Abstract
Facultatively social species exhibit behavioral plasticity in response to changes in ecological conditions and social environment, and thus provide a natural experiment to compare solitary and social behaviors in a single genome. Such species can therefore provide empirical insights into the evolution of eusociality. The small carpenter bees (genus Ceratina) and sweat bees (Halictidae) are of special interest because they exhibit rich behavioral plasticity. Species range from solitary to eusocial, and both groups benefit from detailed behavioral research and well-established phylogenies. As such, small carpenter and sweat bees are poised to further comparative sociogenomic studies which emphasize the necessity of a molecular phylogeny for understanding the evolution of molecular architecture underlying social phenotypes and organizational complexity. Here, we review behavioral, transcriptomic and genomic data in bees across the social spectrum, highlighting the importance of simple societies and facultatively social taxa to examine the genetic basis of cooperative traits and social evolution.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Phenotypic plasticity, an organism’s ability to modify physiological or behavioral traits following environmental cues, has been comprehensively studied over the past four decades (West-Eberhard 1989; Nijhout 2003). In some cases, variation in the regulation of a conserved set of genes can lead to the expression of multiple, highly discrete phenotypes (Whitman and Agrawal 2009). Such polyphenisms are relatively common in the insects (Emlen and Nijhout 2000; Simpson et al. 2011) and are informative targets for addressing questions regarding the origins and elaboration of derived phenomena (e.g., caste determination, Evans and Wheeler 2001; Fjerdingstad and Crozier 2006) including the paradox of eusociality. Eusociality is considered the most complex form of social organization, and eusocial species are often ecologically dominant (Michener 1969; Wilson 1971). However, despite its biological prevalence and ecological success (ants alone represent as much as 25% of all terrestrial biomass, Schultz 2000), the emergence of eusociality remains a rare and highly derived event (Wilson and Hölldobler 2005; Nowak et al. 2010). Determining the molecular and environmental factors underlying the origins of eusocial behavior has consequently become a central focus in the field of social evolution (Michener 1974).
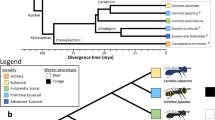
Eusociality has emerged independently only a handful of times in nature, but has done so within bees more than any other lineage (likely four independent origins, Figure 1; Cardinal and Danforth 2011; Gibbs et al. 2012; Rehan et al. 2012). The repeated evolution of eusociality within so widely diversified a monophylum has made eusocial bees an exemplary group for comparative research (Fischman et al. 2011). Accordingly, studies in socially complex bees have generated well-founded theories regarding the origins of eusociality and have contributed to the emergence of the field of sociogenomics, which integrates the disciplines of molecular biology and behavioral genetics with insights from phylogenetics and behavioral ecology to investigate the molecular architecture of social behavior and organization (Robinson 2002; Robinson et al. 2005).
Phylogeny of major bee groups and social lineages. Black nodes indicate independent origins of sociality. Branch colors indicate lineages containing species which are as follows: gray, solitary; blue, solitary through primitively eusocial; green, primitive eusocial; yellow, primitive through advanced eusocial; orange, advanced eusocial. Branches hashed with yellow indicate lineages containing facultatively social species. Letters indicate published resources for that lineage (G genome; T transcriptome; M methylome).
Advanced eusocial bees are considered to have transitioned past an evolutionary “point of no return” (Wilson 1971; Wilson and Hölldobler 2005), and thus provide mainly inferential insights regarding the origins of their complex sociality (Schwarz et al. 2007; Boomsma 2009). By contrast, it is thought that the vast majority of the more than 20,000 bee species worldwide (as many as 94%, Kocher and Paxton 2014) are either solitary or lack clearly defined castes (Michener 2007). Considered alongside their eusocial relatives, bees can thus be seen to comprise a natural gradient of social complexity (Kocher and Paxton 2014), formally termed the “social spectrum” (Rehan and Toth 2015; Table I). There is strong evidence that non-eusocial species are capable of moving along this social spectrum in either direction, “towards” and “away from” social complexity (West-Eberhard 2003; Field et al. 2010; Rehan and Toth 2015). Advanced eusociality thus does not represent a necessary evolutionary eventuality for social species (West-Eberhard 2003), but rather indicates the far extreme of fundamentally flexible social organization. Studies which look across this social spectrum have consequently spurred a concerted push to better unify theory (Rehan and Toth 2015; Toth and Rehan 2017) and to resolve semantic confusion within the field of social evolution (Dew et al. 2016). To fully appreciate the value of comparative research across the social spectrum, it is necessary to first briefly conceptualize this gradient, and to articulate the defining qualities of its major classes.
In solitary bees, reproductive females independently establish a nest and singly forage for their offspring, which require no care during their maturation (Michener 2007). Once provisioning is complete, females often either abandon their brood (e.g., Colletes and Osmia; Černá et al. 2013) or die before their offspring reach adulthood (Michener 2007). Though solitary species are not burdened by the increased risk of disease transmission associated with social living (Fu et al. 2015), they are extremely susceptible to parasitism and predation (Wcislo and Cane 1996). Increased parasitism and predation rates are thus considered primary ecological drivers towards social organization (Lin and Michener 1972; Wcislo et al. 2004; Rehan et al. 2011).
In a handful of seemingly solitary bees, the reproductive female will remain at her nest to guard and clean the developing brood (e.g., Ceratina japonica, Sakagami and Maeta 1984; Lasioglossum laticeps, Plateaux-Quénu 2008; and Ceratina calcarata, Rehan and Richards 2010a), and this extended parental care defines them as subsocial in their behavior (Michener 2007). Subsociality is the simplest form of social organization along the social spectrum (Rehan and Toth 2015) and is considered a necessary precondition for the evolution of more complex social groups (Wilson 1971; Michener 1974). Although subsocial bees are nest-loyal, and at least some are capable of nestmate recognition (Rehan and Richards 2013), they remain effectively casteless and independent as adults.
Like solitary and subsocial species, incipiently social bees remain totipotent throughout adulthood and are able to found nests of their own (Sakagami and Maeta 1989; Rehan et al. 2010b). They are distinguished from their subsocial cousins by their capacity to nest cooperatively under a simple division of labor to rear a single brood (West-Eberhard 1987). As they represent species at the origins of social organization, subsocial through incipiently social bees may be collectively referred to as early-stage social species (Table I; Rehan and Toth 2015).
On the other end of the social spectrum are bees representative of late stage sociality: the primitively eusocial species. In these groups, a reproductive female continues to lay eggs after initially founding her nest (either alone or with nestmate workers), and may do little else once the first generation of her offspring reach maturity (Wilson 1971; Michener 2007). Individuals of primitively eusocial species are typically monomorphic, though a division of labor based on age and body size is common (e.g., Halictus rubicundus, Soucy and Danforth 2002; Megalopta genalis, Smith et al. 2003; Halictus scabiosae, Brand and Chapuisat 2012). Worker-type offspring in primitively eusocial colonies may maintain a capacity for reproductive behavior, though ovary development is often inhibited (Michener 1974). These offspring are rarely limited to a single role, and may switch tasks over their lifetime. For instance, in both sweat bees and small carpenter bees, workers may rapidly develop into reproductives upon removal of a dominant reproductive nestmate (Eickwort 1986; Rehan et al. 2014b). The workers of advanced eusocial species, by contrast, develop into functionally sterile castes, often with distinct behaviors and morphologies (e.g., Apis and Melipona bees, Wilson 1971; Michener 1974, 2007). This distinct caste differentiation is considered a defining feature of advanced eusocial species, which are no longer capable of reverting to simpler forms of social organization (Wilson and Hölldobler 2005).
Intriguingly, early-stage social bees may demonstrate “facultative sociality,” or a capacity to express more than one form of social organization (Figure 1; Smith et al. 2003; Rehan et al. 2011, 2014a, b). The crepuscular sweat bee (M. genalis), for instance, nests either solitarily or in small, primitively eusocial colonies (Smith et al. 2003; Wcislo et al. 2004; Kapheim et al. 2013; Jones et al. 2017). The subsocial small carpenter bee (C. calcarata), by comparison, is facultatively incipiently social and may rear a worker-like offspring to aid in brood care and feeding (Rehan and Richards 2010a, b; Rehan et al. 2014a). The behaviors characteristic of early-stage social bees are theorized to be necessarily antecedent to advanced eusociality, and thus provide a tractable opportunity to empirically test social evolutionary theory (e.g., Wilson 1971; Michener 1974; Seger 1983; Rehan and Toth 2015). Those which are also facultatively social, however, provide particularly informative insights, as the costs, benefits, and molecular mechanisms underlying multiple classes of social organization may be explored concurrently within a single species (West-Eberhard 2003; Rehan and Toth 2015). To make evolutionarily informed conclusions about the molecular origins of complex social behavior across the social spectrum, well-resolved phylogenetic histories and species delineations must first be established (Danforth et al. 2013). Fortunately, considerable efforts have been made to resolve the phylogenies of two socially diverse bee groups: the sweat bees (Halictidae) and carpenter bees (Xylocopinae).
