Abstract
Cancer continues to pose a global threat despite potent anticancer drugs, often accompanied by undesired side effects. To enhance patient outcomes, sophisticated multifunctional approaches are imperative. Small extracellular vesicles (EVs), a diverse family of naturally occurring vesicles derived from cells, offer advantages over synthetic carriers. Among the EVs, the exosomes are facilitating intercellular communication with minimal toxicity, high biocompatibility, and low immunogenicity. Their tissue-specific targeting ability, mediated by surface molecules, enables precise transport of biomolecules to cancer cells. Here, we explore the potential of exosomes as innovative therapeutic agents, including cancer vaccines, and their clinical relevance as biomarkers for clinical diagnosis. We highlight the cargo possibilities, including nucleic acids and drugs, which make them a good delivery system for targeted cancer treatment and contrast agents for disease monitoring. Other general aspects, sources, and the methodology associated with therapeutic cancer applications are also reviewed. Additionally, the challenges associated with translating exosome-based therapies into clinical practice are discussed, together with the future prospects for this innovative approach.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
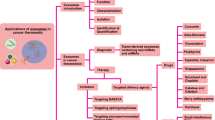
Cancer represents “an abnormal mass of tissue, the growth of which exceeds and is uncoordinated with that of the surrounding normal tissues and persists in the same excessive manner after cessation of the stimuli which caused the change” as described by Willis [1]. This malignancy encompasses a group of diseases characterized by the unregulated multiplication and spread of aberrant cells, often resulting in the formation of tumors or the invasion of other tissues. The perilous potential of these rogue cells to metastasize through the bloodstream or lymphatic system poses significant life-threatening risks [2]. Cancer can affect any part of the body and may arise from various factors, including genetic mutations, environmental exposures, and lifestyle choices. In 2011, Hanahan and Weinberg identified several hallmarks of cancer, including autonomous cell proliferation, insensitivity to antiproliferative signals, evasion of destruction by the immune system, stimulation of inflammation, resistance to cell death, replicative immortality, induction of angiogenesis, activation of tissue invasion and metastasis, reprogramming of energy metabolism, and genomic instability and mutation (Fig. 1A) [3]. These shared characteristics make cancer a formidable disease, driving the quest for innovative anti-cancer treatments.
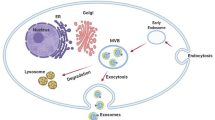
Hallmarks of cancer, timeline and exosome biogenesis. A The hallmarks of cancer. Adapted from Hanahan and Weinberg [3]. B Exosome biogenesis with a size ranging from 30 to 150 nm. Adapted from Colombo et al. [14]. C Timeline of the more representative events in the EV field. Adapted from Couch et al. [12]
The field of cancer nanomedicine seeks alternatives to conventional cancer diagnostics and therapies, accelerating the development of technologies like nanoparticle-based drug delivery systems among others. The growing enthusiasm in this field has led to the study of small extracellular vesicles (EVs), particularly exosomes (30–150 nm) (Fig. 1B), as a promising alternative to synthetic nanoparticles (NPs) [4,5,6].
The term “exosome” was coined by Johnstone et al. to describe that the release of these small EVs during reticulocyte maturation, was leading to the loss of the transferrin receptor [7]. Nowadays, exosomes are defined as nanovesicles secreted by various cell types into the extracellular medium. They are formed within endocytic compartments called multivesicular bodies (MVBs) through inward budding of the membrane during endosome maturation [8]. In 1946, Chargaff and West first reported a “particulate fraction” sedimenting at 31,000 x g, displaying clotting potential like a ‘thromboplastic protein’ [9]. However, the first observations of vesicles surrounding mammalian cells in tissues or fluids came in the late 1960s [10, 11]. Subsequent research described these vesicles initially as “waste disposal” and later recognized their role in cell-to-cell communication [12]. It was not until 2011 that the term “extracellular vesicles” was proposed as a general term encompassing all extracellular structures enclosed by lipid bilayers, coinciding with the establishment of the International Society for Extracellular Vesicles (ISEV) [13] (Fig. 1C).
Exosomes serve as carriers for both water-soluble and non-water-soluble therapeutic agents and possess intrinsic organotrophic and tumor-targeting capabilities [15, 16]. The ability to obtain exosome populations with these intriguing properties paves the way for studying these complex, multifaceted vesicles as biomarkers, therapeutic agents, cancer vaccines or delivery systems while circumventing complicated manufacturing processes. We have discussed the different steps in using exosomes in cancer therapy, from source selection to large-scale manufacturing, to fulfill the exosomes clinical potential. Nonetheless, the main challenges for exosome-based applications in cancer revolve around clearance, limited production and their acceptance for clinical use.
In summary, this review provides a comprehensive and innovative examination of exosome-based cancer therapies. We offer a broader perspective on the role of exosomes in liquid biopsies and their importance in immunotherapy, focusing on the most recent clinical trials that we have selected based on their relevant information on identified biomarkers and as cancer vaccines. We also reviewed recent advances in the use of using exosomes as delivery systems where classical cargo types such as nucleic acids, metabolites and proteins are examined in detail but also new approaches that combine them with nanoparticles, drug or imaging agents.
We discuss classical and novel strategies to address the existing challenges in the field, presenting a state-of-the-art overview that advances the understanding and potential application of exosome-based nanomedicine in cancer therapy. Altogether, we offer a comprehensive exploration of exosome biology to uncover how we can fulfill the great promise of exosome-based nanomedicine for the detection, diagnosis and treatment of cancer patients.
2 Biology of exosomes
The exosomes biogenesis is closely linked to their physiological function and/or the pathological condition of the parent cell [17]. This process involves four main steps: cargo sorting into multivesicular bodies (MVBs), the formation of MVBs, the transport of MVBs, and the fusion of MVBs with the plasma membrane. Among these steps, MVB formation is particularly critical in small EV biogenesis, with a focus on the generation of intraluminal vesicles (ILVs) and membrane budding [18, 19]. Several mechanisms have been proposed for ILVs generation, but the general mechanisms consist of the endosomal sorting complex required for transport (ESCRT)-dependent and ESCRT-independent pathways (Fig. 1B) [19].
In the ESCRT-dependent pathway, ILVs are created through the action of ESCRT complexes, including ESCRT-0, -I, -II, and -III [20]. These complexes facilitate the inward budding of the endosomal membrane, leading to the formation of ILVs within MVBs. Eventually, these ILVs are released as small EVs when MVBs fuse with the plasma membrane [21].
Conversely, the ESCRT-independent pathway relies on the involvement of tetraspanins and lipids. Small EVs often exhibit an enrichment of specific lipids, including cholesterol, sphingolipids, phosphatidylserine, and ceramide, resembling the composition of membrane lipid rafts. Furthermore, certain proteins found in exosomes, such as flotillins, caveolins, and tetraspanins, are also components of these lipid rafts [22, 23]. Lipid rafts are known to play roles in protein sorting, membrane curvature, and vesicle budding, suggesting their relevance to exosome formation and release [19].
