Abstract
In the present work, we used doxycycline hyclate (DX) and tetramethylguanidine (TMG) for the synthesis of silver–silver chloride nanoparticles (DX-Ag–AgCl NPs) as a simple method for the production of nanoparticles (NPs). A new synthesis method for DX-Ag–AgCl NPs in aqueous medium, using the DX and TMG as a reducing and stabilizing agent, is reported. DX-Ag–AgCl NPs were characterized by transmission electron microscopy. The elemental composition and the crystalline nature of synthesized nanoparticles were determined by energy-dispersive X-ray (EDX) spectroscopy and X-ray diffraction (XRD) analysis, respectively. From EDX, it is confirmed that synthesized NPs contain elemental silver and chloride, and their crystalline nature was confirmed by XRD. The Fourier transform infra-red spectra showed the DX present in surface of DX-Ag–AgCl NPs. Synthesized NPs analyzed for antibacterial activity by agar well method against Gram-positive and Gram-negative pathogens. Synthesized DX-Ag–AgCl NPs were spherical shaped and in the range of 10–40 nm of size was observed from TEM images. In the crystalline nature, XRD peaks indicate the presence of silver and silver chloride in produced NPs. Synthesized NPs show potential antibacterial activity on Bacillus cereus KCCM 11773, Bacillus subtilis KCCM 11316, Staphylococcus aureus KCCM 40050, Escherichia coli KCCM 11234, Klebsiella pneumoniae KCCM 11418, and Proteus vulgaris KCCM 40211.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nanotechnology is emerging as the fastest growing technology and interdisciplinary field for manufacturing of material on the nanoscale. In the recent years, interest in the synthesis of noble metal nanoparticles (NPs) has increased due to their significant applications in different fields [1, 2]. Nanoparticle synthesis is currently being focused on in the scientific community for the process is simple, rapid, and eco-friendly [3, 4]. Nowadays, nanotechnology is an active domain for the synthesis of nanoparticles through the use of natural resources like plants and microorganisms [5, 6]. Nanoparticles have significant importance in biomedical applications including antimicrobial, antioxidant, anticancer activities, and many more [7,8,9,10]. The silver nanoparticles (AgNPs) have emerged in recent decades as a useful chemical tool, due to their broad applications in microbiology [11, 12] and biomedicine [13, 14].
Different antibiotics have been reported for the synthesis of AgNPs such as, ampicillin [15], tetracycline [16], tobramycin [17], and amoxicillin [18]. Drug resistance is a major challenge, needed to develop new biomedicine to treat drug resistance pathogens without harming the normal cells [19]. Inorganic nanomaterials in the form of nanoparticles such as Ag, silver chloride (AgCl), and silver–silver chloride (Ag–AgCl) could be alternative drug to control pathogenic microbial infections. Ag NPs are metal NPs containing only silver [20], AgCl NPs are inorganic NPs containing silver and chloride ions [21], and Ag–AgCl NPs are composite NPs containing Ag and AgCl NPs [22]. The antibacterial activity of AgNPs has been well established [11, 12]. AgNPs target different sites in bacteria and inhibit their growth [5]. From the literatures, AgNPs arrest microbial growth by targeting different sites such as direct binding to cell surface which causes membrane damage, changes in cell membrane permeability, leakage of cell constituents [11, 12, 23], formation of reactive oxygen species [24, 25], and inhibition of vital proteins [26]. All these interactions of AgNPs with microorganisms are responsible for the inhibition or killing of microorganisms. Metal nanoparticles with antibiotics show enhanced bactericidal activity, and can be the alternative hybrid drug for the overcoming of multidrug-resistant bacterial infection [16, 17].
Here, we report a simple, one-pot, and rapid method for the synthesis of DX-Ag–AgCl NPs using doxycycline hyclate (DX) and tetramethylguanidine (TMG). The synthesized Ag–AgCl NPs were characterized using different techniques such as the UV–visible spectrophotometer, transmission electron microscope (TEM), energy-dispersive X-ray diffraction (EDX), X-ray diffraction (XRD), and Fourier transform infra-red spectroscopy (FT-IR). The synthesized NPs were evaluated for antibacterial activity on Gram-positive (Bacillus cereus KCCM 11773, Bacillus subtilis KCCM 11316, and Staphylococcus aureus KCCM 40050) and Gram-negative (Escherichia coli KCCM 11234, Klebsiella pneumoniae KCCM 11418, and Proteus vulgaris KCCM 40211) pathogenic bacteria by agar well-diffusion and broth microdilution method.