Represented by solitary, facultatively social, and eusocial species, the wide behavioral diversity of the halictids has made them an informative system for studies of behavioral plasticity and social evolution (Schwarz et al. 2007; Kocher and Paxton 2014). Multiple revisions of halictid phylogeny (Danforth 2002; Brady et al. 2006; Gibbs et al. 2012) have come to posit two origins of eusociality within Halictidae: one in the tribe Augochlorini (Danforth and Eickwort 1997; Danforth et al. 2013) and another for the tribe Halictini (Figure 1; Halictus and Lasioglossum, Gibbs et al. 2012). This research has also provided evidence of multiple reversions from social to solitary behaviors in this group (Danforth et al. 2003).
The carpenter bees (subfamily Xylocopinae) collectively represent the entirety of the social spectrum across their four tribes: the solitary Manueliini, incipiently social Xylocopini, solitary to eusocial Ceratinini, and incipiently social to eusocial Allodapini (Figure 1). This rich diversity, combined with a well-resolved phylogeny (Rehan et al. 2010a, 2012; Rehan and Schwarz 2015) and phylogeography (Dew et al. 2016; Shell and Rehan 2016), makes the xylocopine bees an exceptional system for sociogenomic research (Rehan and Toth 2015). Further, as the Xylocopinae and Apinae are sister subfamilies (Cardinal and Danforth 2011), research involving carpenter bees may directly inform our understanding of the dynamics underlying advanced eusociality in corbiculate species (Figure 1; Rehan and Toth 2015; Toth and Rehan 2017). Phylogenetic work within the Xylocopinae has revealed a single origin of sociality followed by as many as four reversions to a solitary lifestyle (Schwarz et al. 2007; Rehan et al. 2012).
Well-resolved phylogenies among social lineages provide much-needed evolutionary frameworks by which to contextualize comparative molecular research (Rehan and Toth 2015; Romiguier et al. 2015; Branstetter et al. 2017; Peters et al. 2017). Such research has helped to reveal that species may evolve along a non-linear trajectory between simple and complex social organization: incipiently social lineages, for instance, frequently revert to solitary life (Wcislo and Danforth 1997; Rehan et al. 2012; Gibbs et al. 2012). Phylogenetic studies also indicate that the evolution of advanced eusociality is rare, and likely checked by prohibitive natural barriers (Rehan et al. 2012). Integrative research which builds off this phylogenetic foundation may now benefit from ongoing advances in genomic and transcriptomic sequencing methods. As such, studies which strive to integrate genomic, transcriptomic, and behavioral data are poised to gain meaningful insights into the molecular signatures of phenotypic plasticity.
In this review, we provide a detailed overview of sociogenomic research in bees of early stage and facultative sociality. The rich behavioral and ecological diversity of these species provides a powerful workspace in which to explore the molecular, developmental, and environmental drivers of social transition (West-Eberhard 1987; Rehan et al. 2014a; Patalano et al. 2015). Despite their informative value, however, there remains a paucity of research in these groups. We thus argue in support of additional sociogenomic research into bees of early-stage and facultative sociality, and highlight the socially diversified Ceratina small carpenter bees and halictid sweat bees as promising model systems for future studies.
2 Facultatively social bees
Facultative sociality is not a distinct social category (like subsociality or eusociality), but rather indicates a species’ capacity for plasticity in its social organization. For example, conspecifics of some facultatively social sweat bees either establish a solitary nest, or found a primitively eusocial colony in which a helper generation assists in the rearing of a second brood of reproductive individuals (e.g., H. rubicundus, Soucy and Danforth 2002; M. genalis, Smith et al. 2003; Lasioglossum calceatum, Davison and Field 2016). Facultatively social nesting in some carpenter bees, by contrast, may involve incipiently social brood care by a cooperative pair of age-matched females (e.g., Ceratina australensis, Rehan et al. 2014b), or production of a single, worker-like offspring alongside future reproductives (e.g., C. calcarata, Rehan et al. 2014a; C. japonica, Sakagami and Maeta 1984). Where advanced eusocial Hymenoptera feature distinctive and often multifaceted reproductive and morphological caste systems, the behavioral phenotypes of facultatively social bees straddle species-specific ranges along the social spectrum. This behavioral diversity, both within and between facultatively social species, provides an exceptional opportunity to explore how phenotypic variation across modes of social organization is reflected at the molecular level (Kocher and Paxton 2014).
Facultatively social bees are particularly tractable models for comparative sociogenomic study be-cause variation in their social phenotype may be reliably elicited through experimental manipulation (Schwarz et al. 2007; Kocher and Paxton 2014; Rehan and Toth 2015). This can be seen, for example, in the induction of cooperative, multi-female nests in otherwise mainly subsocial species (e.g., C. japonica; Sakagami and Maeta 1984, 1987; Lasioglossum spp., Jeanson et al. 2005, 2008). Sakagami and Maeta (1995) elicited reproductive division of labor in the small carpenter bees, C. japonica (Sakagami and Maeta 1984) and C. okinawana (Sakagami and Maeta 1989), by constraining conspecific females to cohabitate. In these induced social nests, one female assumed queen-like behavior, laying eggs and defending the nest, while the second primarily foraged. In all cases of social nesting, the reproductive role was reliably adopted by the larger of the two females, in age-matched and even mother-daughter cooperative pairs (Sakagami and Maeta 1984, 1989, 1995). In similar fashion, Holbrook et al. (2009) observed that individuals of the normally solitary sweat bee (Lasioglossum NDA-1) assumed dedicated guarding or digging behaviors when forced to nest in pairs, though paired individuals secured deeper and better-guarded nests compared to their solitary counterparts. Holbrook et al. (2009) noted that such arrangements rarely occur in wild populations. Considerable phenotypic plasticity may thus be widespread in bees of early-stage sociality, even in those species which rarely demonstrate facultative sociality in the wild.
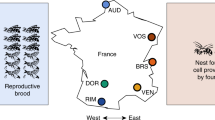
Environmental factors, including local climate, resource availability, predation, and parasite pressure, have long been theorized to comprise the primary ecological drivers of social evolution (Lin and Michener 1972; Evans 1977; Strassman and Queller 1989; Kocher et al. 2014). Accordingly, social complexity in facultatively social bees is often reliably predicted by ecological condition (Richards and Packer 1996; reviewed in Purcell 2011). For example, brood parasitism in the Australian small carpenter bee (C. australensis, Rehan et al. 2010b) appears to be a key selective pressure favoring social nesting (Rehan et al. 2011). The influence of environment on social organization has also been particularly well-documented in the sweat bees (Plateaux-Quénu et al. 2000; Schwarz et al. 2007; Davison and Field 2016). In H. rubicundus (Soucy and Danforth 2002), solitary or social reproduction appears to be largely dependent on geographic location: solitary H. rubicundus females produce a single brood in cooler regions (Eickwort et al. 1996), and social females produce two broods in warmer regions (Yanega 1989, 1993; Soucy 2002). Field et al. (2010) demonstrated this organizational lability by inducing the offspring of solitary or social populations to switch behavioral type via translocation between cool and warm locations. In other species of sweat bees, social phenotype has been reliably predicted specifically by altitude (L. calceatum, Sakagami and Munakata 1972) or even micro-habitat composition. For instance, in the sweat bee, L. balecium, social nests were consistently observed in warmer, sunnier sites, while solitary nests were typically found in adjacent, cooler, well-shaded locations (Hirata and Higashi 2008). In some cases, however, variation in social organization is neutrally or otherwise not predictably affected by environment (e.g., Lasioglossum apristum, Miyanaga et al. 1999; L. malachurum, Richards 2000; Wyman and Richards 2003), supporting the role of genetic architecture as primary in determining social phenotype (Soucy and Danforth 2002; Soro et al. 2010).
3 Behavioral and genetic mechanisms of social evolution
A suite of major theoretical frameworks has been put forward to explain the mechanisms by which complex social behavior may have evolved. Here, we provide an overview of some of these hypotheses which, though distinct in their viewpoints, are not mutually exclusive, and have been recently brought together in theoretical syntheses (Rehan and Toth 2015; Toth and Rehan 2017). A tendency to reuse a nesting substrate among kin groups is a well-supported precondition for the emergence of social behavior (Wild and Koykka 2014). As it may increase the opportunity for inclusive fitness benefits, reuse or inheritance of a “family” nest is expected to promote group nesting, even with some nestmate competition. Accordingly, nest reuse has been observed across facultatively social Ceratina (Sakagami and Maeta 1987; Rehan et al. 2009, 2014b, 2015) and in other basically social bees (e.g., H. rubicundus, Yanega 1990). By contrast, those species which are known to frequently disperse prior to the establishment of a new nest (e.g. C. calcarata and C. flavipes) are less likely to nest socially (Sakagami and Maeta 1987; Rehan and Richards 2010a, b). Along with nest reuse, multi-voltinism, the capacity to produce more than one brood during a reproductive season, has been suggested as antecedent to eusocial behavior (Seger 1983). The ability to produce multiple generations of offspring in a single breeding season establishes a scenario in which the offspring of mated, overwintering females have an opportunity to remain at the nest as helpers in the following reproductive season. This proposition has found demonstration in primitively eusocial bees (e.g., Halictus sexcinctus, Richards 2001) and is supported by research in bivoltine Ceratina (e.g., C. okinawana, Sakagami and Maeta 1995). The prolonged maternal care behavior often seen in multivoltine species has also been suggested as prerequisite to more complex social structuring (West-Eberhard 1987) and is likely readily reinforced through selection (Wade 2001).