Exosomes produced by tumors, commonly referred to as tumor-derived exosomes (TDEs), play a pivotal role in various cancer-related processes, including the remodeling of the tumor microenvironment (TME), angiogenesis, invasion, metastasis, and the development of resistance to treatment [24]. These TDEs facilitate intercellular communication, acting as messengers that transport crucial information among tumor cells, the TME, and healthy cells, thereby triggering those processes [25, 26]. Notably, malignant tumor cells release TDEs containing mRNA molecules associated with cell migration and metastasis. These are taken up by less aggressive cancer cells within the same tumor and even in distant metastatic sites, leading to alterations in cell behavior [27].
Angiogenesis, the formation of new blood vessels, is crucial for tumor growth, expansion, and metastasis [19]. This process is enhanced by TDEs, which carry major angiogenic stimulatory factors such as Vascular endothelial growth factor (VEGF), fibroblast growth factor (FGF), platelet-derived growth factor (PDGF), basic fibroblast growth factor (bFGF), transforming growth factor (TGF), tumor necrosis factor (TNF), and interleukin-8 (IL-8) [28]. For instance, glioblastoma (GBM), an angiogenic tumor type, releases TDEs containing high levels of miR-221, proteoglycans glypican-1, and syndecan-4, which promote the growth of endothelial cells and the development of tubules, thereby enhancing angiogenesis [29]. Moreover, TDEs are implicated in cancer cell migration, invasion, and metastasis. TDEs from cervical carcinomas and breast cancer cells, for example, are enriched in MALAT1, a long non-coding RNA associated with metastasis and invasion in lung cancer and hepatocellular carcinoma [30, 31]. Pancreatic ductal adenocarcinoma (PDAC) is another example where TDEs regulate various aspects of tumor behavior, including angiogenesis (miR-27a, VEGFC), cancer-associated thrombosis (TF, ANXA1), immune evasion (H2AF, PKM2), stromal reprogramming, chemoresistance, metastasis, and tumor growth (miR-27a, mir-194-5p, ANXA1, miR-222) through the delivery of specific cargo [32].
Furthermore, exosomes contribute to the development of drug resistance in cancer. This phenomenon is orchestrated by drug-resistant cancer cells endowed with the capability to encapsulate chemotherapy agents within exosomes, facilitating their transport out of tumor cells and subsequent delivery to sensitive cells [33]. Notably, exosomes enriched with molecules like miR-25-3p, can suppress genes such as Dickkopf WNT signaling pathway inhibitor 3 (DKK3) in osteosarcoma cells. This molecular manipulation promotes cancer growth in vitro and amplifies resistance to a spectrum of chemotherapy drugs, including methotrexate, cisplatin, doxorubicin, and docetaxel [34].
Conversely, TDEs offer compelling advantages for cancer treatment. They carry tumor-specific antigens and oncogenic cargo, endowing them with the capability to serve as potent stimulators of anti-tumor immune responses [35]. Multiple research projects have confirmed the potential of TDEs as biomarkers for cancer diagnosis and disease monitoring at various stages [36]. Moreover, their capacity to modulate the tumor microenvironment and deliver therapeutic cargo, such as microRNA (miRNAs) or chemotherapeutic agents, positions them as promising candidates for targeted therapy [15, 16].
3 Liquid biopsies based on exosomes
Liquid biopsies are used for tumor molecular profiling, which is transforming cancer diagnosis and treatment. EVs can be extracted from a variety of human-derived samples (Table 1), but body fluids such as urine, saliva, cerebrospinal fluid, and mainly blood can be used as liquid biopsies to extract tumor-derived information, providing a comprehensive approach to cancer monitoring with a minimally invasive technique [55]. Circulating extracellular nucleic acids (cell-free DNA; cfDNA), circulating tumor DNA (ctDNA), circulating tumor cells (CTCs), RNA, and TDEs can be isolated from these fluids [56]. Conventional tumor profiling relies on the acquisition of resected tumor samples by invasive surgical procedures, which can be difficult since it might be hard to obtain an adequate quantity and quality of tumor tissue. Furthermore, there are some challenges (e.g. invasive surgeries, quantity and quality of samples, multiple biopsies) associated with the regular use of invasive techniques to track tumor response and recurrence during therapy, which makes them less useful for continuous tumor profiling [57]. In order to identify tumor-derived components, oncology research has recently shifted its attention from studying complete tissues to investigating different bodily fluids. Given the difficulties associated with standard biopsies, liquid biopsies, especially those based on TDEs have gained attention.
3.1 Blood
Blood-derived EVs, especially circulating TDEs, have been a major focus because of their accessibility and rich cargo, which is predominantly composed of miRNAs and proteins that serve as valuable biomarkers for cancer diagnosis and prognosis [58, 59]. It is estimated that hundreds of billions of TDEs are present in a single milliliter of plasma given that tumor cells release tens of thousands of vesicles per day [60]. The high concentration of TDEs in the blood makes it an ideal candidate for liquid biopsies, which allows for the non-invasive collection of vital tumor-related data.
In advanced gastric cancer, researchers developed a methodology based on a 6-exosome-RNA panel (let-7i-5p, miR-1307-3p, LZIC, SRSF6, lncFTH1–211 and lncPTMA-209), obtained from a plasma liquid biopsy, that could robustly identify patients who respond to fluorouracil-based neoadjuvant chemotherapy, as opposed to those who do not [61]. These biomarkers may enable personalized precision administration of neoadjuvant chemotherapy for patients with advanced gastric cancer. A significant challenge is the early detection of ovarian cancer (OC), and the study conducted by Li et al. led to the identification of novel biomarkers by analyzing preoperative peripheral blood exosomes in patients with OC and controls. The Zinc Finger Protein 587B (ZNF587B) was found decreased in OC, confirmed by mRNA, protein levels, and exosomes. The expression levels of this protein and its correlation with the tumor stage show the potential as a novel biomarker for early liquid biopsy in OC screening [62]. In addition, studies on PDAC patients suggest that exosomes may be capable of identifying the earliest stages of the disease. Using mass spectrometry analyses, glypican-1 (GPC1) was identified as being enriched on cancer-cell-derived exosomes. GPC1+ circulating exosomes (crExos) were found in the serum of PDAC patients with complete specificity and sensitivity, distinguishing healthy individuals and those with benign pancreatic illness from those with early- and late-stage PDAC [63]. In the absence of abnormal endoscopic ultrasonography results, GPC1+ crExos may serve as a biomarker to support the diagnosis and categorization of PDAC precursor lesions and identify individuals with a hereditary susceptibility [64]. Additionally, GPC1+ crExos levels allow differential diagnosis between PDAC and chronic pancreatitis including its mass-forming presentation [65]. GPC1+ crExos may function as a biomarker and non-invasive diagnostic and screening tool for pancreatic cancer early detection. Recently, a project called “PANLIPSY” was published to detect pancreatic cancer early by liquid biopsy. In this study, researchers used a combination approach using blood samples and Artificial Intelligence (AI) where they identify circulating biomarkers in samples (CTCs, ctDNA, exosomes, circulating immune system, circulating cell-free nucleosomes, proteins, and microbiota) [66]. In breast cancer (BrCa), several promising biomarkers for early detection that are enriched in plasma exosomes have been identified, including human epidermal growth factor receptor (HER2), CD24, GPC1, and long noncoding RNA (lncRNA) H19 [67,68,69]. All these examples of exosome biomarkers found in patients’ plasma demonstrate the potential of TDEs as useful liquid biopsy for diagnosing and monitoring patients with different types of cancer.
