Abstract
Background and Objective
Human epidermal growth factor receptor 2 (HER2) protein overexpression is one of the most significant biomarkers for breast cancer diagnostics, treatment prediction, and prognostics. The high accessibility of HER2 inhibitors in routine clinical practice directly translates into the diagnostic need for precise and robust marker identification. Even though multigene next-generation sequencing methodologies have slowly taken over the field of single-biomarker molecular tests, the copy number alterations such as amplification of the HER2-coding ERBB2 gene are hard to validate on next-generation sequencing platforms as they are characterized by chromosomal structural heterogeneity, polysomy, and genomic context of ploidy. In our study, we tested the approach of using whole genome sequencing instead of next-generation sequencing panels to determine HER2 status in the clinical set-up.
Methods
We used a large dataset of 876 patients with breast cancer whole genomes with curated clinical data and an additional set of 551 patients’ external genomic data. We used the decision-tree-based algorithm for optimization of the diagnostic tool for HER2 status assessment by whole genome sequencing.
Results
The most efficient approach to assess HER2 status in whole genome sequencing data was the ploidy-corrected copy number, utilizing ERBB2 copy number and mean tumor ploidy. The classifier achieved sensitivity of 91.18% and specificity of 98.69% on the internal validation dataset and 89.86% and 96.06% on the external data, which is similar to other next-generation sequencing methods, currently tested in the clinic.
Conclusions
We provide evidence that the HER2 status may be reliably determined by whole genome sequencing and is applicable across different laboratory protocols and pipelines. We suggest using the ploidy-corrected copy number for diagnostic purposes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Whole genome sequencing is a reliable method for clinical human epidermal growth factor receptor 2 status assessment. |
The ploidy-corrected copy number value is the most accurate biomarker for genetic testing with high concordance to gold standard immunohistochemistry and fluorescence in situ hybridization. |
Short-read whole genome sequencing for human epidermal growth factor receptor 2 assessment is consistent across different platforms and wet lab protocols. |
1 Introduction
Human epidermal growth factor receptor 2 (HER2) is an important biomarker for targeted therapy in breast cancer (BC). Patients with an overexpression of the receptor were considered the worst prognosis group before HER2 inhibitors were introduced into clinical practice [1]. Currently, the first and second generation of these drugs slow down disease progression, improving the outcomes in HER2-positive subgroups of BCs. Therefore, it is crucial to pinpoint the HER2-overexpression status accurately and precisely [2].
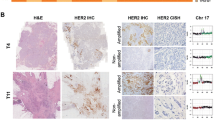
The molecular mechanism of HER2 overexpression is, in most cases, amplification of a 17q12 chromosome region containing the HER2 coding ERBB2 gene. The reference method for the assessment of ERBB2 amplification is immunohistochemistry (IHC) coupled with fluorescence in situ hybridization (FISH) [3]. Currently, diagnostic companies and medical services are beginning to offer novel next-generation sequencing (NGS) assays, detecting dozens of actionable biomarkers in a single test. They are trying to incorporate the ERBB2 copy number (ERBB2 CN) into their portfolio as well. Unfortunately, ERBB2 amplification status cannot be easily determined by establishing a simple threshold for negative and positive values, as the genomic context of chromosome 17 copy number and tumor ploidyFootnote 1 are interrelated with ERBB2 CN [4]. First, duplication or triplication of the whole chromosome set (polyploidy) or just a subset of chromosomes (aneuploidy) is a common feature of BC [5, 6]. However, changes in ploidy may not be associated with an overexpression of the ERBB2 gene, as average global transcript levels remain unchanged. On the contrary, Newcombe et al. noted a decrease in HER2 expression in recurrent polyploid BC cells [7]. Second, the isolated deletion or duplication events of chromosome 17 may influence the ERBB2 transcription [8, 9]. The gain of an additional copy of chromosome 17, called polysomy, is correlated with tumor ploidy and is considered its surrogate in the FISH test, but discrepancies between these parameters are in part the reason for inaccuracy in ERBB2 amplification detection [10].
As it is not feasible to determine ploidy in conventional FISH, the ratio between ERBB2 CN and chromosome 17 centromeric probe (CEP17) CN serves as a diagnostic criterion in dual-probe assays, recommended by the official American Society of Clinical Oncology/College of American Pathologists (ASCO/CAP) clinical practice guidelines for diagnostics of HER2 in patients with BC [3].
In contrast, whole genome sequencing (WGS) is capable of acquiring absolute ERBB2 CN, CEP17 CN, and mean ploidy of tumor cells simultaneously. Moreover, WGS can confirm the presence of the neoplastic cell in the sample, providing quality control of the material for analyses [11]. As WGS is mainly based on polymerase chain reaction-free methodology, it preserves the original proportions of DNA fragments, in contrast to enrichment or PCR-based NGS panels, which may distort the original proportions of DNA fragments and skew the quantification [12].
The purpose of this study was to determine the feasibility of accurately distinguishing between HER2-positive and HER2-negative cases of BC based on matched tumor-normal WGS. To date, there have been only a few studies evaluating the clinical utility of NGS testing of ERBB2 gene status, including the WGS method [4, 11, 13,14,15]. Some of them directly address the clinical need to verify the relevance of their findings for patient management, reporting the overall concordance between IHC/FISH and NGS at about a 90% level.