West-Eberhard (1996) also considered the evolutionary necessity of parental care behavior in her proposition of the ovarian ground-plan hypothesis, which offers that extended maternal care behaviors seen in solitary and subsocial Hymenoptera may have gradually differentiated into the dedicated forager and egg-layer roles seen in incipiently social and primitively eusocial Hymenoptera (Figure 2). The reproductive ground-plan hypothesis builds on this prediction by suggesting that such specialized roles could gradually lead to the emergence of task-specific worker castes, as seen in advanced eusocial species (Amdam et al. 2004, 2006). In many incipiently social bees, however, reproductive and foraging behavior is expressed in tandem; as such, a decoupling of foraging and reproductive behavior is not expected to have initiated advancements in social complexity in most lineages (Schwarz et al. 2011; Rehan et al. 2014a).
Nestedness and interconnectedness of major social evolutionary theories. Genes underlying core developmental and behavioral traits may become available for social roles following chance cooption, duplication, or change in regulation (heterochrony). Evolution of division of labor may then be underpinned by differences in hormone titers and gene expression levels associated with ovarian development (ovarian ground-plan) and/or temporal variations in the expression of genes associated with maternal care behavior (maternal heterochrony).
It might be expected that the initial co-option of maternal care genes into relatively queen-like and worker-like phenotypes would be hindered, or outright prevented, by strict pleiotropic constraints on those genes (Gadagkar 1997). In consideration of this, Gadagkar (1997) proposed the concept of genetic release, in which a gene duplication event or significant variation in the timing of gene expression could free key genes from the effects of stabilizing selection. Variation in gene expression, in combination with the evolution of separate developmental programs, could then allow for the mutual decoupling of strictly reproductive activities (e.g., mating, egg laying) from brood care and nest maintenance tasks. Directional selection through inclusive fitness could then reinforce the expression of increasingly queen-like and worker-like behaviors and physiologies. The maternal heterochrony hypothesis builds on this concept by proposing that a shift in the timing of expression (heterochrony) of maternal care genes may account for the evolution of precocious, sib-social care behavior seen in advanced eusocial species (Linksvayer and Wade 2005). From this approach, the emergence of queen- and worker-type behaviors could result from a slight variation in the regulation of an otherwise conserved set of genes.
Rehan et al. (2014a) directly assessed the maternal heterochrony hypothesis through transcriptomic analysis of the subsocial small carpenter bee, C. calcarata. In C. calcarata, mothers may rear a worker-like daughter to aid in brood care and foraging, and thus enter an incipiently social stage (Rehan and Richards 2010b). Rehan et al. (2014a) compared brain gene expression of individuals from five focal stages in C. calcarata’s reproductive season, and, as predicted by the maternal heterochrony hypothesis, gene expression patterns of worker-like daughters most closely mirrored those of post-reproductive mothers, and included upregulation of genes implicated in maternal care. Further support for the underlying role of molecular heterochrony in emergent brood care behavior has also been found through similar transcriptomic works in other Hymenoptera (e.g., Polistes metricus, Toth et al. 2007, 2010).
Complementary to both heterochrony and ground-plan hypotheses is the idea of the role of genetic “toolkits”—highly conserved genes which play a focal role in the developmental pathways of core physiological and behavioral traits across taxa (Toth and Robinson 2007). The genetic toolkit hypothesis suggests that modifications to these key genes, or the molecular pathways regulating their expression (i.e., transcription factors), could underlie the emergence of behaviors leading to social organization (Figure 2; True and Carroll 2002; Carroll 2005). Expanded by Bloch and Grozinger (2011), the functionality of socially co-opted toolkit genes or pathways could become sensitive to signals from the social environment, and thus become socially regulated. The genetic toolkit hypothesis has found support in widely conserved genetic signaling across both taxonomic and social lineages within bees (Woodard et al. 2011) and among social Hymenoptera more broadly (Toth et al. 2014; Berens et al. 2014; Woodard et al. 2014; Morandin et al. 2016). For instance, two species of primitively eusocial bumble bee, Bombus terrestris and B. impatiens, were found to share many genes and gene regulatory elements with both P. metricus and A. mellifera, including those involved in the regulation of social behaviors (Sadd et al. 2015). In a comparison of transcriptomes across hymenopteran lineages (i.e., bees, ants, and wasps), Berens et al. (2014) determined that, despite high variation in expressed genes, metabolic pathways and molecular functions were relatively well-conserved across distant social lineages. They thus suggested support for a “loose” genetic toolkit: even when relatively few genes are expressed in common, the expression of complex sociality appears to involve highly conserved transcription factor and molecular functional networks.
In contrast to the highly conserved nature of genetic toolkits, novel or taxonomically restricted genes (i.e., those unique to clades or lineages) have also gained support as likely to play a key role in the evolution of complex social behavior (Johnson and Tsutsui 2011; Sumner 2014). With the evolution of eusociality, it may be expected that group behavioral and physiological coordination would come to rely on specialized genes and gene expression networks (Johnson and Linksvayer 2010). Evidence of caste-specific expression in genes unique to bees (e.g., A. mellifera, Harpur et al. 2014) and other social Hymenoptera (e.g., Polistes wasps, Ferreira et al. 2013; Temnothorax ants, Feldmeyer et al. 2014) suggests positive selection on novel genes may underlie derived worker phenotypes (Kapheim et al. 2015). Accordingly, detection of lineage-specific variations in gene families, such as a bias towards gustatory chemoreceptors in Bombus (Sadd et al. 2015) or expansion in odorant receptor genes in the clonal raider ant (Ooceraea biroi; McKenzie et al. 2016), have helped to reveal how changes at the molecular level may dictate the evolutionary trajectory of social Hymenoptera. Despite many novel genes between eusocial lineages, a large body of research in toolkit genes indicates that complex gene expression networks and metabolic pathways remain highly conserved (Simola et al. 2013; Berens et al. 2014; Morandin et al. 2016). Therefore, in theory, (i) eusociality may evolve from a highly diverse suite of antecedent genetic profiles, but (ii) the composition of genes and functional regulatory networks underlying social phenotypes may be increasingly constrained as species approach advanced forms of social complexity. It remains unclear, however, whether the apparent acceleration in rates of positive selection detected among eusocial lineages may instead indicate relaxed selection operating on relatively reduced effective population sizes in these taxa (Harpur and Zayed 2013; Romiguier et al. 2014).
4 Sociogenomics: a comparative approach to social evolution
Broadly, sociogenomic research addresses the feedback circuit between genetics and environmental cues: (i) how molecular functions and expression affect behavior and social organization, and (ii) how social and environmental cues further modify signaling pathways, development, physiology, and behavior (Robinson 2002; Robinson et al. 2005). The publication of the A. mellifera genome represented a critical foundation for concerted sociogenomic research (The Honeybee Genome Sequencing Consortium 2006) and remains a valuable resource for comparative studies of behavioral plasticity and social evolution (Fischman et al. 2011; Dolezal and Toth 2014). As genes are generally conserved across genera, particularly within taxonomic clades, the expanding wealth of honey bee sociogenomic research can act as a guide for explorations into the molecular architecture of species considered antecedent to the eusocial form (Page and Amdam 2007; Fischman et al. 2011; Woodard et al. 2011).
The determination of the molecular signals underlying the origins of caste determination in advanced eusocial Hymenoptera represents a central endeavor in the field of sociogenomics (Toth et al. 2007; Toth and Robinson 2009), and has been astutely explored (Barchuk et al. 2007; reviewed in Berens et al. 2014). As with other complex traits of interest, caste determination can be traced to suites of genes whose functions are tied to key developmental (e.g., vitellogenin), hormonal (e.g., juvenile hormone), and metabolic pathways (e.g., insulin pathway genes; reviewed in Page and Amdam 2007; Corona et al. 2016). As the expression of these critical pathways remains highly sensitive to external cues (such as pheromone signaling, Grozinger et al. 2003; and nutritional intake, Ament et al. 2008), they can be considered as highly responsive to factors within the social environment. It has thus been theorized that gene pathways in eusocial taxa, which are involved in the ontogenetic determination of caste fate during the earliest stages of development, may have arisen through the co-opting of genes previously involved in core physiological processes (i.e., genetic toolkit hypothesis, see also Toth et al. 2007; Toth and Robinson 2009). We can use the power of comparative sociogenomic research across the social spectrum to trace the function of candidate genes across species of varying degrees of sociality (reviewed in Robinson et al. 2005; Smith et al. 2008).
5 Sociogenomics across the social spectrum
Continued advancements in the accuracy and accessibility of sequencing technologies havemade the acquisition and analysis of genomic and transcriptomic data an increasingly achievable undertaking, and have greatly expanded the molecular resources available for social bee lineages (Table I; Figure 1). Comparisons of differentially expressed genes (DEGs) within A. mellifera, for instance, have revealed clear distinctions in brain gene expression between scout, non-scout (Liang et al. 2012), and recruit worker behavioral variants (Southey et al. 2016). As genomic and transcriptomic resources are developed for species of early-stage and facultative sociality (e.g., Lasioglossum albipes; Kocher et al. 2013; C. calcarata; Rehan et al. 2014a, 2016), extended comparative studies within and between Hymenoptera of distinct social organizations and lineages will be made possible (reviewed in Berens et al. 2014; Kapheim 2016). In the genome of C. calcarata, for instance, key transcription factor and chemosensory gene groups were identified as significantly expanded (Rehan et al. 2016), directly in line with expectations from observed gene family expansions in A. mellifera (Kapheim et al. 2015).
