Abstract
Introduction
The availability of new classes of antiretroviral drugs is critical for treatment-experienced patients due to drug resistance to and unwanted side effects from current drugs. Our aim was therefore to evaluate the anti-HIV-1 activity of a new set of antivirals, dipeptides (WG-am or VQ-am) combined with a single-stranded oligonucleotide (ssON). The dipeptides were identified as naturally occurring and enriched in feces and systemic circulation in HIV-1-infected elite controllers and were proposed to act as entry inhibitors by binding to HIV-1 gp120. The ssON is DNA 35-mer, stabilized by phosphorothioate modifications, which acts on the endocytic step by binding to cell host receptors and inhibiting viruses through interference with binding to nucleolin.
Methods
Chou–Talalay’s Combination Index method for quantifying synergism was used to evaluate the drug combinations. Patient-derived chimeric viruses encoding the gp120 (env region) were produced by transient transfection and used to evaluate the antiviral profile of the combinations by drug susceptibility assays.
Results
We found that the combination WG-am:ssON or VQ-am:ssON had low combination index values, suggesting strong antiviral synergism. Of the two combinations, WG-am:ssON (1 mM:1 μM) had high efficacy against all prototype or patient-derived HIV-1 isolates tested, independent of subtype including the HIV-1-A6 sub-subtype. In addition, the antiviral effect was independent of co-receptor usage in patient-derived strains.
Conclusion
WG-am and ssON alone significantly inhibited HIV-1 replication regardless of viral subtype and co-receptor usage, and the combination WG-am:ssON (1 mM:1 μM) was even more effective due to synergism.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Drug resistance, side effects, and increasing issues with pharmacokinetic interactions remain a challenge in HIV-1 infected patients. |
New antiretroviral drugs such as novel compounds which target early steps of the viral cycle are needed. |
The dipeptides and single-stranded oligonucleotide (ssON) on their own and in combination are effective against all HIV-1 subtypes evaluated. The antiviral effect is independent of co-receptor usage. |
The two compounds represent two new categories of anti-HIV-1 compounds. |
Introduction
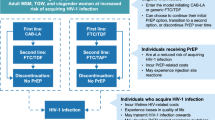
Despite improved HIV care due to combination antiretroviral therapy (cART), including the use of the new class of integrase strand-transfer inhibitors (INSTI), problems remain with drug resistance, occurrence of side effects, and increasing issues with pharmacokinetic interactions in elderly HIV-1 patients. Thus there is a clear need for new antiretroviral drugs such as novel compounds which can target the early steps of the viral cycle. HIV-1 cellular entry is mediated by the interaction between the envelope glycoproteins (gp120, gp41), the cellular CD4 receptor, and the co-receptors CCR5 or CXCR4. Recently, an attachment inhibitor, fostemsavir, and a post-attachment inhibitor ibalizumab have been approved for patients with multi-drug resistance [1]. In addition, small molecule inhibitors of the viral entry can interfere with CD4/gp120 binding [2, 3], co-receptor binding [4, 5] or gp41 six-helix bundle formation [6, 7] have been available for several years. However, all of these approved entry inhibitors are seldom used in clinical care.
Several viruses utilize endocytic pathways for uptake in target cells [8], triggering toll-like receptors (TLRs) located in endosomes. Since these host factors are employed by multiple viruses, the ability to achieve broad antiviral coverage is possible. Targeting virus entry is attractive, as this inhibition prevents subsequent replication [9]. In HIV-1 infected patients, TLR agonists are relevant due to their potential dual effects as immunomodulatory compounds and latency-reversing agents (LRAs) [10]. TLR3/7 are located in the membrane of the intracellular endosomes, whereas TLR4 is expressed on the cell surface [11], although TLR4 can also be found in the endosomes in some cell types [12, 13]. Certain single-stranded oligonucleotides (ssONs) inhibit, in a concentration-dependent manner, endocytic pathways used by cargo destined for TLR3/4/7 signaling endosomes [12, 14]. The selected non-coding 35-mer ssON used here is not an antisense or mimic but instead has the capacity to bind extracellularly and inhibit endocytosis of cargo [12, 14]. We previously showed that there is a length requirement of at least 25 bases for ssONs to have the capacity to inhibit endocytic uptake [12, 14]. The ssON used here is composed of 35 bases, and such long oligonucleotides are normally not taken up by cells unless, for example, transfection reagents are used. Our initial screening of ssONs activity against a broad variety of viruses showed that ssONs possess a broad potent antiviral activity, including HIV-1 (unpublished data), influenza A (23), and respiratory syncytial virus (RSV) [15].
