Abstract
Purpose
Genetic and genomic testing can provide valuable information on individuals’ risk of chronic diseases, presenting an opportunity for risk-tailored disease screening to improve early detection and health outcomes. The acceptability, uptake and effectiveness of such programmes is dependent on public preferences for the programme features. This study aims to conduct a systematic review of discrete choice experiments assessing preferences for genetic/genomic risk-tailored chronic disease screening.
Methods
PubMed, Embase, EconLit and Cochrane Library were searched in October 2023 for discrete choice experiment studies assessing preferences for genetic or genomic risk-tailored chronic disease screening. Eligible studies were double screened, extracted and synthesised through descriptive statistics and content analysis of themes. Bias was assessed using an existing quality checklist.
Results
Twelve studies were included. Most studies focused on cancer screening (n = 10) and explored preferences for testing of rare, high-risk variants (n = 10), largely within a targeted population (e.g. subgroups with family history of disease). Two studies explored preferences for the use of polygenic risk scores (PRS) at a population level. Twenty-six programme attributes were identified, with most significantly impacting preferences. Survival, test accuracy and screening impact were most frequently reported as most important. Depending on the clinical context and programme attributes and levels, estimated uptake of hypothetical programmes varied from no participation to almost full participation (97%).
Conclusion
The uptake of potential programmes would strongly depend on specific programme features and the disease context. In particular, careful communication of potential survival benefits and likely genetic/genomic test accuracy might encourage uptake of genetic and genomic risk-tailored disease screening programmes. As the majority of the literature focused on high-risk variants and cancer screening, further research is required to understand preferences specific to PRS testing at a population level and targeted genomic testing for different disease contexts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Genetic and genomic risk-tailored disease screening programmes present the opportunity to improve early detection and health outcomes |
A large number of programme features were found to influence preferences, leading to estimated uptake rates ranging from no participation to almost full participation |
Careful communication of potential survival benefits and likely genetic/genomic test accuracy might encourage uptake of genetic and genomic risk-tailored disease screening programmes |
1 Introduction
Genetic and genomic testing can provide valuable information on individuals’ risk of specific diseases, enabling risk-tailored prevention and early detection to improve health outcomes. Such risk information can also be combined with risk due to other factors and used to tailor existing or new national chronic disease screening programmes, for example, through an earlier starting age or more frequent disease screening for people at high risk [1,2,3]. The principle of risk-tailored disease screening is already incorporated into many clinical guidelines through consideration of family history [4, 5], and including genetic and genomic information could reduce screening-associated harms, while improving the efficiency of resource utilisation [6, 7]. However, the potential benefits of risk-tailored programmes depend on the specific approaches to genetic/genomic testing and subsequent disease screening, the specific healthcare context and population and uptake rates among the population. Understanding public preferences towards the features of genomics-informed cancer screening is essential to ensure patient-centred care and to ensure uptake is not compromised.
In particular, tests for rare, highly penetrant variants that strongly predispose to disease can identify individuals at markedly high risk. In some countries, specific gene panel tests are already available for individuals who fulfil certain criteria, for example, testing for pathogenic BRCA1/2 variants for those with a strong family history of breast cancer, or for women with ovarian cancer [8]. Even if a person is already known to have the disease, such testing can enable further tailored prevention and/or treatment, as well as cascade testing of close relatives to enhance their disease prevention or screening [9].
Large-scale studies are now underway to assess population-wide carrier screening for rare, highly penetrant variants in selected genes, including DNAscreen in Australia [10]. Some studies are also considering use of population-wide whole genome or exome sequencing to test for rare, highly penetrant variants in a wider range of genes and thus diseases, such as MyCode in the USA [11]. In parallel, there are several trials considering the use of polygenic risk scores (PRS) for disease risk assessment, such as WISDOM for breast cancer and a pilot trial for including heart disease PRS into routine risk assessments in the UK National Health Service [12]. PRS combine risk information across many hundreds or even thousands of genetic variants that each only have a modest effect on disease risk, but together can capture a substantial proportion of disease predisposition [13]. PRS can thus be used to stratify the wider population into different risk groups.
To ensure genetic or genomic risk assessment and risk-tailored disease screening can effectively and efficiently improve health outcomes for a population in practice, the design and delivery of any such programme needs to incorporate public preferences in the specific setting. This includes processes, from the pre-test information and consultation to the genetic/genomic test itself, provision of risk information and disease screening, as well as costs, data privacy and health or life insurance impacts [14]. Such preferences are also likely to depend on whether the programme was offered population-wide or targeted to specific population subgroups (e.g. those with personal or family history of disease) and whether the test considered rare, high-risk variants or PRS.
Discrete choice experiments (DCEs) are a stated preference method that can be used to characterise preferences and understand the trade-offs people make between programme features [15]. In DCEs, participants are typically presented with a series of choice tasks within a survey, where they are asked to choose between two or more alternatives. In each choice task, the alternatives are hypothetical scenarios, in this case specific genetic and/or genomic tests to inform screening and are made up of attributes (e.g. cost and test accuracy) with varying levels. By having each participant complete a number of these choice tasks, researchers are then able to quantify how important attributes and levels are relative to each other. When cost is included as one of the attributes, willingness to pay for particular features of the programme can be calculated. For example, if test accuracy is also included, you can calculate how much people are willing to pay for a test with higher accuracy. The results can also be used to estimate potential uptake. Uptake estimates are important not just to ensure patient-centred care and to maximise health outcomes but can also have a significant impact on the cost effectiveness of an intervention [16].
A recent systematic review identified DCEs and conjoint analysis studies (a broader type of quantitative patient preference study) on genetic and genomic testing in all contexts (screening, diagnostic testing, carrier testing and pharmacogenetics) and provided an overview of the different methodological approaches [17]. Although the review provides useful information for those conducting a DCE, preferences for the implementation of genetic and genomic testing in the context of screening are not presented separately for different types of tests (e.g. rare, high-risk variants versus PRS). These different testing approaches have implications for the complexity and interpretation of the test results and consequences of testing, whereby these risks display different inheritance patterns [13]. In addition, it is important to consider how preferences differ between contexts of population-wide screening versus targeted testing of high-risk subgroups, which impacts the implementation, uptake and feasibility of the programme.