3.2 Saliva
Saliva stands out as a valuable source for studying exosomes with potential diagnostic applications, but the efficacy of the collection process is contingent upon several considerations, including sampling location, timing, instructions, and technique. Different gland sources may affect analyte levels; therefore, standardization is needed for interstudy comparisons. Most exosome analyses are conducted on whole saliva, but gland-specific studies may be vital due to variations in production and composition [70,71,72,73]. Factors like timing, diet, smoking, and stressors can influence saliva composition and standardized protocols often include abstaining from smoking, eating, and strenuous activity before collection. Techniques for saliva collection range from stimulated to passive sampling, with stimulation’s effects on exosomes being less known [47, 71, 72]. Saliva as a liquid biopsy is used in head and neck squamous cell carcinomas (HNSCCs) and has (≥95%) sensitivity and specificity to identify true positive or negative cases [74]. According to recent findings, salivary exosomes from HNSCC patients had a notably elevated expression of miR-24-3p, which was related to increased proliferation when contrasted with normal subjects [75].
3.3 Urine
The second most utilized biofluid in clinical diagnostics is urine. It has garnered attention as a potential source for early biomarkers of kidney pathophysiology [76, 77]. In the study of urine-derived exosomes, the term urinary extracellular vesicles (uEVs) is widely used, therefore in this section although we talk about exosomes the term used will be uEVs. EVs derived from various cell types, including nephrons, kidney parenchyma, tubular, prostate, and bladder cells, offer valuable systemic information, particularly concerning the urinary system [78, 79]. While plasma EVs are infrequent under healthy conditions due to their limited ability to traverse the glomerular filtration barrier, uEVs play a crucial role in detecting renal damage and function as carriers for systemic markers [77]. Challenges in uEV research arise from the inherent variability in urine composition, necessitating standardized techniques for sample comparison. Influential factors, such as exercise, hydration, and electrolyte concentration, impact on uEVs purification yield, complicating result interpretation in the absence of established normality range [80]. Proteases and pigments in urine also pose a challenge in downstream analyses, requiring meticulous processing steps for accurate diagnostics [81]. Podocyte marker Wilms’ tumor 1 (WT-1), a reporter of renal damage, has been found in uEVs and the use of uEVs has proven to detect internalized molecules that, otherwise would have problems of limited detection due to its low levels [80, 82]. These uEVs are also a source of genetic markers, leading to their clinical use for diagnostics, as ExoDx™ Prostate Intelliscore, which detects prostate cancer gene signature of exosomal RNA: ERG, PCA3, SPDEF via RT-qPCR with a high sensitivity and negative predictive value [83]. In thyroid cancer, was recently identified thyroglobulin as a new biomarker found in patients’ uEVs (NCT02862470) (Table 2). Using this biomarker researchers could track thyroid cancer and eliminate the suspicion of recurrence and remove patients from long-term follow-up [85].
3.4 Other bodily fluids
Breast milk, isolated from both human and cow’s milk, constitutes an invaluable source of essential nutrients and protective elements for newborns due to its bioactive components like maternal cells, antibodies, and enzymes. However, the substantial lipid content and the presence of milk fat globules, which bear a resemblance to EVs, can introduce complexity to the isolation of exosomes from milk [87, 88]. The methods employed for milk collection and storage can have significant implications for both milk composition and exosome content. Additionally, the process of freezing and the choice of storage temperatures have the potential to impact the integrity of both cells and exosomes. Breast milk also can be used as a liquid biopsy, researchers found TGFβ2 significantly upregulated in breast milk exosomes during weaning/early involution and could be useful in breast cancer detection [89].
Cerebrospinal fluid (CSF) is primarily secreted by cells within the choroid plexus and its exosomes, although present in low concentrations, influence functions like synaptic communication, synaptic strength, and nerve regeneration [90,91,92]. The identification of protein structures and modifications within CSF exosomes is essential for gaining insights into RNA-protein interactions in the context of neurodegenerative diseases. Comprehensive guidelines for CSF biobanking are in place, specifying the collection process from the lumbar region according to established protocols. Acquiring healthy CSF control samples can be challenging, but potential sources include commercial providers and patients with various alternative disorders or those undergoing specific medical procedures, who can serve as suitable controls for research purposes. CSF exosomes show significant potential as biomarkers for neurological disorders and cancer, opening avenues for the detailed analysis of their associated proteome and lipidome [48, 93]. In low and high-grade gliomas, exosomes isolated from CSF patients show IDH1-R132H mutations and reflect the degree of disease progression [94]. Besides, the EGFRvIII mutation found in CSF patient-derived exosomes proved to have 98% specificity as a biomarker for patients with GBM [95]. Other biomarkers found in CSF related to GBM are miR-21, IDH1-mutant mRNA, miR-9, miR-1298-5p, MYO1C, miR-30b-3p, which are useful for diagnostic, monitoring, and/or therapeutic approaches [96,97,98,99,100].
A recent survey of 116 studies on ClinicalTrials.gov (https://clinicaltrials.gov/) revealed that exosomes have been primarily investigated as biomarkers (50%), with exosome therapy representing only 28.44% of the studies. Additionally, 14.66% of the studies were focused on basic exosome analysis, while drug delivery system studies constituted 5.17% of the registered clinical trials [101]. Table 2 focuses on the most recent trials where exosomes were used to identify cancer-related biomarkers. Only the clinical trials that have identified the biomarkers they focused on have been included in this table.
4 Exosome isolation and loading methods
While small EVs can be obtained from a wide range of biological materials, spanning body fluids to various cell cultures, the establishment of standardized protocols for their isolation and purification from each source is imperative [102]. Moreover, the choice of source and isolation method plays a pivotal role in determining the outcomes in terms of EV population type, purity, yield, and composition. Given the enormous research potential of small EVs in the biomedical field, numerous research projects are focused on refining techniques for separating small EVs from various cellular components and interfering substances. Various methods are available for isolating small EVs from body fluids or cell cultures, depending on their source and size [103]. These methods encompass centrifugation, size-based separation, immunoaffinity capture, polymer precipitation, and microfluidics techniques [104], as outlined in Table 3.