Elite controllers (EC) are HIV-1 infected individuals who have a natural ability to control HIV-1 replication and disease progression in the absence of cART [16, 17]. Understanding these natural control mechanisms in the ECs could identify new unique therapeutic strategies. We recently used metabolomics to report the presence of certain dipeptides in blood plasma and feces of ECs, therapy-naïve HIV-1 progressors, and HIV-1 negative controls, which were enriched in the ECs compared to therapy-naïve HIV-1 progressors and HIV-1-negative controls. Valylglutamine (VQ) and tryptophylglycine (WG) showed the highest anti-HIV-1 activity [18]. While the exact mechanism is still unknown, molecular modeling predicts that VQ and WG can interfere with the gp120 binding to CD4. In contrast to naturally occurring ssONs, our specific ssON has phosphorothioate modification that increases its stability. The ssON-mediated interference of endocytosis does not act via antisense mechanisms or sequence complementarity. Although our specific ssON has a phosphorothioate modification that increases its stability, this is not essential for inhibition but is needed to retain activity over a longer period [12]. While expanded preclinical and clinical assessment of toxicity and pharmacokinetics is needed, the available data suggest that the naturally occurring dipeptides and oligonucleotides both possess anti-HIV activity and have potential for limited side effects. Therefore, we evaluated the efficacy of the combination of dipeptides and ssON against HIV-1 infection in vitro.
Methods
Compounds
Amide forms of valylglutamine (VQ-am) and tryptophylglycine (WG-am) were synthesized and purchased from Pepscan (purity of > 95%; Pepscan Presto, Lelystad, Netherlands). A 35-base-long phosphorothioate (PS)-modified oligonucleotide, designated ssON, with sequence 5′-GAAGTTTTGAGGTTTTGAAGTTGTTGGTGGTGGTG-3′, was purchased from Integrated DNA Technologies (Coralville, Iowa, USA).
Clinical Specimens
Stored plasma samples (n = 30) from patients who were given cART were randomly selected from the HIV-1 cohort at Karolinska University Hospital, Stockholm, Sweden. The HIV-1 cohort at Karolinska includes all (100%) diagnosed patients in Stockholm since 1995. The patients are followed in clinical care regularly 1–3 times a year at the Department of Infectious Diseases, Karolinska University Hospital. The pol region of the virus from all patients is sequenced at diagnosis and at treatment failure, when technically possible. The selection of the HIV-1 isolates was done to cover a large number of different HIV-1 subtypes and were randomly chosen among those available.
Subtyping using the pol region revealed the following: HIV-1A, n = 11; HIV-1B, n = 4; HIV-1C, n = 5; HIV-1D, n = 3; HIV-1F, n = 3; HIV-1G, n = 4. Of these, A1, 01_AE, and 02_AG were grouped as A-like viruses (n = 5), while A6 (n = 6) was categorized as an independent group HIV-1A6 sub-subtype. The co-receptor usage of the chimeric viruses was predicted using Genp2Pheno 2.5 [19]. Ten of 30 samples had at least one drug resistance mutation (DRM) (L74I, L74M, and/or Q148R) causing resistance to integrase strand inhibitors (INSTI), two of 30 samples had DRM (V82A and I50V) to protease inhibitors (PI) and 14 of 30 samples had DRM to reverse transcriptase inhibitors (RTI). The DRM were identified using our MiDRMpol pipeline [20]. A portion of the same virus isolates and resistance data were used in our earlier manuscript which evaluated other antiretroviral compounds [21].
Cells and Viruses
TZM.bl cell line (National Institutes of Health [NIH] AIDS Research and Reference Reagent Program, USA) and HEK-293T cells (American Type Culture Collection [ATCC], USA) were cultured in medium consisting of Dulbecco’s modified Eagle medium (DMEM; Sigma, USA) supplemented with 10% fetal bovine serum (FBS), penicillin/streptomycin (100 IU/mL and 50 mg/mL, respectively), and 2 mM l-glutamine. Peripheral blood mononuclear cells (PBMCs) were isolated on a Ficoll–Paque Plus density gradient (Merck SL, Madrid, Spain). Prior to treatment with the antiviral compounds, PBMCs were stimulated with the mitogen phytohemagglutinin (PHA) for 48 h (2 µg/mL; Thermo Fisher Scientific, Waltham, MA, USA). Antiviral compounds were added extracellularly without the use of a transfection reagent. Viral stocks of CCR5-tropic R5-HIV-1NLAD8 and CXCR4-tropic X4-HIV-1NL4.3 laboratory strains were obtained by transient transfection of pNL (AD8) and pNL (4.3) plasmids, respectively (NIH AIDS Research and Reference Reagent Program) in HEK-293T cells (ATCC, Manassas, VA, USA). Supernatants were collected at 48 h and 72 h. Viral stocks were clarified by centrifugation prior to evaluating the viral titer by HIV-1 p24 Gag ELISA kit (INNOTEST® Innogenetics/Fujirebio, Belgium).
Recombinant Virus Production
The chimeric viruses were generated as described previously [21]. Briefly, viral RNA was extracted using the QIAamp viral RNA extraction kit (Qiagen, Hilden, Germany) from 140 μL of plasma following the manufacturer’s protocol. The env region encoding for the gp120 fragments (HXB2: 6443–8439) was cloned into pMN plasmid following restriction digestion with ngoMIV and Mlu1 (New England Biolabs, USA), following ligation using T4 DNA ligase (New England Biolabs), as described previously [21]. The chimeric viruses were produced by transient transfection of the plasmids into the HEK-293T cell line using FuGENE HD Transfection Reagent (Promega, USA) and harvested 48 h and 72 h later by collection of the cell-free supernatant via centrifugation; aliquots were stored at −80 °C.
