Abstract
Background and Objective
Voriconazole is an important broad-spectrum anti-fungal drug with nonlinear pharmacokinetics. The aim of this single centre fixed-sequence open-label drug–drug interaction trial in healthy participants (N = 17) was to determine whether microdosed probe drugs for CYP3A and CYP2C19 reliably predict voriconazole clearance (CLVRZ).
Methods
At baseline, a single oral microdose of the paradigm substrates midazolam (CYP3A) and omeprazole (CYP2C19) were given to estimate their clearances (CL). Thereafter, a single oral dose of voriconazole was administered (50, 100, 200 or 400 mg), followed by the microdosed probe drugs.
Results
The clearances of midazolam (CLMDZ 790–2790 mL/min at baseline; 248–1316 mL/min during voriconazole) and omeprazole (CLOMZ 66.4–2710 mL/min at baseline; 30.1–1420 mL/min during voriconazole) were highly variable. CLMDZ [geometric mean ratio (GMR) 0.586 at 50 mg voriconazole decreasing to GMR 0.196 at 400 mg voriconazole] and CLOMZ (GMR 0.590 at 50 mg decreasing to GMR 0.166 at 400 mg) were reduced with higher voriconazole doses. CLMDZ was linearly correlated with CLVRZ (slope 1.458; adjusted R2 0.528) as was CLOMZ (slope 0.807; adjusted R2 0.898). Multiple linear regression resulted in an adjusted R2 of 0.997 for the relationship CLVRZ ~ log CLOMZ + log CLMDZ using data during voriconazole treatment and an adjusted R2 of 0.997 for the relationship CLVRZ ~ log CLOMZ + log CLMDZ + voriconazole dose, using baseline data for CLMDZ and CLOMZ.
Conclusion
Microdosed midazolam and omeprazole accurately described and predicted total CLVRZ
Trial Registration
EudraCT No: 2020-001017-20, registered on March 5th, 2020. DRKS: DRKS00022547, registered on August 6th, 2020.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
This in vivo study aimed to investigate the contribution of CYP2C19 and CYP3A activity to oral voriconazole clearance using microdosed probe drugs for CYP2C19 (omeprazole) and CYP3A (midazolam) and different voriconazole doses. |
Multiple linear regression demonstrated that omeprazole and midazolam clearance accurately describe voriconazole clearance when given together, and that their clearances together with voriconazole dose can predict voriconazole clearance precisely. |
After oral administration, voriconazole clearance exclusively depends on the activity of CYP2C19 and to a lesser extent on CYP3A, indicating that other enzymes have no or only a minor contribution to its metabolism in vivo in healthy volunteers. |
1 Introduction
Voriconazole is a second generation triazole antifungal agent that is on the World Health Organization (WHO) essential medicines list [1, 2]. Its broad spectrum of activity, high tissue penetration and particular efficacy against Aspergillus make voriconazole a cornerstone of prevention and treatment of invasive fungal infections in vulnerable patient populations, such as patients with neutropenia or immunosuppression after stem cell or solid organ transplantation [3,4,5,6].
However, due to the complex pharmacokinetics of voriconazole, it is a challenge in clinical practice to reliably and quickly achieve therapeutic antifungal exposures in individual cases [7]. Voriconazole has non-linear and highly variable pharmacokinetics; it is metabolised by cytochrome P450 (CYP) isozymes and is therefore susceptible to numerous drug–drug interactions, some of which are also genotype dependent [8]. CYP2C19 and CYP3A appear to play a major role, but the extent of their involvement in vivo and possible contributions of CYP2C9 and the family of flavin-dependent monooxygenases (FMO3) are quantitatively uncertain [7]. At the same time, voriconazole and its major N-oxide metabolite are inhibitors of CYP3A and CYP2C19 and likely cause concomitant enzyme induction, because increasing doses of voriconazole are required over time to maintain stable plasma concentrations [9,10,11]. Furthermore, CYP2C19 polymorphisms substantially modulate the metabolism of voriconazole and its interaction potential, and co-morbidities such as systemic inflammation further modulate voriconazole clearance (CLVRZ) [8, 12,13,14,15].
