Abstract
Bombus terrestris is a bumblebee with a wide geographic range, with subspecies showing a variety of local adaptations. Global export of commercially-reared B. terrestris started in the 1980s; the bees are a mixture of subspecies bred for ease of rearing, bivoltinism and large nests. This paper investigated whether the increase in bivoltinism in UK resident B. terrestris audax populations was related to introgression with imported foreign subspecies. Workers were collected from wild populations in London and Bristol, as well as two commercial suppliers. Fourteen microsatellite loci were used to study population structure, hybridisation and introgression. No introgression with commercial B. t. dalmatinus was detected in wild populations. Hence, the increase in winter activity appears unrelated to introgression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
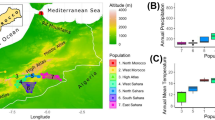
Bombus terrestris (Linnaeus 1758) is a large, highly abundant bumblebee with a wide distribution across Europe (Figure 1), where it can be found in a range of habitats, excluding arctic, alpine and desert regions (Rasmont et al. 2008). Nine distinct subspecies have been described, with physiological and behavioural adaptations appropriate to their local environment, such as differences in colour pattern, pheromones and aggression (Rasmont et al. 2008; Coppée 2010). Despite such variation, the vast majority of B. terrestris populations are univoltine, with queens emerging from diapause in the spring, producing one colony over the summer, and with new queens initiating diapause at the end of summer (Løken 1973; Rasmont et al. 2008). A few exceptions do exist; B. terrestris exhibits facultative bivoltinism in areas where climatic conditions can support colonies over winter, such as the Mediterranean Basin (Beekman et al. 1999; Rasmont et al. 2008). In warmer Mediterranean regions, B. terrestris queens aestivate in response to the summer heat (Gurel et al. 2008).
B. terrestris subspecies distribution across Europe, adapted from data in Rasmont et al. (2008) and Lecocq et al. (2016). Hybrid zones are indicated by striped regions. Note that one subspecies, B. t. canariensis, is not indicated on the map, but can be found solely in the Canary Islands. The blank map of Europe was created by Lvcvlvs and edited under the Creative Commons BY-SA 3.0 license.
Chosen for their ability to pollinate a wide range of plants and to thrive in a variety of environmental conditions, B. terrestris has been bred commercially since the 1980s for agricultural pollination services (Velthuis and Van Doorn 2006; Rasmont et al. 2008). Through a behaviour known as buzz pollination, bumblebees are capable of pollinating several crops (e.g. aubergine, tomato and blueberry) which cannot be effectively pollinated by honeybees, and are thus a cheaper alternative to mechanical pollination in which humans use mechanical, hand-held devices to manually pollinate such crops (Velthuis and Van Doorn 2006). Commercial breeding practices initially relied on the collection of tens of thousands of field-caught queens (Velthuis and Van Doorn 2006). Although fewer field-caught queens are caught today, the numbers are still estimated to be in the hundreds (Velthuis and Van Doorn 2006). Initial stocks are thought to have been built from a mixture of central European B. t. terrestris, eastern European B. t. dalmatinus, and Iberian B. t. lusitanicus. Over time, B. t. dalmatinus has become the market favourite subspecies due to its ease of rearing and development of large colonies (Velthuis and Van Doorn 2006). As well as colony size and domesticity, commercial populations were also bred to discourage diapause, leading to colony initiation immediately after mating, which has a known genetic component (Beekman et al. 1999; Velthuis and Van Doorn 2006). These mixed, domesticated colonies have been distributed worldwide, often with no regard for the impact on local, native bumblebee species. For example, escaped bumblebees have established feral, invasive B. terrestris populations in New Zealand, Chile and Japan (Donovan and Wier 1978; Ruz and Herrera 2001; Inoue et al. 2008; Rasmont et al. 2008; Aizen et al. 2018). Colony import has also been linked to the introduction and spread of parasites to wild bee populations (Meeus et al. 2011; Graystock et al. 2013; Graystock et al. 2016; Chandler et al. 2019). In the UK, regulations controlling B. terrestris subspecies import were only implemented in 2015, but non-native subspecies may still be imported with a licence (England. Natural England 2015). These domesticated bumblebee species could cause negative effects on local species and subspecies within their native range such as spreading disease, disrupting local genetic adaptation or outcompeting local populations (Ings et al. 2006; Lecocq et al. 2016; Chandler et al. 2019).