Cell Viability Assay (Membrane Integrity Assay)
Cell membrane integrity was measured by the CytoTox 96® Non-Radioactive Cytotoxicity (Promega, Germany) lactate dehydrogenase (LDH) assay, following the manufacturer’s instructions. Briefly, 104 TZM.bl cells were seeded in 96-well plates. At 24 h after seeding, cells were incubated with the compounds alone and the combinations for 48 h. Then, cells were lysed for 30 min at 4 °C and 50 μL of LDH reagent (Promega, Germany) was added for 30 min at room temperature, protected from light absorbance, and read in a Berthold plate reader at 490 nm. Three independent experiments were performed in triplicate. The 50% cytotoxicity concentration (CC50) for the combinations was calculated using nonlinear regression analysis (GraphPad Prism, version 8.0.1; GraphPad Software, La Jolla, CA, USA).
Drug Susceptibility Assay (DSA)
DSA was performed to determine the antiviral activity of ssON, WG-am, and VQ-am alone and in different combinations against the reference viruses (R5-HIV-1AD8 or X4-HIV-1NL4.3) and the chimeric viruses derived from 30 ART-treated patients. Drugs were serially diluted in culture medium, ranging from 1 mM to 1 µM for dipeptides and from 1 µM to 1 nm for ssON, and then added in triplicate in 96-well plates that had been seeded 24 h before the start with 104 TZM.bl cells/well. The viruses were added to each well at 100 TCID50 [50% tissue culture infectious dose]/well. In addition, DSA was performed in PHA-activated PBMCs for 48 h (2 × 105 cells/well in 200 µL). PBMCs pre-activated with PHA were seeded in round-bottom 96-well plates and treated with the compound combinations and challenged with HIV-1 infection. The antiviral compounds were added extracellularly without the use of a transfection reagent. Three days later, supernatants were collected and 100 µL was added to TZM.bl cells which had been seeded in 96-well plates at a density of 104 cells/100 µL per well the day before. HIV-1 replication was determined after quantification of luciferase activity (relative light units) using the Bright-Glo Luciferase Assay System (Promega, USA) 48 h post-infection. Drug concentrations required for inhibiting virus replication by 50% (EC50) were calculated from a dose–response curve using nonlinear regression analysis (GraphPad Prism, version 8.0.1; GraphPad Software, Inc., La Jolla, CA, USA). The synergistic profile of the compounds was determined using CalcuSyn software (Biosoft, Cambridge, UK), based on the median-effect principle [22]. Chou–Talalay’s combination indices (CIs) were calculated using methods derived from the mass-action principle, which is based on the following equation, CI = (fa/(1-fa))1/(fa/(1-fa))C +(fa/(1-fa))2/(fa/(1-fa))C, where fa is the fractional inhibition caused by the drug relative to the no-drug control; 1 and 2 represent the individual action of the drugs, and C represents the combined action of the drug combination. CI values < 0.9, 0.9 < CI < 1.,1 and CI > 1.1 indicate synergy, additivity, and antagonism, respectively.
The DSA experiments were performed with three technical replicates for each virus with the specified dynamic concentration range of the drug, and at least three independent analyses (biological replicates) were performed. The reproducibility of the DSA was assessed based on the 95% CI obtained for the drug EC50 and the degree of correlation between technical replicates. The output for the drug EC50 results was used to compute the fold change value for each virus relative to NL4.3 before being exported to GraphPad Prism.
Statistical Analysis
Statistics are presented as means ± standard deviation (SD) unless otherwise noted. Parametric and/or nonparametric statistical tests were performed as appropriate and are listed in the respective figure legends and tables. Statistical significance was accepted when P < 0.05. Statistical analysis was performed using GraphPad Prism 6 (GraphPad Software, Inc.) and CalcuSyn (Biosoft).
Ethics Statement
Ethics clearance for the study was obtained from the Regional Ethics Committee of Stockholm, Sweden (Dnr 2014/928-31/2 and 2013/1944-31/4). All participants gave informed consent.
Results
Evaluation of Cytotoxicity
Previously, the cytotoxicity of the dipeptides and ssON have been evaluated individually [12, 14, 15]. Here, we conducted cell viability assays using TZM.bl cells and PBMCs to evaluate the biocompatibility of these compounds individually (Fig. 1a) and their combination (Fig. 1b). The ssON was used together with WG-am or VQ-am at a 1:1000 ratio and WG-am with VQ-am at a 1:1 ratio. The standard deviation (SD) in the viability assay of untreated cells was around 13%; therefore, a cutoff of 80% was selected as non-cytotoxic measurement. As previously reported, WG-am and VQ-am were not toxic up to 5 mM, and ssON was not toxic at any of the concentrations tested, just reaching borderline values at 20 μM in PBMCs. WG-am or VQ-am in combination with ssON was non-cytotoxic in TZM.bl cells and in PBMCs up to 1 mM:1 μM (WG-am/VQ-am:ssON). VQ-am in combination with ssON, 1 mM:1 μM, was slightly toxic in PBMCs, showing viability of 76%. The lowest CC50 was seen with the combination WG-am:ssON (CC50: 9.5 mM:9.5 μM) and PBMCs (CC50: 5.6 mM:5.6 μM) (Table 1). We selected 1 mM:1 μM as the maximum non-cytotoxic concentration for the subsequent experiments.