In this trial, we aimed to mechanistically investigate the relative contribution of CYP2C19 and CYP3A to the clearance of voriconazole using an in vivo approach with microdosed probe drugs to quantify CYP activities [16, 17]. Because the pharmacokinetics of microdosed midazolam and omeprazole are linear, they reflect isozyme activities at therapeutic doses well [16,17,18], without causing interactions and with minimal risk of adverse events (AE) [19]. In this trial, different doses of voriconazole were used and participants were enrolled regardless of their CYP2C19 genotype to cover a spectrum of CLVRZ values as broad as possible and to investigate its association with CYP3A and CYP2C19 activity.
2 Materials and Methods
The trial protocol was approved by the competent authority (BfArM, Bonn) in Germany (EudraCT No: 2020-001017-20) and the responsible ethics committee of the Medical Faculty of Heidelberg University on 13 July 2020. The trial was conducted in the DIN EN ISO9001-certified Clinical Research Unit (KliPS) of the Department of Clinical Pharmacology and Pharmacoepidemiology at Heidelberg University Hospital according to the standards of good clinical practice (as defined in the ICH E6 Guideline for Good Clinical Practice) and in compliance with the Declaration of Helsinki and all specific legal requirements in Germany. Participants were only enrolled into the trial after they had given their written informed consent.
2.1 Study Population
All participants were physically and mentally healthy as confirmed by a thorough medical history, physical exam, blood pressure measurement, a 12-lead electrocardiogram and standard laboratory analyses, including a urine drug screen and a pregnancy test (in women of childbearing potential). Participants were required to consent to use a highly effective method of contraception during the trial.
Participants were excluded if any of the following criteria were met: clinically relevant abnormalities in the medical history, physical examination or laboratory evaluation as assessed by the investigator, any medical disorder that may require significant treatment, or make the participant unlikely to fully complete the trial, or any condition that presents undue risk from the investigational medicinal products or trial interventions, clinically relevant ongoing or past history of physical or psychiatric illness as judged by the investigator, pregnancy or breast feeding, any acute or chronic illness or clinically relevant finding known or expected to modify the absorption, distribution, metabolism, or excretion of voriconazole, omeprazole, or midazolam, including the use of any co-medications or consumption of known inducers (including St. John’s Wort) in the past 2 weeks or inhibitors of the CYP of interest such as grapefruit, and finally any known allergies to the specific trial medication, triazole derivatives in general, or additives.
2.2 Genotyping
Prior to assigning the voriconazole dose group, CYP2C19 genotyping was performed for CYP2C19*2 (rs4244285), CYP2C19*3 (rs4986893), and CYP2C19*17 (rs12248560) as described previously [20]. The presence of two wild-type alleles was assumed if none of the tested polymorphisms was present.
2.3 Study Design
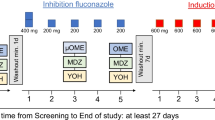
This was a single centre fixed sequence open-label four-arm phase I trial in healthy volunteers. The trial included a screening visit, two treatment visits 3–7 d apart, and an end-of-trial visit.
At baseline, participants were all administered oral microdoses of midazolam (10 µg) and omeprazole (100 µg). For the preparation of midazolam, 0.01 mL Dormicum® V 5 mg/5 mL (Cheplapharm Arzneimittel GmbH, Greifswald, Germany) was added to 100 mL of tap water [21]. Because uncoated omeprazole (OMEP® 40 mg HEXAL powder for solution for infusion, Hexal AG, Holzkirchen, Germany) is subject to degradation in gastric acid, the powder was dissolved in 100 mL of normal saline and 250 µL of the solution were further diluted in 100 mL of sodium bicarbonate buffer (4.2 %, w/v). In addition, 10 min prior to oral administration of the omeprazole microdose, participants drank 100 mL of sodium bicarbonate buffer [17]. Blood was collected in 4.9 mL lithium–heparin tubes before and 0.25, 0.5, 0.75, 1, 1.5, 2, 2.5, 3, 4, 5, 6, 8 and 10 h after drug administration. Midazolam and omeprazole were given in a fasted state. Fluid intake was prohibited during the first 2 h after administration of the trial medication, and food for the first 4 h. Thereafter, participants were served a breakfast.