Materials and methods
Materials
Silver nitrate (AgNO3) was purchased from Fisher Scientific (New Jersey, USA). Doxycycline hyclate (DX), 1,1,3,3-tetramethylguanidine (TMG), and iodonitrotetrazolium chloride (INT) were purchased from Sigma-Aldrich (USA). Mueller–Hinton Broth (MHB) and Mueller–Hinton Agar (MHA) were obtained from BD Diagnostic, France. The 96-well plate was supplied by SPL Life science, Gyeonggi, Korea.
Synthesis of DX-Ag–AgCl NPs
AgNO3 (1 mM), DX (50 mg mL−1) and TMG (50 mg mL−1) solutions were prepared in triple distilled water. In a 10 mL of AgNO3 solution, 20 µL of DX and 40 µL of TMG were added, and the reaction was performed at 80 °C for 5 min with continuous stirring. Immediately after the reaction, the solution was cooled under running tap water. Separation of prepared NPs was performed by high-speed centrifugation (12,000 rpm, 30 min) and unbounded materials were removed by washing with distilled water twice. For characterization, powder form of DX-Ag–AgCl NPs obtained by lyophilization.
Characterization of DX-Ag–AgCl NPs
DX-Ag–AgCl NPs were observed visually, while the absorbance spectra were recorded by Jasco V-670 UV–Vis spectrophotometer (Japan) in the range of 300–800 nm during all reactions. Characterization of synthesized NPs was performed as per procedure reported previously [27, 28]. In brief, the morphological features such as the size, shape, and distribution of DX-Ag–AgCl NPs were observed through TEM, and, for the sample preparation, colloidal suspension was dropped on a copper disk and then dried at room temperature. TEM (Hitachi, H7500, Tokyo, Japan) images captured at 120 kV. Lyophilized DX-Ag–AgCl NPs were used to determine the elemental composition and crystalline nature by EDX spectroscopy and XRD, respectively. XRD (Philips X’Pert-MPD Diffractometer, Netherland) was operated at 40 kV and 30 mA using Cu Kα radiations. Fourier transform infra-red (FT-IR) spectra were obtained in transmission mode on powdered DX and lyophilized DX-Ag–AgCl NPs that were mixed with dry KBr (1:100) and compressed into a thin pellet film. The NPs zeta potential and particle-size analysis (PSA) were analyzed by electrophoretic light scattering spectrophotometer (ELS-8000, OTSUKA Elec., Osaka, Japan). Samples for zeta potential and PSA analysis were prepared/diluted in tripled distilled water [23]. FT-IR (Nicolet iS10, Thermo Electron Scientific Inst. LLC, Fitchburg, WI, USA) spectra in the range of 4000–500 cm−1 were obtained to determine the involvement of functional groups [28].
Antibacterial activity
The pathogenic Gram-positive (B. cereus KCCM 11773, B. subtilis KCCM 11316, and S. aureus KCCM 40050) and Gram-negative (E. coli KCCM 11234, K. pneumoniae KCCM 11418, and P. vulgaris KCCM 40211) bacterial strains used in this study were purchased from the Korean Culture Center of Microorganisms (KCCM), Republic of Korea. Agar well-diffusion method, minimum inhibitory concentration (MIC), and minimum bactericidal concentration (MBC) for DX and DX-Ag–AgCl NPs were performed as per previously published report [28].
Agar well-diffusion method
A single colony of bacterial strain was inoculated in 20 mL of the MHB broth and grown at 37 °C overnight. An overnight culture, 0.5 McFarland-standard bacterial suspension (1 × 108 CFU mL−1), was prepared by adjusting optical density (OD) to around 0.08–0.1, and then used for agar well-diffusion method. OD600 was obtained using a spectrophotometer (Libra S22, Biochrom Ltd, Cambridge, England). 100 µL of bacterial suspension spreaded over MHA plate with the help of sterile glass spreader, while the wells were prepared using 8 mm cork borer. 100 µL of DX and DX-Ag–AgCl NPs stock solutions (1 mg mL−1) were dropped in wells, and plates were incubated at 37 °C for 24 h. After incubation, the zone of inhibition (ZOI) diameter was measured in mm. These experiments were performed in triplicate.