To specifically isolate exosomes, the ultracentrifugation approach is commonly used due to its ability to separate vesicles based on size and density. Ultracentrifugation, a widely employed technique, involves multiple centrifugation steps, starting at low speed (1500 x g) for around 10 min to discard any cell or debris and progressively increasing in intermediate centrifugation steps to high-speed centrifugations (100,000–150,000 x g) for at least 70 min up to 3 hours to isolate exosomes from large EVs in an EV enriched medium, frequently filtered through 0.22 µm PES syringe filters [105, 115]. Additionally, density gradient ultracentrifugation, where the addition of sample particles to inert media with a density gradient (such as sucrose and cesium chloride) can be used to further purify exosomes by separating them based on their buoyant density [116]. Ultrafiltration and size exclusion chromatography (SEC) are size-based isolation techniques that rely on the isolation of exosomes passing through physical barriers depending on the size of the particle [107, 108]. Since it is challenging to differentiate between exosomes and microvesicles due to their similar sizes (100–150 nm) and densities (1.08–1.19 g/ml), exosomes are frequently identified by the presence of endosome-associated proteins, such as tetraspanins CD9, CD63, and CD81 [117]. Immunoaffinity-based techniques, which target exosome-specific surface markers such as those mentioned before, offer an alternative approach for isolating highly pure exosomes. The immunoaffinity chromatography approach is based on the affinity between proteins and their corresponding antibodies; for this purpose, antibodies are mounted on matrices such as magnetic beads [118]. The polymer precipitation method is focused on the use of polyethylene glycol and is frequently available in isolation kits [112]. Finally, microfluidic chip techniques are innovative approaches that exploit distinctions in the biochemical and physical properties of small EVs. These techniques are primarily categorized into three groups: (a) immune-affinity approach, (b) sieving (nanoporous membranes), and (c) trapping exosomes into porous structures (nanowire-on-micropillars) [119]. To enhance the precision of exosome isolation, researchers often combine two methods. For instance, to obtain a greater level of purity in urine-derived EVs, it may be essential to employ a dual approach, starting with ultracentrifugation followed by the SEC isolation method [36]. These methods ensure a more selective isolation of exosomes, distinguishing them from other EVs and allowing for more precise downstream analyses.
Given that the most homogeneous fraction of small EVs are exosomes, it could be considered that what is described in the literature referring to small EVs is mainly related to exosome loading [120]. Exosomes can encapsulate a diverse range of cargo within a protective lipid bilayer, a feature that enhances their stability in vivo and prolongs their circulation time in the bloodstream [121]. This characteristic makes exosomes suitable for drug loading, both within the lipid bilayer and in the aqueous space. Loading drugs into exosomes facilitates the precise and efficient delivery of therapeutic substances to specific cells or tissues, minimizing potential adverse effects on healthy cells [122]. Several loading methods are available for modifying exosomes, including incubation, sonication, electroporation, transfection, extrusion, freeze-thaw cycles, thermal shock, saponin-assisted loading, pH gradient methods, and hypotonic dialysis. The principal loading approaches are outlined in the following Table 4.
All methods described in Table 4, confirmed that exosomes could be easily modified to improve their drug delivery characteristics. Among these techniques, incubation methods stand out as the simplest, albeit yielding a relatively modest drug loading rate. Moreover, incubation methods are inexpensive and extremely safe since they do not compromise the stability of the exosome membranes. The potential for improving the initial drug-loading rate depends on measures like elevating drug concentration and introducing stirring during the incubation process. As a result, the incubation approach holds significant potential across various medical applications, encompassing therapy and drug delivery [130].
5 Exosomes as a delivery system in cancer therapy
Exosomes can be administered through four different routes: intranasal, intravenous, intraperitoneal, and intracranial. This indicates the high flexibility and compatibility of exosome-based drug delivery [137, 138]. Studies and even clinical trials are being developed for its use in oral delivery methods, for example in the case of curcumin for colonic cancer, as it’s been seen that curcumin can act as an antiproliferative molecule in colorectal cancer [139], some other therapeutic lines widely studied implies EVs derived from embryonic stem cells that are capable of conveying early pluripotent transcription factors to hematopoietic progenitors, thereby fostering both proliferation and survival [140].
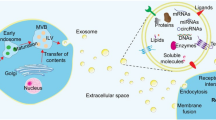
Exosomes can enter recipient cells through a variety of mechanisms, including micropinocytosis, lipid raft-mediated uptake, phagocytosis, and membrane fusion [141]. Hisaaki Hirose et al. assessed EV internalization and fusion by using the NanoBiT system, demonstrating that large EVs showed potential superiority in cargo delivery to recipient cells over small EVs. The introduction of the fusogenic protein VSV-G, however, enhanced EV-cell membrane fusion [142]. As mentioned before, the composition and cargo of exosomes are directly related to their parent cell. They are surrounded by a phospholipid bilayer and contain messenger RNA (mRNA), microRNA (miRNA), long non-coding RNA (lncRNA), DNA fragments, and metabolites from their parental cells in addition to proteins, and lipids [143,144,145]. The ExoCarta exosome database (http://www.exocarta.org) currently contains 9769 proteins, 3408 mRNAs, 2838 miRNAs, and 1116 lipids that have been discovered in exosomes from different cell types and organisms.
Naturally, EVs contain nucleic acids, proteins, lipids, metabolites, and even organelles from parental cells. This led to the initial thought that EVs were mainly used as a cellular mechanism to discard cell waste (Fig. 1C). Today EVs, mainly exosomes, have been shown to have major roles in regulating intercellular communications in both physiological and pathological conditions [146, 147]. Multiple researchers have shown that exosomes can be selectively loaded with the desired cargo using physical/chemical/biological methods [148, 149].
5.1 Nucleic acids
Loading nucleic acids in exosomes has a great potential for therapeutic genome editing and different researchers have reported the benefits of loading CRISPR/Cas9 expression vectors in exosomes [150,151,152,153]. For instance, Kim et al. utilized exosomes derived from cancer cells to effectively transport CRISPR/Cas9 plasmids, aiming for the specific inhibition of poly (ADP-ribose) polymerase-1 (PARP-1). This approach triggers apoptosis in ovarian cancer cells and heightens their responsiveness to cisplatin [150]. EV-mediated vehiculation of suicide genes mRNA (i.e CD-UPRT) has been recently studied by Mizrak et al., they designed a mRNA/protein complex loaded into small EVs, to enhance the expression of CD-UPRT, in HEK-293 T cells resulting in their apoptosis [154, 155]. Kanada et al. and Altanerova et al. also used this approach using exosomes, enabling effective prodrug conversion and tumor cell death in breast cancer and inducing tumor cell death by converting prodrug 5-FC to 5-FU in glioma, respectively [156, 157].