Insights into the molecular signals of ontogenetic variation between social lineages have also been gained through DEG research in facultatively social bees. Analysis of transcriptomic data recently developed for M. genalis revealed that while both functional gene groups and associated expression levels varied greatly over the course of an offspring’s development, there were consistent shifts in primary functional processes during each stage of maturation (Jones et al. 2015). As female M. genalis develop from egg to adult, expression patterns consistently shift from predominantly cellular development and differentiation, to establishment of neurological systems and specialized metabolic pathways at adulthood (Jones et al. 2015). These results contrast with similar DEG analyses of offspring caste determination in eusocial species, in which early nutritional signals lead to increasingly distinct developmental gene expression pathways towards adulthood as either a sterile worker or reproductive queen (as seen in A. mellifera, Barchuk et al. 2007; Polistes wasps, Hunt et al. 2010; and Formica ants, Morandin et al. 2015). More recently, however, research investigating differences in brain and abdominal gene expression among solitary and social M. genalis revealed stark differences between reproductive and non-reproductive roles. The abdominal gene expression patterns of workers that became replacement queens mirrored those of reproductive foundresses, while those of non-reproductive worker daughters stood apart. Differences in brain gene expression patterns among castes were not nearly as strong, however, suggesting that changes in reproductive functionality may precede changes in behavior (Jones et al. 2017). The caste-biased differences in abdominal gene expression detected in M. genalis were found to overlap with those of obligately eusocial taxa, suggesting there may be considerable conservation of the regulatory mechanisms underlying eusocial organization across evolutionary lineages. Further research comparing the transcriptomic profiles of offspring developmental stages and adult reproductive strategies (solitary vs. social) in other bees of early-stage sociality more broadly is a necessary next step.
Genomic and transcriptomic resources have also enabled the exploration of evolutionary developmental (evo-devo) theories as to how highly complex behavioral traits may emerge from a genetic functional architecture, which can be at once widely conserved (Rittschof et al. 2014) and pleiotropically constrained (Carroll 2008; Chen et al. 2013). Transcriptional regulatory networks (TRNs) are the highly complex and interconnected molecular signaling pathways (between transcription factors (TFs) and their target genes) underlying both gene expression and environmentally responsive regulation (Luscombe et al. 2004). TRN analysis in bees was pioneered by Chandrasekaran et al. (2011), who combined the brain transcriptomic profiles from 853 honey bees, representative of 48 distinct behavioral phenotypes, to assemble a brain TRN for A. mellifera. Their work indicated that brain gene expression would reliably predict behavioral phenotype, and that disparate behavioral phenotypes frequently rely on both shared and targeted TFs. A more recent population genomic study of the A. mellifera brain TRN revealed that regulatory and coding sequences throughout the TRN are under purifying selective pressure (Molodtsova et al. 2014). Proteins on the periphery of this regulatory network were most likely to be free of pleiotropic constraint and thus available for adaptive selection. Accordingly, this work provided additional support for the genetic toolkit theory within the evo-devo framework, highlighting the role of alterations to TFs regulating gene expression, rather than changes in the genes themselves, as likely critical in the emergence of complex social behaviors.
The importance of TF functionality in social evolution was further highlighted through comparative genomic studies at a phylogenetic scale. Kapheim et al. (2015) compared the genomic data of 10 bee species (representing three families and containing two independent origins of eusociality) to investigate conserved molecular dynamics across social lineages. In contrast to species of early-stage sociality, those of greater social complexity featured an increased frequency of transcription factor binding sites and putatively methylated genes. Additional evidence of accelerated evolution among eusocial taxa was also seen in similar studies (Woodard et al. 2011; Simola et al. 2013; Harpur and Zayed 2013; Harpur et al. 2014; Romiguier et al. 2014), and suggests that a capacity to regulate gene expression may become increasingly essential in the later stages of social organization. However, large sets of distinct and often lineage-specific genes were also detected in these studies (Woodard et al. 2011; Simola et al. 2013; Kapheim et al. 2015). Thus, despite considerable convergence in social organization and gene network functionality among eusocial taxa, there appears to be great variability in the possible evolutionary routes to complex social organization.
The increased achievability of sociogenomic methods opens doors to the integrative study of previously under-researched taxa. In the past 4 years, the A. mellifera genome has been joined by the draft genomes of two advanced eusocial (Apis florea and Melipona quadrifasciata; Kapheim et al. 2015) and two primitively eusocial bees (B. terrestris and B. impatiens, Sadd et al. 2015); three solitary (Megachile rotundata, Dufourea novaeangliae, and Habropoda laboriosa; Kapheim et al. 2015) and two facultatively social bees (L. albipes, Kocher et al. 2013; C. calcarata, Rehan et al. 2016); and one putatively facultatively social bee (Eufriesea mexicana, Kapheim et al. 2015; Table I, Figure 1). As genomic and transcriptomic data continue to be developed in additional social and solitary bees, we will continue to gain more comprehensive insights into how novel genes, genetic release from pleiotropic constraints, and convergent gene regulation underlie the foundational transitions in social form (Rehan and Toth 2015).
6 Epigenetics and phenotypic plasticity
Variation in gene expression via methylation is a well-appreciated mechanism of phenotypic plasticity across many taxa (Goldberg et al. 2007; Glastad et al. 2011). DNA methylation data from social Hymenoptera (Yan et al. 2015) indicate that a capacity for epigenetic response to ecological or social environment may play a role in the emergence of some complex traits (Weiner and Toth 2012; Kapheim et al. 2015). Studies investigating the epigenetics of phenotypic plasticity in A. mellifera (Lyko et al. 2010; Foret et al. 2012; Li-Byarlay et al. 2013) and bees of early-stage sociality (Kocher et al. 2013; Rehan et al. 2016) have detected conserved expansions in possible methylation sites (CpG densities), and have consequently proposed this expanded capacity for DNA methylation as putatively central in the evolution of social complexity (Glastad et al. 2011; Weiner and Toth 2012). Though CpG densities are indicators for the capacity of methylation (Weiner and Toth 2012; Bewick et al. 2017), only by examining an organism’s total methyl-modified genome can one empirically explore changes in epigenetic architecture across behavioral states and taxa (e.g., bees, A. mellifera, Lyko et al. 2010; C. calcarata, Rehan et al. 2016; Polistes wasps, Patalano et al. 2015; Standage et al. 2016; and the clonal raider ant, Cerapachys biroi, Libbrecht et al. 2016; reviewed in Hunt et al. 2013). Nevertheless, studies exploring methylation activity in primitively eusocial Polistes wasps found little to no evidence of gene methylation in this group (P. canadensis, Patalano et al. 2015; P. dominula, Standage et al. 2016). Thus, though some form of epigenetic regulation of gene expression via DNA methylation appears widespread in many species, its exact function in the evolution of social organization remains irregular and unclear (Bewick et al. 2017). Despite a large body of literature regarding its implications in Hymenoptera, published methylome data in bees is currently limited to four species: A. mellifera (Lyko et al. 2010), C. calcarata (Rehan et al. 2016), L. albipes, and D. novaeangliae (Glastad et al. 2017). Additional resources for bees of early-stage and facultative sociality could help to further illuminate how methylation affects phenotype, and whether epigenetic modifications influence cooperative behavior at the origins of social organization (Weiner and Toth 2012).
7 The eco-evo-devo approach
As genomic and transcriptomic data provide unprecedented insights into the molecular drivers of phenotypic plasticity, the role of environmental factors in affecting gene expression pathways is becoming increasingly well appreciated (Gilbert 2012; Weiner and Toth 2012; Schlichting and Wund 2014). Social species are affected by signals from both their ecological habitat (e.g., temperature or predator pressure) and their social environment (Bloch and Grozinger 2011; Toth and Rehan 2017). Social environmental signals include pheromones (Conte and Hefetz 2008), maternal manipulation of provisions for developing larvae (Rehan and Richards 2010b; Lawson et al. 2016), or reproductive policing by nestmates (Olejarz et al. 2016), any of which may affect changes in an individual’s development or behavior. By integrating measurements in the social and ecological environment with ontogenetic, phylogenetic, and genomic datasets, researchers now have the chance to capture an unprecedentedly holistic portrait of behavioral plasticity and social evolution (Toth and Rehan 2017). This revelation has formed the basis of the emerging “eco-evo-devo” approach, under which the flexible molecular architecture underlying an individual’s developmental pathways is continuously influenced by environmental signal factors (reviewed in Abouheif et al. 2014; Gilbert et al. 2015; Toth and Rehan 2017).
Central to the eco-evo-devo framework is the concept of genetic assimilation, in which an initially plastic phenotypic response to local environmental pressure may gradually, through accommodating mutations and selective reinforcement, lead to canalization for discrete phenotypes (Waddington 1961). Evidence for genetic assimilation has been clearly demonstrated in the tobacco hornworm (Manduca sexta, Suzuki and Nijhout 2006), in which experimental alteration of environmental factors was shown to lead to novel phenotypic responses, which may then be readily reinforced at the genetic level through accommodating mutation. To date, comparative research in Apis (Woodard et al. 2011; Harpur et al. 2014) and among social bee lineages (Kapheim et al. 2015) has suggested that genetic assimilation may play a similar role in the evolution of insect sociality.