Determination of MIC and MBC
The MIC and MBC were performed by broth microdilution and spread plate method, respectively. A 0.5 McFarland-standard adjusted bacterial suspension (1 × 108 CFU mL−1) was diluted with 1:100 ratios in MHB to obtain 1 × 106 CFU mL−1. In broth microdilution method, each well of 96-well plate contained 50 µL of DX and DX-Ag–AgCl NPs with different concentrations (0.5, 1, 2, 4, 6, 8, 10, 12, 14, and 16 µg mL−1) and 50 µL of each bacterial strain suspension (1 × 106 CFU mL−1) were added separately. Two sets of 96-well plate were prepared: one for MIC and another for MBC determination. Negative control was prepared, which contained only media; while the positive control contained bacterial strains without DX or DX-Ag–AgCl NPs. Plates were incubated for 24 h at 37 °C. After that, in one 96-well plate, 40 µL of INT (0.2 mg mL−1) solution was added to each well. The plate was further incubated for 30 min, to interaction of INT with enzyme dehydrogenase. Pink color formation can be seen in the presence of live cells, while there will be no color change in dead cells. The lowest dilution without color change will be indicated as the MIC. From the second plate, 100 µL of MIC concentrations was plated on MHA plate and spread with sterile glass spreader. These plates were incubated for 24 h at 37 °C. After incubation, MBC will be recorded as the lowest concentration of DX or DX-Ag–AgCl NPs that resulted in no bacterial growth.
Results and discussion
Characterization of DX-Ag–AgCl NPs
UV–visible spectroscopy
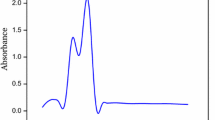
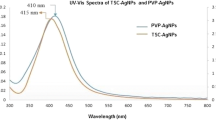
The reactions were carried out at boiling temperature to promote AgNO3 reduction. A typical reaction solution contained 10 mL of AgNO3 (1 mM), 20 µL of DX (50 mg mL−1), and 40 µL of TMG (50 mg mL−1) solution. Reaction was performed at 80 °C for 5 min. During reaction, the formation of DX-Ag–AgCl NPs was observed by change in the color of the solution. After reaction, brown color formation was observed and characteristic sharp peak at 407 nm in UV–visible spectroscopy was observed, as shown in Fig. 1. AgNPs shows characteristic peak, and the surface plasmon resonance (SPR) absorption band is strongly depending on the NPs size, surrounding chemicals and dielectric medium [29, 30]. The SPR peak towards shorter wavelength indicates decreased size of NPs [29]. Our findings are in good agreement with these reports.
Elemental and crystalline nature of nanoparticles
The DX-Ag–AgCl NPs synthesized by addition of 40 µL of TMG and 20 µL of DX in 10 mL of 1 mM AgNO3 solution, and reaction was performed at 80 °C for 5 min with continuous stirring; TMG and DX solutions were prepared in distilled water with concentration of 50 mg mL−1. Synthesized NPs was lyophilized for further characterization.
Figure 2 indicates the EDX spectrum, obtained from lyophilized NPs by EDX on FE-SEM. The EDX analysis confirms the presence of elements in the synthesized DX-Ag–AgCl NPs. The EDX spectrum shows two signals at 2.7 and 3.0 keV, which correspond to the Cl and Ag, respectively. The characteristic presence of element, Cl and Ag, in EDX spectrum confirms the composition of the Ag–AgCl NPs as being consistent with the previous reports [23, 28].
Figure 3 illustrates the XRD diffraction patterns for DX-Ag–AgCl NPs. Nine peaks were detected on the XRD pattern, with five peaks corresponding to AgCl and four to AgNPs. The diffraction peaks at 2θ = 27.96°, 32.35°, 46.26°, 54.97°, and 57.61° correspond to the diffractions from {111}, {200}, {220}, {311}, and {222} planes of a face-centered cubic (FCC) AgCl, respectively. Furthermore, diffraction peaks at 2θ = 38.29°, 44.39°, 64.55°, and 77.69° correspond to the diffractions from {111}, {200}, {220}, and {311} planes of a FCC Ag, respectively. The bars (FCC planes) in the graph are from the Joint Committee on Powder Diffraction Standards (JCPDS) reference diffraction files for Ag (#87-0720) and AgCl (#01-1013). The crystalline nature of DX-Ag–AgCl NPs was confirmed by XRD.
