Abstract
Cardiovascular disease (CVD), the leading cause of mortality worldwide, results from a complex interplay between genetic and environmental factors. Genetic studies identified genetic variants providing insights into the pathogenesis and treatment of the disease. However, the mechanisms linking the genotypic and phenotypic expression remain to be elucidated. Gene–diet interaction studies attempt to elucidate how a modifiable factor interacts with the genetic background. The knowledge gained thus far confers to small increments of CVD risk and cannot explain the molecular mechanisms of the disease. Epigenetic studies attempt to elucidate the molecular pathways affected by an environmental stimulus, such as dietary exposure. The epigenomic changes and their link to gene–diet interactions remain a challenging area for research. Understanding the complex interplay among the epigenome, genome, and dietary exposure should lead to accurate prediction, prevention, or treatment of the disease.
Similar content being viewed by others
Introduction
Cardiovascular disease (CVD) is a class of disorders of the heart and blood vessels and includes coronary artery disease (CAD), cerebrovascular disease (stroke), peripheral artery disease, rheumatic heart disease, congenital heart disease, and heart failure. CVD is the leading cause of mortality worldwide, with more than 80 % of deaths taking place in low- and middle-income countries. In a recent report, the World Health Organization (WHO, 2011) estimated that more than 17 million people died from CVD in the year 2008, thus accounting for 30 % of all global deaths. By the year 2030, it is estimated that CVD will account for 23.6 million deaths worldwide [1••].
Τhe underlying cause of these manifestations is atherosclerosis. The atherosclerotic process accounts for many types of CVD, predominantly CAD and stroke, and is advanced by the time heart problems are detected. Genetic predisposition, behavioral, metabolic, and other risk factors are responsible for the onset of atherosclerosis, which finally leads to CVD (Table 1) [1••].
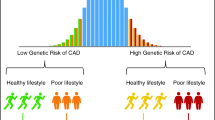
Among the established risk factors of CVD, there is strong scientific evidence that genetic background in response to dietary exposure plays a key role in the onset of the disease [2, 3]. Thus far, implemented guidelines and strategies in primary and secondary prevention of CVD mainly focus on a healthy lifestyle and endeavor to behavioral, social, and environmental changes. Nonetheless, further understanding of the pathogenesis and etiology of the disease requires research into the complex interaction between genetic and environmental factors.
Nutrigenetic and nutrigenomic research intends to explore the mechanisms of the interplay between diet and genetic variants in diet-related diseases (eg, CVD). This will lead to even more specialized guidelines according to individual genotypic architecture. The aim is to reduce the prevalence of chronic diseases, to prevent them, or even to cure them. For the purpose of the current review, outcomes regarding the effects of genetic variation on responsiveness to diet in CVD—specifically CAD—are examined.
Genetic Determinants for Cardiovascular Disease
Individuals differ in their susceptibility to CVD, and genetic variants play a key role in this. Indicatively, in a twin study, it has been shown that there is a 57 % and 38 % heritability for CAD in men and women, respectively [4]. In younger individuals, it has been demonstrated that genetic variants may contribute to a 20 % to 60 % increase in CAD risk [5]. Therefore, research to identify these variants is extended through the past two decades [6, 7].
The candidate gene approach is based on an a priori hypothesis of the plausible involvement of the selected gene in the pathogenesis and process of the disease under investigation. Many studies examined single nucleotide polymorphisms (SNPs) in single candidate genes, while others focused on more SNPs in the same gene. Historically, the success of the first candidate gene study in identifying a genetic variant in the susceptibility of CVD was published in Nature in 1992. Specifically, the study explored a variant found in the gene encoding angiotensin-converting enzyme (ACE) and showed that homozygotes for a deletion in the ACE gene were at higher myocardial infarction (MI) risk [7]. Another example reflecting the success of this approach is the identification of variants in the apolipoprotein E (APOE) gene, which has a major role in cholesterol metabolism [8]. On the other hand, animal and human studies have provided strong evidence regarding the pro- and antiatherogenic roles of the cholesterol ester transfer protein (CETP) gene by playing a key role in reverse cholesterol transport [9]. The breakthrough of the candidate gene studies was the knowledge obtained in the pathogenesis and the treatment of CVD through the identification of variants in genes associated with rare and monogenic forms of CVD—that is, mendelian CVD. However, candidate gene studies appeared unsuccessful in identifying the genetic background of polygenic CVD, including CAD, mainly due to limited tested polymorphisms and small sample sizes. Furthermore, a significant number of SNPs involved in disease pathways were left aside.