Here, we aim to summarise methodological approaches and provide a synthesis of the results of DCEs examining preferences for genetic and genomic testing in the context of adult population-based chronic disease screening. Further, we aim to distinguish preferences for genomics-risk–informed programmes focusing on rare, high-risk genomic variants and PRS.
2 Materials and Methods
2.1 Review Type
This systematic review was reported following the Preferred Reporting Items in Systematic Reviews and Meta-analyses (PRISMA) statement [18] and has been registered with the International Prospective Register of Systematic Reviews (PROSPERO) [CRD42021284701]. An internal protocol was prepared and accepted by all authors prior to beginning the review, no further amendments were made.
2.2 Information Sources
Systematic searches were conducted in three databases: EMBASE, Medline and EconLit. The PROSPERO database and the Cochrane Library were also searched to ensure there were no existing or planned systematic reviews in this area. The search span covered database inception until October 2023, with initial searches conducted in October 2021 and then updated in December 2022 and October 2023. To supplement the literature search, backward and forward citation searching was used to ensure no studies were missed.
The systematic search strategy was based on three concepts: (i) DCEs, (ii) genetic/genomic testing and (iii) chronic disease screening and early detection. During initial scoping searches, it was observed that many DCEs were being coded as conjoint analysis. Consequently, search terms for conjoint analysis were included. The PubMed search included the following search terms:
(‘discrete choice experiment’ [All Fields] OR ‘discrete choice experiments’ [All Fields] OR ‘discrete choice modeling’ [All Fields] OR ‘discrete choice modeling’ [All Fields] OR ‘discrete choice conjoint experiment’ [All Fields] OR ‘stated preference’[All Fields] OR ‘part worth utilities’[All Fields] OR ‘functional measurement’ [All Fields] OR ‘paired comparisons’ [All Fields] OR ‘pairwise choices’ [All Fields] OR ‘conjoint analysis’ [All Fields] OR ‘conjoint measurement’ [All Fields] OR ‘conjoint studies’ [All Fields] OR ‘conjoint choice experiment’ [All Fields] OR ‘conjoint choice experiments’ [All Fields]) AND (‘genetic’ [All Fields] OR ‘genomic’ [All Fields] OR ‘polygenic’ [All Fields] OR ‘pharmacogenetic’ [All Fields]) AND (‘screening’ [All Fields] OR ‘early detection’ [All Fields] OR ‘prevent*’ [All Fields] OR ‘risk’ [All Fields] OR ‘predict*’ [All Fields] OR ‘diagno*’ [All Fields])
Similar search terms were used for Embase, EconLit and the Cochrane Library and can be found within the electronic supplementary material (ESM).
2.3 Eligibility Criteria
Discrete choice experiments measuring patient or general preferences for genetic or genomic testing for risk of chronic diseases were eligible for inclusion in this review. Reasons for exclusion were classified as the following: ‘Does not use DCE’, ‘Is not written or published in English’, ‘Does not focus on genetic or genomic information’, ‘Does not relate to chronic disease screening’, ‘Focuses on the return of incidental findings’, ‘Does not measure patient or public preferences’, ‘Conference proceeding’, ‘Abstract only’.
2.4 Study Selection and Data Extraction
Studies were selected following PRISMA reporting guidelines [18]. Two reviewers screened all titles and abstracts against the inclusion/exclusion criteria (JC and AS). Where disagreements occurred, a third reviewer (AP) was consulted. Full texts were obtained for eligible studies based on the title/abstract screen and data from included studies were double extracted (JC and AS). A data extraction template was created within Microsoft Excel (JC) and was based on three prior systematic reviews of DCEs in health economics [17, 19, 20]. Additional data extraction fields were added that were not included in the previous systematic reviews but were contextually relevant to the research objective (e.g. disease(s), genetic variant type, population for genetic testing, genetic testing technology, opt-out rates, significant and non-significant attributes). The data extraction template can be found within the ESM. The following information was extracted from each study: (i) basic characteristics including study location, population surveyed, number of respondents and disease; (ii) information on the genetic test including the genetic variant type, population for genetic testing and genetic test technology; (iii) DCE characteristics including number of options, included attributes, range of levels for included attributes, inclusion of an opt-out option, experimental design, analysis method, outcomes reported (WTP or uptake); and (iv) DCE results, significance of included attributes (based on study-specific p-value), most and least important attributes (based on methods used by study author, including part worth utilities, marginal rates of substitution and rankings).
2.5 Quality Assessment
Quality assessment was performed using the Mandeville checklist [21]. The Mandeville checklist is based on the Lancsar and Louviere checklist (designed for DCEs within a general healthcare context) [22], as well as areas of concern highlighted by a review of DCEs in healthcare [20]. The Mandeville checklist focuses on quality assessment, rather than reporting standards, by limiting the criteria to those considered a substantive threat to the validity of results. Each criterion was marked, and colour coded as either Yes (dark green), Assumed Yes (light green), Partially (orange), No (red) or Unsure (grey). Given the Mandeville checklist focuses on key areas of quality, studies with more than half the criteria coded as red, orange or grey were considered low quality. Due to the low quality of these studies, the results were less likely to reflect the true preferences of participants and were excluded from the analysis.
2.6 Data Analysis
2.6.1 Basic Characteristics
Summary statistics (such as number and range) were created for (i) basic characteristics, including sample size, study location, diseases and population surveyed; and (ii) genomic test characteristics, including type of genetic variant, population tested and technology used. Studies were grouped by the type of genetic variant, the population tested and the genomic technology used.
