Abstract
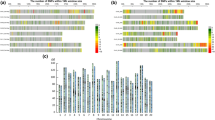
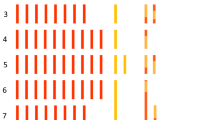
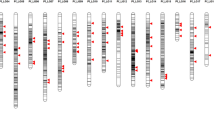
Traditional breeding methods usually involve field tests carried out by experienced breeders. However, such methods are costly and time-consuming. Recently, with the development of Next-Generation Sequencing (NGS) technology, molecular markers are being utilized for selection processes in breeding. To implement a high-throughput system using molecular markers in Chinese cabbage (Brassica rapa subsp. pekinensis) breeding, we developed single nucleotide polymorphism (SNP) marker sets for background selection and testing F1 purity using Fluidigm genotyping assays. SNPs were generated using NGS technology on 209 varieties of Chinese cabbage collected from around the world. Those with minor allele frequency ≥ 5% and polymorphism information content ≥ 0.3 were screened, and then based on the physical distribution among the 10 chromosomes, 177 SNPs were selected and synthesized for testing. To obtain marker sets with high selection efficiency, we tested 192 SNPs on 45 types of inbred lines and 29 types of F1 hybrids. Among the 192 SNPs, we selected 96 markers sets for background selection and 24 marker sets for F1 purity testing according to the following criteria; the genotype of the parents was homozygous, and the F1 follows the parents’ genotypes. These SNP sets are suitable for high-throughput systems using the 96.96 and 192.24 integrated fluidic circuit platforms of Fluidigm genotyping assays. These SNP marker sets are not only efficient for selecting of early fixed lines as background selection but are also useful for testing the purity of F1 hybrids.


Similar content being viewed by others
References
An SJ, Kwon JK, Yang HB, Choi HJ, Jeong HJ, Kim YJ, Choi GJ, Kang BC (2010) SNP marker development for purity test of oriental melon and melon. Korean J Breed Sci 42:397–406
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314
Cheng F, Liu S, Wu J, Fang L, Sun S, Liu B, Li P, Hua W, Wang X (2011) BRAD, the genetics and genomics database for Brassica plants. BMC Plant Biol 11:136. https://doi.org/10.1186/1471-2229-11-136
Collard BCY, Mackill DJ (2008) Marker-assisted selection: an approach for precision plant breeding in the twenty-first century. Philos Trans R Soc Lond B Biol Sci 363:557–572. https://doi.org/10.1098/rstb.2007.2170
Davey JW, Hohenlohe PA, Etter PD, Boone JQ, Catchen JM, Blaxter ML (2011) Genome-wide genetic marker discovery and genotyping using next-generation sequencing. Nat Rev Genet 12:499–510. https://doi.org/10.1038/nrg3012
Duran C, Appleby N, Clark T, Wood D, Imelfort M, Batley J, Edwards D (2009) AutoSNPdb: an annotated single nucleotide polymorphism database for crop plants. Nucleic Acids Res 37:D951–953. https://doi.org/10.1093/nar/gkn650
Edwards D, Batley J (2010) Plant genome sequencing: applications for crop improvement. Plant Biotechnol J 8:2–9. https://doi.org/10.1111/j.1467-7652.2009.00459.x
Frisch M, Melchinger AE (2005) Selection theory for marker-assisted backcrossing. Genetics 170:909–917. https://doi.org/10.1534/genetics.104.035451
Hasan MM, Rafii MY, Ismail MR, Mahmood M, Rahim HA, Alam MA, Ashkani S, Malek MA, Latif MA (2015) Marker-assisted backcrossing: a useful method for rice improvement. Biotechnol Biotechnol Equip 29:237–254. https://doi.org/10.1080/13102818.2014.995920
Jang I, Moon JH, Yoon JB, Yoo JH, Yang TJ, Kim YJ, Park HG (2004) Application of RAPD and SCAR markers for purity testing of F1 hybrid seed in chili pepper (Capsicum annuum). Mol Cells 18:295–299
Jeong HS, Jang S, Han K, Kwon JK, Kang BC (2015) Marker-assisted backcross breeding for development of pepper varieties (Capsicum annuum) containing capsinoids. Mol Breed 35:226
Jiang GL (2013) Molecular markers and marker-assisted breeding in plants. In: Andersen SB (ed) Plant breeding from laboratories to fields. InTech, Rijeka, Croatia, pp 45–83
Kawamura K, Saeki N, Ebe Y, Abe H, Shimizu M, Osabe K, Okazaki K, Kawanabe T, Fujimoto R (2014) Screening of DNA markers suitable for purity test of inbred lines in Brassica rapa. Bull Fac Agric Niigata Univ 66:111–124
Kim J, Kim D-S, Park S, Lee H-E, Ahn Y-K, Kim JH, Yang H-B, Kang B-C (2016) Development of a high-throughput SNP marker set by transcriptome sequencing to accelerate genetic background selection in Brassica rapa. Hortic Environ Biotechnol 57:280–290
Kim H, Yoon JB, Lee J (2017a) Development of fluidigm SNP type genotyping assays for marker-assisted breeding of chili pepper (Capsicum annuum L.). Hortic Sci Technol 35:465–479. https://doi.org/10.12972/kjhst.20170050
Kim YY, Oh SH, Pang WX, Li XN, Ji SJ, Son E, Han S, Park S, Soh E et al (2017b) A review of the scientific names of chinese cabbage according to the international codes of nomenclature. Hortic Sci Technol 35:165–169. https://doi.org/10.12972/kjhst.20170019