On the second trial day, participants were given a single dose of either 50, 100, 200, or 400 mg of voriconazole (Voriconazol Hexal® 50 mg film tablets, Hexal AG, Holzkirchen, Germany). Participants were sequentially assigned to voriconazole doses with four participants per dose, regardless of the CYP2C19 genotype, except for CYP2C19 poor metabolisers who were always assigned 400 mg voriconazole. One hour after voriconazole, participants were given midazolam 10 µg and omeprazole 100 µg as on the baseline day. Blood samples were collected in 7.5 mL lithium-heparin tubes before and 0.25, 0.5, 0.75, 1, 1.25, 1.5, 1.75, 2, 2.5, 3, 3.5, 4, 5, 6, 7, 9, 11 and 24 h after voriconazole administration. As on day 1, participants arrived fasted, were allowed to drink water 2 h after the microdoses, and were given a breakfast after 4 h. All samples were centrifuged within 30 min at 2500g and 4 °C for 10 min, distributed to 2 aliquots per substance and sample, and stored at − 20 °C until analysis.
2.4 Quantification of Midazolam, Omeprazole, and Voriconazole
Midazolam and omeprazole plasma concentrations were quantified by ultra-high performance liquid chromatography coupled to tandem mass spectrometry with a lower limit of quantification (LLOQ) of 1 pg/mL for midazolam [22] and 10 pg/mL for omeprazole [20]. Voriconazole concentrations were quantified using a validated high-performance liquid chromatography coupled to tandem mass spectrometry method, with a LLOQ of 1 ng/mL [23]. All methods were validated according to the current US Food and Drug Administration (FDA) and European Medical Agency (EMA) standards [24, 25].
2.5 Data Analysis
The primary endpoint was the correlation of CLMDZ and CLOMZ with CLVRZ after a single oral voriconazole dose of 50, 100, 200 or 400 mg. No formal sample size calculation was performed. Secondary endpoints were the pharmacokinetics of midazolam, omeprazole and voriconazole, and the frequency, severity, seriousness, relatedness, expectedness and outcome of AE.
Standard non-compartmental pharmacokinetic parameters for all substances were determined using Phoenix WinNonlinTM version 8.3 (Certara Inc., Princeton, NJ, USA). This included maximum plasma concentration (Cmax), time to reach Cmax (tmax), terminal elimination half-life (t1/2), area under the plasma concentration–time curve extrapolated to infinity (AUC0–∞) and apparent oral clearance (CL/F). Descriptive statistics of pharmacokinetic parameters were calculated. Paired t tests of the geometric mean ratios on log-transformed data were performed to assess pharmacokinetic differences at baseline and under different doses of voriconazole.
Linear regression analysis was used to individually assess the relationships between CLVRZ and clearance of the probe drugs at baseline and during voriconazole treatment. To analyse skewed non-normally distributed data, the analysis was carried out using a log–log linear equation.
Multiple linear regression analysis was used to assess whether a better fit could be achieved when the variables were combined. CLVRZ was defined as the dependent variable and CLMDZ, CLOMZ and voriconazole dose as independent variables as described in Eq. (1), where β1, β2 and β3 are the regression coefficient estimates resulting from the analysis.
The analysis was performed first with CLOMZ alone, then after systematically including the other variables. The regression was also performed after log transformation of CLVRZ, CLOMZ and CLMDZ. The most suitable models were selected by comparing the adjusted R2 and Akaike’s information criterion corrected for sample size (AICc). To assess predictive performance, predictive R2 (i.e., the correlation coefficient between predicted and observed values) was calculated in-sample and out-of-sample via enhanced non-parametric bootstrap from 1000 bootstrap samples. All statistical analyses were performed using Prism Version 9.0.1 (GraphPad Software Inc., La Jolla, CA, USA), R Version 4.0.5 (R Foundation for Statistical Computing, Vienna, Austria), and a p value < 0.05 was considered statistically significant.
3 Results
Seventeen participants [8 females; all Caucasians; 1 ultra-rapid (*17/*17), 7 rapid (*1/*17), 5 normal (*1/*1), 3 intermediate (*1/*2, *2/*17), and 1 poor metaboliser (*2/*2)] were enrolled and included in all analyses; each dose group was assigned four participants irrespective of the genotype, one poor metaboliser was assigned to the 400 mg group, which therefore had five participants. Their mean age was 31 years (range: 22–50) and body mass index was 24.6 kg/m2 (range 19.3–29.2). All participants completed the trial.