Our study operates on the large population-based cohort of 876 BCs from publicly available databases and additional external secondary data from 551 patients, supplied with the final clinical HER2 status based on ASCO/CAP guidelines and targeted treatment information, which serves to validate metastatic sample status. We analyzed the whole cohort of patients, aiming to establish the criteria for WGS ERBB2 status assessment as close to the gold standard as possible, optimized for both sensitivity and precision with a bias-free machine learning approach. We also provide the proof of concept that genomic data, acquired on different platforms with different chemistry, yield sufficiently uniform results for molecular diagnostics of ERBB2 amplification by WGS.
2 Materials and Methods
2.1 Sample Choice
Matched tumor-normal genomes from 876 patients with BC sequenced within three large Genomic Consortia (119, International Cancer Genome Consortium; 70, The Cancer Genome Atlas; 688, Hartwig Medical Foundation [HMF]) were downloaded from controlled-access databases after meeting formal criteria [13, 16,17,18]. The samples were sequenced using a low PCR amplification or PCR-free library preparation protocols and paired-end 100–150 base-pair Illumina reads with a 350–550 base-pair insert size (for details, see Table 1 of the Electronic Supplementary Material [ESM]). For analyses of primary tumor samples, we included the datasets with clinical HER2 status described as positive or negative, according to ASCO/CAP guidelines 2007–18 (depending on the year the original study was conducted, see Table 1 of the ESM). For metastatic/advanced tumor samples from the HMF database, metadata on HER2 status were available only for primary tumors, the IHC/FISH status for sequenced sample from the second biopsy was not provided. Because of the high rate of conversion from HER2-negative to HER2-positive status (and vice versa) during the cancer evolution [4, 11], in metastatic cancers, we have taken into consideration also the patients’ treatment metadata and discarded all samples for which treatment history (pre-biopsy and post-biopsy) was discordant with initial HER2 status (e.g., if trastuzumab was included in any line of treatment even though HER2 status was reported negative). For details on discarded samples, see the ESM.
Additional genomic data from an external pipeline were also used for validation purposes. Secondary data, derived from 560 BC genomes, were previously published by Nik-Zainal et al. [13]. From these data, we have extracted the complete information about clinical HER2 status, ploidy, purity, ERBB2 CN, and CEP17 CN of 551 patients. These data are also available in the ESM.
As there were no new tissue/DNA/RNA samples processed, the written consent of each subject is in possession of data providers. The primary data were collected in accordance with the standards set by the Declaration of Helsinki and the highest data security standards of ISO 27001.
2.2 Whole-Genome Data Processing
The files downloaded from HMF, The Cancer Genome Atlas, and the International Cancer Genome Consortium were analyzed using publicly available, open-source software embedded within an in-house pipeline (Fig. 1) implemented using Ruffus [19]. The analysis started with FASTQ file extraction from the BAM/CRAM files using Broad Institutes’ Picard tools [20]. Tumor samples with coverage exceeding 75× were downsampled with Seqtk version 1.3-r106 [21] to approximately 60× mean coverage. Next, all reads were trimmed using cutadapt version 2.10 [22] and mapped to the GRCh37 genome using Sanger’s Cancerit CGPMAP pipeline version 3.0.0 [23]. Samples with uniquely mapped read coverage below 20× for either tumor or normal genomes were excluded from the analysis [24, 25]. Mean tumor sample coverage across all datasets after downsampling was 48×, reference blood/EBV-transformed lymphocyte sample mean coverage was 36× (detailed data are provided in Table 2 and Fig. 1 of the ESM).
Variant calling was performed using Sanger’s Cancerit CGPWGS pipeline version 2.0.1 [23], and specifically copy number variants, purity, and ploidy were identified with ascatNgs [26]. Identified variants were annotated using Ensembl VEP version 102 [27].
The external validation data of Sanger’s Institute were processed by the Wellcome Sanger Institute as described in the original study, with a key step of copy number variant calling performed with ascatNgs [26]. The samples with incomplete/missing/inconsistent clinical data, failed processing, or a low depth of coverage were discarded (see Fig. 2 and the ESM).
2.3 Analyzed Parameters and Method Validation
In the study, we used clinical data on HER2 status according to ASCO/CAP recommendations. In the case of HMF metastatic/advanced tumors, pathomorphological evaluation data were available only at the point of diagnosis and were not supplied for actual WGS biopsies. HER2 status may shift in metastatic cancers, and without the information about the latest HER2 assessment, samples from HMF could have been wrongly labeled. To tackle this problem, we have also evaluated the metadata of the presence of targeted treatment with HER2 inhibitors in these samples and excluded all discrepant instances, in which therapy of metastatic cancer was not in compliance with initial HER2 status.