Genome-wide association studies (GWAS) enabled the scanning of the entire genome in order to seek out associations between hundreds of thousands of SNPs and diseases. The first GWAS on CVD identified some SNPs on 9p21.3 loci associated with CAD [10–13]. Reviewing the data up to 2011, 35 variants associated with CAD are reported [14••]. A meta-analysis performed by the CARDIoGRAM Consortium found that rs1333049 on the 9p21 region confers a 29 % increase in MI risk per allele [15••]. Schunkert et al. [16••] identified 13 new loci associated with CAD. Specifically, the risk alleles for the new loci were associated with an increase in CAD risk ranging from 6 % to 17 % per allele. The C4D Consortium identified five novel loci, including the lipase (LIPA) gene [17]. The CARDIoGRAMplusC4D Consortium raised the number of variants associated with CAD to 46 in the Caucasian population. In total, all the genetic variants identified thus far explain approximately 10 % of the cardiovascular risk in humans (unpublished data). Although the genome-wide approach led to the discovery of many SNPs associated with CVD, mechanisms linking the genotypic and phenotypic expression remain to be elucidated. More insights will be gained by the investigation of gene–environment interactions that will help us further understand the molecular pathways of the disease and explain the interindividual variability. Diet is one of the environmental factors that plays a major role in CVD.
Dietary Risk Factors for Cardiovascular Disease
Many researchers have tried to explore the relationship between dietary factors and CVD for almost half a century. The effects of many nutrients, foods, and dietary patterns on CVD have been evaluated by numerous studies. The importance of diet as a risk factor for CVD was first demonstrated by the Seven Countries Study [18–21]. Specifically, the Mediterranean-type diet was inversely associated with the incidence of CAD in Southern Europe when compared with the United States and Northern Europe, after adjusting for confounding factors.
The Seven Countries Study paved the way for other large-scale cohorts to follow, which examined the effects of dietary food groups or dietary patterns on CVD risk. Only a few of the major findings of some studies are highlighted in the next lines, as this is not the purpose of the current review. The CARDIO2000 study, a multicenter case-control study, showed that exclusive olive oil consumption was associated with a 0.55 times lower likelihood of having acute coronary syndrome (ACS) among hypercholesterolemic subjects [22]. Another case-control study, the INTERHEART study, demonstrated that a healthy dietary pattern characterized by increased consumption of fruits and vegetables has a protective role against MI [23]. The protective effect of vegetable intake in CAD has been also demonstrated by the Physicians’ Health Study in male subjects. Specifically, in this study, men who consumed at least 2.5 servings/d of vegetables had a relative risk (RR) of 0.77 (95 % CI, 0.60–0.98) for CAD compared with men with vegetable consumption of less than 1 serving/d [24]. The WHO (2009) attributes about 11 % of ischemic heart disease deaths to insufficient intake of fruits and vegetables [25].
Furthermore, prospective studies have examined the role of dietary patterns in relationship to CAD risk. Specifically, the Multi-Ethnic Study (MESA) demonstrated a negative association between the prevalence of CVD and the consumption of fruits, vegetables, whole grains, and low-fat content dairy after a 4-year follow-up. On the contrary, high consumption of red meat, fried food, and sweets increased the prevalence of CAD [26]. Along the same lines, the ATTICA study demonstrated a protective role of fruits, vegetables, and exclusive olive oil use against CAD. On the other hand, high consumption of red meat, margarine, sweets, cheese, and alcohol was positively associated with the prevalence of CΑD [27].
Regarding micronutrients, the consumption of foods rich in antioxidant compounds has been associated with decreased MI markers by exerting protective effects against low-density lipoprotein cholesterol (LDL-C) oxidation and endothelial vascular dysfunction. However, the benefit of their clinical use requires further investigation [28].