Silencing oncogenes via the administration of siRNA or miRNA loaded exosomes has also been reported as an effective approach for cancer therapy. In a recent study, Kamerkar et al., used exosomes to deliver siRNAs targeting oncogenic KRASG12D, to treat pancreatic cancer, which demonstrated greater efficacy compared to the other administration methods tested [158]. To enhance the precision of targeted siRNA delivery, Pi et al., enhanced EV-mediated siRNA delivery by incorporating folate and RNA aptamers as targeting ligands, which specifically bind to receptors overexpressed on cancer cells. The modified exosomes proved successful in delivering survivin siRNAs to prostate and breast cancer cells, exhibiting reduced endosome trapping and heightened delivery efficiency [159, 160]. The delivery of miRNAs (e.g., miR-206, 26a, 122, 126, 146b, 124a, and let-7a) by MSC-derived exosomes presented promising anti-tumor effects in osteosarcoma, hepatocellular carcinoma, non-small cell lung cancer, breast cancer, and glioma [161,162,163,164,165,166,167]. Moreover, treating cancer cells with exosomes loaded with miRNA inhibitors (anti-miR-9, anti-miR-214, anti-miR-374) has also shown antitumor results [168,169,170].
5.2 Proteins
Systemic administration of functional proteins presents great challenges for clinical application, due to protein degradation and poor cellular intake; however, exosome-mediated delivery presents an alternative approach. Although loading proteins into exosomes has shown some technical issues, authors have reported the creation of technology platforms to generate small exosomes expressing therapeutic proteins. One of them is the EXosomes for Protein Loading via Optically Reversible protein–protein interaction (EXPLOR), that can load non-anchored free-from proteins into exosomes [171]. Engineered exosomes called (Exo-pYSTAT3 IB) were produced using EXPLOR, these small exosomes were loaded with an mCherry-tagged intracellular antibody (intrabody, IB) targeting a Tyr705-phosphorylated signal transducer and activator of transcription 3 (pYSTAT3). The use of Exo-pYSTAT3 IB, by intraperitoneal injection resulted in a tumor growth inhibition of 41.6% in a CT-26 mouse colon cancer syngeneic model [172, 173]. On the other hand, Aspe et al. reported the obtention of exosomes containing Survivin-T34A by transfecting melanoma cells with dominant-negative mutant (Survivin-T34A) to produce the protein of interest and carry it in their EVs. These exosomes were then used for the treatment of pancreatic adenocarcinoma, exhibiting promising results as they induced cell apoptosis and inhibited tumor growth [174].
5.3 Metabolites
The metabolite cargo of small EVs, mainly lipids, varies depending on the origin and metabolic state of the cells. In pancreatic cancer patients, lipidomic analysis of exosome serum revealed that 270 lipids were dysregulated between patients and controls. Specifically, Lysophosphatidylcholine (LysoPC), phosphatidylcholine (PC), and phosphatidylethanolamine (PE) were associated with tumor stage, tumor markers (CA19-9 and CA242), and tumor diameter. Additionally, there was a strong correlation between PE and overall patient survival. These dysregulated lipids may serve as diagnostic biomarkers or may suggest a pathogenic correlation between the progression of pancreatic cancer and small EVs [175]. In other types of cancer, such as breast cancer, there is evidence that exosomes derived from human mesenchymal stem/stromal cells (hMSCs) can enhance proliferation and metastasis in this type of cancer. A metabolomic study has revealed the presence of glutamate and lactate. Lactate may contribute to cancer cells being more resistant to hypoxia and starvation conditions, whereas glutamate may supply precursors for the main macromolecular classes through carbon and nitrogen trafficking [176]. Other metabolites, such as glycerophosphoglycerol (PG), glycerophosphatidylserine (PS), triacylglycerol (TG), and glycerophosphorylcholine, were found in exosomes derived from melanoma stem cells, similar to those derived from patients with malignant melanoma [177]. The transfer of these metabolites among cancer cells could support their proliferation and survival and promote tumor growth, angiogenesis, immune evasion, and metabolic reprogramming in recipient cells [178]. These findings open the door to the discovery of clinically useful biomarkers in cancer by metabolomic characterization of exosomes.
5.4 Drugs
Multiple studies claim that EV-mediated delivery improves the stability of drugs in the bloodstream, leading to greater drug accumulation in recipient cells [179, 180]. Our group investigated the efficiency of exosomes for GBM and PDAC treatment. Two chemotherapeutic drugs (TMZ and EPZ015666) were loaded into RWP1-derived and GBM-derived exosomes, by direct incubation method. Results claimed the greater efficiency of EV-mediated treatment compared to the direct administration of the drugs, where TMZ-loaded EVs showed the best antiproliferative effects on both PDAC and GBM [16, 181]. Furthermore, a lower drug dosage was required in exosome-mediated treatments, suggesting the specificity of the delivery system. Moreover, similar studies also claim the strong antiproliferative effects of exosome-mediated PTX chemotherapy on human PDAC cells [180, 182, 183]. Kim et al. further improved this treatment for pulmonary metastatic lung cancer, developing aminoethylanisamide-polyethylene glycol (AA-PEG) exosomes, which were loaded with PTX (AA-PEG-exoPTX), these exosomes exhibit a higher loading capacity and better anti-cancer effect in a mouse model [184].
5.5 Nanoparticles
It has been shown that the combination of exosomes with nanoparticles can improve their targeting ability and therapeutic efficacy [133, 185]. Along this line, Qi et al. developed an exosome-based superparamagnetic nanoparticle cluster (SPMNs) by linking superparamagnetic NPs with blood-derived exosomes via transferrin and its receptor. This device presents enhanced cancer targeting and tumor growth inhibition under the effects of an external magnetic field in murine cancer models [179]. Moreover, Silva et al. developed a macrophage-derived nanovector loaded with citrate-coated magnetic NPs and m-THPC photosensitizer to generate vesicles with magnetic and optical responsiveness allowing therapeutic and imaging functions. This nanovector is valuable in both cancer diagnosis and treatment due to its dual-mode imaging [186]. Furthermore, enclosing iron oxide NPs and different therapeutic agents into macrophage-derived EVs, which can be manipulated by magnetic force for targeted delivery of drugs, also exhibited antiproliferative cancer results [187]. Additionally, Liu et al. designed a tumor-cell-derived exosome-based delivery system, where high sono-activatable sinoporphyrin sodium (DVDMS) was loaded onto exosomes. Using this system led to enhanced cargo release under ultrasound exposure and accumulation in the targeted location, suggesting that exosomes are promising delivery vehicles for nanosensitizers [188]. Dumontel et al. loaded zinc oxide nanocrystals (ZnO NCs) onto B-cell EVs, enhancing specificity to lymphoid cancer cells by incorporating anti-CD20 monoclonal antibodies on their surface. Notably, subsequent EV engineering rendered them responsive to high-energy ultrasound shock waves, facilitating the on-demand activation of cytotoxic cargo release against specific lymphoid cancer cells [189]. On the other hand, Dumontel et al., use mesoporous silica nanoparticles (MSNs) in combination with TDEs from GBM and HeLa cells. This hybrid nanosystem demonstrated particular cytotoxic efficacy against specific cancer cells as well as an increased and selective intracellular accumulation of doxorubicin, suggesting that it is a promising candidate for the development of novel targeted treatments [190].