Based on ASCAT copy number alteration calling, ERBB2 (NC_000017.10:37844167_37886679) and the uniquely mapped 8250 bp sequence adjacent to CEP17 (NC_000017.10:22236000_22244250), copy numbers were extracted along with ploidy and purity estimation for all the tumor samples. The data were used to create three features for HER2 status assessment: absolute ERBB2 CN, ERBB2 CN-n (ploidy-adjusted ERBB2 CN), and ERBB2 CN/CEP17 CN ratio. Based on these features, a machine learning-based classifier was constructed, which determined the best approach for HER2 status discrimination. Six hundred and fourteen samples from the datasets were used as a training set (discovery cohort), the remaining 264 samples served as a validation hold-out set for the classifier and were not analyzed a priori. An additional external dataset processed by Wellcome Sanger Institute consisted of 551 samples, for which the same coordinates were used for extracting CEP17 CN and ERBB2 CN.
A decision tree-based classifier was chosen after comparing the effectiveness of logistic regression, random forest, and decision-tree models. After tuning the hyperparameters of each classifier, all three approaches achieve nearly identical performance, with the most robust decision tree model performing the best on average (for more details, see the ESM).
For the decision-tree-based modeling, the discovery cohort was randomly split into a training (75%) and a test set (25%). As the number of samples in IHC/FISH HER2-positive and HER2-negative groups was unbalanced (there were almost eight times less HER2-positive samples than HER2-negative samples), we added class weights (8:1) for compensation. After constructing the model, we measured its performance on 264 samples from the validation set. We used accuracy, precision, and recall along with the F1 score. Cohen’s Kappa score was estimated to evaluate the non-randomness of classification.
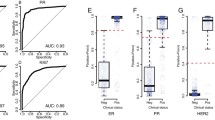
To show how each of the three features influences the classifier’s performance alone, we have established the same parameters independently for each of them as well and compared all the approaches with random data classification methods (Fig. 3). To further test the validity of our results, we decided to evaluate whether differences in tumor purity, heterogeneity of ploidy, or differences in mean depth of coverage had any deteriorative effects on the correctness of the results. For these experiments, we divided the samples into two near-equinumerous groups for each comparison and evaluated the differences in the tests’ performance.
Training set cross-validation accuracy comparison between three features used to determine human epidermal growth factor receptor 2 (HER2) amplification status in whole genome sequencing (WGS) data. Because of a class imbalance, the plot also includes a reference classifier assigning all samples to the majority class (DummyMostFrequent), which serves as a simple baseline
As the most simplistic model with one feature and a predetermined threshold was optimal, it was then used instead of ML to establish HER2 status for the test set and the external data. For analytical validation, we have determined the overall predictive value, positive predictive value (PPV), and negative predictive value (NPV) with confidence intervals (CIs) separately for the test set as well as for an external Sanger dataset of 551 patients (Table 1).
3 Results
In the analyzed dataset, 159 patients were categorized as triple-negative BC (18%), among HER2-negative patients, ER+/HER− accounted for 599 (88%). One hundred and ten samples (13%) were identified by clinical testing as HER2 positive, among them: 74 ER+/HER2+ (8%), 36 ER−/HER2+ (4%). For eight patients, ER status was unavailable.
HER2 positivity was slightly underrepresented in favor of triple-negative BC in comparison with statistics for the Caucasian population (18%), which may be an accidental or sampling bias related to the Genomic Consortia’s sample collection process, or an effect of discarding datasets with incomplete clinical data. The decision-tree machine learning approach has demonstrated the best discrimination between HER2-positive and HER2-negative cases based on a single-parameter, ploidy-corrected ERBB2 CN with a threshold of 2.265 (Fig. 3). The decision tree algorithm was evaluated in a three-fold cross-validation repeated ten times to estimate the mean value and standard deviation for each metric. The results were as follows: accuracy = 96.7% (± 0.87%), precision = 86% (± 5%), recall = 89% (± 6%), Cohen’s Kappa = 85% (± 3.7%), and F1 = 87% (± 3%). A high value of Cohen’s Kappa strongly indicates that our model classifies samples in a non-random manner.
The learning curve displayed no further improvement with sample numbers exceeding 150 instances; therefore, we believe the results display the best reflection of the biological phenomenon of HER2 amplification we could extract from genomic data. Moreover, a principal component analysis of the dataset (Fig. 4) has shown a very good and robust separation of data into two groups, representing differences in HER2 status.
As data distribution across depths of coverage, tumor purities, and ploidies was not normal (Tables 2–3 and Figs. 1–2 of the ESM), we decided to compare the accuracy distributions for these parameters with the Wilcoxon signed-rank test. The evaluation of results across data coverages has shown no significant differences (p > 0.05) between groups.
The comparison of low vs high purity also has not yielded significant differences (p > 0.05). However, there is a significant decrease in the mean accuracy of the test from 0.97 to 0.94, dependent on an increased tumor ploidy above two (p = 5.1 × 10-6) (Fig. 5).
As the best classification of samples was achieved by a single feature approach, the final classifier was reduced to a single feature. The use of the ML approach was therefore not necessary for further analyses of HER2 status in the analytical validation step, thus a simple threshold was used instead. The analytical validation of the ploidy-corrected ERBB2_CN method gave the diagnostic sensitivity of 91.18% (95% CI 76.32–98.14) and specificity of 98.69% (95% CI 96.22–99.73) for the hold-out dataset. In the external dataset, sensitivity was 89.86% (95% CI 80.21–95.82) and specificity was 96.06% (95% CI 93.91–97.61). For details, see Table 1.