3.1 Midazolam
At baseline, CLMDZ was highly variable ranging from 790 to 2790 mL/min. Coadministration of voriconazole dose-dependently increased midazolam plasma concentrations over time and prolonged its terminal half-life in all participants (Fig. 1, Table 1). Midazolam AUC0–∞ showed an increase by a factor of 1.58, 2.42, 3.0 and 5.24 compared with baseline. Terminal elimination half-life was dose dependently prolonged by voriconazole with a 2.1-fold increase at 400 mg. During voriconazole, CYP3A was inhibited and CLMDZ was reduced in all participants ranging from 248 to 1316 mL/min. Concurrently, CLMDZ were reduced to 58.6% of the baseline value (50 mg), 42.6% (100 mg), 34.1% (200 mg) and 19.6% (400 mg) (Fig. 2; Table 2).
3.2 Omeprazole
The pharmacokinetics of microdosed omeprazole was highly variable with CLOMZ ranging from 66.4 to 2710 mL/min at baseline. Dependent on the voriconazole dose CLOMZ decreased in every participant (Fig. 3; Table 1). With the 50 mg voriconazole dose omeprazole AUC0–∞ was increased by a factor of 1.3 compared with baseline and by a factor of 12.1 with the 400 mg dose (Table 1). Correspondingly, CLOMZ during 400 mg voriconazole was reduced to only 8.3 % compared with baseline (Fig. 2; Table 2). Interestingly, the CYP2C19 poor metaboliser had a an AUC0–∞ of 25.1 h ng/mL and a CLOMZ of 66.4 mL/min at baseline, which was altered by 400 mg voriconazole to 55.4 h ng/mL and 30.1 mL/min. The ultra-rapid metaboliser showed an AUC0–∞ of 0.62 h ng/mL and a CLOMZ of 2712 mL/min at baseline, which was altered by 100 mg voriconazole to 2.22 h ng/mL and 711 mL/min, respectively.
Semilogarithmic plot of mean (± standard deviation) plasma concentration–time curves of omeprazole 100 µg at baseline (grey) and after co-administration of a single oral dose of 50, 100, 200 or 400 mg voriconazole (VRZ). The concentration–time curves were capped at 5 h because thereafter most concentrations (especially in the control group and the low-dose voriconazole groups) were below the limit of quantification
3.3 Voriconazole
The plasma concentration–time curves of the four voriconazole dose groups and the corresponding pharmacokinetics are shown in Fig. 4, Table 1 and Supplementary Table S1. Voriconazole was rapidly absorbed (tmax 0.66 h). Cmax and AUC0–∞ showed a disproportionate increase after the three stepwise dose doublings, with Cmax increasing twofold, 6.7-fold and 1.6-fold for each doubling of the dose from 50 to 400 mg, while AUC0–∞ showed an increase by a factor of 2.2, 6.2 and 4.3. Compared with the next lower dose group, CLVRZ decreased by 9.4%, 67.8% and 54%, respectively, and CLVRZ at 400 mg of voriconazole was reduced to 13.3% of CLVRZ at 50 mg. Furthermore, t1/2 increased by 15.4%, 23.4%, and 27.4% with each dose step. CLVRZ values varied 44-fold across the different dose groups with the lowest clearance observed in the CYP2C19 PM with 200 mL/min and the highest in the 50 mg dose group (8860 mL/min). The ultra-rapid metaboliser had a clearance of 3602 mL/min after 100 mg voriconazole.
3.4 Bivariate Clearance Relationships
Linear regression analyses with the clearance data collected during voriconazole coadministration resulted in a slope of 1.458 (p < 0.001) with an adjusted R2 of 0.528 for the relationship of CLVRZ and CLMDZ, and a slope of 0.807 (p < 0.0001) with an adjusted R2 of 0.898 for CLVRZ and CLOMZ. Using the clearance data for midazolam and omeprazole at baseline revealed slopes of 0.612 (p = 0.488) (CLMDZ) and 0.846 (p < 0.001) (CLOMZ) with adjusted R2 values of − 0.032 (CLMDZ) or 0.617 (CLOMZ).