Recent sociogenomic research in M. genalis, however, offers the first empirical assessment of this idea in a facultatively eusocial species (Jones et al. 2017). Jones et al. (2017) found that genes with worker-biased expression in M. genalis overlapped with genes shown to be rapidly evolving in ten other bee species. Similarities in the patterns of evolutionary rate and caste-biased gene expression between M. genalis and advanced eusocial species (e.g., Apis) thus suggest that strong positive selection through genetic accommodation may have driven the elaboration of social phenotype (i.e., a worker caste) in the ancestors of modern eusocial taxa (Jones et al. 2017). The developmentally constrained caste systems of advanced eusocial bees may thus be the result of sustained environmental pressures operating on the flexible “substrate for selection” seen in bees of early and facultative sociality (Kapheim et al. 2015; Toth and Rehan 2017).
8 Conclusions
Phenotypic plasticity, particularly within the context of social evolution, is a multifaceted and highly dynamic phenomenon. Advanced eusocial species have provided a wealth of insight into the molecular mechanisms underlying highly elaborated insect societies; however, the complex nature of their sociality avails mainly inferential conclusions regarding the evolutionary origins of division of labor. Bees of early-stage sociality are most plausibly antecedent to the eusocial form and, in many cases, remain responsive to a wide range of social, biotic, and abiotic factors. As such, species of early-stage and facultative sociality represent particularly informative models towards a better understanding of the diverse origins and drivers of behavioral plasticity and social organization. Future comparative sociogenomic studies will benefit from the ongoing expansion of molecular resources for bees across the social spectrum, and through an integrative application of the emerging eco-evo-devo approach.
Change history
18 September 2017
The above mentioned article, written by Wyatt A. Shell and Sandra M. Rehan, was originally published electronically on the publisher’s internet portal (currently SpringerLink) on 24 July 2017 without open access.
References
Abouheif E., Favé M.J., Ibarrarán-Viniegra A.S., Lesoway M.P., Rafiqi A.M. (2014) Eco-evo-devo: the time has come. In: Landry, C.R., Aubin-Horth, N. (Eds.) Ecological Genomics: Ecology and the Evolution of Genes and Genomes. Springer, Netherlands. pp. 107–125
Amdam G.V., Norberg K., Fondrk M.K., Page R.E. (2004) Reproductive ground plan may mediate colony-level selection effects on individual foraging behavior in honey bees. Proc. Natl. Acad. Sci. U.S.A. 101 (21), 11250–11255
Amdam G.V., Csondes A., Fondrk M.K., Page R.E. (2006) Complex social behaviour derived from maternal reproductive traits. Nature 439 (7072), 76–78
Ament S.A., Corona M., Pollock H.S., Robinson G.E. (2008) Insulin signaling is involved in the regulation of worker division of labor in honey bee colonies. Proc. Natl. Acad. Sci. U.S.A 105 (11), 4226–4231
Barchuk A., Cristino A.S., Kucharski R., Costa L.F., Simões Z. L. P., Maleszka R. (2007) Molecular determinants of caste differentiation in the highly eusocial honeybee Apis mellifera. BMC Dev. Biol. 7 (70), 1–19
Berens A.J., Hunt J.H., Toth A.L. (2014) Comparative transcriptomics of convergent evolution: different genes but conserved pathways underlie caste phenotypes across lineages of eusocial insects. Mol. Biol. Evol. 32 (3), 690–703
Bewick A.J., Vogel K.J., Moore A.J., Schmitz R.J. (2017) The evolution of DNA methylation and its relationship to sociality in insects. Mol. Biol. Evol. In press
Bloch G., Grozinger C.M. (2011) Social molecular pathways and the evolution of bee societies. Phil. Trans. R. Soc. B. 366, 2155–2170
Boomsma J.J. (2009) Lifetime monogamy and the evolution of eusociality. Philos. T. Roy. Soc. B. 364, 3191–3207
Brady S.G., Sipes S., Pearson A., Danforth B.N. (2006) Recent and simultaneous origins of eusociality in halictid bees. Proc. R. Soc. B. 273, 1643–1649
Brand N., Chapuisat, M. (2012) Born to be bee, fed to be worker? The caste system of a primitively eusocial insect. Front. Zool. 9 (35), 1–9
Branstetter M.G., Danforth B.N., Pitts J.P., Faircloth B.C., Ward P.S., Buffington M.L., Gates M.W., Kula R.R., Brady S.G. (2017) Phylogenomic insights into the evolution of stinging wasps and the origins of ants and bees. Curr. Biol. 27, 1019–1025
Cardinal S., Danforth, B.N. (2011) The antiquity and evolutionary history of social behavior in bees. PLOS One 6 (6), e21086
Carroll S.B. (2005) Endless forms most beautiful. W. W. Norton
Carroll S.B. (2008) Evo-devo and an expanding evolutionary synthesis: a genetic theory of morphological evolution. Cell 134, 25–36
Černá K., Zemenová M., Macháčková L., Kolínová Z., Straka J. (2013) Neighborhood society: nesting dynamics, usurpations and social behaviour in solitary bees. PLOS One 8 (8), e73806
Chandrasekaran S., Ament S.A., Eddy J.A., Rodriguez-Zas S.L., Schatz B.R., Price N.D., Robinson G.E. (2011) Behavior-specific changes in transcriptional modules lead to distinct and predictable neurogenomic states. Proc. Natl. Acad. Sci. U.S.A. 108 (44), 18020–18025
Chen S., Krinsky B.H., Long M. (2013) New genes as drivers of phenotypic evolution. Nature Rev. Genet. 14, 645–660
Conte, Y.L., Hefetz, A. (2008) Primer Pheromones in Social Hymenoptera. Annu. Rev. Entomol. 53 (1), 523–542
Corona M., Libbrecht R., Wheeler D.E. (2016) Molecular mechanisms of phenotypic plasticity in social insects. Curr. Opin. Insect Sci. 13, 55–60
Craik, F.I.M. & Lockhart, R.S. (1972). Levels of processing: A framework for memory research. J. Verbal Learn. Verbal Behav., 11(6), 671-684. doi:10.1016/S0022-5371(72)80001-X
Danforth B.N. (2002) Evolution of sociality in a primitively eusocial lineage of bees. PNAS. 99 (1), 286–290
Danforth B.N., Eickwort G.C. (1997) The evolution of social behavior in the augochlorine sweat bees (Hymenoptera: Halictidae) based on a phylogenetic analysis of the genera. in: Choe J. C., Crespi B. J. (Eds.) The Evolution of Social Behavior in Insects and Arachnids. Cambridge University Press, Cambridge. pp. 270–292
Danforth B.N., Conway L., Shuqing J. (2003) Phylogeny of eusocial Lasioglossum reveals multiple losses of eusociality within a primitively eusocial clade of bees (Hymenoptera: Halictidae). Syst. Biol. 52 (1), 23–36
Danforth B.N., Cardinal S., Praz C., Almedia E.A.B., Michez D. (2013) The impact of molecular data on our understanding of bee phylogeny and evolution. Annu. Rev. Entomol. 58, 57–78
Davison P.J., Field J. (2016) Social polymorphism in the sweat bee Lasioglossum (Evylaeus) calceatum. Insectes Soc. 63, 327–338
Dew R.M., Tierney S.M., Schwarz M.P. (2016) Social evolution and casteless societies: needs for new terminology and a new evolutionary focus. Insectes Soc. 63, 5–14
Dolezal A.G., Toth A.L. (2014) Honey bee sociogenomics: a genome-scale perspective on bee social behavior and health. Apidologie 45, 375–395
Eickwort G.C. (1986) First steps into eusociality: the sweat bee Dialictus lineatulus. Fla. Entomol. 69, 742–754
Eickwort G.C., Eickwort J.M., Gordon J., Eickwort M.A. (1996) Solitary behavior in a high-altitude population of the social sweat bee Halictus rubicundus (Hymenoptera: Halictidae). Behav. Ecol. Sociobiol. 38 (4), 227–233
Emlen D.J., Nijhout H.F. (2000) The development and evolution of exaggerated morphologies in insects. Annu. Rev. Entomol. 45, 661–708
Evans H.E. (1977) Commentary: extrinsic versus intrinsic factors in the evolution of insect sociality. Bioscience 27 (9), 613–617
Evans J.D., Wheeler D.E. (2001) Gene expression and the evolution of insect polyphenisms. Bio Essays 23, 62–68
Feldmeyer B., Elsner D., Foitzik S (2014) Gene expression patterns associated with caste and reproductive status in ants: worker-specific genes are more derived than queen-specific ones. Mol. Ecol. 23, 151–161