4 Discussion
Decreasing NGS prices and increasing availability of this technology in medical practice have encouraged the transition from conventional cytogenetic and molecular methods to NGS in contemporary oncology. However, the evidence on the reliability of NGS techniques in the clinical use for copy number detection is still very limited. As the HER2 protein is one of the most significant biomarkers for BC diagnostics, targeted treatment response prediction, and prognostics, there were several attempts to show the applicability of NGS techniques in this indication.
The largest analytical validation study was conducted by Memorial Sloan Kettering on their proprietary MSK-IMPACT Assay [4]. This hybrid-capture-based panel NGS test was analyzed in 213 BC samples and evaluated in a clinical setting on further 599 samples. The cut-off for a positive result was established based solely on ERBB2 CN, adjusted to the background and normal signal of diploid genomes (defined as a ‘fold change’ of 1.5). The group reported 95% specificity and 100% sensitivity on > 10% of tumor content, with IHC/FISH evaluated by the newest2018 guidelines, and a dual-probe FISH assay [4]. In 2020, a continuation of the study exploited the borderline cases with excellent concordance [15]. Several other studies have also proven the clinical value of panel NGS for HER2 testing in BC and other solid tumors, with the same strategy of fold change determination, using either Illumina [1,2,3, 8] or Ion Torrent short-read methodology [9]. Another approach to define the ERBB2 amplification by panel NGS was using a cut-off of 2 standard deviations from the median depth of coverage across on-target data in a pool of samples [28, 29]. The question remains how universal this strategy is depending on the size of the panel used by different groups. The ‘resolution’ determined by the number of on-target sites and the presence of additional single nucleotide variant (SNV) targeting probes scattered across the genome (SNV ‘backbone’) greatly influence the ability to properly call the copy number alterations and normalize the data. The choice of the ‘panel of normals’ for data normalization may also influence the output, as different populations vary in inborn copy number variants. Hence, panel NGS strategies for normalizing any copy number variation in cancer should be cautiously validated for different methodologies, pipelines, and populations.
As new long-read sequencing technologies gain popularity, new methods of copy number detection in cancer are emerging. Nattestad et al. have described the high efficiency of PacBio SMRT technology in ERBB2 amplification detection in cell lines [30]. The main advantage of using this technique is the ability to explain the structure of a particular rearrangement. However useful, this feature may not be crucial from the diagnostic point of view. Moreover, the technology was not tested in patient samples yet, especially in degraded formalin-fixed paraffin embedded (FFPE) material [30].
There are also attempts to use cell-free/plasma DNA as starting material for copy number determination in NGS [31], and approaches to use targeted RNA sequencing on tumor tissue derived from FFPE [32]. The drawbacks of these methods are the quantity and/or quality of initial material—plasma DNA is lacking stability and FFPE blocks present uneven degradation of RNA. In terms of routine diagnostics, these methods are still too inaccurate and hard to standardize to embrace them at present.
However, data on clinical WGS utility for HER2 status assessment are scarce. There have only been two small clinical validation studies with direct comparison to the orthogonal methods. The first, released by Hartwig Medical Foundation, was a part of a WGS pan-cancer validation study. The ERBB2 status was evaluated on only 16 samples with the overall concordance of 93%. The HMF group compared ploidy and chromosome 17 CN with absolute CN of ERBB2 but did not draw any conclusions because of the small sample size [11]. The second, performed by King’s College Hospital in London, was performed on 145 BC samples with only 27 positives for amplified HER2. With the four discrepant samples, the sensitivity in the UK cohort was 88% and specificity was 98% [28]. The method itself has a big advantage over panel NGS as it may be universally used for different biomarkers in different cancers by using adequate bioinformatics pipelines. Moreover, library preparation is straightforward and reproducible, as is variant calling. The vast amounts of information extracted from the WGS analysis may serve as both a foundation for basic cancer science and a universal pan-cancer diagnostic tool. The main drawback of the method is the necessity to use fresh tissue, preserving big quantities (> 1 µg) of high-molecular-weight DNA. Second, the accurate variant calling in WGS requires also using a germline reference, a non-cancerous patient material, e.g., DNA isolated from the peripheral blood sample. Last, the analysis of big data generated from WGS experiments is challenging in terms of computational power and storage.
We attempted to systematically determine the criteria for WGS of ERBB2 CN in matched-normal tumor samples. Our strategy was to gather publicly available BC datasets with reliable clinical metadata and analyze them uniformly with a minimal 20× depth of coverage. We have chosen such a sensitivity threshold based on AscatNGS algorithm doccumentation, recommending at least t for accurate CN alterations calling (https://github.com/cancerit/ascatNgs).
Our machine learning approach, based on the decision tree classifier, agnostically chose the optimal approach of HER2 status assessment to be ploidy-corrected ERBB2 CN over ERBB2 CN/CEP17 CN ratio and absolute ERBB2 CN. To measure the test’s reliability, we used Cohen’s kappa coefficient. The high value of 85% rules out the possibility of the data agreement occurring by chance. To further test the superiority of the ploidy-corrected approach, we compared the results for the three features in a separate external dataset from the Wellcome Sanger Institute, which confirmed our observations: the accuracy and precision were 0.952813 and 0.765432, vs 0.724138 and 0.306977, vs 0.945554 and 0.724138 for absolute ERBB2 CN, ERBB2 CN/CEP17 CN ratio, and ploidy-corrected ERBB2 CN, respectively (for further details, see the project’s GitHub repository).