Multiple linear regression All evaluated equations provided good fits to the data as shown by very high adjusted R2 values exceeding 0.99 (Supplementary Table S2). Using the data collected during voriconazole administration, CLVRZ was best predicted (adjusted R2 0.997; AICc − 53.2) by including the predictors CLMDZ and CLOMZ during voriconazole administration only (log CLVRZ ~ β1 log CLOMZ + β2 log CLMDZ) (Fig. 5). Adding voriconazole dose did not improve the model, likely because dose is reflected in the clearance values. With the final equation, an in-sample predictive R2 of 0.861 and an out-of-sample predictive R2 of 0.849 was calculated.
Best-fit multiple regression models: plot of actual voriconazole clearance (CLVRZ) versus predicted CLVRZ during voriconazole (using CLMDZ and CLOMZ during VRZ) and at baseline (using CLMDZ and CLOMZ at baseline without VRZ) with different single doses of voriconazole (red 400 mg; grey 200 mg; blue 100 mg; green 50 mg). Open circles depict the CYP2C19 poor metaboliser (red) and ultrarapid metaboliser (blue). Both plots include the line of identity
Using the baseline clearance data of midazolam and omeprazole, CLVRZ was best predicted (adjusted R2 0.997; AICc −54.53) with all predictors included (log CLVRZ ~ β1 log CLOMZ + β2 log CLMDZ + β3 voriconazole dose) (Fig. 5) yielding a predictive R2 of 0.891 (in-sample) and 0.852 (out-of-sample).
3.5 Safety and Tolerability
All trial medications were well tolerated, and no serious AE occurred. A total of 22 AE in 11 participants occurred. Most AE were mild [common terminology criteria for AE (CTCAE) grade 1], while two cases of headache were treated with ibuprofen (CTCAE grade 2). Two cases of visual disturbances (photopsia, xanthopsia) were deemed probably related to the investigational medicinal products (voriconazole) and eight AE (headache, bloating, diarrhoea) possibly related to the study. The remaining AE were considered unrelated to the trial procedures and interventions or their relationship was not assessable.
4 Discussion
This trial demonstrated that combining microdose pharmacokinetics of midazolam and omeprazole as surrogates of current CYP3A and CYP2C19 activity can reliably predict CLVRZ in healthy volunteers. Recently, contributions of CYP2C19 and CYP3A to voriconazole N-oxide formation in vitro were estimated to be 63% and 30%, respectively [26], which is consistent with our in vivo observations that these enzymes are responsible for almost all metabolic clearance. Interestingly, a reduction of CLVRZ by ritonavir (which is only a weak CYP2C19 inhibitor [27]) by only approximately 150 mL/min was observed, corresponding to about one-third of the CLVRZ in CYP2C19 normal metabolisers [8], confirming the important but not dominant role of CYP3A in voriconazole metabolism. Major factors known to affect CLVRZ include the dose and metaboliser status for CYP2C19 [7, 14, 28, 29].
To evaluate a possible relationship between CLVRZ and the clearances of midazolam and omeprazole, a large range of voriconazole doses (and thus CYP activities) was tested by administering four different doses of voriconazole to a variety of CYP2C19 ultra-rapid, rapid, normal and intermediate metaboliser, and one poor metaboliser treated with 400 mg voriconazole to decrease CLVRZ to a (potential) minimum.
The clearances were highly variable for all substances and spanned a range of about 1.5–2 orders of magnitude. CYP3A activity, as indicated by CLMDZ, was increasingly suppressed with each increase in voriconazole exposure. This was very similar for CPY2C19 activity with the strongest inhibition with 400 mg voriconazole. With this voriconazole dose, a clearance of less than 20% of baseline was observed for both midazolam and omeprazole. Since all drugs were given orally, it is not possible to distinguish the contribution of the small intestine and the liver to the resulting overall inhibition. However, because Cmax of omeprazole and midazolam increased dose-dependently, it can be assumed that at the intestinal level, both CYP isozymes are inhibited with increasing voriconazole doses.