KOSA (2019) 2018 Export status by vegetable seed crop. Korea Seed Association. Available via http://www.kosaseed.or.kr/bbs/board.php?bo_table=seed_out&wr_id=53. Accessed 22 Jan 2019
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25:1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Li X, Ramchiary N, Choi SR, Van Nguyen D, Hossain MJ, Yang HK, Lim YP (2010) Development of a high density integrated reference genetic linkage map for the multinational Brassica rapa Genome Sequencing Project. Genome 53:939–947. https://doi.org/10.1139/G10-054
Li X, Ramchiary N, Dhandapani V, Choi SR, Hur Y, Nou IS, Yoon MK, Lim YP (2013) Quantitative trait loci mapping in Brassica rapa revealed the structural and functional conservation of genetic loci governing morphological and yield component traits in the A, B, and C subgenomes of Brassica species. DNA Res 20:1–16. https://doi.org/10.1093/dnares/dss029
Li L, Liu L, Zhang DS, Wu P, Zhang FL, Xu XL (2017) Hybrid purity testing of Brassica rapa using SSR marker technology. HortScience 52:1342–1348. https://doi.org/10.21273/Hortsci12023-17
Liu L, Liu G, Gong Y, Dai W, Wang Y, Yu F, Ren YJH (2007) Evaluation of genetic purity of F1 hybrid seeds in cabbage with RAPD, ISSR, SRAP, and SSR markers. HortScience 42:724–727
Livingstone DS, Motamayor JC, Schnell RJ, Cariaga K, Freeman B, Meerow AW, Brown JS, Kuhn DNJMB (2011) Development of single nucleotide polymorphism markers in Theobroma cacao and comparison to simple sequence repeat markers for genotyping of Cameroon clones. Mol Breed 27:93–106
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S et al (2010) The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303. https://doi.org/10.1101/gr.107524.110
Pang W, Li X, Choi SR, Dhandapani V, Im S, Park MY, Jang CS, Yang MS, Ham IK et al (2015) Development of a leafy Brassica rapa fixed line collection for genetic diversity and population structure analysis. Mol Breed. https://doi.org/10.1007/s11032-015-0221-9
Ramchiary N, Nguyen VD, Li XN, Hong CP, Dhandapani V, Choi SR, Yu G, Piao ZY, Lim YP (2011) Genic microsatellite markers in Brassica rapa: development, characterization, mapping, and their utility in other cultivated and wild brassica relatives. DNA Res 18:305–320. https://doi.org/10.1093/dnares/dsr017
Ribaut JM, Hoisington D (1998) Marker-assisted selection: new tools and strategies. Trends Plant Sci 3:236–239. https://doi.org/10.1016/S1360-1385(98)01240-0
Thomson MJ (2014) High-throughput SNP genotyping to accelerate crop improvement. Plant Breed Biotechnol 2:195–212
Wang J, Lin M, Crenshaw A, Hutchinson A, Hicks B, Yeager M, Berndt S, Huang WY, Hayes RB et al (2009) High-throughput single nucleotide polymorphism genotyping using nanofluidic Dynamic Arrays. BMC Genome. https://doi.org/10.1186/1471-2164-10-561
Ye S, Wang Y, Huang D, Li J, Gong Y, Xu L, Liu LJSH (2013) Genetic purity testing of F1 hybrid seed with molecular markers in cabbage (Brassica oleracea var. capitata). Sci Hortic 155:92–96
Zhang L, Cai X, Wu J, Liu M, Grob S, Cheng F, Liang J, Cai C, Liu Z et al (2018) Improved Brassica rapa reference genome by single-molecule sequencing and chromosome conformation capture technologies. Hortic Res 5:50. https://doi.org/10.1038/s41438-018-0071-9
Acknowledgements
This work was supported by Korea Institute of Planning and Evaluation for Technology in Food, Agriculture, and Forestry (IPET) through Golden Seed Project (213006-05-3-SB110, 213006-05-3-SBD30), funded by Ministry of Agriculture, Food and Rural Affairs (MAFRA), Ministry of Oceans and Fisheries (MOF), Rural Development Administration (RDA) and Korea Forest Services (KFS).
Author information
Authors and Affiliations
Contributions
SRC designed the experiment. SHO, HAK, IJ, and JJR performed the experiment. SRC, SHO, and VD analyzed the data. SRC and SHO wrote the manuscript. CSJ and CHA provided the plant materials. YPL supervised the experiment and provided valuable suggestion.
Corresponding author
Ethics declarations
Conflict of interest
The authors can declare that they have no conflict of interest.
Data availability
All data generated or analyzed during this study are included in this published article [and its supplementary information files].
Additional information
Communicated by Inhwa Yeam.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Choi, S.R., Oh, S.H., Dhandapani, V. et al. Development of SNP markers for marker-assisted breeding in Chinese cabbage using Fluidigm genotyping assays. Hortic. Environ. Biotechnol. 61, 327–338 (2020). https://doi.org/10.1007/s13580-019-00211-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13580-019-00211-y