This is, to the best of our knowledge, the first and the largest study of its type utilizing machine learning approaches to evaluate diagnostic criteria of a genomic test. In the context of traditional HER2 status determination, first proof-of-concept ML-based solutions for robust FISH and IHC assessment are already tested in a clinical set-up [33, 34].
The ML approach is an emerging field of medicine, improving the efficiency of pathomorphological assessment [35], radiology [36], and clinical chemistry [37]. In the field of BC diagnostics, genomics and transcriptomics ML is being applied to distinguish between intrinsic BC subtypes with different prognoses [38], identify new potential biomarkers, or repurpose the existing biomarkers. These strategies perform well in the scientific environment but may only be used in the clinical setting after well-planned validation, showing concordance and stability of the tests. The potential limitation of using ML-based classifiers in medicine lies in the complexity and explainability of the results in the clinical set-up. This complexity led to the formulation of strict European Union and US Food and Drug Administration regulations for ML-based medical devices, which requires adherence to a set of tightly supervised rules. Nonetheless, ML classifiers are now widely used in medical genomics in biomarker discovery and refinement, providing unbiased solutions that can be then evaluated in the traditional validation process.
Our results prove that WGS may be a reliable method for HER2 diagnostics. We hypothesize that it could be implemented as a stand-alone test or in combination with IHC instead of FISH or other NGS-based methods in routine clinical practice. With a diagnostic sensitivity of 91.18% and a specificity of 98.69%, determined on unselected and heterogeneous groups of patients and validated on additional external data, we conclude that the technology is mature enough for prospective, multicenter, analytical, and clinical validation.
Our results do not deviate relevantly from those reported by other groups focusing on ERBB2 NGS testing; however, the diagnostic sensitivity is still not optimal. It can be improved by optimizing the threshold value, which might increase the sensitivity by compromising the specificity. In this study, we aimed for a reduction in the false-positive error rate; however, balancing between type I and type II errors should be left to the decisions of oncologists and regulatory bodies after careful assessment of clinical consequences of both strategies.
The limitation of the study in the context of sensitivity is also the large CI, as there were few HER2+ samples in the test set. However, the external validation results strongly support the high sensitivity of WGS.
Several other factors might contribute to a slightly lower analytical sensitivity of our solution in comparison with panel NGS sensitivity reported by other groups. We suspect that heterogenous evaluation of IHC/FISH results, made on the basis of different issues of ASCO/CAP guidelines, may have contributed to the discrepancy in the results. Additionally, there was some heterogeneity in the WGS raw data, acquired on different equipment by different genomic consortia. There were differences in the tumor sample collection, DNA extraction, and library preparation methods. All these pre-analytical and analytical factors must have contributed to the greater variation in HER2 results than in the single-facility method with uniform IHC/FISH evaluation methodology and a single laboratory protocol for sample management. Even so, the WGS method exhibits robustness and effectiveness, which is a great advantage, allowing for a low-cost external, even worldwide quality-control assessment program to be held out in the near future.
Other factors lowering the sensitivity are changes in HER2 status, which could have occurred in metastatic tumors from the HMF dataset. In these instances, we could not directly evaluate the correctness of IHC/FISH data because they came from the primary biopsy, not the biopsy corresponding with the sample used for WGS. The shift between IHC-positive and IHC-negative status is reported in up to 11.5% of HER2-negative cancers (conversion to HER2 positive) and in 37% of those initially positive (conversion to HER2 negative in the presence of selective pressure of trastuzumab) [4, 11].
The metastatic nature of HMF samples may have also contributed to lower than population numbers of HER2-positive cases in our cohort, as they tend to have better outcomes and might have been recruited less often to the study. The algorithm of patients’ qualification to first-line targeted therapy in all genomic consortia mentioned above could also have been somehow related to this under-representation.
Some of the discrepancies may have come from tumor subclonality, which is a common serious diagnostic issue. The signal from a small proportion of HER2 amplified cells may be below the resolution of WGS at 30–60× depth of coverage [13].
The spatial intra-tumor heterogeneity may have also contributed to false-negative/positive results when there were differences in sampling locations between tissue collected for FFPE blocks and WGS (e.g., different distant metastases are sampled). In addition, overexpression of HER2 is not always based on ERBB2 amplification, as an estimated 3–10% of non-amplified tumors exhibit high overexpression [39]. Even though there is currently no known genomic background of this phenomenon, WGS could potentially detect alterations in HER2 regulatory pathways, leading to overexpression, which could further improve WGS diagnostic power.
We may not forget that our results are compared to the BC diagnostic gold standard (IHC and FISH) and some issues of these tests may influence our results. There will always be a ‘diagnostic gray zone’ in which the prediction of response is not optimal, even though ASCO/CAP struggles to refine and evaluate their guidelines systematically [3]. Our classification tends to gradually worsen when ERBB2 CN is decreasing below 6 copies especially in tumors with higher ploidies. We cannot provide the data on the patient outcomes in these cases, but some authors made observations that polysomic tumors are not a homogenous group and they display different responses to trastuzumab and different clinicopathological features [40, 41]. As classical methods do not distinguish between polysomy and polyploidy, it might be reasonable to evaluate whether these two entities differ in terms of their properties and anti-HER2 drug response.