The clearances of the probe drugs midazolam and omeprazole closely reflect actual CYP isozyme activities and, when administered in microdoses, do not exert any perpetrator effects. Voriconazole, however, as an inhibitor of both CYP2C19 and CYP3A, affects the clearances of both probe drugs, which were therefore expected to be related to CLVRZ and to its exposure. It is therefore not surprising that CLVRZ correlated with CLMDZ or CLOMZ during voriconazole treatment but also with baseline clearances before inhibitor administration. While baseline CLOMZ was correlated with CLVRZ, there was no significant relationship between baseline CLMDZ and CLVRZ, confirming that the contribution of CYP2C19 to voriconazole metabolism is more substantial. This is also in line with the considerable impact of the CYP2C19 genotype on voriconazole clearance and thus exposure [15].
According to the relative contributions of the two isozymes to the total CLVRZ, CLOMZ achieved a better correlation than CLMDZ, but an almost perfect correlation with CLVRZ was only achieved when both clearances were considered together. Accounting for voriconazole doses did not improve the regression, probably because voriconazole exposure (and thus dose) is already accounted for by CLMDZ or CLOMZ via voriconazole’s perpetrator characteristics. This is consistent with the assumption that CYP3A and CYP2C19 almost exclusively determine CLVRZ in healthy volunteers. In vitro, the involvement of CYP2C9 and FMO3 in the metabolism of voriconazole has also been observed [30, 31], but our findings clearly suggest that the metabolic contribution of other enzymes is minor and likely not clinically relevant in adult patient populations. This might be different in a paediatric population [32]. Moreover, the activities of FMO3 cannot be induced by xenobiotics [33], and almost all inducers of CYP2C19 are also and often stronger inducers of CYP3A, suggesting that even in the presence of CYP inducers other enzymes are unlikely to participate in voriconazole metabolism. Finally, in addition to genetic variants of CYP2C19 as the strongest modulators of CLVRZ, an indirect modulation of voriconazole concentrations in inflammatory states has also been reported [30, 31] with exposure increases caused by inflammatory states that down-regulate CYP3A and possibly also CYP2C19 [34].
5 Limitations
This trial was conducted in healthy participants and it remains to be shown whether the suggested exclusive dependence of CLVRZ from CYP3A and CYP2C19 activities can be generalised to patient populations. This appears likely because the major known alterations of voriconazole pharmacokinetics beyond genetics in patients (inflammatory states, drug–drug interactions) could all be mediated through modulation of the activities of the same isozymes. Furthermore, the low prevalence of CYP2C19 poor metabolisers in the local population resulted in the recruitment of only one poor metaboliser, making the results of the lowest evaluated clearance range less certain. However, its results fit well into the regression of the whole group, well supporting the concept. Finally, this trial used a single oral dose of voriconazole, for the moment limiting interpretation to acute voriconazole effects. However, it appears likely that neither intravenous administration nor multiple dosing of voriconazole will change these findings, except in rare occasions in which autoinduction of the metabolism has been suggested [9].
6 Conclusions
In healthy volunteers, the CLVRZ exclusively depends on the activity of CYP2C19 and CYP3A, indicating that other enzymes metabolising voriconazole in vitro (CYP2C9, FMO3) do not contribute to its metabolism in vivo. Whether chronic treatment, enzyme inducing comedication or comorbidities can recruit additional enzymes remains to be studied, but appears unlikely.
References
Fachinformation. Voriconazol HEXAL® 50 mg Filmtabletten HA, Germany, October 2016.
World Health Organization. WHO Model List of Essential Medicines. 2019(21).
Nett JE, Andes DR. Antifungal agents: spectrum of activity, pharmacology, and clinical indications. Infect Dis Clin N Am. 2016;30(1):51–83.
Cadena J, Thompson GR 3rd, Patterson TF. Invasive aspergillosis: current strategies for diagnosis and management. Infect Dis Clin N Am. 2016;30(1):125–42.
Patterson TF, Thompson GR 3rd, Denning DW, Fishman JA, Hadley S, Herbrecht R, et al. Practice guidelines for the diagnosis and management of aspergillosis: 2016 update by the Infectious Diseases Society of America. Clin Infect Dis. 2016;63(4):e1–60.
Rosanova MT, Bes D, Serrano Aguilar P, Sberna N, Lede R. Efficacy and safety of voriconazole in immunocompromised patients: systematic review and meta-analysis. Infect Dis (Lond). 2018;50(7):489–94.
Schulz J, Kluwe F, Mikus G, Michelet R, Kloft C. Novel insights into the complex pharmacokinetics of voriconazole: a review of its metabolism. Drug Metab Rev. 2019;51(3):247–65.