Bombus terrestris audax is the resident subspecies of the British Isles, and like most B. terrestris populations, has historically been considered univoltine (Alford 1969). However, records of winter active colonies can be found as early as 1991, when queens would typically undergo diapause, especially in the South and Midlands of England (Robertson 1991; Stelzer et al. 2010). Reports of winter active bumblebees have been increasing in frequency, with a number of proposed mechanisms (Stelzer et al. 2010). Given the dependence of diapause induction and termination on temperature cues, the warmer winters associated with climate change (and an abundance of introduced winter-flowering plants in urban areas) are thought to be facilitating bivoltinism (Alford 1969; Stelzer et al. 2010). Owen et al. (2013) suggested that imported B. t. dalmatinus, which are more fecund and more inclined to bivoltinism, may be displacing native B. t. audax (Ings et al. 2006). Additionally, Kraus et al. (2011) found that in Poland, commercial colonies frequently escape containment and interbreed with local populations. In Spain and Portugal, Seabra et al. (2018) and Cejas et al. (2020) both found evidence of commercial B. terrestris introgression with wild populations of local subspecies B. t. lusitanicus. Assuming the same is true for the UK, introduction of more bivoltine-inclined bumblebees may encourage winter activity in hybrid populations.
In this paper, we aim to investigate whether hybridisation and consequent introgression with imported, non-native subspecies is underpinning the recent trend towards bivoltinism in British B. terrestris, by using microsatellite analysis to detect differences and similarities between wild winter and summer populations, and commercial, imported bumblebees. To do so, wild B. t. audax workers were collected from London and Bristol across consecutive summers and winters (2009-2011) and were genotyped with 14 microsatellite loci. Genetic profiles of wild bumblebees were compared with mass-reared workers of B. t. audax and B. t. dalmatinus.
Several different scenarios may be detectable through this method. If introgression is absent, bivoltinism will presumably be the consequence of behavioural plasticity. If this is true, then the summer and winter populations would look genetically similar, wild populations would be strongly differentiated to commercial populations, and no signs of introgression would be present. If introgression is present but not linked to winter activity, then summer and winter populations would not be distinct, and both should show medium to high similarity to commercial stocks. Finally, if winter activity is induced by introgression, then we would expect to see one bivoltine population present in both summer and winter, and a distinct, un-introgressed summer population, with the winter population showing higher similarity to commercial stocks.
2 Materials and methods
2.1 Specimen collection
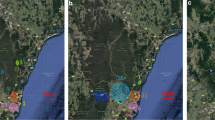
Between 2009 and 2011, 317 presumably wild B. terrestris audax workers were collected during summer and winter from London and Bristol areas (Figure 2). Other B. terrestris subspecies are not expected to be present at these urban locations, as they are associated with agricultural use of colonies (Kraus et al. 2011; Moreira et al. 2015; Bartomeus et al. 2020). Individuals were later genotyped.
Samples were frozen whole and stored in individual vials at − 20 °C. Two specimens were identified as B. lucorum by RFLP analysis, as workers of these two species cannot be easily distinguished (Murray et al. 2008), and excluded from further analyses. Thus, 315 wild B. t. audax workers remained. Thirteen workers were sampled in Bristol in the winter of 2009; 15 workers were collected from London in the winter of 2009; 136 from the summer of 2010; 39 from the winter of 2010, and 112 from the summer of 2011 (Supplementary Table 1). In addition to the wild bees, individual workers were also sampled from different commercially-reared colonies: 29 B. t. dalmatinus from Syngenta (Weert, the Netherlands) and 14 B. t. audax from Agralan Ltd. (Swindon, UK). OpenStreetMap data were used to indicate sampling site locations (OpenStreetMap 2017).
2.2 DNA extraction and microsatellite protocol
DNA extractions were performed using one hind leg per sample, using the Promega Wizard SV 96 Genomic DNA Purification System, and automated using a Biomek Liquid Handling Platform. Quality control was performed using a Nanodrop spectrophotometer.