2.6.2 DCE Development
The frequency of inclusion of each attribute within a study was calculated. Attributes were grouped together where feasible; for example, accuracy encompasses false negatives, true positives and the general term ‘accuracy’. Similarly, survival and mortality were grouped together. Attributes were allocated to the following categories: personal characteristics (attributes relating to the individual taking the test e.g. genetic predisposition and perceived risk), process of testing (attributes related to the procedures, methods, or protocols involved in the testing or screening process itself), testing characteristics (attributes related to the genetic test e.g. sensitivity and specificity) and outcomes of testing/screening (attributes related to the results or consequences of the testing or screening process). These categories were based on those presented by a recent systematic review [17]. Allocation was based on the judgement of the research team and there may be conceptual overlap between the categories. The number of attributes in each category was calculated.
Summary statistics were calculated for DCE development characteristics, including the range and most common number of alternatives, the range and most common number of levels and the number of studies which incorporated an opt-out option.
2.6.3 Preference Results
The following preference results were calculated: number of studies which found an attribute to be statistically significant (based on the study-specific p-value); and the number of studies which found an attribute to be most important or least important (based on the measure used by the study such as marginal rates of substitution and part worth utilities). Where diverse preferences were found across studies, individual studies were explored in more detail and a descriptive analysis conducted.
A descriptive analysis was conducted for studies that assessed uptake and/or WTP, aiming to synthesise findings across studies and provide contextualisation for the results.
3 Results
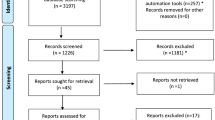
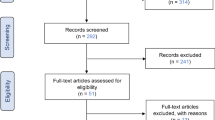
The database search yielded 469 records published from inception until October 2023 (See PRISMA diagram, Fig 1). After removing duplicates, 271 studies were included in the title/abstract screen. Two additional records were identified through scanning included studies in relevant systematic reviews and were excluded due to ineligible intervention, as genomics-informed chronic disease screening was not the focus of the study [23,24,25]. Nineteen articles were obtained for full-text review, and of these, 12 met the inclusion criteria (Table 1). The primary reasons for exclusion were ineligible intervention (n = 4), ineligible publication type (n = 2) and ineligible study type (n = 1). Studies excluded at the full-text screen can be found in the ESM.
3.1 Quality Assessment
No studies were marked as poor quality and thus all studies were included within the analysis. The quality of studies was particularly high in the analysis section, where the majority of studies considered the panel nature of data (where participants are answering multiple questions) and accounted for the heterogeneity of preferences, through the choice of DCE analysis model (e.g. the mixed MNL model or latent class model, which both account for different preferences across the sample) or subgroup analyses. A large proportion of studies used a Market Research Company for recruitment and the sample was reported to be representative of their intended population. Weaknesses were noted in the experimental design. Using pilot data to pick choice sets that are likely to provide the most information was not done in the majority of studies (i.e. to obtain priors for the D-efficient design), risking biased estimates of preferences. Lastly, a small proportion of studies did not undergo qualitative work with the target population, either through focus groups to obtain attributes and levels, or through qualitative piloting of the DCE (think aloud interviews). Results from the quality assessment of included studies are presented in the ESM.
3.2 Study Location, Population and Intervention
All studies were conducted in high-income countries: Australia (n = 3) [26,27,28], USA (n = 3) [29,30,31], the Netherlands (n = 3) [32,33,34], UK (n = 1) [35], Singapore (n = 1) [36], Sweden (n = 1) [25] and Germany (n = 1) [37] (Table 1). Ten of twelve studies were conducted in the context of cancer screening, colorectal cancer (n = 6) [29,30,31,32,33,34], breast cancer (n = 2) [26, 36] and multiple cancers (n = 2) [28, 35]. Plöthner et al. used broad disease classifications (all disease, treatable and serious hereditary) [37] and Goranitis et al. assessed a range of selected adult conditions for symptomatic adults (complex neurological conditions, mitochondrial conditions, cardiac arrythmias and Lynch syndrome) and at-risk adults (Huntington’s disease, familial cardiomyopathy and Lynch syndrome) [27]. Ten studies focused on testing for rare, high-risk variants (e.g. pathogenic variants in BRCA1/2 genes, or Lynch syndrome). Of these studies, six considered gene panel testing in a targeted population [26, 29, 30, 33,34,35, 38]; Knight et al. also considered gene panels, but it was not clear what population would undergo testing [30]; two studies explored preferences for using whole genome sequencing (WGS, in a wide range of genes) [23], or whole exome sequencing (WES, in a limited number of genes) [27], also in a targeted population; Plöthner et al. explored preferences for population-wide testing using WGS [37]. The remaining two studies explored preferences for population-wide testing using PRS and a targeted PRS assay [28, 36]. The populations surveyed varied from general population [27,28,29,30, 32,33,34,35, 37, 39] to targeted patient groups [26, 31, 34, 35] and did not always align with the population who would undergo testing. Eleven of twelve studies explored how preferences differed in the population: five studies tested the significance of selected participant characteristics using regression analysis [26, 27, 29, 31, 32], three studies assessed preferences in different population subgroups [34,35,36] and three studies used latent class analysis, where classes are defined using participant characteristics [28, 33, 37]. The sample size ranged from 115 [35] to 1045 [32] participants.
3.3 DCE Development
3.3.1 Included Attributes
Twenty-six attributes were identified (Table 2). The attributes were grouped into the following categories: (i) personal (n = 3); (ii) process (n = 13); (iii) testing (n = 5); and (iv) outcomes (n = 5). Attributes could cover multiple aspects, for example, Venning et al. used an ‘overall process’ attribute that specified both the healthcare professional providing the test and in-person versus telehealth consultation [28]. Peacock et al. included attributes from one category [26], five studies included attributes from two categories [32,33,34,35,36], three studies included attributes from three categories [28, 29, 37] and Goranitis et al. included attributes from all four categories [27]. Among all included studies, cost was most commonly included (n = 8). Among the ten studies that assessed rare, high-risk variants, the most commonly included attributes were cost (n = 6) and disease penetrance probability (n = 6), followed by genetic predisposition (n = 4) and privacy (n = 4). Among the two studies that assessed polygenic risk scores [28, 40], the only attribute included in both studies was cost. Half the attributes (n = 13) were only included within one study. It should be noted that Veldwijk et al. conducted three DCEs using the same attributes and levels, with different research questions. As a result, there is biased overlap between the attributes included within these studies (gene predisposition, probability of disease, screening impact and survival/mortality).
