bousquet PF, Ghoreishi-Haack NS (2006) Preclinical activity of ABT-869, a multitargeted receptor tyrosine kinase inhibitor. Mol Cancer Ther 5:995–1006
Ortiz LD, Syro LV, Scheithauer BW, Ersen A, Uribe H, Fadul CE, Rotondo F, Horvath E, Kovacs K (2012) Anti-VEGF therapy in pituitary carcinoma. Pituitary 15:445–449
Wright CJM, McCormack PL (2013) Trametinib: first global approval. Drugs 73:1245–1254
Robert C, Karaszewska B, Schachter J, Rutkowski P, Mackiewicz A, Stroiakovski D, Lichinitser M, Dummer R, Grange F, Mortier L (2015) Improved overall survival in melanoma with combined dabrafenib and trametinib. N Engl J Med 372:30–39
Walters DM, Lindberg JM, Adair SJ, Newhook TE, Cowan CR, Stokes JB, Borgman CA, Stelow EB, Lowrey BT, Chopivsky ME (2013) Inhibition of the growth of patient derived pancreatic cancer xenografts with the MEK inhibitor trametinib is augmented by combined treatment with the epidermal growth factor receptor HER2 inhibitor lapatinib. Neoplasia 15(2):143-IN10
Planchard D, Besse B, Groen HJM, Souquet P-J, Quoix E, Baik CS, Barlesi F, Kim TM, Mazieres J, Novello S (2016) Dabrafenib plus trametinib in patients with previously treated BRAFV600E-mutant metastatic non-small cell lung cancer: an open-label, multicentre phase 2 trial. Lancet Oncol 17:984–993
Corcoran RB, Atreya CE, Falchook GS, Kwak EL, Ryan DP, Bendell JC, Hamid O, Messersmith WA, Daud A, Kurzrock R (2015) Combined BRAF and MEK inhibition with dabrafenib and trametinib in BRAF V600–mutant colorectal cancer. J Clin Oncol 33:4023
Borthakur G, Popplewell L, Boyiadzis M, Foran JM, Platzbecker U, Vey N, Roland WB, Olin RL, Raza A, Giagounidis A (2012) Phase I/II trial of the MEK1/2 inhibitor trametinib (GSK1120212) in relapsed/refractory myeloid malignancies: evidence of activity in patients with RAS mutation-positive disease. Blood. https://doi.org/10.1182/blood.V120.21.677.677
Sollfrank L, Lettmaier S, Erdmann M, Uslu U (2019) Panniculitis under successful targeted inhibition of the MAPK/ERK signaling pathway in a patient with BRAF V600E-mutated spindle cell oncocytoma of the pituitary gland. Anticancer Res 39:3955–3959
Brastianos PK, Shankar GM, Gill CM, Taylor-Weiner A, Nayyar N, Panka DJ, Sullivan RJ, Frederick DT, Abedalthagafi M, Jones PS (2016) Dramatic response of BRAF V600E mutant papillary craniopharyngioma to targeted therapy. JNCI J Natl Cancer Inst. https://doi.org/10.1093/jnci/djv310
Boer R, Grassegger A, Schudt C, Glossmann H (1989) (+)-Niguldipine binds with very high affinity to Ca2+ channels and to a subtype of α1-adrenoceptors. Eur J Pharmacol Mol Pharmacol 172:131–145
Klöckner U, Isenberg G (1989) The dihydropyridine niguldipine modulates calcium and potassium currents in vascular smooth muscle cells. Br J Pharmacol 97:957
Lee AR, Seo MJ, Kim J, Lee DM, Kim IY, Yoon MJ, Hoon H, Choi KS (2019) Lercanidipine synergistically enhances bortezomib cytotoxicity in cancer cells via enhanced endoplasmic reticulum stress and mitochondrial Ca2+ Overload. Int J Mol Sci 20:6112
Wee S (2015) Identification of compounds that target glioma initiating cells. Inst för fysiologi och farmakologi/Dept of Physiology and Pharmacology
Kenny B, Ballard S, Blagg J, Fox D (1997) Pharmacological options in the treatment of benign prostatic hyperplasia. J Med Chem 40:1293–1315
Kulig K, Malawska B (2006) Trends in the development of new drugs for treatment of benign prostatic hyperplasia. Curr Med Chem 13:3395–3416
Gibson RC, Fenton M, da Coutinho E, SF, Campbell C, (2004) Zuclopenthixol acetate for acute schizophrenia and similar serious mental illnesses. Cochrane Database Syst Rev. John Wiley, Chichester
Khalifa AE (2004) Pro-oxidant activity of zuclopenthixol in vivo: differential effect of the drug on brain oxidative status of scopolamine-treated rats. Hum Exp Toxicol 23:439–445
Fond G, Macgregor A, Tamouza R, Hamdani N, Meary A, Leboyer M, Dubremetz J-F (2014) Comparative analysis of anti-toxoplasmic activity of antipsychotic drugs and valproate. Eur Arch Psychiatry Clin Neurosci 264:179–183
Bourdoulous S, Faure C (2020) Zuclopenthixol hydrochloride derivatives and Ebselen derivatives as ErbB2 inhibitors.Google Patents. https://patents.google.com/patent/WO2017121755A1/en.