Fourteen microsatellite loci were selected from published B. terrestris microsatellite lists (Estoup et al. 1996; Funk et al. 2006; Stolle et al. 2009): BT06, BT09, BL13, BTMS0044, BT119, BT20, BTMS0114, BT118, BT116, BTMS0067, BT100, BT132, BTMS0106 and BT30. A Qiagen Type-It kit was used to amplify samples and optimise amplification (Supplementary Table 2). Markers were organised into 3 panels of 4–5 markers according to size, fluorescent labels and optimisation results (Supplementary Table 2). Samples were diluted 1:100 using the Biomek platform before having ROX 400 and HiDi size standards added, after which the samples were denatured at 95 °C for 5 min for fragment size analysis on the ABI 3730xl platform. All samples were manually scored with PeakScanner v2.0 (Applied Biosystems).
2.3 Data validation
Locus BTMS0106 was found to be monomorphic and excluded from further analysis. Individuals with data missing at three or more loci were also excluded. Colony version 2.0.6.5 (Wang 2004) was used to detect siblings via the combined full likelihood and pairwise likelihood score (FPLS) algorithm. Upon full sibling detection, the individual with the most missing data was removed; if equal, one was randomly chosen. After validation, 248 individuals with data for 13 loci remained (Supplementary Table 3).
The presence of null alleles was determined using Microchecker version 2.2.3 (Van Oosterhout et al. 2004) using the Bonferroni 1000× analysis option. GenAlEx 6.503 (Peakall and Smouse 2006) was used to check for deviations from the Hardy-Weinberg equilibrium, after which Bonferroni correction was manually applied. FSTAT 2.9.4 (Goudet 1995) was used to check for linkage disequilibrium between loci, which also underwent Bonferroni correction. Reported p values are post-Bonferroni correction.
2.4 Genetic diversity and population structure
GenAlEx 6.503 was also used to calculate expected and observed heterozygosity (HE and HO), as well as allelic differentiation measure DEST, and fixation measures FST and GST (Jost 2008). Rarification and calculation of allelic richness (normalised to 10 genes) was performed with HP-Rare version June-6-2006 (Kalinowski 2004). Averages of genetic diversity (but not allelic differentiation or fixation) were weighted according to the number of individuals in each population.
BAPS 5.4 (Corander et al. 2003) was used to indirectly infer the degree of introgression using the ‘population mixture analysis trained clustering’ method. Using the two commercial populations as clusters of known origin, the algorithm assigns individuals of unknown origin (here, the wild samples) into clusters. Wild bees with a high degree of similarity to the commercial populations will be assigned in the same cluster. Fifty replicates of this algorithm were used to generate a mean and standard deviation to infer introgression, a method previously used for detecting introgression in B. terrestris by Kraus et al. 2011.
STRUCTURE 2.3.4 (Pritchard et al. 2000) was used to find evidence of population structure by geography, season and origin (i.e. wild or commercial). K was run from 1 to 10, with 3 iterations for each with 250,000 MCMC burn-in reps and 750,000 after burn-in. STRUCTURE harvester 0.6.94 (Earl and vonHoldt 2012) was used to find the most likely value of K. CLUMPP mac version 1.1.2 (Jakobsson and Rosenberg 2007), and DISTRUCT version 1.1 (Rosenberg 2004) were used to visualise STRUCTURE results.
3 Results
3.1 Data validation
Twenty-four individuals were removed from the dataset due to excessive missing data (Supplementary Table 3). Colony detected the presence of multiple full sibling pairs, leading to the exclusion of 12 bees from further analysis (Supplementary Table 3). In some cases, three siblings were collected from the same colony, thus two were removed. Microchecker detected three null alleles at allele frequencies above 10%, leading to the exclusion of three further loci (BT119, BTMS0067, BT132). FSTAT detected no significant (p = 0.0014) disequilibrium between loci over 78 pairwise comparisons when using Bonferroni correction for multiple comparisons. Five significant (p = 0.0007) deviations from the Hardy-Weinberg equilibrium were detected after adjusting for multiple comparisons, four in the summer 2010 population (BT06, BT09, BT20 and BTMS0014) and one in the summer 2011 population (BT30).
3.2 Genetic diversity
Overall, wild populations showed a marginally higher average allelic richness (AR) and observed heterozygosity (HO), but lower private allelic richness, than the commercial populations (Table I). Average AR for the commercial populations was 4.34, but 4.64 for wild populations. For private allelic richness, wild populations averaged 0.40, while commercial populations averaged 0.45. HO was very similar for the wild (0.680 ± 0.070) and commercial (0.660 ± 0.080) populations. Expected heterozygosity was also very similar for both wild (0.700 ± 0.130) and commercial (0.650 ± 0.080) populations.