3.3.2 Number of Alternatives, Attributes and Attribute Levels
The number of attributes ranged from 4 to 8, with 4 being the most common (n = 8). The maximum number of levels ranged from 3 to 6, with 3 being the most common (n = 7). The number of alternatives presented to participants ranged from 2 to 3, with 3 being the most common (n = 10). Nine studies included an opt-out option within their DCE. Five of these studies used a two-stepped approach [32,33,34, 36, 37], where participants were presented with two questions, one as a forced choice and one with an opt-out option. The remaining used a one-step approach [27, 29,30,31], where participants were presented one question, which included an opt-out option. Griffith et al. presented an option for indifference between pairs of scenarios [35].
3.4 Preference Results
3.4.1 Significance of Attributes
The majority of attributes were found to significantly impact preferences (Table 2; with study-specific definition of statistical significance). Across studies, mixed preferences were found for privacy, impact on screening and availability of testing. Test location and number of tests were not found to be significant [31, 36].
Among the five studies which included privacy (who has access to the test results) [28,29,30, 37, 41], Venning et al. did not find it to be significant [28]. In contrast to the other studies, Venning et al. focused on PRS testing (not rare, high-risk variants). Additionally, ‘disclosure to insurance companies’ was a separate attribute rather than a level within privacy. Among the four studies which included impact on screening [28, 32,33,34], Venning et al. did not find it to be significant [28]. In contrast to the other studies, Venning et al. framed the attribute differently. The attribute focused on the number of people that would be impacted and how they would be impacted (more or less screening), rather than different frequencies of screening (every year, every 2 years, every 5 years). Griffith et al. found mixed results for availability of testing (for all patients or high-risk patients only) based on the actual risk level of the population [35]. People with low and moderate risk preferred testing to be offered to all risk levels as opposed to high-risk subgroups only, while people with high risk did not have a significant preference between the two approaches.
3.4.2 Most and Least Important Attributes
Among studies assessing testing for rare, high-risk variants, survival was most commonly reported as the most important attribute (n = 3) [32,33,34], followed by test accuracy (n = 2) [28, 31, 42] and impact on screening (n = 2) [32, 33]. The following attributes were reported as least important: genetic predisposition (the likelihood that a person is genetically predisposed to develop the disease, n = 2) [33, 41], probability of developing disease (n = 2) [32, 34], decision making (n = 1) [26], accuracy (n = 1) [30], number of tests (n = 1) [31] and privacy (n = 1) [37].
Mixed results were found within two of the Veldwijk studies. Veldwijk et al. tested whether positive or negative framing of an attribute (survival vs mortality) influenced their importance [32]. When framed positively, the most important attribute was survival; for negative framing, impact on screening was the most important (rather than mortality). In a separate study, Veldwijk et al. used a latent class model to separate out individuals with different preference types (‘classes’). For class 1, the most important attribute was survival while in class 2 it was impact on screening. For both classes, genetic predisposition was the least important attribute. Respondents with a higher educational level, who had no experience with genetic screening tests and who were less worried and anxious about their genetic predisposition were more likely to belong to class 1 than class 2.
Among studies assessing testing for PRS, type of healthcare provider (specialist doctor vs non-specialist doctor) [36] and test accuracy [24] were reported as the most important attributes. Sample type [36] and impact on screening [28] were reported as the least important attributes.
3.4.3 Uptake Rates and WTP Estimates for Genetic and Genomic Testing
Five studies calculated uptake rates or preferences for testing over no testing and WTP estimates [27, 29,30,31, 36]. Three studies only calculated uptake rates [32,33,34] and three studies only calculated WTP estimates [28, 35, 37]. Across all studies, uptake ranged from 0% [36] to almost full participation (97%) [29]. Studies reported uptake and WTP estimates across different clinical contexts and used different methods of calculation and reporting, limiting further synthesis across all studies.
The clinical context where multiple studies estimated uptake/WTP was rare, high-risk variant testing for a targeted population, using a targeted gene panel for colorectal cancer (n = 5). Knight et al. [30] calculated uptake rates for different scenarios of testing, assuming a test cost of USD500. Almost all participants (97%) would undergo genetic testing in the best scenario (no false negatives and results disclosed to primary care physician). In contrast, only 41% would undergo genetic testing in the worst scenario (20% false negatives and results disclosed to insurance company). The other studies reported different types of uptake including the proportion of participations who never chose to opt-out (88.9% of the general Dutch population) [32], the proportion of participants who always chose to opt-out (3.5% of the general population and 2.4% of high-risk patients within the Dutch national CRC screening programme) [34] and the significance of the opt-out option (not significant relative to genetic testing in CRC screening within the US) [29]. Outside the DCE context, Veldwijk et al. also asked participants whether they would participate in genetic testing within the Dutch national CRC screening programme and 89.1% said yes [33].
Knight et al. calculated the marginal WTP (mWTP) for testing compared with no testing, and each attribute level [30]. The mWTP of testing relative to no testing was USD622 and the largest mWTP was USD999 for sharing results with primary care doctors compared with insurance companies. Kilambi et al. calculated the ‘value of accurate test information’, finding the conditional value of accurate test information was USD1800 [29].
4 Discussion
This study summarises the discrete choice experiment literature on preferences for genetic and genomic testing in chronic disease screening programmes. Twelve studies were identified, all conducted in high income countries and the majority focused on cancer screening (largely colorectal cancer). Most studies explored preferences for testing of rare, high-risk variants in a targeted population, and only a small number of studies explored preferences for population-wide testing (either for high-risk variants or PRS). Due to the small number of studies assessing PRS, limited comparisons could be made between the different testing approaches. Multiple attributes were found to be important when considering genetic and genomic informed risk-tailored screening programmes, with survival being most frequently reported as most important. Genetic and genomic risk-tailored screening programmes were generally well received, although the extent of estimated participation varied significantly depending on different features of the programme, with predicted uptake rates ranging from zero to almost complete participation.