Mikus G, Schowel V, Drzewinska M, Rengelshausen J, Ding R, Riedel KD, et al. Potent cytochrome P450 2C19 genotype-related interaction between voriconazole and the cytochrome P450 3A4 inhibitor ritonavir. Clin Pharmacol Ther. 2006;80(2):126–35.
Hsu AJ, Dabb A, Arav-Boger R. Autoinduction of voriconazole metabolism in a child with invasive pulmonary aspergillosis. Pharmacotherapy. 2015;35(4):e20–6.
Ferguson MJ, Randles ML, de Freitas DG. A suspected case of autoinduction of voriconazole metabolism in a patient with cerebral aspergillosis. Drug Healthc Patient Saf. 2017;9:89–91.
Farrokh S, Avdic E. Voriconazole autoinduction and saturable metabolism after cessation of rifampin in a patient with invasive central nervous system Aspergillus: importance of therapeutic drug monitoring. J Pharm Pract. 2019;32(5):589–94.
Vreugdenhil B, van der Velden W, Feuth T, Kox M, Pickkers P, van de Veerdonk FL, et al. Moderate correlation between systemic IL-6 responses and CRP with trough concentrations of voriconazole. Br J Clin Pharmacol. 2018;84(9):1980–8.
Aiuchi N, Nakagawa J, Sakuraba H, Takahata T, Kamata K, Saito N, et al. Impact of polymorphisms of pharmacokinetics-related genes and the inflammatory response on the metabolism of voriconazole. Pharmacol Res Perspect. 2022;10(2): e00935.
Hassan A, Burhenne J, Riedel KD, Weiss J, Mikus G, Haefeli WE, et al. Modulators of very low voriconazole concentrations in routine therapeutic drug monitoring. Ther Drug Monit. 2011;33(1):86–93.
Weiss J, Ten Hoevel MM, Burhenne J, Walter-Sack I, Hoffmann MM, Rengelshausen J, et al. CYP2C19 genotype is a major factor contributing to the highly variable pharmacokinetics of voriconazole. J Clin Pharmacol. 2009;49(2):196–204.
Mikus G. Probes and cocktails for drug-drug interaction evaluation: the future is microdosing? Clin Pharmacol Ther. 2019;105(6):1335–7.
Elbe A, Foerster KI, Blank A, Rose P, Burhenne J, Haefeli WE, et al. Evaluation of CYP2C19 activity using microdosed oral omeprazole in humans. Eur J Clin Pharmacol. 2022;78(6):975–87.
Hohmann N, Haefeli WE, Mikus G. CYP3A activity: towards dose adaptation to the individual. Expert Opin Drug Metab Toxicol. 2016;12(5):479–97.
Hohmann N, Blank A, Burhenne J, Suzuki Y, Mikus G, Haefeli WE. Simultaneous phenotyping of CYP2E1 and CYP3A using oral chlorzoxazone and midazolam microdoses. Br J Clin Pharmacol. 2019;85(10):2310–20.
Mahmoudi M, Foerster KI, Burhenne J, Weiss J, Mikus G, Haefeli WE. Application of microdosed intravenous omeprazole to determine hepatic CYP2C19 activity. J Clin Pharmacol. 2021;61(6):789–98.
Hohmann N, Kocheise F, Carls A, Burhenne J, Haefeli WE, Mikus G. Midazolam microdose to determine systemic and pre-systemic metabolic CYP3A activity in humans. Br J Clin Pharmacol. 2015;79(2):278–85.
Burhenne J, Halama B, Maurer M, Riedel KD, Hohmann N, Mikus G, et al. Quantification of femtomolar concentrations of the CYP3A substrate midazolam and its main metabolite 1′-hydroxymidazolam in human plasma using ultra performance liquid chromatography coupled to tandem mass spectrometry. Anal Bioanal Chem. 2012;402(7):2439–50.
Huppertz A, Ott C, Bruckner T, Foerster KI, Burhenne J, Weiss J, et al. Prolonged-release tacrolimus is less susceptible to interaction with the strong CYP3A inhibitor voriconazole in healthy volunteers. Clin Pharmacol Ther. 2019;106(6):1290–8.