Membrane integrity assay of WG-am or VQ-am and ssON. LDH assays were performed on TZM.bl cells and PBMCs after 48 h. Cells were treated with WG-am (1 μM to 20 mM), VQ-am (1 μM to 20 mM), and ssON (1 nM to 20 μM) alone (a) or in combination (b). Untreated cells were used as cell viability control and were set as 100% viability, and the values below 80% (red dotted line) were regarded as reduced viability based on the SD of untreated cells. Data shown represent the mean of three independent experiments
Evaluation of the Antiviral Inhibitory Profiles of the Combinations
Next, we evaluated the antiviral combination profiles of the compounds using the non-cytotoxic concentrations. The combinations of WG-am:ssON, VQ-am:ssON, and WG-am:VQ-am were evaluated in the DSA using TZM.bl cells and expressed as the combination index. All combinations tested significantly inhibited HIV-1 replication against the prototype CCR5 and CXR4 strains in TZM.bl cells and PBMCs.
To understand whether the combinations had synergistic, additive, or antagonistic effects, the combination index (CI) values of the compounds were calculated at different effective doses (ED): CI > 1.3 indicates antagonism, CI = 1.1–1.3 moderate antagonism, CI = 0.9–1.1 additive effect, CI = 0.8–0.9 slight synergism, CI = 0.6–0.8 moderate synergism, CI = 0.4–0.6 synergism, and CI = 0.2–0.4 strong synergism [23].
Based on low CI values, WG-am showed strong synergism with ssON at ED50, ED75, and ED90 against the NL4.3X4 and JRFLR5 strains (Table 1). Notably, the WG-am:ssON combination had a lower EC50 against CCR5-tropic HIV-1 JRFL (0.00065 mM:μM) compared to CXCR4-tropic NL4.3 X4 (0.00253 mM:μM). The VQ-am:ssON combination also showed synergism, which was weak at ED50 (0.840) but strong at ED75 (0.461) and ED90 (0.262) (Table 1). However, when the WG-am:VQ-am combination was evaluated, the CI values were high, indicating an antagonist effect (ED50: 11.425).
Because WG-am showed more effective anti-HIV-1 inhibition together with ssON than did VQ-am, and since WG-am and VQ-am had an antagonistic effect on TZM.bl cells, we only evaluated the antiviral activity of WG-am alone, ssON alone, and their combination on PBMCs. Similar to data obtained with TZM.bl cells, we found that WG-am:ssON had a strong synergistic effect and that both compounds, as well as their combination, had somewhat more potent anti-HIV-1 activity against the CCR5-tropic prototype HIV-1 JRFL strain (0.0024 mM:μM) compared to CXCR4 prototype NL4.3 X4strain (0.00339 mM:μM) (Fig. 2c, d).
Drug susceptibility assays of the combinations of ssON with WG-am or VQ-am. TZM.bl (a, b) and PBMCs (c, d) were treated with ssON (1 nM to 1 μM) and WG-am or VQ-am (1 μM to 1 mM) and then infected with HIV-1, JRFL (a, c) or NL4.3 (b, d) strains. Non-infected cells were used as infectivity control and defined as 100% of infection. The Log (concentration) refers to the nM concentration for ssON and μM concentration for dipeptides
Selectivity Index Calculation
The selectivity index (SI) was determined for all combinations. The SI is a ratio that measures the window between cytotoxicity and antiviral activity by dividing the given CC50 value by the EC50 value (EC50/CC50). A higher SI is indicative of a more effective and safer drug during in vivo treatment for a viral infection. SI greater than 1.000 is considered indicative of exceptional efficacy/safety [24]. WG-am:ssON resulted in the most effective treatment, achieving SI values of over 14.000 and 1.000 in TZM.bl cells against CCR5 and CXCR4 strains, respectively. The combination also showed a high selectivity index in PBMCs.
The VQ-am:ssON combination was also highly effective, albeit to a lesser extent than WG-am:ssON, in TZM.bl cells. Despite the high CC50, the VQ-am:WG-am combination resulted in a low SI index (28.1 and 33.32) due to their high EC50, 75.9 μM for the CCR5 strain and 64.01 μM for the CXCR4 strain, due to their antagonism.
WG-am:ssON Inhibits HIV-1 Replication Independently of the Subtype and Co-receptor Usage
Finally, we evaluated the inhibitory potential of WG-am alone and in combination with ssON towards a panel of patient-derived viruses comprising a wide variety of subtypes and drug resistance patterns, defined based on the analysis of the pol sequence. The evaluation of WG-am and ssON against chimeric viruses (n = 30) derived from ART-treated patients showed that 1 mM WG-am alone and 1 μM ssON alone significantly inhibited the replication of all viruses, independently of the subtype, but they did not completely abrogate the replication (Fig. 3a–i). In contrast, WG-am:ssON (1 mM:1 μM) inhibited all isolates > 95% (Fig. 3j) due to high synergism.