These results highlight that the design of genetic and genomic risk-tailored screening programmes requires careful consideration of preferences regarding their various features. The review identified a large number of attributes across the following categories: personal characteristics, process of testing, testing characteristics and outcomes of testing/screening. The largest number of attributes were identified in the process category. The large number can be partly attributed to the extensive qualitative research conducted in this area [43]. This qualitative literature highlights the importance of education and support at every stage of the screening programme (i.e. pre-test information and when receiving the results). Similarly, there is consensus that genomic tests have impacts extending beyond clinical outcomes, including both process impacts and intermediate outcomes. Consequently, there has been a drive for research to adequately capture these impacts through stated preference methods [44]. Across all categories, most attributes were found to impact on preferences, highlighting the large range of attributes which could be considered when designing genomics-informed screening programmes. Mixed preferences were found for privacy, which may be attributed to the different testing approaches (PRS vs identifying rare, high-risk variants) or the inclusion of different levels within the attribute (specifically the absence of insurance disclosure). Given the cancer risk associated with PGS can be as high as for single rare, high-risk variants, these findings suggest the main concern in regard to privacy is disclosure to insurance companies.
There was large variation between studies in terms of the attributes which participants found to be most and least important. This is likely due to the large number of attributes which impact on preferences and the resulting small number of attributes which overlap in the included studies. The attributes most frequently reported as most important were survival, followed by accuracy and impact on screening. These results align with previous reviews of stated preference studies in cancer screening [45, 46]. However, they contrast with the findings of Knight et al., where accuracy was reported as least important [30]. This may be due to the different ways accuracy was described and understood (e.g. false positives, false negatives, positive predictive values). Similarly, mixed preferences were found for impact on screening, which is likely due to the different ways the attribute was framed. This is consistent with the findings of Veldwijk et al., demonstrating how the importance of an attribute can vary depending on its framing [38]. These results emphasise the crucial role of the qualitative piloting stage in the DCE development, ensuring that participants have an appropriate understanding of the attributes and levels.
A large number of studies explored preferences for targeted testing for CRC risk. Population-based screening programmes for colorectal cancer are recommended and widely implemented in many high-income countries [47]. Three studies explored preferences for including genetic testing within current screening programmes, offering testing to those who receive a positive faecal immunochemical test (FIT). Overall, the public have positive attitudes towards genetic testing within this context, however their choice to undergo testing depends on the different features of the programme. Survival and frequency of colonoscopies appear to be the most important features. Interestingly, preferences for frequency of colonoscopies were found to differ based on participant characteristics. Those with lower education and experience with genetic testing preferred a higher number of colonoscopies, regardless of risk. This is likely due to a belief that more frequent colonoscopies will improve survival. Additionally, respondents tended to prefer testing that was more accurate, identified a higher proportion of individuals with high CRC risk and involved shorter wait time for results. To encourage uptake of genetic-tailored screening for colorectal cancer, it is important to communicate relevant aspects of survival and accuracy and consider the procedural features. As increased survival relies on participation in follow-up disease screening, effective communication of risk-appropriate screening and adherence to the recommendations is also crucial to realising the potential improvements in health outcomes.
While numerous large-scale trials are investigating the application of PRS in chronic disease screening programmes [12, 13], only two have evaluated preferences within this context. Venning et al. focused on multiple cancers and the Australian population [28]. This study found an overall preference for a PRS test that is highly accurate, tests for multiple cancers, enables non-invasive risk reduction measures and is performed through the GP (rather than getting the information online and doing a test at home). An unexpected result of this study was the lack of a clear preference for the chance the test would change screening recommendations, that is, no preference for tests that could help tailor subsequent screening for a larger number of people. This could reflect concerns about reduced screening, especially where screening programmes are already well established [48, 49]. Potential reduced screening of individuals with low risk is important from a policy perspective, as it influences the cost–benefit ratio of PRS testing and risk-tailored screening at a population level. Further research is required to understand how the public would respond to screening recommendations in this context. Wong et al. focused on incorporating genetic testing into the already established breast screening programme in Singapore, focusing on key process and testing characteristics [36]. The study suggests the most likely feasible option for implementation is a saliva test, which involved a pre-test discussion with a trained nurse educator at the hospital. In a realistic base-case scenario (saliva test, specialist doctor, hospital and testing cost of $175), uptake of testing was 3.7%, which increased to 88.9% in the most preferred scenario (saliva test, specialist doctor, private family clinic, testing cost of $50). These studies demonstrate incorporating PRS testing into cancer screening programmes has the potential to achieve high levels of acceptance, aligning with a recent UK DCE that finds a public preference for cancer screening programmes which consider both age and genetic risk scores, rather relying solely on the age-based criteria [24]. However, key aspects of implementation should be tailored to suit the particular context and demographic to ensure the benefits of genomics-informed screening are maximised.
DCEs need to be designed carefully to minimise the cognitive burden on participants, whilst also maximising the value of the information gained. In principle, there are no restrictions on the number of attributes that can be included within a DCE. However, in practice, most DCEs have contained fewer than 10 attributes to reduce the complexity of choice tasks and avoid ‘attribute non-attendance’, in which respondents simplify the task before answering by ignoring a subset of attributes [50]. Within this review, the most common number of attributes was 4, which is slightly lower than typically seen in DCEs of genetic and genomic testing [17] or more generally in health economics [19]. Given the large number of important attributes identified within this review and minimal overlap between included attributes, the median number of attributes included within an individual DCE appears relatively low. Interestingly, Venning et al. included the largest number of attributes and identified the largest number of attributes without a significant result. Whilst the number of attributes will depend on the research context, this review suggests there may be complex relationships between attribute preferences that remain unexplored within the current evidence base. D-efficient factorial designs and mixed MNL models were the most common methods of design and analysis, consistent with both best practice guidelines [51] and current practice [17] [19, 52]. A small proportion of studies did not include an opt-out option, these may have biased preference results as they include participants who in reality may not choose to participate.