To tackle this issue, a prospective WGS study with an up-to-date pathomorphological HER2 evaluation according to the newest ASCO/CAP guidelines should be performed. It might not only evaluate the single biomarker concordance but would analyze the treatment response and try to seek new meaningful correlations with a vast number of genomic alterations, e.g., ploidy or complexity of chromosome 17 rearrangements.
The current knowledge on ERBB2 genetics is still based on methods established in the 20th century. We believe new potent tools such as ML and WGS present a vast array of solutions and opportunities to help improve the diagnostics and treatment of HER2-positive BC, and allow us to move into the 21st century.
5 Conclusions
We provide evidence that the ERBB2 status can be reliably determined by WGS methodology, which may be included in a comprehensive test for BC diagnostics. The 20% of tumor purity and 20× depth of coverage are sufficient to ensure good quality of genomic data in most instances. Given good concordance of a WGS with routinely used methods, we suggest that assessment by a WGS method may be an alternative to other NGS-based methods as well as FISH-based diagnostic tools. Hence, it should be subjected to evaluation by ASCO/CAP in the future updates of the HER2 testing recommendations. In our work, we have also proven that short-read WGS technology bears great potential for establishing a harmonized global quality assessment program for ERBB2 detection, as the outputs of heterogeneous data gathered from four different genomic consortia show a high degree of concordance between methodologies and pipelines.
Notes
In the context of this study, we use the term ‘ploidy’ to describe the mean number of whole chromosome sets.
References
King CR, Kraus MH, Aaronson SA. Amplification of a novel v-erbB-related gene in a human mammary carcinoma. Science. 1985;229:974–6. https://doi.org/10.1126/science.2992089.
Kunte S, Abraham J, Montero AJ. Novel HER2-targeted therapies for HER2-positive metastatic breast cancer. Cancer. 2020;126:4278–88. https://doi.org/10.1002/cncr.33102.
Wolff AC, Hammond MEH, Allison KH, et al. Human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline focused update. J Clin Oncol. 2018;36:2105–22. https://doi.org/10.1200/JCO.2018.77.8738.
Ross DS, Zehir A, Cheng DT, et al. Next-generation assessment of human epithelial growth factor receptor 2 (ERBB2) amplification status. J Mol Diagn. 2017;19:244–54. https://doi.org/10.1016/j.jmoldx.2016.09.010.
Cornelisse CJ, van de Velde CJ, Caspers RJ, et al. DNA ploidy and survival in breast cancer patients. Cytometry. 1987;2:225–34. https://doi.org/10.1002/cyto.990080217 (PMID: 3582068).
Burkard ME, Choudhary A, Robert FL, et al. Abstract 2526: Polyploidy: A new breast cancer subtype and a lead compound that targets it with high selectivity. Cancer Res. 2014;74(19 Suppl.):2526. https://doi.org/10.1158/1538-7445.AM2014-2526.
Newcomb R, Dean E, McKinney BJ, et al. Context-dependent effects of whole-genome duplication during mammary tumor recurrence. Sci Rep. 2021;11:14932. https://doi.org/10.1038/s41598-021-94332-z.
Hansen TVO, Vikesaa J, Buhl SS, et al. High-density SNP arrays improve detection of HER2 amplification and polyploidy in breast tumors. BMC Cancer. 2015;15:35. https://doi.org/10.1186/s12885-015-1035-1.
Niu D, Li L, Yu Y, et al. Evaluation of next generation sequencing for detecting HER2 copy number in breast and gastric cancers. Pathol Oncol Res. 2020;26:2577–85. https://doi.org/10.1007/s12253-020-00844-w.
Halilovic A, Verweij DI, Simons A, et al. HER2, chromosome 17 polysomy and DNA ploidy status in breast cancer; a translational study. Sci Rep. 2019;9:11679. https://doi.org/10.1038/s41598-019-48212-2.
Roepman P, de Bruijn E, van Lieshout S, et al. Clinical validation of whole genome sequencing for cancer diagnostics. J Mol Diagn. 2021;23(7):816–33. https://doi.org/10.1016/j.jmoldx.2021.04.011.
Arora K, Shah M, Johnson M, et al. Deep whole-genome sequencing of 3 cancer cell lines on 2 sequencing platforms. Sci Rep. 2019;9:19123. https://doi.org/10.1038/s41598-019-55636-3.
Nik-Zainal S, Davies H, Staaf J, et al. Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature. 2016;534:47–54. https://doi.org/10.1038/nature17676.
Pfarr N, Penzel R, Endris V, et al. Targeted next-generation sequencing enables reliable detection of HER2 (ERBB2) status in breast cancer and provides ancillary information of clinical relevance. Genes Chromosomes Cancer. 2017;56:255–65. https://doi.org/10.1002/gcc.22431.