3.3 Population differentiation by origin
Three different metrics of population differentiation were tested, namely fixation indexes FST and GST, and the allelic differentiation measure DEST, comparing the commercial B. t. audax, commercial B. t. dalmatinus and wild B. t. audax populations (Table II). Pairwise comparisons between the commercial and wild B. terrestris populations resulted in the wild population being most similar to the commercial B. t. audax by GST (0.006 versus 0.014 for B. t. dalmatinus, p = 0.041 and 0.001, respectively), DEST (0.027 versus 0.070 for B. t. dalmatinus, p = 0.041 and 0.001), and FST (0.019 versus 0.021 for B. t. dalmatinus, p = 0.044 and 0.001). Not surprisingly, the strongest differentiation was observed between the two commercial populations, with an FST of 0.037, a GST of 0.018 and a DEST of 0.089 (p = 0.002 for each).
STRUCTURE and STRUCTURE harvester were also used to study and visualise population differences (Figure 3). STRUCTURE harvester suggested that the most likely number of populations (K) when separated by origin was 4. However, this result is probably a consequence of a limitation of the program, which cannot detect the value of K if it is equal to 1 (Earl and vonHoldt 2012). Visualisation of the structure results indicates the approximately equal contribution of each population (K) in Figure 3, implying that the true value of K is indeed 1, with no structuring over origin.
Population structure results obtained in STRUCTURE and visualised using CLUMPP and DISTRUCT. Each plot shows the proportion of the genome, for each individual, likely belonging to each cluster (K). When data was analysed by season, the results of K = 2 are seen in A, and K = 5 in B. When analysed by origin, K = 2 in C, and K = 4 in D. Labels of ‘Winter’ or ‘Summer’ are referring to wild populations.
3.4 Population differentiation by season
FST, GST, and DEST were also tested between commercial populations and wild populations sorted by season (Table III). FST-values between seasons were very low, with an average 0.010, with varying levels of significance (Table III). Although the average genetic difference between wild B. terrestris grouped by season and both commercial populations was slightly higher (average FST of 0.030), these pairwise measurements showed a low but significant genetic differentiation (p < 0.05; Table III). DEST and GST were in agreement with the observed pattern of FST; seasons were not differentiated, and medium (DEST) or low (GST) but significant differentiation was present between season and both commercial B. t. audax and B. t. dalmatinus populations.
Here, STRUCTURE harvester suggested that the most likely number of populations (K) when separated by season was 5 (Figure 3). However, the structure results indicated, by the approximately equal contribution of each population in all bees (see Figure 3), that K is 1. Similar to the results of separation by origin, no structuring was thus observed for season by STRUCTURE.
3.5 Introgression
BAPS was used to infer the presence of introgression by attempting to assign the 216 wild individuals to commercial B. t. dalmatinus, commercial B. t. audax or their own cluster. After 50 repeats, BAPS consistently assigned all wild individuals to the same cluster as the commercial B. t. audax, with identical results for all 50 repeats.
4 Discussion
4.1 Seasonal population structure
Despite the known genetic component of bivoltinism (Beekman et al. 1999), no population structure was apparent between different seasons of wild bees, implying there is little genetic difference between summer and winter populations (if any), in contrast to the significant, low or medium differentiation between seasons and commercial populations. These results, including the STRUCTURE results, are in agreement with Moreira et al. (2015), who found that wild B. t. audax across the island of Great Britain were one population. Furthermore, the BAPS analysis did not assign wild B. t. audax of any season to the commercial B. t. dalmatinus cluster, which excludes the scenario in which winter activity is due to introgression, as the winter population showed no higher similarity to commercial stocks than the summer populations.
4.2 Introgression
The BAPS results showed no evidence of hybridisation or introgression between the commercial B. t. dalmatinus and wild B. t. audax. However, this does not necessarily dictate that introgression is not occurring between these two populations, as introgression with imported commercial bumblebees and local subspecies appears widespread across other parts of Europe (Kraus et al. 2011; Seabra et al. 2018; Bartomeus et al. 2020; Cejas et al. 2020). This may be a consequence of the limitations of microsatellites, as discussed below.