An additional finding from this review was the importance of choosing the appropriate population to survey. Within included studies exploring preferences for targeted testing, the population surveyed was either the general adult population (either adults or adults of screening age) or those who would be offered testing (i.e. the targeted population). It has been argued by decision-making bodies that preferences from the general adult population should be sought, given their status as taxpayers who fund the public health system [53]. One the other hand, within the context of genetic and genomic testing, it is often argued that those with greater understanding are more likely to provide informed and considered responses. Veldwijk et al. compared how preferences for genetic testing within colorectal cancer screening differed between a more general population and those who would be offered testing (patients with a positive FIT) [25]. There were very different outcomes when looking at which attributes were more important to respondents [54]. Respondents in the targeted population focused on attributes related to outcomes (i.e. survival), while the general population focused on the more abstract attributes, genetic predisposition and CRC risk.
4.1 Limitations
This review focuses on overall preferences and does not systematically explore how preferences differ between population subgroups. For potential future genetics and genomics-informed screening programmes to equitably improve population health, it will be crucial to ensure that these programmes meet the needs of communities who have been or are disadvantaged, including culturally and linguistically diverse and rural and remote populations. A large proportion of the identified studies focused on colorectal cancer and targeted testing; therefore, the results may not be easily generalisable to other disease contexts or population-wide testing. The most commonly reported most and least important attributes are dependent on the attributes and levels included within the DCEs and may not be reflective of the attribute which is truly most or least important to respondents. Due to different categorisations of attributes, the results related to the types of attributes may not be comparable between studies. Lastly, only published studies were included, as a grey literature search was not conducted.
5 Conclusions
This systematic review synthesised the methods and results of 12 discrete choice experiments (DCEs) exploring preferences for genetic and genomic testing in chronic disease screening programmes. The overall findings suggest the public are interested in undergoing genetic and genomic testing for chronic disease screening. However, the uptake of potential programmes would strongly depend on specific programme features and the disease context.
The results of this review may offer insights for the development of acceptable, effective and efficient genomics-informed screening programmes aimed at improving health outcomes and meeting community needs. In particular, careful communication of potential survival benefits and likely genetic/genomic test accuracy might encourage uptake of genetic and genomic risk-tailored disease screening programmes. Further research is needed to determine how individuals would respond to specific risk-tailored screening recommendations and to understand preferences for incorporating polygenic risk scores into population-wide screening programmes, and preferences for targeted testing of highly penetrant variants in contexts other than colorectal cancer.
This study offers valuable insights for guiding future discrete choice experiments to explore preferences for genetic or genomic testing by presenting the broad scope of attributes which may impact on preferences. The review highlights the need for careful consideration of which attributes to include, selecting the appropriate population to survey and the importance of attribute framing and the resulting role of qualitative piloting.
Data Availability
The data used to support the findings of this study are available from the corresponding author upon reasonable request.
Code availability
Not applicable.
References
Jeon J, Du M, Schoen RE, et al. Determining Risk of colorectal cancer and starting age of screening based on lifestyle, environmental, and genetic factors. Gastroenterology (New York, NY 1943). 2018; 154: 2152-64.e19. https://doi.org/10.1053/j.gastro.2018.02.021
Weigl K, Thomsen H, Balavarca Y, et al. Genetic Risk Score Is Associated With Prevalence of Advanced Neoplasms in a Colorectal Cancer Screening Population. Gastroenterology (New York, NY 1943). 2018; 155: 88-98.e10. https://doi.org/10.1053/j.gastro.2018.03.030
Guo F, Weigl K, Carr PR, et al. Use of polygenic risk scores to select screening intervals after negative findings from colonoscopy. Clin Gastroenterol Hepatol. 2020;18:2742-51.e7. https://doi.org/10.1016/j.cgh.2020.04.077.
Shaukat A, Kahi CJ, Burke CA, et al. ACG clinical guidelines: colorectal cancer screening 2021. Am J Gastroenterol. 2021;116:458–79. https://doi.org/10.14309/ajg.0000000000001122.
Parkin C, Bell S, Mirbagheri N. Colorectal cancer screening in Australia: an update. Aust J General Practitioners. 2018;47:859–63.
Dixon P, Keeney E, Taylor JC, Wordsworth S, Martin RM. Can polygenic risk scores contribute to cost-effective cancer screening? A systematic review. Genet Med. 2022;24:1604–17. https://doi.org/10.1016/j.gim.2022.04.020.
Wang L, Liu C, Wang Y, Du L. Cost-effectiveness of risk-tailored screening strategy for colorectal cancer: a systematic review. J Gastroenterol Hepatol. 2022;37:1235–43. https://doi.org/10.1111/jgh.15860.
Forbes C, Fayter D, De Kock S, Quek RGW. A systematic review of international guidelines and recommendations for the genetic screening, diagnosis, genetic counseling, and treatment of BRCA-mutated breast cancer. Cancer Manag Res. 2019;11:2321–37. https://doi.org/10.2147/CMAR.S189627.
Reid S, Spalluto LB, Lang K, Weidner A, Pal T. An overview of genetic services delivery for hereditary breast cancer. Breast Cancer Res Treat. 2022;191:491–500. https://doi.org/10.1007/s10549-021-06478-z.
Lacaze PA, Tiller J, Winship I, et al. Population DNA screening for medically actionable disease risk in adults. Med J Aust. 2022;216:278–80. https://doi.org/10.5694/mja2.51454.
Carey DJ, Fetterolf SN, Davis FD, et al. The Geisinger MyCode community health initiative: an electronic health record-linked biobank for precision medicine research. Genet Med. 2016;18:906–13. https://doi.org/10.1038/gim.2015.187.
Eklund M, Broglio K, Yau C, et al. The WISDOM Personalized Breast Cancer Screening Trial: Simulation Study to Assess Potential Bias and Analytic Approaches. JNCI cancer spectrum. 2018; 2: pky067-pky67. https://doi.org/10.1093/JNCICS/PKY067
Khera AV, Chaffin M, Aragam KG, et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat Genet. 2018;50:1219–24. https://doi.org/10.1038/s41588-018-0183-z.