5.6 Imaging agents
Imaging techniques as single photon emission computed tomography (SPECT), positron emission tomography (PET), optical imaging, magnetic resonance imaging (MRI), and magnetic resonance spectroscopy (MRS), are very important in cancer diagnosis. These techniques use imaging agents to monitor structural, functional, and molecular changes in cancer tissues clinically and pre-clinically [191]. Imaging agents are tools used to visualize and track changes within the body, and exosomes can be engineered or naturally used for this purpose. To obtain high-quality contrast images for disease diagnosis and monitoring the treatment response, a contrast agent is needed in MRI. In this sense, researchers have used macrophage cell-derived EVs reconstructed with a Gd-conjugated liposomal system herein called gadolinium-infused hybrid EVs (Gd-HEVs), which showed excellent contrast enhancement in the blood vasculature with a higher retention time compared to its commercial one, Magnevist®. This novel imaging agent effectively demonstrated how adding Gd to EVs can improve contrast ability, which may serve as a platform for the formulation of safer MRI contrast agents [192]. As another example, the authors describe a new method for SPECT using the radioisotope labeling of erythrocyte-derived EVs with the (99 m) Tc-tricarbonyl complex. his method allowed for the assessment of in vivo biodistribution studies in a mouse model, with the notable advantage of the radioactive label being only partially separated from the EVs [193]. Exosome-based imaging agents represent a promising frontier in non-invasive cancer diagnostics, combining the specificity of molecular markers with the advanced capabilities of modern imaging technologies.
6 Exosome-based immunotherapy
Immunotherapy is rapidly advancing as a key approach in cancer treatment employing several strategies such as targeting different immune checkpoints, developing cancer vaccines, and enhancing the natural immune response. Immunotherapy offers a clever solution to the limitations of traditional cancer therapies, including the need for greater specificity and the ability to generate more durable responses.
6.1 Immunosuppression caused by TDEs
When it comes to TDEs, their dual behavior in immune response must be considered to understand their therapeutical possibilities. They facilitate communication between the tumor and surrounding cells, favoring the invasion, angiogenesis, and metastasis and thus contributing to tumor progression [194].
It has been described that TDEs cause immunosuppression, for instance due to the presence of immunosuppressive factors such as TGF-β1 and PD-L1, thus interfering with immunotherapy (Fig. 2) [195]. TDEs have also been suggested to interfere with NK cells, modulating their activation by the binding of MICA and MICB ligands to NKG2D receptors [196, 197]. To overcome this immunosuppressive profile, the use of drugs such as tunicamycin A has been proposed to alter the TDEs N-glycosylation profile that leads to an enhanced immunogenicity, dendritic cell activation, and IFN-γ mediated CD8+ T-cells activation thus delaying tumor recurrence. The study by Han and colleagues revealed that only 17% of tumor growth was detected in mice treated with immunogenic TDEs compared to those treated with non-modified TDEs [198]. Another possibility is the clearance of these TDEs immunosuppressors of the circulatory system, especially in the case of liquid tumors. Promising technologies that offer this possibility are currently in development. One example is the hemodialysis device called Aethlon ADAPT™ (adaptive dialysis-like affinity platform technology) which is now in a clinical trial (NCT04453046) [199].
Overview of engineered exosome constructs and their role in modulating antitumor immune responses. Different cells are the sources (blue box) for engineer exosomes involved in immunosuppression or targeting immune checkpoints while others act as a cancer vaccine (green box). The immune response generated by each type of exosome is indicated by arrows in a variety of colors (brown box). TDEs acts cause immunosuppression while activation of DCs by the engineer exosomes initiates a cascade that stimulates T lymphocytes, amplifying the antitumor immune response. Acting on immune checkpoints throughout exosomes led to M2 to M1 macrophage transition
6.2 Cancer vaccines
Despite the immune-inhibitory behavior of TDEs on the immune system, there is a whole field of research exploring the ability of loaded TDEs to enhance immune responses toward the tumor. F. Huang et al. recently used PD-L1 silenced leukemia-derived exosomes (LEX–si) to enhance the immune response against leukemia. Leukemia cells were transfected with a lentiviral vector containing anti-PDL1 shRNA. This strategy led to the maturation of DCs expressing CD86, CD80 and MHC-II on their surface, and secreting proinflammatory cytokines (IL-12p70 and TNF-α) and resulted in a higher T cell activation, proliferation and Th1 cytokine release. In vivo assays in mice also demonstrated a high potency of this vaccine based in exosomes reducing the tumor progression [200]. W. Zhou et al. also demonstrated the plausible use of TDEs as a cancer vaccine for pancreatic cancer treatment (spMEXO) (Fig. 2). They used pancreatic TDEs loaded with CCL22 siRNA and with a conjugation of CP05 and MART-1 immunogenic peptides. CCL22 is a ligand that mediates the Treg cells activation pathway by the DCs. Silencing of this gene results in a depletion of this pathway, resulting in the inhibition of Treg expansion. This mechanism proved to be effective in vivo and in vitro as a cancer prophylactic vaccine [201].
Other authors have also explored the engineering of cancer vaccines consisting of hybrid exosomes from tumoral and non-tumoral sources, through vesicle fusion. This vaccine, called Lipo@HEV is the product of combining B16–F10, 4T1 and CT26 (epithelial, breast and colon) tumor cell lines with the biocompatible and coadjuvant outer membrane vesicle from Akkermansia muciniphila (Akk-OMV) and loaded with gene therapy plasmid encoding PD-L1 binding domain in a trimeric form that acts as a trap. The effect shown for this therapy was synergistic, preventing the CTL exhaustion by gene therapy PD-L1 means and penetrating the lymph node, promoting DCs maturation and CTL activation. It reached a significant increase of the survival rate in the treated mice with Lipo@HEV as well as a tumor reduction [202]. Another recent vaccine strategy against breast cancer was recently developed by L. Huang et al., exploring the use of engineered breast cancer-derived exosomes (HELA-exos) that overexpressed α-lactalbumin and loaded them with ICD inducer human neutrophil elastase (ELANE) and a TLR3 agonist (Hiltonol). These exosomes induced immunogenic cell death (ICD) in tumor, activated dendritic cells (cDC1s) and primed tumor reactive CD8+ T cells. HELA-Exos also proved inhibiting growth in an immunocompetent mouse model and patient-derived organoids, showing a promising application as immunotherapy (Fig. 2) [203].