Ferreira P.G., Patalano S., Chauhan R., Ffrench-Constant R., Gabaldón T., Guigó R., Sumner S. (2013) Transcriptome analyses of primitively eusocial wasp reveal novel insights into the evolution of sociality and the origin of alternative phenotypes. Genome Biol. 14 (2), 1–14
Field J., Paxton R.J., Soro A., Bridge C. (2010) Cryptic plasticity underlies a major evolutionary transition. Curr. Biol. 20 (22), 2028–2031
Fischman B.J., Woodard S.H., Robinson G.E. (2011) Molecular evolutionary analyses of insect societies. Proc. Natl. Acad. Sci. U.S.A. 108 (Supplement 2), 10847–10854
Fjerdingstad E.J., Crozier R.H. (2006) The evolution of worker caste diversity in social insects. Am. Nat. 167 (3), 390–400
Foret S., Kucharski R., Pellegrini M., Feng S., Jacobsen S.E., Robinson G.E., Maleszka R. (2012) DNA methylation dynamics, metabolic fluxes, gene splicing, in alternative phenotypes in honey bees. Proc. Natl. Acad. Sci. U.S.A. 109 (13), 4968–4973
Fu F., Kocher S.D., Nowak M.A. (2015) The risk-return trade-off between solitary and eusocial reproduction. Ecol. Lett. 18, 74–84
Gadagkar R. (1997) The evolution of caste polymorphism in social insects: genetic release followed by diversifying evolution. J. Genet. 76 (3), 167–179
Gibbs J., Brady S.G., Kanda K., Danforth B.N. (2012) Phylogeny of halictine bees supports a shared origin of eusociality for Halictus and Lasioglossum (Apoidea: Anthophila: Halictidae). Mol. Phylogenet. Evol. 65, 926–939
Gilbert S.F. (2012) Ecological developmental biology: environmental signals for normal animal development. Evol. Dev. 14 (1), 20–28
Gilbert S.F., Bosch T. C.G., Ledón-Rettig C. (2015) Eco-evo-devo: developmental symbiosis and developmental plasticity as evolutionary agents. Nature Rev. Genet. 16, 611–622
Glastad K.M., Hunt B.G., Yi S.V., Goodisman M.A.D. (2011) DNA methylation in insects: on the brink of the epigenetic era. Insect Mol. Biol. 20 (5), 553–565
Glastad, K.M., Arsenault, S.V., Vertacnik, K.L., Geib, S.M., Kay, S., Danforth, B.N., Rehan, S. M., Linnen, C.R., Kocher, S.D., Hunt, B.G. (2017) Variation in DNA methylation is not consistently reflected by sociality in Hymenoptera. Genome Biol. Evol. doi:10.1093/gbe/evx128
Goldberg A.D., Allis C.D., Bernstein E. (2007) Epigenetics: a landscape takes shape. Cell 128, 635–638
Grozinger, C.M., Sharabash, N.M., Whitfield, C.W., Robinson, G.E. (2003) Pheromone-mediated gene expression in the honey bee brain. Proc. Natl. Acad. Sci. 100 (Supplement 2), 14519–14525
Harpur B.A., Zayed A. (2013) Accelerated evolution of innate immunity proteins in social insects: adaptive evolution or relaxed constraint? Mol. Biol. Evol. 30 (7), 1665–1674
Harpur B.A., Kent C.F., Molodtsova D., Lebon J.M.D., Alqarni A.S., Owayss A.A., Zayed A. (2014) Population genomics of the honey bee reveals strong signatures of positive selection on worker traits. Proc. Natl. Acad. Sci. U.S.A. 111 (7), 2614–2619
Hirata M., Higashi S. (2008) Degree-day accumulation controlling allopatric and sympatric variations in the sociality of sweat bees, Lasioglossum (Evylaeus) baleicum (Hymenoptera: Halictidae). Behav. Ecol. Sociobiol. 62, 1239–1247
Holbrook C.T., Clark R.M., Jeanson R., Bertram S.M., Kukuk P.F., Fewell J.H. (2009) Emergence and consequences of division of labor in association of normally solitary sweat bees. Ethology 115, 301–310
Honeybee Genome Sequencing Consortium (2006) Insights into social insects from the genome of the honeybee Apis mellifera. Nature 443, 931–949
Hunt J.H., Wolschin F., Henshaw M.T., Newman T.C., Toth A.L., Amdam G.V. (2010) Differential gene expression and protein abundance evince otogenetic bias toward castes in a primitively eusocial wasp. PLoS One 5 (5), e10674
Hunt B.G., Glastad K.M., Yi S.V., Goodisman M.A.D. (2013) The function of intragenic DNA methylation: insights from insect epigenomes. Integr. Comp. Biol. 53 (2), 319–328
Jeanson R., Kukuk P.F., Fewell J.H. (2005) Emergence of division of labour in halictine bees: contributions of social interactions and behavioural variance. Anim. Behav. 70, 1183–1193
Jeanson R., Clark R., Holbrook C., Bertram S., Fewell J., Kukuk P. (2008) Division of labour and socially induced changes in response thresholds in associations of solitary halictine bees. Anim. Behav. 76 (3), 593–602
Johnson B.R., Linksvayer T.A. (2010) Deconstructing the superorganism: social physiology, groundplans, and sociogenomics. Q. Rev. Biol. 85 (1), 57–79
Johnson B.R., Tsutsui N.D. (2011) Taxonomically restricted genes are associated with the evolution of sociality in the honey bee. BMC Genomics. 12 (164), 1–10
Jones B.M., Wcislo W.T., Robinson G.E. (2015) Developmental transcriptome for a facultatively eusocial bee, Megalopta genalis G3 Genses Genom. Genet. 5, 2127–2135
Jones B.M., Kingwell C.J., Wcislo W.T., Robinson G,E. (2017) Caste-biased gene expression in a facultatively eusocial bee suggests a role for genetic accommodation in the evolution of eusociality. Proc. R. Soc. B. 284 (1846), 1–9
Kapheim K.M. (2016) Genomic sources of phenotypic novelty in the evolution of eusociality in insects. Curr. Opin. Insect Sci. 13, 24–32
Kapheim K.M., Smith A.R., Nonacs P., Wcislo W.T., Wayne R.K. (2013) Foundress polyphenism and the origins of eusociality in a facultatively eusocial sweat be, Megalopta genalis (Halictidae). Behav. Ecol. Sociobiol. 67, 331–340
Kapheim K.M., Pan H., Li C., Salzber S.L., Puiu D., et al. (2015) Genomic signatures of evolutionary transitions from solitary to group living. Science. 348 (6239), 1139–1143
Kocher S.D., Paxton R. (2014) Comparative methods offer powerful insights into social evolution in bees. Apidologie 45, 289–305
Kocher S.D., Li C., Yang W., Tan H., Yi S.V., et al. (2013) The draft genome of a socially polymorphic halictid bee, Lasioglossum albipes. Genome Biol. 14 (12), 1–14
Kocher S.D, Pellisier L., Veller C., Purcell J., Nowak M.A. et al. (2014) Transitions in social complexity along elevational gradients reveal a combined impact on season length and development time on social evolution. Proc. R. Soc. B. 281, 1–8
Lawson S.P., Ciaccio K.N., Rehan S.M. (2016) Maternal manipulation of pollen provisions affects worker production in a small carpenter bee. Behav. Ecol. Sociobiol. 70 (11), 1891–1900
Liang Z.S., Nguyen T., Mattila H.R., Rodriguez-Zas S.L., Seeley T.D., Robinson G.E. (2012) Molecular determinants of scouting behavior in honey bees. Science 335, 1225–1228
Libbrecht, R., Oxley, P.R., Keller, L., Kronauer, D.J.C. (2016) Robust DNA Methylation in the Clonal Raider Ant Brain. Curr. Biol. 26 (3), 391–395
Li-Byarlay H., Li Y., Stroud H., Feng S., Newman T.C. (2013) RNA interference knockdown of DNA methyl-transferase 3 affects gene alternative splicing in the honey bee. Proc. Natl. Acad. Sci. U.S.A. 110 (31), 12750–12755
Lin N., Michener C.D. (1972) Evolution of sociality in insects. Q. Rev. Biol. 47 (2), 131–159
Linksvayer T.A., Wade M.J. (2005) The evolutionary origin and elaboration of sociality in the aculeate Hymenoptera: maternal effects, sib-social effects, and heterochrony. Q. Rev. Biol. 80 (3), 317–336
Luscombe, N.M., Madan Babu, M., Yu, H., Snyder, M., Teichmann, S.A, Gerstein, M. (2004) Genomic analysis of regulatory network dynamics reveals large topological changes. Nature 431 (7006), 308–312
Lyko F., Foret S., Kucharski R., Wolf S., Falckenhayn C., Maleszka R. (2010) The honey bee epigenomes: differential methylation of brain DNA in queens and workers. PLoS Biol. 8 (11), e1000506
McKenzie S.K., Fetter-Pruneda I., Ruta V., Kronauer D.J.C. (2016) Transcriptomics and neuroanatomy of the clonal raider ant implicate an expanded clade of odorant receptors in chemical communication. P. Natl. Aca. Sci. USA. DOI:10.1073/pnas.1610800113
Michener C.D. (1969) Comparative social behavior of bees. Annu. Rev. Entomol. 14, 299–342
Michener C.D. (1974) The social behaviour of the bees: a comparative study. Harvard University Press, Cambridge, MA
Michener C.D. (2007) The bees of the world, 2nd edn. Baltimore, MD: John Hopkins University Press.