Drug susceptibility assays of the combination of ssON with WG-am. TZM.bl cells were treated with WG-am alone (1 μM to 1 mM), ssON alone (1 nM to 1 μM), and the combination (a–g) and then infected with ART-treated patient-derived chimeric viruses. N represents the number of viral isolates corresponding to each subtype. Maximum nontoxic concentrations of the compounds alone, 1 mM WG-am (h) and 1 μM ssON (i), and their combination, 1 mM:1 μM WG-am:ssON (j) are represented. Non-infected cells were used as infectivity control. The Log (concentration) refers to the nM concentration for ssON and μM concentration for dipeptides
We evaluated the co-receptor usage using Geno2pheno software, which showed that 16 of 30 patient-derived HIV-1 strains evaluated were CCR5-tropic, 12 of 30 were CXCR4-tropic, and the remaining two strains were not possible to predict with certainty. When we evaluated the differences in sensitivity to WG-am, ssON, and WG-am:ssON treatment, respectively, there were no significant differences according to CCR5 or CXCR4 usage (Fig. 4).
Drug susceptibility assays of WG-am, ssON, and ssON combined with WG-am according to the co-receptor usage. TZM.bl cells were treated with WG-am alone (1 μM to 1 mM) (a), ssON alone (1 nM to 1 μM) (b), and with the combination WG-am:ssON (c) and then infected with CCR5-tropic (n = 16) or CXCR4-tropic (n = 12) patient-derived chimeric viruses. N represents the number of viral isolates of each tropism. Non-infected cells were used as infectivity control. The Log (concentration) refers to the nM concentration for ssON and μM concentration for WG-am
Discussion
In this study, antiviral profiles were evaluated for the combination of two dipeptides, WG-am or VQ-am, with ssON against a variety of HIV-1 strains. These dipeptides have been identified as a naturally occurring enrichment in feces and blood of HIV-1 infected elite controllers. When tested earlier in vitro, the dipeptides were found to exhibit high anti-HIV-1 activity.
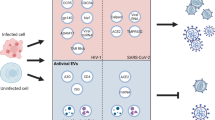
The 35-mer ssON temporarily inhibits certain endocytic pathways by interacting with the cell surface and thereby modulates the uptake of ligands destined for TLR3/4/7 signaling endosomes [8, 12]. Moreover, we previously showed that extracellular addition of ssON (no transfection reagents added) prevents RSV from binding to the cell surface and exhibits antiviral effects through interference with binding to nucleolin in the cell membrane [15]. Notably, nucleolin has been implicated as an HIV-1 cell surface binding receptor on CD4+ T cells [25]. Thus, both of these novel categories of compounds exhibit anti-HIV-1 activity on their own by inhibiting the entry process [18]. It was therefore of interest to evaluate their anti-HIV-1 efficacy and cytotoxicity in combination.
Based on their natural occurrence, the dipeptides and oligonucleotides are unlikely to have high toxicity. Indeed, the CC50 values for the combination of the dipeptides and the PS-stabilized ssON showed low toxicity, making them good candidates for further experiments. The Chou–Talalay Combination Index [23] method for quantifying synergism/antagonism has been increasingly used in the last two decades for a wide variety of drug combinations [26, 27], and we used it here for the WG-am:ssON, VQ-am:ssON, and WG-am:VQ-am combinations. Dipeptides and ssON use two different mechanisms to halt early steps of HIV-1 replication, the first being proposed as interfering with the gp120 attachment to the CD4 T-cell receptor and the latter acting on the endocytic step during entry by binding to other cell host receptors and/or preventing binding to the cell surface. Therefore, it was not surprising that WG-am:ssON and VQ-am:ssON had low CI at ED50, ED75, and ED90, respectively, suggesting strong antiviral synergism. WG-am EC50 was 7.8 µM, and in combination WG-am:ssON the EC50 was 0.65 µM/0.65 nM against the CCR5-tropic HIV-1 JRFL strain and completely halted the infection in TZM.bl cells and PBMCs at 1 mM/1 µM concentration. VQ-am:ssON showed a similar profile despite being slightly less effective. No differences in the synergistic profile were found when comparing different subtypes or CCR5- and CXR4 using strains.
Since both dipeptides act on the entry process of HIV-1, we were not surprised to find an antagonistic effect with a CI higher than 1.1 and an ED50 of 11.425 when combined, although the combination was still able to completely inhibit HIV-1 replication in both CCR5 (EC50 = 75.9)- and CXCR4 (EC50 = 64.01)-tropic strains. This is probably explained by the ability of dipeptides to bind to the same binding pocket which can block viral gp120 binding to CD4 on T cells.