The use of exosomes derived from dendritic cells (DEX) is another current trend in cancer treatment based on exosomes. Researchers demonstrated tumor-specific immune response for Hepatocellular carcinoma (HCC) using DEXP&A&N. This personalized exosomal therapy consisted of a conjugate coupling DEX with several peptides [HCC-targeting peptide (P47-P), an α-fetoprotein epitope (AFP212-A2) and a functional domain of high mobility group nucleosome-binding protein 1 (N1ND-N)] able to recruit and activate DC, as well as recognize HCC-antigens. Tumor-specific CD4+ and CD8+ T cell response and augmented tumor antigen presentation were found in mice with HCC when treated with DEXP&A&N, resulting in significant tumor growth retardation [204]. Similarly, DEX presenting epitopes of tumor-specific neoantigens, called Exo-OVA vaccine, was used to treat melanoma. Exo-OVA vaccine stimulated tumor-specific CD4+ and CD8+ T cell immune response resulting in a significant reduction in tumor growth and an increased survival rate in a mice model (Fig. 2). Additionally, long-term immunological memory was developed, mediating both cellular and humoral immunity, leading to delayed tumor recurrence in B16F10 melanoma models and the elimination of lung metastasis in MC-38 models. Furthermore, no relevant side effects or IL-17-mediated autoimmune responses were found systemically after the administration of this vaccine [205]. Moreover, DEX conjugated with a MUC-1 glycopeptide antigen was tested in vivo in a murine model as a therapeutic approach for breast carcinoma. MUC-1 is overexpressed and aberrantly glycosylated in a wide variety of carcinomas, playing a crucial role in disease progression [206]. Treatment with this conjugate resulted in strong CD8+ T cells-mediated cytotoxicity against MUC1-positive tumor cells, inhibiting tumor growth and prolonging survival [207].
6.3 Targeting of immune checkpoints
The efficacy of cancer immunotherapy often relies on the presence of tumor-associated macrophages (TAMs) with an immunosuppressive M2 phenotype, which allows the tumor to evade the immune system. Recent immunotherapeutic strategies aim to reprogram these TAMs toward a proinflammatory M1 phenotype to promote antitumor immunity [208]. A recent clinical trial (NCT05375604) focused on silencing the expression of STAT6, a key transcription factor to promote the M2 phenotype, to stimulate the expression of nitric oxide synthase 2 (NOS2) and the transition towards the proinflammatory M1 phenotype. Thus, reshaping the TME and generating an adaptive immune response, mediated by CD8+ T cells. This trial proposed the utilization of a conjugate known as exoASO-STAT6 consisting of exosomes purified from human embryonic kidney (HEK) cells, loaded with an antisense oligonucleotide (ASO) selectively silencing STAT6 (Fig. 2). Preliminary results suggested exoASO-STAT6 administration had tumor growth inhibitory effects, greater than 90%, and 50–80% complete remissions in colorectal cancer (CRC) and HCC models with minimal side effects [209]. Another key immune checkpoint for the regulation of inflammatory responses is the stimulator of the interferon genes (STING) pathway. Cyclic dinucleotides (CDN), agonists of the STING pathway, have been shown to activate the immune response and eliminate tumors in preclinical models [210]. The use of a conjugate called ExoSting, consisting of CDN-loaded exosomes purified from HEK293 cells or cells overexpressing Prostaglandin F2 receptor negative regulator (PTGFRN), is currently under study in a clinical trial (NCT04592484). Initial results indicate that exoSting increases CDN potency, and activates antigen presentation in the TME, thus enhancing local Th1 responses, CD8+ T cell recruitment, and generating systemic anti-tumor specific immunity after intratumoral injection (Fig. 2) [211].
Overall, exosome-mediated antitumor immunotherapies offer a promising strategy for cancer treatment by activating immune responses and inducing protective immunity against specific antigens on tumor cells.
7 Challenges and future perspectives
As mentioned before, the use of exosomes as drug carriers holds significant potential for advancing cancer treatment. However, a great number of obstacles must be overcome to translate this promising strategy from the bench to the bedside.
In the 90s begun the use of NPs for drug delivery, since their nanoscale size allows for efficient cellular uptake and distribution of cargo molecules [212, 213]. Several types of NPs are being used like liposomes, micelles, dendrimers, mesoporous silica NPs, gold NPs, superparamagnetic iron oxide NPs (SPION), carbon nanotubes, and quantum dots [36]. The most used NPs in nanomedicine are liposomes, which consist of up to four different lipid types and two or more drugs or medicinal compounds in a spherical bilayer [214]. Nevertheless, there are disadvantages to using NPs, including early elimination, bioincompatibility, long-term toxicity, and undesirable biological effects [36]. In clinical trials, the use of NPs targeting has failed, BIND-014 a polymeric NP that contains surface ligands is an example. It binds to the prostate-specific membrane antigen (PSMA), but the complex interaction between NPs and the biological environment including the development of a protein corona that can conceal surface ligands and prompt immune detection, affects their function [214].
The unique properties of exosomes, make them more suitable for drug delivery and targeted therapy than liposomes. They have higher binding affinity, greater targeting abilities, are more stable in body fluids, and are highly biocompatible because of their endogenous origins [215,216,217,218,219]. Moreover, they can be modified to express targeting ligands on their surface, enabling specific delivery to target cells or tissues and enhancing the therapeutic efficacy while reducing off-target effects of the delivered molecules [121, 220]. However, is still challenging the isolation, characterization, and scale-up of pure populations of exosomes, have hampered basic and translational research studies [221].
Scaling up production and creating standardized manufacturing processes are two of the main obstacles to using exosomes for cancer treatment. To upscale the release of exosomes from cells various approaches have been explored, including not only 3D cultures but also chemical and physical stimulation, physiological alteration, and genetic manipulation of source cells [222].
Preconditioning media, genetic modification of stem cells, or conjunction of exosomes with biomaterials are some strategies used to increase exosome production that also led to the improvement of the therapeutic potential of exosomes [223,224,225]. Among chemical strategies, norepinephrine and N-methyldopamine can be used to promote exosome secretion from MSCs without changing their ability to induce angiogenesis, polarize macrophages to an anti-inflammatory phenotype, or downregulate collagen expression [226]. Other strategies include incubation of cells under hypoxia, exposure to an acidic medium, and the use of lipopolysaccharides, although the characteristics of these exosomes need to be evaluated.
Among the scaling-up production methods, 3D culture systems, which encompass bioreactors and cell spheres have gained prominence, with bioreactors being the most commonly used technique [227] (Fig. 3). Zha et al., have reported the establishment of a microfluidic platform to culture human fallopian tube epithelium (hFTE) tissue for small EV collection, they characterize the proteomic profile of the EVs that can be useful for studying if the fallopian tube shifts its small EV cargo during ovarian cancer carcinogenesis and its role in the disease development [228].