Miyanaga R., Maeta Y., Sakagami S.F. (1999) Geographical variation of sociality and size-linked color patterns in Lasioglossum (Evylaeus) apristum (Vachal) in Japan (Hymenoptera, Halictidae). Insectes Soc. 46, 224–232
Molodtsova D., Harpur B.A., Kent C.F., Seevananthan K., Zayed A. (2014) Pleiotropy constrains the evolution of protein but not regulatory sequences in a transcription regulatory network influencing complex social behaviors. Front. Genet. 5 (431), 1–7
Morandin C., Dhaygude K., Paviala J., Trontti K., Wheat C., Helanterä H. (2015) Caste-biases in gene expression are specific to developmental stage in the ant Formica exsecta. J. Evol. Biol. 28, 1705–1718
Morandin C., Tin M.M.Y., Abril S., Gómez C., Pontieri L., et al. (2016) Comparative transcriptomics reveals the conserved building blocks involved in parallel evolution of diverse phenotypic traits in ants. Genome Biol. 17 (43), 1–19
Nijhout H.F. (2003) Development and evolution of adaptive polyphenisms. Evol. Dev. 5 (1), 9–18
Nowak M.A., Tarnita C.E., Wilson E.O. (2010) The evolution of eusociality. Nature 466, 1057–1062
Olejarz J.W., Allen B., Veller C., Gadagkar R., Nowak M.A. (2016) Evolution of worker policing. J. Theor. Biol. 339, 103–116
Page Jr. R.E., Amdam G.V. (2007) The making of social insect: development architectures of social design. Bio Essays 29 (4), 334–343
Patalano S., Vlasova A., Wyatt C., Ewels P., Camara F., et al. (2015) Molecular signatures of plastic phenotypes in two eusocial insect species with simple societies. Proc. Natl. Acad. Sci. U.S.A. 112 (45), 13970–13975
Peters R.S., Krogmann L., Mayer C., Donath A., Gunkel S., et al. (2017) Evolutionary history of the Hymenoptera. Curr. Biol. 27, 1–6
Plateaux-Quénu C. (2008) Subsociality in halictine bees. Insectes Soc. 55, 335–346
Plateaux-Quénu C., Plateaux I., Packer L. (2000) Population-typical behaviours are retained when eusocial and non-eusocial forms of Evylaeus albipes (F.) (Hymenoptera, Halictidae) are reared simultaneously in the laboratory. Insectes Soc. 47, 263–270
Purcell J. (2011) Geographic patterns in the distribution of social systems in terrestrial arthropods. Biol. Rev. 86, 475–491
Rehan, S.M., Richards, M.H. (2010a) Nesting biology and subsociality in Ceratina calcarata (Hymenoptera: Apidae). Can. Entomol. 142 (1), 65–74
Rehan S.M., Richards M.H. (2010b) The influence of maternal quality on brood sex allocation in the small carpenter bee, Ceratina calcarata. Ethology 116 (9), 876–887
Rehan S.M., Richards M.H. (2013) Reproductive aggression and nestmate recognition in a subsocial bee. Anim. Behav. 85 (4), 733–741
Rehan S.M., Schwarz M.P. (2015) A few steps forward and no steps back: long-distance dispersal patterns in small carpenter bees suggest major barriers to back-dispersal. J. Biogeogr. 42 (3), 485–494
Rehan S.M., Toth A.L. (2015) Climbing the social ladder: the molecular evolution of sociality. Trends Ecol. Evol. 30 (7), 426–433
Rehan S.M., Richards M.H., Schwarz M.P. (2009) Evidence of social nesting in Ceratina of Borneo. J. Kans. Entomol. Soc. 82, 194–209
Rehan S.M., Chapman T.W., Craigie A., Richards M.H., Cooper S.J., Schwarz M.P. (2010a) Molecular phylogeny of the small carpenter bees (Hymenoptera: Apidae: Ceratinini) indicates early and rapid global dispersal. Mol. Phylogenet. Evol. 55, 1042–1054
Rehan S.M., Richards M.H., Schwarz M.P. (2010b) Social polymorphism in the Australian small carpenter bee, Ceratina (Neoceratina) australensis. Insectes Soc. 57 (4), 403–412
Rehan S.M., Schwarz M.P., Richards M.H. (2011) Fitness consequences of ecological constraints and implications for the evolution of sociality in an incipiently social bee. Biol. J. Linnean Soc. 103 (1), 57–67
Rehan S.M., Leys R., Schwarz M.P. (2012) A mid-Cretaceous origin of sociality in Xylocopine bees with only two origins of true worker castes indicates severe barriers to eusociality. PLoS ONE 7 (4), e34690
Rehan S.M., Berens A.J., Toth A.L. (2014a) At the brink of eusociality: transcriptomic correlates of worker behaviour in a small carpenter bee. BMC Evol. Biol. 14 (260), 1–10
Rehan, S.M., Richards, M.H., Adams, M., Schwarz, M.P. (2014b) The costs and benefits of sociality in a facultatively social bee. Anim. Behav. 97, 77–85
Rehan S.M., Tierney S.M., Wcislo W.T. (2015) Evidence for social nesting in Neotropical creatinine bees. Insectes Soc. 62 (2), 465–469
Rehan S.M., Glastad K.M., Lawson S.P., Hunt B.G. (2016) The genome and methylome of a subsocial small carpenter bee, Ceratina calcarata. Genome Biol. Evol. 8 (5), 1401–1410
Richards, M.H. (2000) Evidence for geographic variation in colony social organization in an obligately social sweat bee, Lasioglossum malachurum Kirby (Hymenoptera; Halictidae). Can. J. Zool. 78, 1259–1266
Richards, M.H. (2001) Nesting biology and social organization of Halictus sexcinctus (Fabricius) in southern Greece. Can. J. Zool. 79 (12), 2210–2220
Richards M.H., Packer L. (1996) The socioecology of body size variation in the primitively eusocial sweat bee, Halictus ligatus (Hymenoptera: Halictidae). Oikos 77 (1), 68–76
Rittschof C.C., Bukhari S.A., Sloofman L.G., Troy J.M., Caetano-Anollés D. et al. (2014) Neuromolecular responses to social challenge: common mechanisms across mouse, stickleback fish, and honey bee. Proc. Natl. Acad. Sci. U.S.A. 111 (50), 17929–17934
Robinson G.E. (2002) Sociogenomics takes flight. Science 297 (5579), 204–205
Robinson G.E., Grozinger C.M., Whitfield C.W. (2005) Sociogenomics: social life in molecular terms. Nat. Rev. Genet. 6 (4), 257–270
Romiguier J., Lourenco J., Gayral P., Faivre N., Weinert L.A., et al. (2014) Population genomics of eusocial insects: the costs of a vertebrate-like effective population size. J. Evol. Biol 27, 593–603
Romiguier J., Cameron S.A., Woodard S.H., Fischman B.J., Keller L., Praz C.J. (2015) Phylogenomics controlling for base compositional bias reveals a single origin of eusociality in corbiculate bees. Mol. Biol. Ecol. 33 (3), 670–678
Sadd M., Barribeau S.M., Bloch G., de Graaf D.C., Dearden P., et al. (2015) The genomes of two key bumblebee species with primitive eusocial organization. Genome Biol. 16 (1), 1–31
Sakagami S.F., Maeta Y. (1984) Multifemale nests and rudimentary castes in the normally solitary bee Ceratina japonica (Hymenoptera: Xylocopinae) J. Kans. Entomol. Soc. 57 (4), 639–656
Sakagami S.F., Maeta Y. (1987) Sociality, induced and/or natural, in the basically solitary small carpenter bees (Ceratina). Anim. Soc. Theor. Fac. 1-16
Sakagami S.F., Maeta Y. (1989) Compatibility and incompatibility of solitary life with eusociality in two normally solitary bees Ceratina japonica and Ceratina okinawana (Hymenoptera: Apoidea), with notes on the incipient phase of eusociality. Jpn. J. Ent. 57 (2), 417–439
Sakagami S.F., Maeta Y. (1995) Task allocation in artificially induced colonies of a basically solitary bee Ceratina (Ceratinidia) okinawana, with a comparison of sociality between Ceratina and Xylocopa (Hymenoptera, Anthophoridae, Xylocopinae). Jpn. J. Ent. 63 (1), 115–150
Sakagami, S.F., Munakata M. (1972) Distribution and bionomics of a transpalaearctic eusocial Halictine bee, Lasioglossum (Evylaeus) calceatum, in northern Japan, with reference to its solitary life cycle at high altitude. J. Fac. Sci. Hokkaido Univ. Ser. VI. Zool. 18 (3), 411–439
Schlichting C.D., Wund M.A. (2014) Phenotypic plasticity and epigenetic marking: an assessment of evidence for genetic accommodation. Evolution 68 (3), 656–672
Schultz T.R. (2000) In search of ant ancestors. Proc. Natl. Acad. Sci. U.S.A. 97 (26), 14028–14029
Schwarz, M.P., Richards, M.H., Danforth B.N. (2007) Changing paradigms in insect social evolution: insights from Halictine and Allodapine bees. Annu. Rev. Entomol. 52, 127–150
Schwarz, M.P., Tierney, S.M., Rehan, S.M., Chenoweth, L.B., Cooper, S.J.B. (2011) The evolution of eusociality in allodapine bees: workers began by waiting. Biol. Lett. 7 (2), 277–280
Seger J. (1983) Partial bivoltinism may cause alternating sex-ratio biases that favour eusociality. Nature 301, 59–62
Shell W.A., Rehan S.M. (2016) Recent and rapid diversification of the small carpenter bees in eastern North America. Biol. J. Linnean Soc. 117, 633–645