A higher SI, the ratio between cytotoxicity and antiviral activity, corresponds to a higher possibility that a drug or combination will reach clinical trials, and as a reference, an SI greater than 1000 is considered exceptional [24, 28, 29]. Our data show that the SI of WG-am:ssON was very high in all cell lines tested and against different HIV-1 isolates, achieving values around 103–104 in PBMCs. On the contrary, the CI of WG-am:VQ-am was ~ 30. These results suggested us to focus our efforts in the WG-am:ssON combination.
Previous studies have suggested that the treatment outcome of cART may be associated with subtype-specific differences and with variability in the evolution of DRM [30,31,32]. Characterization of anti-HIV-1 agents should therefore adequately describe the antiviral efficacy not only for HIV-1B, but also for the globally dominant non-B subtypes. For example, for the recently approved attachment inhibitor fostemsavir, CRF01_AE isolates have naturally occurring resistance [33]. Also, natural subtype polymorphisms may influence the ease with which drug resistance develops, which has been emphasized for sub-subtype A6 and the use of INSTI [34, 35]. Thus, a combination of ≥ 2 factors, including pretreatment rilpivirine DRM, HIV-1 subtype A6/A1, and high body mass index (≥ 30 kg/m2), is associated with increased risk of treatment failure on long-acting cabotegravir and rilpivirine therapy [36]. The WG-am:ssON combination used here was highly effective against all subtypes including the A6 sub-subtype. Therefore, this combination could be a relevant candidate for treatment of any subtype strain, appearing after cART, including INSTI-based therapy.
Since dipeptides and ssON act on cell entry, it was important to evaluate the efficacy on HIV-1 strains with different co-receptor usage. The combination was found to be effective on both prototype CCR5- and CXCR4-tropic strains, although the CCR5 strain was inhibited somewhat more efficiently. We also determined the co-receptor usage of 30 patient-derived strains, as subtypes may differ in co-receptor usage. For example, HIV-1 subtype C predominantly uses the CCR5 receptor [37]. No significant difference was found between CCR5-tropic (n = 16) and CXCR4-tropic (n = 12) patient-derived strains. Thus, WG-am and ssON alone significantly inhibited HIV-1 replication regardless of subtype and co-receptor usage. However, their combination, WG-am:ssON (1 mM:1 μM), was even more effective due to the high synergism.
This study has limitations that merit comments. The chimeric viruses were generated to only carry the env region encoding for the gp120 fragments (HXB2: 6443–8439); therefore, only the ability to block the entry and attachment from different subtypes was tested. However, as pointed out before, both compounds tested targeted viral entry by targeting either gp120/CD4 binding or co-receptors. In addition, the low numbers and unevenness of some chimeric virus subtypes may seem problematic. However, we want to emphasize the wide range of subtypes evaluated and the total number of chimeric viruses produced.
Conclusions
The dipeptides and ssON on their own and in combination are effective against all HIV-1 subtypes, independent of co-receptor usage, and show low cytotoxicity and high synergism. We postulate that compounds from these new classes of antiviral drugs may inhibit HIV-1 replication as well as other viruses in vivo. If so, the compounds can be used as part of antiretroviral regimens in all HIV-1-infected patients including those with multidrug HIV-1 resistance.
References
Giroud C, Du Y, Marin M, et al. Screening and functional profiling of small-molecule HIV-1 entry and fusion inhibitors. Assay Drug Dev Technol. 2017;15(2):53–63. https://doi.org/10.1089/adt.2017.777.
Relano-Rodriguez I, Munoz-Fernandez MA. Emergence of nanotechnology to fight HIV sexual transmission: the trip of G2–S16 polyanionic carbosilane dendrimer to possible pre-clinical trials. Int J Mol Sci. 2020;21(24):9403. https://doi.org/10.3390/ijms21249403.
Si Z, Madani N, Cox JM, et al. Small-molecule inhibitors of HIV-1 entry block receptor-induced conformational changes in the viral envelope glycoproteins. Proc Natl Acad Sci USA. 2004;101(14):5036–41. https://doi.org/10.1073/pnas.0307953101.
Baba M, Nishimura O, Kanzaki N, et al. A small-molecule, nonpeptide CCR5 antagonist with highly potent and selective anti-HIV-1 activity. Proc Natl Acad Sci USA. 1999;96(10):5698–703. https://doi.org/10.1073/pnas.96.10.5698.
Dorr P, Westby M, Dobbs S, et al. Maraviroc (UK-427,857), a potent, orally bioavailable, and selective small-molecule inhibitor of chemokine receptor CCR5 with broad-spectrum anti-human immunodeficiency virus type 1 activity. Antimicrob Agents Chemother. 2005;49(11):4721–32. https://doi.org/10.1128/AAC.49.11.4721-4732.2005.
Pang W, Wang RR, Gao YD, et al. A novel enzyme-linked immunosorbent assay for screening HIV-1 fusion inhibitors targeting HIV-1 Gp41 core structure. J Biomol Screen. 2011;16(2):221–9. https://doi.org/10.1177/1087057110393333.
Stewart KD, Huth JR, Ng TI, et al. Non-peptide entry inhibitors of HIV-1 that target the gp41 coiled coil pocket. Bioorg Med Chem Lett. 2010;20(2):612–7. https://doi.org/10.1016/j.bmcl.2009.11.076.