The crystalline size of DX-Ag–AgCl NPs was calculated by Debye–Scherrer’s equation as follows:
where D is the crystalline size in nm, K is the Scherrer’s constant (K = 0.94), λ is the wavelength of incident X-ray, β is the full width at half maximum (FWHM) of the diffraction peak, and 2θ is the diffraction angle. The major five diffraction peaks, viz, 32.35, 38.29, 46.26, 64.55, and 77.69 corresponding to (200), (111), (220), (200), and (311) hkl planes, were considered for the determination of average particle size of NPs [31]. XRD spectra revealed highly crystallinity, and the average crystalline particle size of DX-Ag–AgCl NPs was found 8.25 nm which was calculated by Debye–Scherrer’s formula. The plausible reaction mechanism for formation of AgCl NPs is presented in Scheme 1. These results are compatible with the previous reports of Ag and AgCl NPs synthesis using biological methods [23, 28] and chemical method [32].
Morphology of nanoparticles
The shape, size, and distribution of the DX-Ag–AgCl NPs were observed by TEM. In Fig. 4, the TEM image reveals the synthesized NPs mostly spherical, and polydisperse. In addition, the particle size was found to be in the range of 10–40 nm for DX-Ag–AgCl NPs. The polydisperse nature of NPs is due to the formation of Ag and AgCl NPs. These patterns were found to be well in agreement with those observed from XRD peaks and demonstrate the crystalline nature.
Zeta potential and particle size of nanoparticles
Zeta potential analysis was applied to determine the surface overall charges on NPs [23]. The zeta potential value of synthesized DX-Ag–AgCl NPs is − 5.01 mV (shown in Fig. 5), which indicates negative charges on the NPs. Distilled water were used as a dispersion medium for zeta potential and PSA analysis. Figure 6 indicates the histogram of synthesized DX-Ag–AgCl NPs’ particle-size distribution. From PSA analysis, it is observed that maximum NPs are in range of 5–20 nm. This observation is compatible with NPs’ size observed in TEM image.
Nanoparticle surface study by Fourier transform infra-red (FT-IR) spectroscopy
The FT-IR measurement was carried out to determine the presence of biomolecules on the nanoparticles copping layer. In Fig. 7, the FT-IR spectra of DX and DX-Ag–AgCl NPs suggested extensive similarities between the samples. From the FT-IR studies, the characteristic bands for functional groups of DX were identified, including O–H group at 3318 cm−1, C=O group at 1683 cm−1, and N–H group at 1573 cm−1. In particular, the characteristic bands of DX at 1650 cm−1 region related to the NH2 group, which is known to interact with HCl, cause a decrease in transmittance intensity. On the other hand, the DX-Ag–AgCl NPs sample had a more intense peak at 1650 cm−1 which corroborate the fact that HCl has been removed from the molecular complex and DX transformed to doxycycline monohydrate [33]. Formation of the doxycycline monohydrate from DX is presented in Scheme 1. The present study results are compatible with these results.
Plausible mechanism of DX-Ag–AgCl NPs formation
The plausible formation of AgCl NPs can be best explained on the basis of Scheme 1. Here, the DX is the source of H+ (via HCl) which, under the reaction conditions, can react with the TMG to produce the Lewis adduct and chloride anion. The chloride anion thus formed reacts with the free Ag+ in aqueous medium forming the AgCl NPs. However, the formation of Ag NPs can be best explained on the basis of hypothetical mechanism reported previously, by the functional groups (–OH) similarity of the DX with quercetin in the leaf extract [34].