Gene–Diet Interactions in Cardiovascular Disease Risk
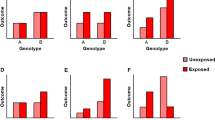
Gene–diet interaction studies are emerging and attempting to elucidate how a modifiable factor—that is, dietary exposure—interacts with genetic background. The new insights from gene–diet interaction studies promise to determine the individual risk of the disease, which will further lead to the implementation of personalized diets.
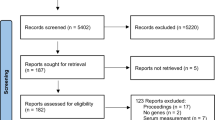
To date, the vast majority of studies that examined gene–diet interactions in CVD are referring mostly to cardiovascular risk factor outcomes. For the purpose of the current review, we searched PubMed, limited to studies on human subjects, with an emphasis on CAD and MI. A combination of keywords, namely genes, SNPs, polymorphisms, diet, dietary patterns, food groups, fat, cardiovascular disease, and coronary artery disease, was used. To provide updated knowledge, papers published during the past 2 years were used. During this period, the amount of information regarding gene–diet interactions in CVD in humans is rather limited. A summary of these studies is provided in Table 2.
A recent study on 9p21 loci, a chromosome region, extensively explored identified interactions between dietary patterns and rs2383206 on CVD risk. The results of the study showed that homozygous individuals (from the INTERHEART study) for the risk allele with a less prudent diet have a 1.6- to 2.0-fold increased risk of MI [29]. A prudent diet was a dietary pattern that consisted of raw vegetables, fruits, leafy green vegetables, nuts, desserts, and dairy products (food items that had factor loadings more than 0.25) [30]. A proxy SNP for rs2383206—that is, rs4977574—was also investigated for genetic association with CVD in a different sample (from the FINRISK study). It was found that the risk allele had an effect on the incidence of CVD among subjects with low and medium consumption of vegetables, fruits, and berries [29]. This result is similar to the one from the INTERHEART analysis. Furthermore, decreased CAD incidence has also been demonstrated in individuals with the BHMT minor allele in a case-control study [31•].
Regarding gene–diet interactions in clinical manifestations of the atherosclerotic process, there is only one study in humans examining the effect of the interaction between the 5-lipoxygenase promoter (ALOX5) genotype and dietary intake on carotid artery intima-media thickness. Although this study was not published within the last two years, we are referring to it as hallmark. The results of the study demonstrated that increased dietary intake of n-6 fatty acids, namely arachidonic and linoleic acid, was associated with increased intima-media thickness only among individuals with two variant alleles of the ALOX5 gene. In contrast, dietary intake of n-3 fatty acids was inversely associated with intima-media thickness only among individuals carrying two variant alleles. These results suggest for the first time that interactions between genes in inflammatory pathways and diet may lead to the onset of the atherosclerotic process in humans. However, these results need to be replicated [32].
Significant genotype–diet interactions have been also reported for the APOE gene. Frequent intake of olive oil and polyunsaturated fatty acids (PUFAs) was associated with an increase in triglyceride and LDL-C levels in E4 allele carriers. On the other hand, when analysis was performed only in male subjects, it was demonstrated that high olive oil intake was associated with low LDL-C levels in E2 carriers. Moreover, high intake of PUFAs was associated with decreased triglyceride levels in E2 male and female carriers [33]. Regarding saturated fatty acids (SFAs), individuals carrying the E4 allele had a greater CAD risk for SFA intake of more than 10 % of energy compared with individuals carrying the E2 allele [34]. In response to a high-fat diet, individuals carrying an allele for rs4246444 on the FASN gene had LDL-C with smaller peak particle diameter (LDL-PPD) compared with non-carriers [35•].
Regarding the CETP gene, a recent study demonstrated no interaction between CETP polymorphisms and dietary factors, namely alcohol, SFA, and adherence to the Mediterranean diet, in determining high-density lipoprotein cholesterol (HDL-C) levels [36]. A study on young Chinese subjects examined the interaction between lipoprotein lipase (LPL) gene variants and high-carbohydrate (HC)/low-fat diet [37•]. The polymorphisms examined modified the lipid traits of the young population in various ways (Table 1). What makes the study noteworthy is that the study population was made up of young individuals, in contrast to middle-aged or older individuals. Furthermore, the Chinese population is documented to have a diet with more carbohydrates and less fat, better lipid profiles, and lower risk of CAD incidence [37•, 38, 39]. Most studies tend to examine the effect of an HC diet on lipids, especially triglycerides, among middle-aged or older subjects.