Hoda RS, Bowman AS, Zehir A, et al. Next-generation assessment of human epidermal growth factor receptor 2 gene (ERBB2) amplification status in invasive breast carcinoma: a focus on Group 4 by use of the 2018 American Society of Clinical Oncology/College of American Pathologists HER2 testing guideline. Histopathology. 2021;78:498–507. https://doi.org/10.1111/his.14241.
Joly Y, Dove ES, Knoppers BM, Bobrow M, Chalmers D. Data sharing in the post-genomic world: the experience of the International Cancer Genome Consortium (ICGC) Data Access Compliance Office (DACO). PLoS Comput Biol. 2012;8(7): e1002549. https://doi.org/10.1371/journal.pcbi.1002549.
Priestley P, Baber J, Lolkema MP, et al. Pan-cancer whole-genome analyses of metastatic solid tumours. Nature. 2019;575:210–6. https://doi.org/10.1038/s41586-019-1689-y.
Campbell PJ, Getz G, Korbel JO, et al. Pan-cancer analysis of whole genomes. Nature. 2020;578:82–93. https://doi.org/10.1038/s41586-020-1969-6.
Goodstadt L. Ruffus: a lightweight Python library for computational pipelines. Bioinformatics. 2010;26:2778–9. https://doi.org/10.1093/bioinformatics/btq524.
Picard toolkit. Broad Institute, GitHub repository. 2019. http://broadinstitute.github.io/picard. Accessed 11 Dec 2021.
Li H. Dana Farber Institute. SeqTk tool, GitHub repository. 2021. https://github.com/lh3/seqtk. Accessed 11 Dec 2021.
Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011;17:10. https://doi.org/10.14806/ej.17.1.200.
Sanger Institute. CancerIt tools, GitHub repository. 2021. https://github.com/cancerit/dockstore-cgpmap. Accessed 11 Dec 2021.
Cibulskis K, Lawrence MS, Carter SL, et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol. 2013;31:213–9. https://doi.org/10.1038/nbt.2514.
Alioto TS, Buchhalter I, Derdak S, et al. A comprehensive assessment of somatic mutation detection in cancer using whole-genome sequencing. Nat Commun. 2015;6:10001. https://doi.org/10.1038/ncomms10001.
Raine KM, Loo P, Wedge DC, et al. ascatNgs: identifying somatically acquired copy-number alterations from whole-genome sequencing data. Curr Protoc Bioinform. 2016;56:15–9. https://doi.org/10.1002/cpbi.17.
McLaren W, Gil L, Hunt SE, et al. The Ensembl variant effect predictor. Genome Biol. 2016;17:122. https://doi.org/10.1186/s13059-016-0974-4.
Echejoh G, Liu Y, Chung-Faye G, et al. Validity of whole genomes sequencing results in neoplasms in precision medicine. J Clin Pathol. 2021;74(11):718–23. https://doi.org/10.1136/jclinpath-2020-206998.
Vives-Usano M, García Pelaez B, Román Lladó R, et al. Analysis of copy number variations in solid tumors using a next generation sequencing custom panel. J Mol Pathol. 2021;2(2):123–34. https://doi.org/10.3390/jmp2020013.
Nattestad M, Goodwin S, Ng K, et al. Complex rearrangements and oncogene amplifications revealed by long-read DNA and RNA sequencing of a breast cancer cell line. Genome Res. 2018;8:1126–35. https://doi.org/10.1101/gr.231100.117.
Lee J, Franovic A, Rich TA, et al. Cell-free DNA (cfDNA) landscape in ERBB2 (HER2)-amplified Asian cancer patient population. Ann Oncol. 2018;29:ix114–5.
Pennock ND, Jindal S, Horton W, et al. RNA-seq from archival FFPE breast cancer samples: molecular pathway fidelity and novel discovery. BMC Med Genom. 2019;12:195. https://doi.org/10.1186/s12920-019-0643-z.
Rawat RR, Ortega I, Roy P, et al. Deep learned tissue “fingerprints” classify breast cancers by ER/PR/Her2 status from H&E images. Sci Rep. 2020;10:7275. https://doi.org/10.1038/s41598-020-64156-4.
Zakrzewski F, de Back W, Weigert M, et al. Automated detection of the HER2 gene amplification status in fluorescence in situ hybridization images for the diagnostics of cancer tissues. Sci Rep. 2019;9:8231. https://doi.org/10.1038/s41598-019-44643-z.
Bera K, Schalper KA, Rimm DL, et al. Artificial intelligence in digital pathology: new tools for diagnosis and precision oncology. Nat Rev Clin Oncol. 2019;16:703–15. https://doi.org/10.1038/s41571-019-0252-y.
Montagnon E, Cerny M, Cadrin-Chênevert A, et al. Deep learning workflow in radiology: a primer. Insights Imaging. 2020;11:22. https://doi.org/10.1186/s13244-019-0832-5.
Ford BA, McElvania E. Machine learning takes laboratory automation to the next level. J Clin Microbiol. 2020;58:e00012-20. https://doi.org/10.1128/JCM.00012-20.
Dawson S-J, Rueda OM, Aparicio S, et al. A new genome-driven integrated classification of breast cancer and its implications. EMBO J. 2013;32:617–28. https://doi.org/10.1186/s13059-014-0431-1.