4.3 Explaining winter activity
In general, our results support the scenario in which winter activity is not due to a genetic similarity to a bivoltinism-inclined commercial population. Indeed, given that the winter bees are not a subset of summer bees, or a consequence of introgression, it seems likely that the winter active populations are the same wild populations exhibiting behavioural plasticity, taking advantage of the warmer weather and abundant urban winter resources (Stelzer et al. 2010). This is supported by the incidence of winter activity occurring mostly in the South of England and climate data showing that the southern regions are also more intensely experiencing climate change, with a severe North-South gradient in the temperature of the hottest days of summer and winter (Stelzer et al. 2010; Stainforth et al. 2013).
However, it could also be the case that the wild and commercial populations are extremely highly introgressed, and have entirely replaced the native wild B. t. audax populations. STRUCTURE and STRUCTURE harvester indicated that the wild and both commercial populations were actually one population, due to high genetic similarity. If extreme introgression was actually the case, we would not be able to tell from our data whether the presence of introgression was related to winter activity, as all populations were bivoltine.
4.4 Comparison of commercial and wild populations
Given that domesticated populations are known to be under strong selective pressure for commercially-valuable traits, the level of differentiation between commercial and wild populations is surprisingly low (Velthuis and Van Doorn 2006; Abadía-Cardoso et al. 2016). Levels of observed and expected heterozygosity were similar to those seen in both British and continental European populations of B. terrestris (Estoup et al. 1996; Moreira et al. 2015), but our own data showed lower allelic richness.
Although the commercial B. t. dalmatinus population showed higher genetic diversity than the commercial B. t. audax, and similar levels to wild audax populations, the level of differentiation was unexpectedly similar to the wild audax population, showing only marginal difference in FST and GST to the values of the commercial audax (Tables I; II). This may be attributable to a number of different causes; a 25-year period of importing B. t. dalmatinus into the UK for commercial reasons is likely to have established some level of introgression, a pattern seen elsewhere in Europe (Kraus et al. 2011; Seabra et al. 2018; Bartomeus et al. 2020; Cejas et al. 2020). The 1980s practices of crossing different subspecies may mean that commercial stocks are hybridised, and even populations labelled as B. t. dalmatinus may contain genetic material from other subspecies such as B. t. audax, contributing to the lack of differentiation (Velthuis and Van Doorn 2006). B. t. audax was also domesticated much more recently than B. t. dalmatinus, and thus the commercial stock has had a shorter time period to diverge from its parent population (Velthuis and Van Doorn 2006). Finally, the intermittent addition of field-caught B. t. audax queens, which may be hybrid, to commercial B. t. audax stocks could also have potentially contributed to the lack of differentiation between commercial B. t. dalmatinus and wild population (Velthuis and Van Doorn 2006).
4.5 Alternative approaches to microsatellites
Microsatellites are a useful and dependable tool for population genetics studies, with the ability to reliably resolve population structure, identify inbreeding and genetic diversity parameters (Maebe et al. 2016, 2018, 2019a, b) and have previously been used to find evidence of introgression in B. terrestris populations (Kraus et al. 2011; Lemopoulos et al. 2019). However, no technique is without limitation. Microsatellites may struggle to find siblings, or to identify the nuances of poorly differentiated populations (Lemopoulos et al. 2019).
In this study, a locus density of less than one microsatellite per chromosome may have limited our ability to uncover fine-scale clues in the dataset. Introgression or hybridisation can be hard to detect using microsatellites as large introgressed regions may lie undetected between sparse markers, which may help to explain why no evidence of introgression was detected through BAPS analysis (Jeffries et al. 2016). Additionally, the genes related to winter activity may not be linked with any of the microsatellite markers used. Such a gene must be in linkage (i.e. physically proximal) with one of the microsatellite loci in order to be detected; increasing the number of microsatellite loci used increases the resolution, thus increasing the likelihood that a gene is linked to a microsatellite locus.
A more thorough genomic surveying technique such as Restriction-site Associated DNA Sequencing (RADSeq), which can provide tens of thousands of informative SNP loci, may answer questions where microsatellites have failed to do so. In particular, RADSeq may help to understand why commercial and wild populations so closely resemble each other.