Antibacterial activity of DX-Ag–AgCl NPs
As a potential application, the DX-functionalized Ag–AgCl NPs were tested as a bactericidal. Antibacterial activity was performed by agar well-diffusion method against Gram-positive (B. cereus KCCM 11773, B. subtilis KCCM 11316, and S. aureus KCCM 40050) and Gram-negative (E. coli KCCM 11234, K. pneumoniae KCCM 11418, and P. vulgaris KCCM 40211) pathogenic bacteria. The antibacterial activity of DX and DX-Ag–AgCl NPs was observed after 24 h. Figure 8 shows the antibacterial activity of synthesized DX-Ag–AgCl NPs compared with pure DX. The variation in the ZOI was noticed with respect to the type of bacterium, which might be due to the difference in the bacterial surface characteristics. Gram-positive and Gram-negative bacteria differ in several ways when looking at the cell wall. Gram-positive bacteria have a thick, multi-layered peptidoglycan (negatively charged), virtually none of lipopolysaccharide and presence of teichoic acid; Gram-negative bacteria have a thin single-layered peptidoglycan, which contains high lipopolysaccharides and absence of teichoic acid. All these different cell surface compositions make differences in the sensitivity of bacteria to NPs and antibiotics [35, 36]. Table 1 shows the MIC and MBC of DX-Ag–AgCl NPs and DX, which was obtained against Gram-positive and Gram-negative pathogens. From these results, it is observed that the growth inhibition of Gram-positive (B. cereus KCCM 11773) bacteria inhibited at lowest concentration of DX-Ag–AgCl NPs (MBC, 1 µg mL−1), while Gram-negative (E. coli KCCM 11234) bacteria inhibited at higher concentration of DX-Ag–AgCl NPs (MBC, 12 µg mL−1). When the antibacterial effect of the DX-functionalized NPs is compared with that of DX alone, the amount of DX required is equal or considerably lower. These findings are in good agreement with the reported studies [16, 37, 38]. AgNPs are effective antibacterial agent, which cause adverse toxic effect on bacterial cells [8]. It is well known that the antibacterial activity of AgNPs is dependent on their size and shape; smaller AgNPs shows more bactericidal activity due to their more surface-to-volume ratio [11, 39].
Conclusions
The study of a simple method for the synthesis of DX-Ag–AgCl NPs using DX and TMG, which has an antibacterial activity against Gram-positive and Gram-negative pathogens, has been described here for the first time. Using the developed methodology, DX-Ag–AgCl NPs are synthesized rapidly, in 5 min. The synthesized NPs were spherical shaped with a size of 10–40 nm, and contained Ag and AgCl. Further studies are required for the synthesis of NPs with improved functions, and controlled shape and size. Applications of NPs based on these findings may lead to valuable discoveries in different fields such as medical devices and antibacterial systems.
References
Jain, P.K., Huang, H., El-Sayed, I.H., El-Sayed, M.A.: Noble on the nanoscale: optical and photothermal properties and some applications in imaging, sensing, biology, and medicine. Acc. Chem. Res. 41, 1578–1586 (2008)
El-Nour, K.M.M.A., Eftaiha, A., Al-Warthan, A., Ammar, R.A.A.: Synthesis and application of silver nanoparticles. Arab. J. Chem. 3, 135–140 (2010)
Kharissova, O.V., Dias, H.V.R., Kharisov, B.I., Perez, B.O., Perez, V.M.J.: The greener synthesis of nanoparticles. Trends Biotechnol. 31, 240–248 (2013)
Hussain, I., Singh, N.B., Singh, A., Singh, H., Singh, S.C.: Green synthesis of nanoparticles and its potential applications. Biotechnol. Lett. 38, 545–560 (2016)
Patil, M.P., Kim, G.-D.: Eco-friendly approach for nanoparticles synthesis and mechanism behind antibacterial activity of silver and anticancer activity of gold nanoparticles. Appl. Microbiol. Biotechnol. 101, 79–92 (2017)