In mammals, peroxisomal proliferator-activated receptor-α (PRAPα) and peroxisomal proliferator-activated receptor-γ (PRAPγ) are transcription factor that regulate the cardiac energy production from fatty streaks. PRAPα promotes fatty acid β-oxidation and PRAPγ promotes triglycerol synthesis [40]. A study on individuals at high cardiometabolic risk investigated the effect of interaction of PRAPα and PRAPγ genes with dietary fatty acids on lipid traits. It was shown that in habitual diet, the PUFA:SFA ratio influenced total cholesterol, LDL-C, and triglyceride levels in white individuals carrying the 12Ala allele. As the ratio increased, plasma lipids decreased [41].
GWAS have identified many genetic variants associated with CVD that provide insights into genetic background. However, the variants identified confer small increments of CVD risk and cannot explain interindividual susceptibility. Thus far, investigated gene–diet interactions cannot give plausible explanations regarding the molecular mechanisms of the disease, and we are far beyond CVD occurrence prediction or personalized diet implementation. Scientific evidence in identical twins demonstrated that the different susceptibility to polygenic disease, such as CVD, is attributable to alterations in the epigenome, followed by different environmental exposure [42, 43]. The question that arises is how environmental exposures, such as diet, could result in different individual susceptibility.
Epigenetics
Epigenetics are heritable changes in the genome that may lead to altered gene expression, but with no alterations in the nucleotide sequence. Histone modification, DNA methylation, and noncoding RNA functions are examples of the epigenetic process. DNA methylation is mostly explored in epigenetic studies. It refers to a biochemical procedure during which a methyl group is added to CpG dinucleotides, and it normally confers silence of gene expression. Epigenome is established during the development of the embryo and changes to regulation mechanisms of gene expression by an environmental stimulus conferring phenotype variability and different susceptibility to chronic diseases. Diet is one of the most important environmental factors that can induce epigenome perturbations. Nutrients such as folic acid, B vitamins, methionine, and choline are methyl-donating components. Figure 1 shows a model presenting the genetic variants, dietary exposure, and epigenome interactions.
Influence of Diet on Epigenome in Cardiovascular Disease
Studies on animal models and humans showed that exposure to imbalanced nutrient supply in fetal life, deficient growth in gestational periods, or during infancy is associated with higher CVD mortality [43, 44•]. Nutrient excess or deficiency results in epigenetic alterations that are carried throughout the life span, are inherited, and may play a major role in phenotype diversity. The Dutch famine (1944–1945) demonstrated strong evidence that maternal nutrient deficiency was associated with epigenetic alterations and increased risk of chronic diseases, including CVD, later in the life of the fetus [45, 46].
A recent study in mice revealed that female offsprings (APOE+/-) were more susceptible to neointima formation when exposed to maternal hypercholesterolemia (APOE-/-) compared with identical offsprings from normocholesterolemic (wild-type) mothers. Furthermore, diet-induced hypercholesterolemia during the postnatal period further enhanced the susceptibility to atherosclerosis in offspring from APOE-/- mothers compared with those from wild-type mothers. The results of the study demonstrate that fetal programming and dietary exposure influence histone methylation and affect the epigenome [47]. A previous study has similarly shown that APOE-/- mice with athero-susceptibility present DNA methylation changes that promote atherosclerosis [48].
Other studies in mice have investigated whether diet can have epigenetic effects on PRAPα and PRAPγ regulation. PRAPα-altered regulation is associated with diabetic cardiomyopathy in animal models [49]. Specifically, in mice, PRAPα methylation was lower in offspring exposed to protein-restricted maternal diet compared with offspring exposed to protein-sufficient maternal diet. Changes in PRAPα methylation exert their effect in mRNA expression only later in life. The results of the study suggest that maternal deficiency leads to alterations in epigenomic background, resulting in health consequences later in life [50].