Simola D.F., Wissler L., Donahue G., Waterhouse R.M., Helmkampf M., et al. (2013) Social insect genomes exhibit dramatic evolution in gene composition and regulation while preserving regulatory features linked to sociality. Genome Res. 23, 1235–1247
Simpson S.J., Sword G.A., Lo N. (2011) Polyphenism in insects. Curr. Biol. 21, 738–749
Smith A.R., Wcislo W.T., O’Donnell S. (2003) Assured fitness returns favor sociality in a mass-provisioning sweat bee, Megalopta genalis (Hymenoptera: Halictidae). Behav. Ecol. Sociobiol. 54, 14–21
Smith C.R., Toth A.L., Suarez A.V., Robinson G.E. (2008) Genetic and genomic analyses of the division of labour in insect societies. Nature Rev. Genet. 9, 735–748
Soro A., Field J., Bridge C., Cardinal S.C., Paxton R.J. (2010) Genetic differentiation across the social transition in a socially polymorphic sweat bee, Halictus rubicundus. Mol. Ecol. 19, 3351–3363
Soucy S.L. (2002) Nesting biology and socially polymorphic behavior of the sweat bee Halictus rubicundus (Hymenoptera: Halictidae). Ann. Entomol. Soc. Am. 95 (1), 57–65
Soucy S.L., Danforth B.N. (2002) Phylogeography of the socially polymorphic sweat bee Halictus rubicundus (Hymenoptera: Halictidae). Evolution 56 (2), 330–341
Southey B.R., Zhu P., Carr-Markell M.K., Liang Z.S., Zayed A., et al. (2016) Characterization of genomic variants associated with scout and recruit behavioral castes in honey bees using whole-genome sequencing. PLoS One 11 (1), e0146430
Standage D.S., Berens A.J., Glastad K.M., Severin A.J. (2016) Genome, transcriptome and methylome sequencing of a primitively eusocial wasp reveal a greatly reduced DNA methylation system in a social insect. Mol. Ecol. 25, 1769–1784
Strassman J.E., Queller D.C. (1989) Ecological determinants of social evolution, in: Breed, M. D., Page, R. E. (Eds.) The genetics of social evolution. Westview Press, Boulder Colorado pp. 81–101
Sumner S. (2014) The importance of genomic novelty in social evolution. Mol. Ecol. 23, 26–28
Suzuki Y., Nijhout H.F. (2006) Evolution of a polyphenism by genetic accommodation. Science 311, 650–652
Toth A.L., Rehan S.M. (2017) Molecular evolution of insect sociality: an eco-evo-devo perspective. Annu. Rev. Entomol. 62, 419–442
Toth A.L., Robinson G.E. (2007) Evo-devo and the evolution of social behavior. Trends Genet. 23 (7), 334–341
Toth A.L., Robinson G.E. (2009) Evo-devo and the evolution of social behavior: brain gene expression analyses in social insects. Cold Spring Harb. Symp. Quant. Biol. 74, 419–426
Toth A.L., Varala K., Newman T.C., Miguez F.E., Hutchison S.K., et al. (2007) Wasp gene expression supports an evolutionary link between maternal behavior and eusociality. Science 318 (5849), 441–444
Toth, A.L., Varala, K., Henshaw, M.T., Rodriguez-Zas, S.L., Hudson, M.E., Robinson, G.E. (2010) Brain transcriptomic analysis in paper wasps identifies genes associated with behaviour across social insect lineages. Proc. R. Soc. B Biol. Sci. 277 (1691), 2139–2148
Toth, A.L., Tooker, J.F., Radhakrishnan, S., Minard, R., Henshaw, M.T., Grozinger, C.M. (2014) Shared genes related to aggression, rather than chemical communication, are associated with reproductive dominance in paper wasps (Polistes metricus). BMC Genomics 15 (1), 75
True J.R., Carroll S.B. (2002) Gene co-option in physiological and morphological evolution. Annu. Rev. Cell. Dev. Biol. 18, 53–80
Waddington C.H. (1961) Genetic assimilation. Adv. Genet. 10, 257–293
Wade M.J. (2001) Maternal effect genes and the evolution of sociality in haplo-diploid organisms. Evolution 55 (3), 453–458
Wcislo W.T., Cane J.H. (1996) Floral resource utilization by solitary bees (Hymenoptera: Apoidea) and exploitation of their stored foods by natural enemies. Annu. Rev. Entomol. 41, 257–286
Wcislo W.T., Danforth B.N. (1997) Secondarily solitary: the evolutionary loss of social behavior. Trends Ecol. Evolut. 12 (12), 468–474
Wcislo W.T., Arneson L., Roesch K., Gonzalez V., Smith A., Fernández H. (2004) The evolution of nocturnal behaviour in sweat bees, Megalopta genalis and M. ecuadoria (Hymenoptera: Halictidae): an escape from competitors and enemies? Biol. J. Linnean Soc. 83, 377–387
Weiner S.A., Toth A.L. (2012) Epigenetics in social insects: a new direction for understanding the evolution of castes. Genet. Res. Int. 2012, 1–11
West-Eberhard M.J. (1987) Flexible strategy and social evolution. Anim. Soc. Theor. Fac. 35-51
West-Eberhard M.J. (1989) Phenotypic plasticity and the origins of diversity. Annu. Rev. Ecol. Syst. 20, 249–278
West-Eberhard M.J. (1996) Wasp societies as microcosms for the study of development and evolution, in: Turillazzi, S. & West-Eberhard, M. J. (Eds.) Natural history and evolution of paper-wasps. Oxford University Press, Oxofrd, pp. 291–317
West-Eberhard M.J. (2003) Developmental Plasticity and Evolution. Oxford University of Press, Oxford
Whitman D.W., Agrawal A.A. (2009) What is phenotypic plasticity and why is it important? In: Whitman, D. W., Ananthakrishnan, T. N. (Eds.) Phenotypic plasticity of insects: mechanisms and consequences pp. 1–63
Wild G., Koykka C. (2014) Inclusive-fitness logic of cooperative breeding with benefits of natal philopatry. Phil. Trans. R. Soc. B. 369, 1–9
Wilson E.O. (1971) The Insect Societies. Belknap Press of Harvard University Press, Cambridge, Massachusetts
Wilson E.O., Hölldobler B. (2005) Eusociality: origin and consequences. P. Natl. Aca. Sci. USA. 102, 13367–13371
Woodard S.H., Fischman B.J., Venkat A., Hudson M.E., Varala K., et al. (2011) Genes involved in convergent evolution of sociality in bees. Proc. Natl. Acad. Sci. U.S.A. 108 (18), 7472–7477
Woodard S.H., Bloch G.M., Band M.R., Robinson G.E. (2014) Molecular heterochrony and the evolution of sociality in bumblebees (Bombus terrestris). Proc. R. Soc. B. 281, 1–8
Wyman L.M., Richards M.H. (2003) Colony social organization of Lasioglossum malachurum Kirby (Hymenoptera, Halictidae) in southern Greece. Insectes Soc. 50, 201–211
Yan H., Bonasio R., Simola D.F., Liebig J., Berger S.L., Reinberg D. (2015) DNA methylation in social insects: how epigenetics can control behavior and longevity. Annu. Rev. Entomol. 60, 435–452
Yanega D. (1989) Caste determination and differential diapause within the first brood of Halictus rubicundus in New York (Hymenoptera: Halictidae). Behav. Ecol. Sociobiol. 24, 97–107
Yanega D. (1990) Philopatry and nest founding in a primitively social bee, Halictus rubicundus. Behav. Ecol. Sociobiol. 27, 37–42
Yanega D. (1993) Environmental influences on male production and social structure in Halictus rubicundus (Hymenoptera: Halictidae). Insectes Soc. 40, 169–180
Acknowledgements
We thank two anonymous reviewers and members of the Rehan lab for constructive feedback on this manuscript. Funding from the University of New Hampshire, NSF-GRF no. 1450271, and NSF-IOS, Behavioral Systems Award no. IOS-1456296, supported this work.
Contributions
WAS and SMR wrote the paper. All authors read and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Manuscript editor: Klaus Hartfelder
Mécanismes comportemental et génétique de l’évolution sociale: indications fournies par des abeilles au comportement social naissant et facultatif
Evolution sociale / abeille charpentière / abeille à sueur / sociogénomique / comportement social primitif / comportement social facultatif / premier niveau de sociabilité / plasticité phénotypique
Verhaltens- und genetische Mechanismen der sozialen Evolution: Einsichten aus der Sicht von Biene mit beginnendem und fakultativem Sozialverhalten
soziale Evolution / Keulhornbienen / Furchenbienen / Soziogenomik / primitiv sozial / fakultativ / beginnende Sozialiät / phänotypische Plastizität
This article was corrected due retrospective Open Access. The erratum is available under doi:10.1007/s13592-017-0544-0
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Shell, W.A., Rehan, S.M. Behavioral and genetic mechanisms of social evolution: insights from incipiently and facultatively social bees. Apidologie 49, 13–30 (2018). https://doi.org/10.1007/s13592-017-0527-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13592-017-0527-1