5 Conclusion
Microsatellite analysis of wild and commercial populations of B. terrestris has not revealed the presence of introgression; thus, introgression is unlikely to be the cause of the observed winter activity in wild B. terrestris populations in the UK. Furthermore, we found a lack of population structure and differentiation, which is consistent with previous literature. Further investigation is however necessary to determine if the import of foreign B. terrestris has had negative effects on the genetic diversity and fitness of native, wild populations.
References
Abadía-Cardoso, A., Pearse, D.E., Jacobson, S., Marshall, J., Dalrymple, D., Kawasaki, F., Ruiz-Campos, G., Garza, JC. (2016) Population genetic structure and ancestry of steelhead/rainbow trout (Oncorhynchus mykiss) at the extreme southern edge of their range in North America. Conserv. Genet. 17 (3), 675–689. doi:https://doi.org/10.1007/s10592-016-0814-9
Aizen, M.A., Smith-Ramírez, C., Morales, C.L., Vieli, L., Sáez, A., et al. (2018) Coordinated species importation policies are needed to reduce serious invasions globally: The case of alien bumblebees in South America. J. Appl. Ecol. 56, 100–106.
Alford, D.V. (1969) A study of the hibernation of bumblebees (Hymenoptera: Bombidae) in southern England. J. Anim. Ecol. 38, 149–170.
Bartomeus, I., Molina, F.P., Hidalgo-Galiana, A., Ortego, J. (2020) Safeguarding the genetic integrity of native pollinators requires stronger regulation on commercial lines. Ecol. Solut. Evid. DOI: https://doi.org/10.1002/2688-8319.12012
Beekman, M., van Stratum, P., Veerman, A. (1999) Selection for non-diapause in the bumblebee Bombus terrestris, with notes on the effect on inbreeding. Entomol. Exp. Appl. 93, 69–75.
Cejas, D., López-López, A., Muñoz, I., Ornosa, C., De la Rúa, P. (2020) Unveiling introgression in bumblebee (Bombus terrestris) populations through mitogenome-based markers. Anim. Genet. 51, 70-77
Chandler, D., Cooper, E., Prince, G. (2019) Are there risks to wild European bumble bees from using commercial stocks of domesticated Bombus terrestris for crop pollination? J. Apic. Res. 58, 665-681.
Coppée, A. (2010) ‘Bombus terrestris (L. 1758): A complex species or a species complex? Intraspecific pheromonal and genetic variations of Bombus terrestris (L.) - Impacts on the speciation’. PhD Thesis, Mons University, Mons.
Corander, J. Waldmann, P. and Sillanpää, M.J. (2003) Bayesian analysis of genetic differentiation between populations. Genetics 163, 367–374.
Donovan, B.J., Wier, S.S. (1978) Development of hives for field population increase, and studies on the life cycles of the four species of introduced bumble bees in New Zealand. N.-Z. J. Agric. Res. 21,733–756.
Earl, D.A., vonHoldt, B.M. (2012) STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 4, 359–361.
England. Natural England (2015) Bumblebees: licence for emergency release of non-native species for pollination or research (CL28) [online] https://www.gov.uk/government/publications/bumblebees-licence-to-release-them-for-pollination-and-research (Accessed on 08 Jan 2020).
Estoup, A., Solignac, M., Cornuet, J.M., Goudet, J., Scholl, A. (1996) Genetic differentiation of continental and island populations of Bombus terrestris (Hymenoptera: Apidae) in Europe. Mol. Ecol. 5, 19–31.
Funk, C.R., Schmid-Hempel, R., Schmid-Hempel, P. (2006) Microsatellite loci for Bombus spp. Mol. Ecol. Notes 6, 83–86.
Goudet, J. (1995) FSTAT: A computer program to calculate F-statistics. J. Hered. 86, 485–486.
Graystock, P., Yates, K., Evison, S.E.F., Darvill, B., Goulson, D., Hughes, W.O.H. (2013) The Trojan hives: pollinator pathogens, imported and distributed in bumblebee colonies. J. Appl. Ecol. 50, 1207–1215.
Graystock, P., Blane, E.J., McFrederick, Q.S., Goulson, D., Hughes, W.O.H. (2016) Do managed bees drive parasite spread and emergence in wild bees? Int. J. Parasitol. Parasites Wild. 5, 64–75.