Timothy WJ, Allen MG, Hadi Y, et al. Comparison of fluorescence in situ hybridization and immunohistochemistry for the evaluation of HER-2/neu in breast cancer. J Clin Oncol. 1999;17(7):1974.
Sun H, Chen H, Crespo J, et al. Clinicopathological features of breast cancer with polysomy 17 and its response to neoadjuvant chemotherapy. Eur J Breast Health. 2021;17(2):128–36. https://doi.org/10.4274/ejbh.galenos.2021.2021-2-9.
Vanden Bempt I, Van Loo P, Drijkoningen M, et al. Polysomy 17 in breast cancer: clinicopathologic significance and impact on HER-2 testing. J Clin Oncol. 2008;26(30):4869–74. https://doi.org/10.1200/JCO.2007.13.4296.
Acknowledgements
We thank the International Genomic Consortia, their researchers, and public funding institutions who have generously donated the genomic data and clinical metadata for this project. This publication and the underlying study have been made possible by the data that the Hartwig Medical Foundation and the Center of Personalized Cancer Treatment have made available. The results published here are also in part based upon data generated by the The Cancer Genome Atlas Research Network: https://www.cancer.gov/tcga. This work was also supported by data obtained from the International Cancer Genome Consortium Breast Cancer Working Group. We also thank the Wellcome Sanger Institute for inspiring research on 560 breast cancer genomes, which was the source for our external validation data. Importantly, we thank the patients and their families for their participation in the genomic projects and the opportunity to use their clinical and genomic data for fundamental cancer research. We would also acknowledge the work of all clinical staff gathering samples and medical records. Last, we thank all members of Adam Mickiewicz University, Poznan Supercomputing and Networking Center, and MNM Diagnostics staff for help with genomic data processing and general support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
All the expenses were covered from the Poznan Supercomputing and Networking Center’s ‘computational grant’ no. 437, titled ‘Molekularny mechanizm interakcji pomiędzy szlakami naprawy DNA w oporności na chemioterapię’. Open access reimbursement will be granted based on Springer’s Read and Publish agreement.
Conflict of interest
Alicja Woźna, Pawel Sztromwasser, Maciej Piernik, and Paweł Zawadzki are share owners in the company MNM Bioscience Inc. Marzena D. Wojtaszewska, Rafal Stępien, Maciej Dąbrowski, and Michał Gniot were employees of MNM Bioscience Inc. Marzena D. Wojtaszewska and Michal Gniot received conference reimbursements/speaker honoraria from Novartis in the past. Other authors declare no conflicts of interest.
Ethics approval
The collection of all primary data used in this study was performed by international Genomic Consortia, which provided de-identified samples. Data collection and transmission was approved by local medical ethics committees during design of the original studies. No additional approvals are needed, and our study was reviewed and deemed exempt by the Adam Mickiewicz University Review Board. The protocols are in accordance with the ethical standards of our institution and with the 1964 Helsinki Declaration and its later amendments.
Consent to participate
The written consent of each participant is in possession of Genomic Consortia. The primary data were collected in accordance with the standards set by the Declaration of Helsinki and the highest data security standards of ISO 27001. All necessary patient/participant consent has been obtained and the appropriate institutional forms have been archived during the design of the original studies.
Consent for publication
The investigators have adhered to controlled-access agreement conditions and obtained consent to publish the study results.
Availability of data and material
The data that support the findings of this study are openly available in the following repositories: Hartwig Medical Foundation at https://www.hartwigmedicalfoundation.nl/, which was acquired under data request number DR-169 and International Cancer Genome Consortium at https://dcc.icgc.org/pcawg/, which was acquired under data request number DACO-6030. The Cancer Genome Atlas data were acquired via dbGaP platform (project phs000178.v11.p8) at: https://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs000178.v11.p8, under data request number #86794-3. The processed genomic data from 551 samples used for external validation were extracted from the publication by Nik-Zainal et al., 2016 (https://doi.org/10.1038/nature17676) [file 1 and 4 of the ESM]. Secondary data that support the findings of this study generated by the authors are available in the ESM.
Code availability
The code for HER2/ERBB2 amplification classification is available at: https://github.com/MNMdiagnostics/her2_analysis.
Author contributions
MW was responsible for the study design, manuscript preparation, and data analysis. AW is the main genomic data administrator and was responsible for the primary data acquisition. MP designed the computational framework for the machine learning approach. PS developed the pipeline and was responsible for bioinformatics support. MD refined the secondary data and helped to write the manuscript. MG conducted statistical analyses and verified the clinical actionability of genomic variants. SS curated medical data and made manuscript corrections. MS coordinated the statistical analysis. PK evaluated the concordance of the molecular and clinical data. RM provided expertise on molecular HER2 status assessment and evaluated the clinical data. PZ was the supervisor of the project, reviewed the methodology, and corrected and revised the manuscript.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Wojtaszewska, M., Stępień, R., Woźna, A. et al. Validation of HER2 Status in Whole Genome Sequencing Data of Breast Cancers with the Ploidy-Corrected Copy Number Approach. Mol Diagn Ther 26, 105–116 (2022). https://doi.org/10.1007/s40291-021-00571-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40291-021-00571-1