The exact molecular pathways that are affected by epigenetic changes need further investigation. More studies need to be employed regarding the influence of dietary exposure to the epigenome in the context of CVD. Therefore, understanding epigenetic disruptions and the relationship between diet and epigenome is undoubtedly one of the most important challenges in recent research.
Conclusions
CVD is a multifactorial, complex disease that involves interaction of genetic and environmental factors. The International Hap Map project enabled GWAS to identify novel genetic variants that contribute to the disease manifestations [51]. Although GWAS offer a powerful contribution to identifying common alleles in disease genes, the SNPs identified thus far provide small increments of disease risk and cannot provide predictive accuracy. In further elucidating the pathways resulting in the disease, the genetics of CVD is explored in relation to environmental factors. Gene–diet interaction studies remain limited to the literature and are still in progress, yet they promise to unravel some of the underlying mechanisms of the disease and determine the individual risk. Studies on epigenome modifications seem to play a key role in understanding diversities in individual susceptibility. DNA methylation and histone modification in response to dietary exposure may help us elucidate the interplay between and diet and disease and understand how this interaction leads to increased CVD risk. Epigenetic disruptions are potentially reversible; thus, the development of epigenomic drugs and functional foods or supplements that can regulate the epigenome will be an upcoming challenge for the drug and food research industry.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
•• World Health Organisation: Global Atlas on cardiovascular disease prevention and control. Available at: http://www.who.int/cardiovascular_diseases/publications/atlas_cvd/en/index/html. Accessed September 2011. This report presents the most current trends in CVD.
Lovegrove JA, Gitau R. Nutrigenetics and CVD: what does the future hold? Proc Nutr Soc. 2008;67:206–13.
Ordovas JM. Nutrigenetics, plasma lipids, and cardiovascular risk. J Am Diet Assoc. 2006;106:1074–81.
Zdravkovic S, Wienke A, Pedersen NL, et al. Heritability of death from coronary heart disease: a 36-year follow-up of 20 966 Swedish twins. J Intern Med. 2002;252:247–54.
Chaer RA, Billeh R, Massad MG. Genetics and gene manipulation therapy of premature coronary artery disease. Cardiology. 2004;101:122–30.
Lee DS, Pencina MJ, Benjamin EJ, et al. Association of parental heart failure with risk of heart failure in offspring. N Engl J Med. 2006;355:138–47.
Cambien F, Poirier O, Lecerf L, et al. Deletion polymorphism in the gene for angiotensin-converting enzyme is a potent risk factor for myocardial infarction. Nature. 1992;359:641–4.
Song Y, Stampfer MJ, Liu S. Meta-analysis: apolipoprotein E genotypes and risk for coronary heart disease. Ann Intern Med. 2004;141:137–47.
Vourvouhaki E, Dedoussis GV. Cholesterol ester transfer protein: a therapeutic target in atherosclerosis? Expert Opin Ther Targets. 2008;12:937–48.
McPherson R, Pertsemlidis A, Kavaslar N, et al. A common allele on chromosome 9 associated with coronary heart disease. Science. 2007;316:1488–91.
Helgadottir A, Thorleifsson G, Manolescu A, et al. A common variant on chromosome 9p21 affects the risk of myocardial infarction. Science. 2007;316:1491–3.
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature. 2007;447:661–78.
Samani NJ, Erdmann J, Hall AS, Hengstenberg C, et al. WTCCC and the Cardiogenics Consortium. Genomewide association analysis of coronary artery disease. N Engl J Med. 2007;357:443–53.
•• Peden JF, Farrall M. Thirty-five common variants for coronary artery disease: the fruits of much collaborative labour. Hum Mol Genet. 2011;20:198–205. This article gives a summary of GWAS and large-scale genecentric genotyping studies that have reported genetic variants associated with CAD.
•• Preuss M, König IR, Thompson JR, et al. CARDIoGRAM Consortium. Design of the Coronary ARtery DIsease Genome-Wide Replication And Meta-Analysis (CARDIoGRAM) Study: A Genome-wide association meta-analysis involving more than 22 000 cases and 60 000 controls. Circ Cardiovasc Genet. 2010;3:475–83. This meta-analysis provides strong evidence regarding the role of 9p21 chromosome in CVD risk.