Gurel, F., Gosterit, A., Eren, Ö. (2008) Life-cycle and foraging patterns of native Bombus terrestris (L.) (Hymenoptera, Apidae) in the Mediterranean region. Insect. Soc. 55, 123 –128.
Ings, T.C., Ward, N.L., Chittka, L. (2006) Can commercially imported bumble bees out-compete their native conspecifics? J. Appl. Ecol. 43, 940–948.
Inoue, M.N., Yokoyama, J., Washitani, I. (2008) Displacement of Japanese native bumblebees by the recently introduced Bombus terrestris (L.) (Hymenoptera: Apidae). J. Insect Conserv. 12, 135–146.
Jakobsson, M., Rosenberg, N.A. (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23, 1801–1806.
Jeffries, D.L., Copp, G.H., Lawson Handley, L., Håkan Olsén, K., Sayer, C.D., Hänfling, B. (2016) Comparing RADseq and microsatellites to infer complex phylogeographic patterns, an empirical perspective in the Crucian carp, Carassius carassius, L, Mol. Ecol. 25, 2997–3018.
Jost, L. (2008) GST and its relatives do not measure differentiation. Mol. Ecol. 17, 4015–4026.
Kalinowski, S.T. (2004) Counting alleles with rarefaction: private alleles and hierarchical sampling designs. Conserv. Genet. 5, 539–543.
Kraus, F.B., Szentgyörgyi, H., Rozej, E., Rhode, M., Moroń, D., Woyciechowski, M., Moritz, R.F.A. (2011) Greenhouse bumblebees (Bombus terrestris) spread their genes into the wild. Conserv. Genet. 12, 187–192.
Lecocq T, Rasmont P, Harpke A, Schweiger O (2016). Improving international trade regulation by considering interspecific variation for invasion risk assessment of commercially traded species: The Bombus terrestris case. Conserv. Lett. 9, 281–289.
Lemopoulos, A., Prokkola, J.M., Uusi-Heikkliä, S., Vasemägi, A., Huusko, A., Hyvärinen, P., Kolionen, M., Koskiniemi, J., Vainikka, A. (2019). Comparing RADseq and microsatellites for estimating genetic diversity and relatedness – Implications for brown trout conservation. Ecol. Evol. 9, 2106–2120.
Løken, A. (1973) Studies on Scandinavian bumblebees (Hymenoptera, Apidae). Norsk Entomol. Tidsskr. 20, 1–219.
Maebe, K., Meeus, I., Vray, S., Claeys, T., Dekoninck, W., Boevé, J.-L., Rasmont, P., Smagghe, G. (2016) Genetic diversity of restricted wild bumblebees was already low a century ago. Sci. Rep. 6, 38289. doi:https://doi.org/10.1038/srep38289
Maebe, K., Golsteyn, L., Nunes-Silva, P., Blochtein, B., Smagghe, G. (2018) Temporal changes in genetic variability in three bumblebee species from Rio Grande do Sul, South Brazil. Apidologie 49(3), 415–429.
Maebe, K., Karise, R., Meeus, I., Mänd, M, Smagghe, G. (2019a) Pattern of population structuring between Belgian and Estonian bumblebees. Sci. Rep. 9, 9651. doi:https://doi.org/10.1038/s41598-019-46188-7
Maebe, K., Karise, R., Meeus, I., Mänd, M, Smagghe, G. (2019b) Level of genetic diversity in European bumblebees is not determined by local species abundance. Front. Genet. 10, 1262. doi:https://doi.org/10.3389/fgene.2019.01262
Meeus, I., Brown, M.J.F., De Graaf, D.C., Smagghe, G. (2011) Effects of Invasive Parasites on Bumble Bee Declines. Conserv. Biol. 25, 662–671.
Moreira, A., Horgan, F.G., Murray, T.E., Kakouli-Duarte, T. (2015) Population genetic structure of Bombus terrestris in Europe: Isolation and genetic differentiation of Irish and British populations. Mol. Ecol. 24, 3257–3268.
Murray, T.E., Fitzpatrick, Ú., Brown, M.J.F., Paxton, R.J. (2008) Cryptic species diversity in a widespread bumble bee complex revealed using mitochondrial DNA RFLPs. Conserv. Genet. 9, 653–666.