Patil, M.P., Kim, G.-D.: Marine microorganisms for synthesis of metallic nanoparticles and their biomedical applications. Colloids Surf. B 172, 487–495 (2018)
Schrofel, A., Kratosova, G., Safarik, I., Safarikova, M., Raska, I., Shor, L.M.: Applications of biosynthesized metallic nanoparticles—a review. Acta Biomater. 10, 4023–4042 (2014)
Kim, J.S., Kuk, E., Yu, K.N., Kim, J.-H., Park, S.J., Lee, H.J., Kim, S.H., Park, Y.K., Park, Y.H., Hwang, C.-Y., Kim, Y.-K., Lee, Y.-S., Jeong, D.H., Cho, M.-H.: Antimicrobial effects of silver nanoparticles. Nanomedicine. 3, 95–101 (2007)
Azizi, M., Sedaghat, S., Tahvildari, K., Derakhshi, P., Ghaemi, A.: Synthesis of silver nanoparticles using Peganum harmala extract as a green route. Green Chem. Lett. Rev. 10, 420–427 (2017)
Pugazhendhi, A., Edison, T.N.J.I., Karuppusamy, I., Kathirvel, B.: Inorganic nanoparticles: a potential cancer therapy for human welfare. Int. J. Pharm. 539, 104–111 (2018)
Morones, J.R., Elechiguerra, J.L., Camacho, A., Holt, K., Kouri, J.B., Ramirez, J.T., Yacaman, M.J.: The bactericidal effect of silver nanoparticles. Nanotechnology. 16, 2346–2353 (2005)
Rai, M., Yadav, A., Gade, A.: Silver nanoparticles as a new generation of antimicrobials. Biotechnol. Adv. 27, 76–83 (2009)
Chaloupka, K., Malam, Y., Seifalian, A.M.: Nanosilver as a new generation of nanoproduct in biomedical applications. Trends Biotechnol. 28, 580–588 (2010)
Wei, L., Lu, J., Xu, H., Patel, A., Chen, Z.-S., Chen, G.: Silver nanoparticles: synthesis, properties, and therapeutic applications. Drug Discov. Today 20, 595–601 (2015)
Junejo, Y., Sirajuddin, S., Baykal, A., Safdar, M., Balouch, A.: A novel green synthesis and characterization of Ag NPs with its ultra-rapid catalytic reduction of methyl green dye. Appl. Surf. Sci. 290, 499–503 (2014)
Djafari, J., Marinho, C., Santos, T., Igrejas, G., Torres, C., Capelo, J.L., Poeta, P., Lodeiro, C., Fernandez-Lodeiro, J.: New synthesis of gold- and silver-based nano-tetracycline composites. ChemistryOpen. 5, 206–212 (2016)
Ullah, S., Ahmad, A., Subhan, F., Jan, A., Raza, M., Khan, A.U., Rahman, A.U., Khan, U.A., Tariq, M., Yuan, Q.: Tobramycin mediated silver nanospheres/graphene oxide composite for synergistic therapy of bacterial infection. J. Photochem. Photobiol. B 183, 342–348 (2018)
Junejo, Y., Guner, A., Baykal, A.: Synthesis and characterization of amoxicillin derived silver nanoparticles: its catalytic effect on degradation of some pharmaceutical antibiotics. Appl. Surf. Sci. 317, 914–922 (2014)
Levy, S.B., Marshall, B.: Antibacterial resistance worldwide: causes, challenges and responses. Nat. Med. 10, S122–S129 (2004)
Omidi, S., Sedaghat, S., Tahvildari, K., Derakhshi, P., Motiee, F.: Biosynthesis of silver nanoparticles with Adiantum capillus-veneris L leaf extract in the batch process and assessment of antibacterial activity. Green Chem. Lett. Rev. 11, 544–551 (2018)
Dhas, T.S., Kumar, V.G., Karthick, V., Angel, K.J., Govindaraju, K.: Facile synthesis of silver chloride nanoparticles using marine alga and its antibacterial efficacy. Spectrochim. Acta Part A 120, 416–420 (2014)
Eugenio, M., Campanati, L., Muller, N., Romao, L.F., Souza, J.D., Alves-Leon, S., Souza, W.D., Sant’Anna, C.: Silver/silver chloride nanoparticles inhibit the proliferation of human glioblastoma cells. Cytotechnology. 70, 1607–1618 (2018)
Patil, M.P., Seo, Y.B., Kim, G.-D.: Morphological changes of bacterial cells upon exposure of silver–silver chloride nanoparticles synthesized using Agrimonia pilosa. Microb. Pathog. 116, 84–90 (2018)
Carlson, C., Hussain, S.M., Schrand, A.M., Braydich-Stolle, L.K., Hess, K.L., Jones, R.L., Schlager, J.J.: Unique cellular interaction of silver nanoparticles: size-dependent generation of reactive oxygen species. J. Phys. Chem. 112, 13608–13619 (2008)
Park, H.-J., Kim, J.Y., Kim, J., Lee, J.-H., Hahn, J.-S., Gu, M.B., Yoon, J.: Silver–ion-mediated reactive oxygen species generation affecting bactericidal activity. Water Res. 43, 1027–1032 (2009)