•• Schunkert H, Konig IR, Kathiresan S, et al. Large-scale association analysis identifies 13 new susceptibility loci for coronary artery disease. Nat Genet. 2011;43:333–8. This analysis is important mainly to the number of new loci identified in relation to CAD.
Kathiresan S, Voight BF, Purcell S, et al. Genome-wide association of early-onset myocardial infarction with single nucleotide polymorphisms and copy number variants. Nat Genet. 2009;41:334–41.
Keys A, Aravanis C, Blackburn HW, et al. Epidemiological studies related to coronary heart disease: characteristics of men aged 40-59 in seven countries. Acta Med Scand Suppl. 1966;460:1–392.
Keys A (Ed). Coronary heart disease in seven countries. Circulation. 1970, 41:1–200.
Keys A, Menotti A, Aravanis C, et al. The seven countries study: 2,289 deaths in 15 years. Prev Med. 1984;13:141–54.
Menotti A, Keys A, Aravanis C, et al. Seven Countries Study. First 20-year mortality data in 12 cohorts of six countries. Ann Med. 1989;21:175–9.
Kontogianni MD, Panagiotakos DB, Chrysohoou C, et al. The impact of olive oil consumption pattern on the risk of acute coronary syndromes: The CARDIO2000 case-control study. Clin Cardiol. 2007;30:125–9.
Iqbal R, Anand S, Ounpuu S. Islam Set al. INTERHEART Study Investigators. Dietary patterns and the risk of acute myocardial infarction in 52 countries: results of the INTERHEART study. Circulation. 2008;118:1929–37.
Liu S, Lee IM, Ajani U, et al. Intake of vegetables rich in carotenoids and risk of coronary heart disease in men: The Physicians’ Health Study. Int J Epidemiol. 2001;30:130–5.
World Health Organisation: Global health risks 2009. Available at http://www.who.int/healthinfo/global_burden_disease/global_health_risks/en/index.html, Accessed December 2009.
Nettleton JA, Polak JF, Tracy R, et al. Dietary patterns and incident cardiovascular disease in the Multi-Ethnic Study of Atherosclerosis. Am J Clin Nutr. 2009;90:647–54.
Panagiotakos DB, Pitsavos C, Chrysohoou C, et al. ATTICA Study. Five-year incidence of cardiovascular disease and its predictors in Greece: the ATTICA study. Vasc Med. 2008;13:113–21.
Kaliora AC, Dedoussis GV. Natural antioxidant compounds in risk factors for CVD. Pharmacol Res. 2007;56:99–109.
Do R, Xie C, Zhang X, et al. The effect of chromosome 9p21 variants on cardiovascular disease may be modified by dietary intake: evidence from a case/control and a prospective study. PLoS Med. 2011;8:e1001106.
Iqbal R, Anand S, Ounpuu S, et al. Dietary patterns and the risk of acute myocardial infarction in 52 countries: results of the INTERHEART study. Circulation. 2008;118:1929–37.
• Singh PR, Lele SS, Mukherjee MS. Gene polymorphisms and low dietary intake of micronutrients in coronary artery disease. J Nutrigenet Nutrigenomics. 2011;4:203–9. This is a recent gene–diet interaction study regarding micronutrients and SNP interactions.
Dwyer JH, Allayee H, Dwyer KM, Fan J, Wu H, Mar R, Lusis AJ, Mehrabian M. Arachidonate 5-lipoxygenase promoter genotype, dietary arachidonic acid, and atherosclerosis. N Engl J Med. 2004;350:29–37.
de Andrade FM, Bulhões AC, Maluf SW, et al. The influence of nutrigenetics on the lipid profile: interaction between genes and dietary habits. Biochem Genet. 2010;48:342–55.
Corella D, Portolés O, Arriola L, Chirlaque MD, et al. Saturated fat intake and alcohol consumption modulate the association between the APOE polymorphism and risk of future coronary heart disease: a nested case-control study in the Spanish EPIC cohort. J Nutr Biochem. 2011;22:487–94.