FDA. Guidance for industry: bioanalytical method validation. https://www.fda.gov/files/drugs/published/Bioanalytical-Method-Validation-Guidance-for-Industry.pdf. Accessed 10 Aug 2022.
EMA. Guideline on bioanalytical method validation. https://www.ema.europa.eu/en/documents/scientific-guideline/guideline-bioanalytical-method-validation_en.pdf. Accessed 10 Aug 2022.
Schulz J, Thomas A, Saleh A, Mikus G, Kloft C, Michelet R. Towards the elucidation of the pharmacokinetics of voriconazole: a quantitative characterization of its metabolism. Pharmaceutics. 2022;14(3):477.
Hossain MA, Tran T, Chen T, Mikus G, Greenblatt DJ. Inhibition of human cytochromes P450 in vitro by ritonavir and cobicistat. J Pharm Pharmacol. 2017;69(12):1786–93.
Moriyama B, Obeng AO, Barbarino J, Penzak SR, Henning SA, Scott SA, et al. Clinical Pharmacogenetics Implementation Consortium (CPIC) guidelines for CYP2C19 and voriconazole therapy. Clin Pharmacol Ther. 2017;102(1):45–51.
Theuretzbacher U, Ihle F, Derendorf H. Pharmacokinetic/pharmacodynamic profile of voriconazole. Clin Pharmacokinet. 2006;45(7):649–63.
Yanni SB, Annaert PP, Augustijns P, Bridges A, Gao Y, Benjamin DK Jr, et al. Role of flavin-containing monooxygenase in oxidative metabolism of voriconazole by human liver microsomes. Drug Metab Dispos. 2008;36(6):1119–25.
van Wanrooy MJ, Span LF, Rodgers MG, van den Heuvel ER, Uges DR, van der Werf TS, et al. Inflammation is associated with voriconazole trough concentrations. Antimicrob Agents Chemother. 2014;58(12):7098–101.
Yanni SB, Annaert PP, Augustijns P, Ibrahim JG, Benjamin DK Jr, Thakker DR. In vitro hepatic metabolism explains higher clearance of voriconazole in children versus adults: role of CYP2C19 and flavin-containing monooxygenase 3. Drug Metab Dispos. 2010;38(1):25–31.
Krueger SK, Williams DE. Mammalian flavin-containing monooxygenases: structure/function, genetic polymorphisms and role in drug metabolism. Pharmacol Ther. 2005;106(3):357–87.
Aitken AE, Morgan ET. Gene-specific effects of inflammatory cytokines on cytochrome P450 2C, 2B6 and 3A4 mRNA levels in human hepatocytes. Drug Metab Dispos. 2007;35(9):1687–93.
Acknowledgements
The authors would like to express their gratitude for the excellent assistance of Katja Gümüs, MD, Simon A. Hermann, MD, Brigitte Tayrouz, RN, Sophie Glockner, RN, Marlies Stützle-Schnetz, MSc, RN, and Florian Michel.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
Open Access funding enabled and organized by Projekt DEAL.
Conflict of interest
No conflicts of interest have been declared by any author.
Ethics approval
The trial protocol was approved by the responsible ethics committee of the Faculty of Medicine of Heidelberg University (Afmo-431/2020) and the German competent authority (BfArM) on 13 July 2020 (EudraCT No: 2020-001017-20, registered on March 5th, 2020).
Availability of data and material
The collected and analysed data can be made available upon reasonable request.
Code availability
Not applicable.
Consent for publication
Not applicable.
Informed consent
Informed consent was obtained from all participants included in this trial prior to carrying out any trial procedures.
Author contributions
G.M. proposed the research topic. A.M., A.B., F.S., G.M. and W.E.H. developed the trial protocol. A.M. and A.B. executed the trial. J.B. and K.I.F. analysed the samples. A.M., G.M. and A.D.M. performed the statistical analyses. A.M. wrote the first draft of the manuscript. All authors discussed and interpreted the results and contributed to the manuscript.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Muhareb, A., Blank, A., Meid, A.D. et al. CYP3A and CYP2C19 Activity Determined by Microdosed Probe Drugs Accurately Predict Voriconazole Clearance in Healthy Adults. Clin Pharmacokinet 62, 1305–1314 (2023). https://doi.org/10.1007/s40262-023-01287-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40262-023-01287-7