Wigginton, N.S., Titta, A.D., Piccapietra, F., Dobias, J., Nesatyy, V.J., Suter, M.J.F., Bernier-Latmani, R.: Binding of silver nanoparticles to bacterial proteins depends on surface modifications and inhibits enzymatic activity. Environ. Sci. Technol. 44, 2163–2168 (2010)
Patil, M.P., Rokade, A.A., Ngabire, D., Kim, G.-D.: Green synthesis of silver nanoparticles using water extract from galls of Rhus chinensis and its antibacterial activity. J. Clust. Sci. 27, 1737–1750 (2016)
Patil, M.P., Palma, J., Simeon, N.C., Jin, X., Liu, X., Ngabire, D., Kim, N.-H., Tarte, N.H., Kim, G.-D.: Sasa borealis leaf extract-mediated green synthesis of silver–silver chloride nanoparticles and their antibacterial and anticancer activities. New J. Chem. 41, 1363–1371 (2017)
Mock, J.J., Barbic, M., Smith, D.R., Schultz, D.A., Schultz, S.: Shape effects in plasmon resonance of individual colloidal silver nanoparticles. J. Chem. Phys. 116, 6755–6759 (2002)
Noginov, M.A., Zhu, G., Bahoura, M., Adegoke, J., Small, C., Ritzo, B.A., Drachev, V.P., Shalaev, V.M.: The effect of gain and absorption on surface plasmons in metal nanoparticles. Appl. Phys. B 86, 455–460 (2007)
Rokade, A.A., Patil, M.P., Yoo, S.I., Lee, W.K., Park, S.S.: Pure green chemical approach for synthesis of Ag2O nanoparticles. Green Chem. Lett. Rev. 9, 216–222 (2016)
Bai, J., Li, Y., Li, M., Wang, S., Zhang, C., Yang, Q.: Electrospinning method for the preparation of silver chloride nanoparticles in PVP nanofiber. Appl. Surf. Sci. 254, 4520–4523 (2008)
Tamimi, F., Torres, J., Bettini, R., Ruggera, F., Rueda, C., Lopez-Ponce, M., Lopez-Cabarcos, E.: Doxycycline sustained release from brushite cements for the treatment of periodontal diseases. J. Biomed. Mater. Res. Part A 85, 707–714 (2008)
Gade, A., Gaikwad, S., Tiwari, V., Yadav, A., Ingle, A., Rai, M.: Biofabrication of silver nanoparticles by Opuntia ficus-indica: in vitro antibacterial activity and study of the mechanism involved in the synthesis. Curr. Nanosci. 6, 370–375 (2010)
Feng, Q.L., Wu, J., Chen, G.Q., Cui, F.Z., Kim, T.N., Kim, J.O.: A mechanistic study of the antibacterial effect of silver ions on Escherichia coli and Staphylococcus aureus. J. Biomed. Mater. Res. 52, 662–668 (2000)
Pal, S., Tak, Y.K., Song, J.M.: Does the antibacterial activity of silver nanoparticles depend on the shape of the nanoparticles? Appl. Environ. Microbiol. 73, 1712–1720 (2007)
Brown, A., Smith, K., Samuels, T.A., Lu, J., Obare, S., Scott, M.E.: Nanoparticles functionalized with ampicillin destroy multiple antibiotic resistant isolates of Pseudomonas aeruginosa, Enterobacter aerogenes and methicillin resistant Staphylococcus aureus. Appl. Environ. Microbiol. 78, 2768–2774 (2012)
Wang, L., Chen, Y.P., Miller, K.P., Cash, B.M., Jones, S., Glenn, S., Benicewicz, B.C., Decho, A.W.: Functionalised nanoparticles complexed with antibiotic efficiently kill MRSA and other bacteria. Chem. Commun. 50, 12030–12033 (2014)
Agnihotri, S., Mukherji, S., Mukherji, S.: Size-controlled silver nanoparticles synthesized over the range 5–100 nm using the same protocol and their antibacterial efficacy. RSC Adv. 4, 3974–3983 (2014)
Acknowledgements
This research was supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (2018R1D1A1B07043388) and also supported by the Research and Education Program at Korea Science Academy of KAIST with funds from the Ministry of Science, ICT and Future Planning (Year 2018).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Patil, M.P., Piad, L.L.A., Bayaraa, E. et al. Doxycycline hyclate mediated silver–silver chloride nanoparticles and their antibacterial activity. J Nanostruct Chem 9, 53–60 (2019). https://doi.org/10.1007/s40097-019-0297-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40097-019-0297-6