Cossart P, Helenius A. Endocytosis of viruses and bacteria. Cold Spring Harb Perspect Biol. 2014;6(8): a016972. https://doi.org/10.1101/cshperspect.a016972.
Vanderlinden E, Naesens L. Emerging antiviral strategies to interfere with influenza virus entry. Med Res Rev. 2014;34(2):301–39. https://doi.org/10.1002/med.21289.
Martinsen JT, Gunst JD, Hojen JF, Tolstrup M, Sogaard OS. The use of toll-like receptor agonists in HIV-1 cure strategies. Front Immunol. 2020;11:1112. https://doi.org/10.3389/fimmu.2020.01112.
Tartey S, Takeuchi O. Pathogen recognition and Toll-like receptor targeted therapeutics in innate immune cells. Int Rev Immunol. 2017;36(2):57–73. https://doi.org/10.1080/08830185.2016.1261318.
Jarver P, Dondalska A, Poux C, et al. Single-stranded nucleic acids regulate TLR3/4/7 activation through interference with clathrin-mediated endocytosis. Sci Rep. 2018;8(1):15841. https://doi.org/10.1038/s41598-018-33960-4.
Kagan JC, Su T, Horng T, Chow A, Akira S, Medzhitov R. TRAM couples endocytosis of Toll-like receptor 4 to the induction of interferon-beta. Nat Immunol. 2008;9(4):361–8. https://doi.org/10.1038/ni1569.
Poux C, Dondalska A, Bergenstrahle J, et al. A single-stranded oligonucleotide inhibits toll-like receptor 3 activation and reduces influenza A (H1N1) infection. Front Immunol. 2019;10:2161. https://doi.org/10.3389/fimmu.2019.02161.
Palsson SA, Dondalska A, Bergenstrahle J, et al. Single-stranded oligonucleotide-mediated inhibition of respiratory syncytial virus infection. Front Immunol. 2020;11: 580547. https://doi.org/10.3389/fimmu.2020.580547.
Lopez-Galindez C, Pernas M, Casado C, Olivares I, Lorenzo-Redondo R. Elite controllers and lessons learned for HIV-1 cure. Curr Opin Virol. 2019;38:31–6. https://doi.org/10.1016/j.coviro.2019.05.010.
Noyan K, Nguyen S, Betts MR, Sonnerborg A, Buggert M. Human immunodeficiency virus type-1 elite controllers maintain low co-expression of inhibitory receptors on CD4+ T cells. Front Immunol. 2018;9:19. https://doi.org/10.3389/fimmu.2018.00019.
Sperk M, Ambikan AT, Ray S, et al. Fecal metabolome signature in the HIV-1 elite control phenotype: enrichment of dipeptides acts as an HIV-1 antagonist but a Prevotella agonist. J Virol. 2021;95(18): e0047921. https://doi.org/10.1128/JVI.00479-21.
Vicenti I, Lai A, Giannini A, et al. Performance of Geno2Pheno[coreceptor] to infer coreceptor use in human immunodeficiency virus type 1 (HIV-1) subtype A. J Clin Virol. 2019;111:12–8. https://doi.org/10.1016/j.jcv.2018.12.007.
Aralaguppe SG, Ambikan AT, Ashokkumar M, et al. MiDRMpol: a high-throughput multiplexed amplicon sequencing workflow to quantify HIV-1 drug resistance mutations against protease, reverse transcriptase, and integrase inhibitors. Viruses. 2019;11(9):806. https://doi.org/10.3390/v11090806.
Njenda DT, Aralaguppe SG, Singh K, et al. Antiretroviral potency of 4′-ethnyl-2′-fluoro-2′-deoxyadenosine, tenofovir alafenamide and second-generation NNRTIs across diverse HIV-1 subtypes. J Antimicrob Chemother. 2018;73(10):2721–8. https://doi.org/10.1093/jac/dky256.
Chou TC, Talalay P. Quantitative analysis of dose-effect relationships: the combined effects of multiple drugs or enzyme inhibitors. Adv Enzyme Regul. 1984;22:27–55. http://www.ncbi.nlm.nih.gov/pubmed/6382953.
Chou TC. Drug combination studies and their synergy quantification using the Chou–Talalay method. Cancer Res. 2010;70(2):440–6. https://doi.org/10.1158/0008-5472.CAN-09-1947.
De Clercq E. Potential antivirals and antiviral strategies against SARS coronavirus infections. Expert Rev Anti Infect Ther. 2006;4(2):291–302. https://doi.org/10.1586/14787210.4.2.291.
Callebaut C, Blanco J, Benkirane N, et al. Identification of V3 loop-binding proteins as potential receptors implicated in the binding of HIV particles to CD4(+) cells. J Biol Chem. 1998;273(34):21988–97. https://doi.org/10.1074/jbc.273.34.21988.
Garcia-Broncano P, Cena-Diez R, de la Mata FJ, Gomez R, Resino S, Munoz-Fernandez MA. Efficacy of carbosilane dendrimers with an antiretroviral combination against HIV-1 in the presence of semen-derived enhancer of viral infection. Eur J Pharmacol. 2017;811:155–63. https://doi.org/10.1016/j.ejphar.2017.05.060.