Tiller JM, Bakshi A, Brotchie AR, et al. Public willingness to participate in population DNA screening in Australia. J Med Genet. 2023;60:662–8. https://doi.org/10.1136/jmg-2022-108921.
McFadden D. Econometric models for probabilistic choice among products. J Business (Chicago, Ill). 1980;53:S13–29. https://doi.org/10.1086/296093.
Fern T-P, Matthew Q, Peter V, Peter V. Parameterising user uptake in economic evaluations: the role of discrete choice experiments: parameterising user uptake in economic evaluations. Health Econ. 2016;25:116–23. https://doi.org/10.1002/hec.3297.
Ozdemir S, Lee JJ, Chaudhry I, Ocampo RRQ. A systematic review of discrete choice experiments and conjoint analysis on genetic testing. Patient. 2022;15:39–54. https://doi.org/10.1007/s40271-021-00531-1.
Page MJ, McKenzie JE, Bossuyt PM, et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. Syst Rev. 2021;10:1–11.
Soekhai V, de Bekker—Grob E, Ellis AR, Vass CM. Discrete choice experiments in health economics: past, present and future. PharmacoEconomics. 2019; 37: 201-26. https://doi.org/10.1007/s40273-018-0734-2
de Bekker-Grob EW, Ryan M, Gerard K. Discrete choice experiments in health economics: a review of the literature. Health Econ. 2012;21:145–72. https://doi.org/10.1002/hec.1697.
Mandeville KL, Lagarde M, Hanson K. The use of discrete choice experiments to inform health workforce policy: a systematic review. BMC Health Serv Res. 2014;14:367–467. https://doi.org/10.1186/1472-6963-14-367.
Lancsar E, Louviere J. Conducting discrete choice experiments to inform healthcare decision making: a users guide. Pharmacoeconomics. 2008;26:661–77. https://doi.org/10.2165/00019053-200826080-00004.
Hall J, Fiebig DG, King MT, Hossain I, Louviere JJ. (2006) What influences participation in genetic carrier testing?. Results from a discrete choice experiment. J Health Econ. 25: 520–37. https://doi.org/10.1016/j.jhealeco.2005.09.002
Dennison RA, Taylor LC, Morris S, et al. Public preferences for determining eligibility for screening in risk-stratified cancer screening programs: a discrete choice experiment. Med Decis Making. 2023;43:374–86. https://doi.org/10.1177/0272989X231155790.
Viber-Johansson J, Langenskiold S, Segerdahl P, et al. Research participants’ preferences for receiving genetic risk information: a discrete choice experiment. Genet Med. 2019;21:2381–9. https://doi.org/10.1038/s41436-019-0511-4.
Peacock S, Apicella C, Andrews L, et al. A discrete choice experiment of preferences for genetic counselling among Jewish women seeking cancer genetics services. Br J Cancer. 2006;95:1448–53. https://doi.org/10.1038/sj.bjc.6603451.
Goranitis I, Best S, Christodoulou J, Stark Z, Boughtwood T. The personal utility and uptake of genomic sequencing in pediatric and adult conditions: eliciting societal preferences with three discrete choice experiments. Genet Med. 2020;22:1311–9. https://doi.org/10.1038/s41436-020-0809-2.
Venning B, Saya S, De Abreu Lourenco R, Street DJ, Emery JD. Preferences for a polygenic test to estimate cancer risk in a general Australian population. Genetics in Medicine. 2022; 24: 2144-54. https://doi.org/10.1016/j.gim.2022.07.011
Kilambi VPC, Johnson FRP, González JMP, Mohamed AFMA. Valuations of genetic test information for treatable conditions: the case of colorectal cancer screening. Value Health. 2014;17:838–45. https://doi.org/10.1016/j.jval.2014.09.001.
Knight SJ, Mohamed AF, Marshall DA, et al. Value of genetic testing for hereditary colorectal cancer in a probability-based US online sample. Med Decis Making. 2015;35:734–44. https://doi.org/10.1177/0272989X14565820.
Weymann D, Veenstra DL, Jarvik GP, Regier DA. Patient preferences for massively parallel sequencing genetic testing of colorectal cancer risk: a discrete choice experiment. Euro J Hum Genet. 2018;26:1257–65. https://doi.org/10.1038/s41431-018-0161-z.
Veldwijk JP, Essers BABP, Lambooij MSP, et al. Survival or mortality: does risk attribute framing influence decision-making behavior in a discrete choice experiment? Value Health. 2015;19:202–9. https://doi.org/10.1016/j.jval.2015.11.004.
Veldwijk J, Lambooij MS, Kallenberg FGJ, et al. Preferences for genetic testing for colorectal cancer within a population-based screening program: a discrete choice experiment. Euro J Hum Genet. 2016;24:361–6. https://doi.org/10.1038/ejhg.2015.117.
Veldwijk J, Groothuis-Oudshoorn CGM, Kihlbom U, et al. How psychological distance of a study sample in discrete choice experiments affects preference measurement: a colorectal cancer screening case study. Patient Prefer Adherence. 2019;13:273–82. https://doi.org/10.2147/PPA.S180994.
Griffith GL, Edwards RT, Williams JMG, et al. Patient preferences and National Health Service costs: a cost-consequences analysis of cancer genetic services. Fam Cancer. 2009;8:265–75. https://doi.org/10.1007/s10689-008-9217-5.
Wong XY, Groothuis-Oudshoorn CGM, Tan CS, et al. Women’s preferences, willingness-to-pay, and predicted uptake for single-nucleotide polymorphism gene testing to guide personalized breast cancer screening strategies: a discrete choice experiment. Patient Prefer Adherence. 2018;12:1837–52. https://doi.org/10.2147/PPA.S171348.