OpenStreetMap (2017) Planet Dump [online] http://planet.osm.org. <http://www.openstreetmap.org/> (Accessed on 5th September 2019)
Owen, E.L., Bale, J.S., Hayward, S.A.L. (2013) Can winter-active bumblebees survive the cold? Assessing the cold tolerance of Bombus terrestris audax and the effects of pollen feeding. PLoS ONE 8(11), e80061. doi:https://doi.org/10.1371/journal.pone.0080061
Peakall, R., Smouse, P.E. (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. Notes 6, 288–295.
Pritchard, J.K., Stephens, M., Donnelly, P. (2000) Inference of population structure using multilocus genotype data. Genetics 155, 945–959.
Rasmont, P., Coppee, A., Michez, D., De Meulemeester, T. (2008) An overview of the Bombus terrestris (L. 1758) subspecies (Hymenopetera: Apidae). Ann. Soc. Entomol. Fr. 44, 243–250.
Robertson, A. (1991) A mid-winter colony of Bombus terrestris L. (Hym., Apidae) in Devon. Entomol. Mon. Mag. 127, 165–166.
Rosenberg, N.A. (2004) DISTRUCT: a program for the graphical display of population structure. Mol. Ecol. Notes 4, 137–138.
Ruz, L., Herrera, R. (2001) Preliminary observations on foraging activities of Bombus dahlbomii and Bombus terrestris (Hym: Apidae) on native and non-native vegetation in Chili. Acta Hortic. 561, 165–169.
Seabra, S.G., Silva, S.E., Nunes, V.L., Sousa, V.C., Martins, J., et al. (2018) Genomic Signatures of introgression between commercial and native bumblebees, Bombus terrestris, in western Iberian Peninsula - Implications for conservation and trade regulation. Evol. Appl. 12, 679–691. doi:https://doi.org/10.1111/eva.127
Stainforth, D.A., Chapman, S.C., Watkins, N.W. (2013) Mapping climate change in European temperature distributions. Environ. Res. Lett. 8, 034031.
Stelzer, R.J., Chittka, L., Carlton, M., Ings, T.C. (2010) Winter active bumblebees (Bombus terrestris) achieve high foraging rates in urban Britain. PLoS One 5(3), e9559. doi:https://doi.org/10.1371/journal.pone.0009559
Stolle, E., Rohde, M., Vautrin, D., Solinac, M., Schmid-Hempel, P., Schmid-Hempel, R., Moritz, R.F.A. (2009) Novel microsatellite DNA loci for Bombus terrestris (Linnaeus, 1758). Mol. Ecol. Resour. 9, 1345–1352.
Van Oosterhout, C., Hutchinson, W.F., Wills, D.P.M., Shipley P. (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol. Ecol. Notes 4, 535–538.
Velthuis, H.H.W., van Doorn, A. (2006) A century of advances in bumblebee domestication and the economic and environmental aspects of its commercialization for pollination. Apidologie 37, 421–451.
Wang, J. (2004) Sibship reconstruction from genetic data with typing errors. Genetics 166, 1963–1979.
Acknowledgements
We would like to thank the Royal Parks and London Borough Councils for permission to sample bees, and Dr. Tomas Murray for advice on microsatellite analysis at the beginning of the project.
Funding
This work was supported by a Leverhulme Early Career Fellowship ECF/2009/0180 awarded to TI, by the Research Foundation-Flanders (FWO-Vlaanderen; G042618N) and the Belgian Science Policy (BELSPO; BR/132/A1/BELBEES) awarded by GS.
Author information
Authors and Affiliations
Contributions
TI planned the experiments and performed the sampling and lab work. GB collected and prepared the genotyping data. AH and KM performed the data analysis. AH wrote the manuscript with feedback from KM, TI and GS.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Manuscript editor: Mathieu Lihoreau
L'activité hivernale de l'espèce de bourdon britannique Bombus terrestris audax est indépendante de l'introgression.
Bombus terrestris / bourdons / introgression / microsatellites / génétique des populations.
Die Winteraktivität der britischen Hummelart Bombus terrestris audax ist unabhängig von Introgression.
Bombus terrestris / Hummeln / Introgression / Microsatelliten / Populationsgenetik.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 61 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hart, A.F., Maebe, K., Brown, G. et al. Winter activity unrelated to introgression in British bumblebee Bombus terrestris audax. Apidologie 52, 315–327 (2021). https://doi.org/10.1007/s13592-020-00822-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13592-020-00822-w