Stier CT (2003) Eplerenone: a selective aldosterone blocker. Cardiovasc Drug Rev 21:169–184. https://doi.org/10.1111/j.1527-3466.2003.tb00114.x
Weinberger MH, Roniker B, Krause SL, Weiss RJ (2002) Eplerenone, a selective aldosterone blocker, in mild-to-moderate hypertension. Am J Hypertens 15:709–716
Pitt B, Remme W, Zannad F, Neaton J, Martinez F, Roniker B, Bittman R, Hurley S, Kleiman J, Gatlin M (2003) Eplerenone, a selective aldosterone blocker, in patients with left ventricular dysfunction after myocardial infarction. N Engl J Med 348:1309–1321
Zannad F, McMurray JJV, Krum H, van Veldhuisen DJ, Swedberg K, Shi H, Vincent J, Pocock SJ, Pitt B (2011) Eplerenone in patients with systolic heart failure and mild symptoms. N Engl J Med 364:11–21
Zhao M, Célérier I, Bousquet E, Jeanny J-C, Jonet L, Savoldelli M, Offret O, Curan A, Farman N, Jaisser F (2012) Mineralocorticoid receptor is involved in rat and human ocular chorioretinopathy. J Clin Investig 122:2672–2679
Epstein M, Williams GH, Weinberger M, Lewin A, Krause S, Mukherjee R, Patni R, Beckerman B (2006) Selective aldosterone blockade with eplerenone reduces albuminuria in patients with type 2 diabetes. Clin J Am Soc Nephrol 1:940–951
Shavit L, Lifschitz MD, Epstein M (2012) Aldosterone blockade and the mineralocorticoid receptor in the management of chronic kidney disease: current concepts and emerging treatment paradigms. Kidney Int 81:955–968
Aydin B, Arslan S, Bayraklı F, Karademir B, Arga KY (2022) MicroRNA-Mediated drug repurposing unveiled potential candidate drugs for prolactinoma treatment. Neuroendocrinology 112:161–173
Aydin B, Caliskan A, Arga KY (2021) Overview of omics biomarkers in pituitary neuroendocrine tumors to design future diagnosis and treatment strategies. EPMA J 12:383–401
Aydin B, Yildirim E, Erdogan O, Arga KY, Yilmaz BK, Bozkurt SU et al (2022) Past, present, and future of therapies for pituitary neuroendocrine tumors: need for omics and drug repositioning guidance. Omi A J Integr Biol 26:115–129
Neou M, Villa C, Armignacco R, Jouinot A, Raffin-Sanson ML, Septier A, Assié G (2020) Pangenomic classification of pituitary neuroendocrine tumors. Cancer Cell 37:123–134
Mete O, Ezzat S, Perry A, Yamada S, Uccella S, Grossman AB, Asa SL (2021) The pangenomic classification of pituitary neuroendocrine tumors: quality histopathology is required for accurate translational research. Endocr Pathol 32(3):415–417
Di Somma C, Scarano E, de Alteriis G, Barrea L, Riccio E, Arianna R, Colao A (2021) Is there any gender difference in epidemiology, clinical presentation and co-morbidities of non-functioning pituitary adenomas? A prospective survey of a national referral center and review of the literature. J Endocrinol Investig 44:957–968
Acknowledgements
The scholarships under the YOK 100/2000 Doctoral Fellowship Program and 2211-C Doctoral Fellowship Program under The Scientific and Technological Research Council of Turkey (TUBITAK), and the financial support provided to Busra Aydin under project number 3629 from the Health Institutes of Turkey (TUSEB) are greatly acknowledged.
Author information
Authors and Affiliations
Contributions
BA designed data analysis framework. BA and HB performed the data analyses. BA, BT interpreted the results. FB and KYA conceived and directed the study. BA drafted the manuscript. BA and BT revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Ethical approval
The authors declare that the datasets used in this study belonged to previously published studies, therefore no ethical statement is required.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
40618_2022_1923_MOESM1_ESM.xlsx
Supplementary file1 Supplementary Information 1: The associations of ncRNAs with target genes are represented as networks. A) The network displayed the interactions of ncRNAs in literature and their known targets. B) The network was composed of the DEncRNAs in NF-PitNET dataset and their target genes that were retrieved from literature (XLSX 11418 KB)
40618_2022_1923_MOESM2_ESM.tif
Supplementary file2 Supplementary Information 2: LncRNA and miRNA target-gene interactions retrieved from various databases (TIF 1648 KB)
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Aydin, B., Beklen, H., Arga, K.Y. et al. Epigenomic and transcriptomic landscaping unraveled candidate repositioned therapeutics for non-functioning pituitary neuroendocrine tumors. J Endocrinol Invest 46, 727–747 (2023). https://doi.org/10.1007/s40618-022-01923-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40618-022-01923-2