Plöthner M, Schmidt K, Schips C, Damm K. Which attributes of whole genome sequencing tests are most important to the general population? Results from a German preference study. Pharmacogenomics Personalized Med. 2018;11:7–21. https://doi.org/10.2147/PGPM.S149803.
Veldwijk J, Essers BAB, Lambooij MS, et al. Survival or mortality: does risk attribute framing influence decision-making behavior in a discrete choice experiment? Value Health. 2016;19:202.
Wong JZY, Chai JH, Yeoh YS, et al. Cost effectiveness analysis of a polygenic risk tailored breast cancer screening programme in Singapore. BMC Health Services Res. 2021;21:379. https://doi.org/10.1186/s12913-021-06396-2.
Wong SMY, Lam BYH, Wong CSM, et al. Measuring subjective stress among young people in Hong Kong: validation and predictive utility of the single-item subjective level of stress (SLS-1) in epidemiological and longitudinal community samples. Epidemiol Psychiatric Sci. 2021;30:195–6. https://doi.org/10.1017/S2045796021000445.
Goranitis I, Best S, Stark Z, Boughtwood T, Christodoulou J. The value of genomic sequencing in complex pediatric neurological disorders: a discrete choice experiment. Genet Med. 2021;23:155–62. https://doi.org/10.1038/s41436-020-00949-2.
Plöthner M, Schmidt K, Damm K, Schips C. Which attributes of whole genome sequencing tests are most important to the general population? Results from a German preference study. Pharmacogenomics and personalized medicine. 2020: 7.
Smit AK, Reyes-Marcelino G, Keogh L, et al. Implementation considerations for offering personal genomic risk information to the public: a qualitative study. BMC Public Health. 2020;20:1–1028. https://doi.org/10.1186/s12889-020-09143-0.
Regier DA, Weymann D, Buchanan J, Marshall DA, Wordsworth S. Valuation of health and nonhealth outcomes from next-generation sequencing: approaches, challenges, and solutions. Value Health. 2018;21:1043–7. https://doi.org/10.1016/j.jval.2018.06.010.
Phillips KA, Van Bebber S, Walsh J, Marshall D, Lehana T. Peer Reviewed: A Review of Studies Examining Stated Preferences for Cancer Screening. Preventing chronic disease. 2006; 3.
Hall MA, Rich SS. Patients’ fear of genetic discrimination by health insurers: the impact of legal protections. Genet Med. 2000;2:214–21. https://doi.org/10.1097/00125817-200007000-00003.
Navarro M, Nicolas A, Ferrandez A, Lanas A. Colorectal cancer population screening programs worldwide in 2016: an update. World J Gastroenterol. 2017;23:3632–42. https://doi.org/10.3748/wjg.v23.i20.3632.
Dunlop K, Rankin NM, Smit AK, et al. Acceptability of risk-stratified population screening across cancer types: qualitative interviews with the Australian public. Health Expect. 2021;24:1326–36. https://doi.org/10.1111/hex.13267.
Koitsalu M, Sprangers MAG, Eklund M, et al. Public interest in and acceptability of the prospect of risk-stratified screening for breast and prostate cancer. Acta Oncol. 2016;55:45–51. https://doi.org/10.3109/0284186X.2015.1043024.
DeShazo JR, Fermo G. Designing choice sets for stated preference methods: the effects of complexity on choice consistency. J Environ Econ Manag. 2002;44:123–43. https://doi.org/10.1006/jeem.2001.1199.
Bridges JFPP, Hauber ABP, Marshall DP, et al. Conjoint analysis applications in health—a checklist: a report of the ispor good research practices for conjoint analysis task force. Value Health. 2011;14:403–13. https://doi.org/10.1016/j.jval.2010.11.013.
Marshall D, Bridges JFP, Hauber B, et al. Conjoint analysis applications in health—how are studies being designed and reported?: An update on current practice in the published literature between 2005 and 2008. The Patient. 2010;3:249–56. https://doi.org/10.2165/11539650-000000000-00000.
NICE. Developing NICE guidelines: the manual. In: Excellence NIfHaC, ed. www.nice.org.uk/process/pmg20 2014.
Lázaro-Muñoz G, Conley JM, Davis AM, et al. Looking for trouble: preventive genomic sequencing in the general population and the role of patient choice. Am J Bioeth. 2015;15:3–14. https://doi.org/10.1080/15265161.2015.1039721.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions. This research has been funded by the Medical Research Future Fund’s grant #2007708 (The Australian Cancer Risk Study). Anne E. Cust is supported by a National Health and Medical Research Council Investigator Grant (#2008454).
Author information
Authors and Affiliations
Contributions
Conceptualisation: A.S., J.C., R.N., A.K.S., A.E.C., C.L., M.C., L.G., K.C., J.S., A.P.; Data curation: A.S., J.C.; Formal analysis: A.S., J.C., A.P.; Funding acquisition: A.E.C., J.S., A.P., K.C., M.C.; Investigation: not applicable; Methodology: A.S., J.C., R.N., A.K.S., A.E.C., C.L., M.C., L.G., K.C., J.S., A.P.; Project administration: A.S., J.S., A.P.; Resources: not applicable; Software: not applicable; Supervision: not applicable; Validation: not applicable; Visualisation: not applicable; Writing–original draft: A.S., J.C., J.S., A.P.; Writing–review & editing: A.S., J.C., R.N., A.K.S., A.E.C., C.L., M.C., L.G., K.C., J.S., A.P.
Corresponding author
Ethics declarations
Conflict of interests
The authors declare no competing interests.
Ethics declaration
This research has been approved by the Human Ethics Committee at the University of Sydney (#2022/844).
Consent to participate
Not applicable.
Consent for publication (from patients/participants)
Not applicable.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Salisbury, A., Ciardi, J., Norman, R. et al. Public Preferences for Genetic and Genomic Risk-Informed Chronic Disease Screening and Early Detection: A Systematic Review of Discrete Choice Experiments. Appl Health Econ Health Policy (2024). https://doi.org/10.1007/s40258-024-00893-1
Accepted:
Published:
DOI: https://doi.org/10.1007/s40258-024-00893-1