Lindstrom G, Jonathan H, Friedman R. The effects of combination treatments on drug resistance in chronic myeloid leukaemia: an evaluation of the tyrosine kinase inhibitors axitinib and asciminib. BMC Cancer. 2020;20(1):397. https://doi.org/10.1186/s12885-020-06782-9.
Theerawatanasirikul S, Kuo CJ, Phetcharat N, Lekcharoensuk P. In silico and in vitro analysis of small molecules and natural compounds targeting the 3CL protease of feline infectious peritonitis virus. Antiviral Res. 2020;174: 104697. https://doi.org/10.1016/j.antiviral.2019.104697.
You HL, Huang CC, Chen CJ, Chang CC, Liao PL, Huang ST. Anti-pandemic influenza A (H1N1) virus potential of catechin and gallic acid. J Chin Med Assoc. 2018;81(5):458–68. https://doi.org/10.1016/j.jcma.2017.11.007.
Hill KJ, Rogers LC, Njenda DT, et al. Strain-specific effect on biphasic DNA binding by HIV-1 integrase. AIDS. 2019;33(3):588–92. https://doi.org/10.1097/QAD.0000000000002078.
Kirichenko A, Lapovok I, Baryshev P, et al. Genetic features of HIV-1 integrase sub-subtype A6 predominant in Russia and predicted susceptibility to INSTIs. Viruses. 2020;12(8):838. https://doi.org/10.3390/v12080838.
van de Vijver DA, Wensing AM, Angarano G, et al. The calculated genetic barrier for antiretroviral drug resistance substitutions is largely similar for different HIV-1 subtypes. J Acquir Immune Defic Syndr. 2006;41(3):352–60. https://doi.org/10.1097/01.qai.0000209899.05126.e4.
Gartland M, Zhou N, Stewart E, et al. Susceptibility of global HIV-1 clinical isolates to fostemsavir using the PhenoSense(R) Entry assay. J Antimicrob Chemother. 2021;76(3):648–52. https://doi.org/10.1093/jac/dkaa474.
Lapovok IA, Lopatukhin AE, Kireev DE, et al. Molecular epidemiological analysis of HIV-1 variants circulating in Russia in 1987–2015. Ter Arkh. 2017;89(11):44–9. https://doi.org/10.17116/terarkh2017891144-49.
Riva C, Romano L, Saladini F, et al. Identification of a possible ancestor of the subtype A1 HIV Type 1 variant circulating in the former Soviet Union. AIDS Res Hum Retroviruses. 2008;24(10):1319–25. https://doi.org/10.1089/aid.2008.0119.
Cutrell AG, Schapiro JM, Perno CF, et al. Predictors of HIV-1 virologic failure to long-acting cabotegravir and rilpivirine: a multivariable analysis across three phase 3 studies. AIDS. 2021. https://doi.org/10.1097/QAD.0000000000002883.
Kalu AW, Telele NF, Gebreselasie S, et al. Monophylogenetic HIV-1C epidemic in Ethiopia is dominated by CCR5-tropic viruses-an analysis of a prospective country-wide cohort. BMC Infect Dis. 2017;17(1):37. https://doi.org/10.1186/s12879-016-2163-1.
Acknowledgements
We acknowledge Dr. Ujjwal Neogi for important intellectual contributions. We thank the participants of the study.
Funding
This work was supported by the Swedish Research Council awarded to A.-L.S (521-2014-6718) and awarded to A.S (2016-01675 and 2020-02129). It was also supported by grants from Region Stockholm (FoUI-955284 and FoUI-953887). The Journal's Rapid Service Fee was funded by the Swedish Research Council awarded to A.S (2020-02129).
Author Contributions
AS and ALS conceived the study. AS has initiated and designed the Swedish Elite controller cohort. RCD, ALS and AS designed the experimental plan. RCD performed the laboratory experiments. RCD and KS performed statistical and bioinformatical analyses. RCD wrote the first draft of the manuscript, with help from AS, KS and ALS which was then reviewed and approved by all the authors.
Disclosures
Anna-Lena Spetz is consultant and shareholder of TIRmed Pharma in possession of IPR regarding ssON. Rafael Cena-Diez, Kamalendra Singh and Anders Sönnerborg all have nothing to disclose.
Compliance with Ethics Guidelines
Ethics clearance for the study was obtained from the Regional Ethics Committee of Stockholm, Sweden (Dnr 2014/928-31/2 and 2013/1944-31/4). All participants gave informed consent.
Data Availability
Data sharing is not applicable to this article as no datasets were generated or analyzed during the current study.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Ceña-Diez, R., Singh, K., Spetz, AL. et al. Novel Naturally Occurring Dipeptides and Single-Stranded Oligonucleotide Act as Entry Inhibitors and Exhibit a Strong Synergistic Anti-HIV-1 Profile. Infect Dis Ther 11, 1103–1116 (2022). https://doi.org/10.1007/s40121-022-00626-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40121-022-00626-8