• Dolley G, Boisclair ME, Lamarche B, Després JP, Bouchard C, Pérusse L, Vohl MC. Interactions between dietary fat intake and FASN genetic variation influence LDL peak particle diameter. J Nutrigenet Nutrigenomics. 2011;4:137–45. Little is known regarding the effect of diet on LDL-PPD. Most studies evaluate the influence on LDL values.
Corella D, Carrasco P, Fitó M, et al. Gene-environment interactions of CETP gene variation in a high cardiovascular risk Mediterranean population. J Lipid Res. 2010;51:2798–807.
• Huang X, Gong R, Lin J, et al. Effects of lipoprotein lipase gene variations, a high-carbohydrate low-fat diet, and gender on serum lipid profiles in healthy Chinese Han youth. Biosci Trends. 2011;5(5):198–204. Not many studies have been conducted on young populations with a low CVD incidence.
Saha N, Heng CK, Mozoomdar BP, et al. Racial variation of factor VII activity and antigen levels and their correlates in healthy Chinese and Indians at low and high risk for coronary artery disease. Atherosclerosis. 1995;117:33–42.
Lee MM, Wu-Williams A, Whittemore AS, et al. Comparison of dietary habits, physical activity and body size among Chinese in North America and China. Int J Epidemiol. 1994;23:984–90.
Bishop-Bailey D. Peroxisome proliferator-activated receptors in the cardiovascular system. Br J Pharmacol. 2000;129:823–34.
AlSaleh A, Sanders TA, O’Dell SD. Effect of interaction between PPARG, PPARA and ADIPOQ gene variants and dietary fatty acids on plasma lipid profile and adiponectin concentration in a large intervention study. Proc Nutr Soc. 2012;71(1):141–53.
Manolio TA, Collins FS, Cox NJ, et al. Finding the missing heritability of complex diseases. Nature. 2009;461:747–53.
Barker DJ, Martyn CN. The maternal and fetal origins of cardiovascular disease. J Epidemiol Community Health. 1992;46:8–11.
• Jackson AA, Burdge GC, Lillicrop KA. Diet, nutrition and modulation of genomic expression in fetal origins of adult disease. J Nutrigenet Nutrigenomics. 2010;3:192–208. This is an interesting review discussing how dietary exposure in early development affects later adult life through epigenome alterations.
Painter RC, de Rooij SR, Bossuyt PM, et al. Blood pressure response to psychological stressors in adults after prenatal exposure to the Dutch famine. J Hypertens. 2006;24:1771–8.
Heijmans BT, Tobi EW, Stein AD, et al. Persistent epigenetic differences associated with prenatal exposure to famine in humans. Proc Natl Acad Sci USA. 2008;105:17046–9.
Alkemade FE, van Vliet P, Henneman P, et al. Prenatal exposure to apoE deficiency and postnatal hypercholesterolemia are associated with altered cell-specific lysine methyltransferase and histone methylation patterns in the vasculature. Am J Pathol. 2010;176:542–8.
Lund G, Andersson L, Lauria M, et al. DNA methylation polymorphisms precede any histological sign of atherosclerosis in mice lacking apolipoprotein E. J Biol Chem. 2004;279:29147–54.
Finck BN, Han X, Courtois M, et al. A critical role for PPARalpha-mediated lipotoxicity in the pathogenesis of diabetic cardiomyopathy: modulation by dietary fat content. Proc Natl Acad Sci U S A. 2003;100:1226–31.
Slater-Jefferies JL, Lillycrop KA, Townsend PA, et al. Feeding a protein-restricted diet during pregnancy induces altered epigenetic regulation of peroxisomal proliferator-activated receptor-α in the heart of the offspring. J Dev Orig Health Dis. 2011;2:250–5.
International HapMap Consortium. The International HapMap Project. Nature. 2003;18(426):789–96.
Disclosure
No potential conflicts of interest relevant to this article were reported.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dimitriou, M.E., Dedoussis, G.V.Z. Gene–Diet Interactions in Cardiovascular Disease. Curr Nutr Rep 1, 153–160 (2012). https://doi.org/10.1007/s13668-012-0020-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13668-012-0020-4