Schematic illustration of small EVs large production steps. The process begins with the production of different types of EVs with a special focus on exosomes, including native, hybrid, and synthetic EVs. These EVs are then subjected to scalable isolation methods to ensure reproducibility. Following isolation, storage protocols are implemented to address batch-to-batch variations, ensuring consistency in the final product. EVs undergo purification to remove contaminants and improve quality. Critical steps also include drug loading and functionalization of exosomes to enhance their therapeutic potential. Before clinical application, these EVs are tested in preclinical studies to assess their efficacy and safety. Finally, rigorous quality control measures are conducted to ensure that the exosomes meet the required standards for clinical use in humans. The color-coded bar indicates the current state of the information and technology known in every production step. Purple color (high) means technological development and detailed examples in the literature, yellow (medium) highlights the need for additional studies, and light blue (low) implies the need for additional basic and comparative research
Kang et al. developed a bioreactor system that achieved a sevenfold increase in the production of MSC-derived small EVs in just one day. In this system, they introduced Ca2+ under flow conditions highlighting the link between small EV biogenesis and this ion concentration [229]. Moreover, Watson et al. used a hollow-fiber bioreactor to produce a daily yield of small EVs that was 40 times higher per volume of conditioned media compared to a conventional 2D cultured flask. They produced human embryonic kidney (HEK) 293-derived small EVs bearing the heterodimeric interleukin-15 and observed that 3D-derived EVs contained fewer serum protein impurities than their 2D counterparts [230]. Additionally, a hollow-fiber bioreactor has been employed to produce exosomes from MSCs, demonstrating that 3D culture had no discernible impact on MSC surface markers when compared to 2D-exosomes. Similarly, 3D culture did not affect MSC shape, size, and exosome markers of 3D-exosomes. The 3D culture approach enhanced overall exosome production up to 19.4-fold when compared to conventional 2D culture, resulting in a higher exosome collection efficiency of the 3D culture technique [231]. However, the current techniques for isolating and purifying exosomes frequently do not scale well and can lead to poor yields and heterogeneity. Therefore, the study of milk as a source of exosomes has been extended lately. Milk stands out as a highly prospective and expandable reservoir of exosomes suitable for large-scale manufacturing. This is due to its widespread availability, cost-effectiveness, and the absence of necessity for cellular cultivation. Numerous investigations into the safety profile of milk-derived exosomes have indicated minimal levels of toxicity and favorable in vivo acceptance [232, 233]. Recently, safe engineered milk-derived EVs are being developed to deliver miRNA interferent to treat cancer, especially for colorectal cancer where its easy absorption has been previously demonstrated. Some preclinical studies in mice have shown that milk-derived EVs, that are rich in functional miRNA-148a, exhibit potential anti-cancer effects in colorectal cancer treatment. These non-toxic nanocarriers influence immunological function, inhibit tumor growth, and show promise as therapeutic agents, although rigorous quality control, standardized isolation methods, and consideration of dietary influences are crucial for their clinical application [234].
Recently, it has been identified for the first time that cell-free tumor DNA can be detected in breast milk, offering a potential new source for liquid biopsy in the early detection of pregnancy and postpartum breast cancer [235]. On the other hand, Quin et al., identified that TGFβ2 was significantly upregulated in breast milk exosomes. These exosomes caused alterations in both benign and malignant breast epithelial cells, which is consistent with the development and spread of breast cancer and suggests that high TGFβ2-expression on breast milk exosomes may determine the risk of breast cancer [89]. Those studies led to the idea that the analysis of exosomes from breast milk could also have breast cancer diagnostic potential.
Another source of EVs that allows scaling the production are the plant-derived EVs, which offer biocompatibility and non-toxicity [236]. They are obtained from plant tissues like stems, leaves, seeds, and fruit juices, and hold the potential for drug delivery [236].
One of the main challenges with using exosomes for drug delivery is how rapidly they leave the bloodstream after being administered in vivo [120]. Creeden et al. developed a “Smart Exosome” that co-displays RGD and CD47p110–130 through CD9 engineering (ExoSmart). This ExoSmart not only greatly decreased the amount of exosomes cleared from the liver and spleen by blocking macrophage phagocytosis, but also increased binding capacity to αvβ3 on PDAC cells, leading to increased cellular absorption [237]. On the other hand, Mu et al. fabricated microneedle patches for loading and delivering exosomes, as a solution for the rapid clearance. These patches prolong the residence time of exosomes at the administration site and maintain the drug stability and concentration. Therefore, these microneedle patches could be useful in cancer therapy [238].
Despite these limitations, exosomes have huge potential as cancer treatment delivery mechanisms. Overall, exosomes hold great promise as targeted delivery systems of therapeutic molecules in nanomedicine due to their unique properties, such as their nanoscale size and ability to be modified. However, further research is needed to overcome the challenges associated with their isolation, modification, and characterization to fully exploit their potential for clinical applications.
8 Conclusions
This study highlights the significance of exosomes in cancer-related processes, drug resistance, and potential therapeutic applications. Exosomes and especially TDEs, play a significant role in cancer-related processes, including TME remodeling, angiogenesis, metastasis, and drug resistance. They facilitate intercellular communication and carry messengers that transport crucial information between tumor cells, the TME and healthy cells, influencing various aspects of cancer progression. Exosomes contribute to drug resistance in cancer by packaging chemotherapy agents and transporting them to sensitive cells. This can promote cancer growth and resistance to treatment. However, they can also be engineered to deliver therapeutic cargo, making them promising candidates for targeted therapy. Moreover, exosomes can be obtained from various sources, including several body fluids, that can be used in liquid biopsies to offer minimally invasive diagnostic and monitoring options for cancer.
Challenges in scaling up production, manufacturing standardization, cargo loading efficiency, and regulatory considerations need to be addressed. Nonetheless, exosomes hold great promise as targeted delivery systems for cancer treatment and other therapeutic applications in nanomedicine. In summary, exosomes represent a promising avenue for cancer diagnosis and treatment, offering the potential for targeted therapy and personalized medicine.
Data availability
No datasets were generated or analysed during the current study.
References
R.A. Willis, Pathology of Tumours (Butterworth & Co., London, 1960)
K. Truskowski, S.R. Amend, K.J. Pienta, Dormant cancer cells: programmed quiescence, senescence, or both? Cancer Metastasis. Rev. 42, 37–47 (2023). https://doi.org/10.1007/S10555-022-10073-Z
D. Hanahan, R.A. Weinberg, Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011). https://doi.org/10.1016/J.CELL.2011.02.013
S. Maitra, S. Sarkar, B. Dhara, Plant-derived exosomes: a new frontier in nano-medicine for cancer and microbial infection therapy. Clin. Transl. Discov. 4, e342 (2024). https://doi.org/10.1002/CTD2.342
J. Kim, Y. Zhu, S. Chen, D. Wang, S. Zhang, J. Xia, S. Li, Q. Qiu, H. Lee, J. Wang, Anti-glioma effect of ginseng-derived exosomes-like nanoparticles by active blood-brain-barrier penetration and tumor microenvironment modulation. J. Nanobiotechnol. 21 (2023). https://doi.org/10.1186/S12951-023-02006-X
G. Liang, Y. Zhu, D.J. Ali, T. Tian, H. Xu, K. Si, B. Sun, B. Chen, Z. Xiao, Engineered exosomes for targeted co-delivery of MiR-21 inhibitor and chemotherapeutics to reverse drug resistance in colon cancer. J. Nanobiotechnol. 18, 1–15 (2020). https://doi.org/10.1186/S12951-019-0563-2
R.M. Johnstone, M. Adam, J.R. Hammond, L. Orr, C. Turbide, Vesicle formation during reticulocyte maturation. Association of plasma membrane activities with released vesicles (exosomes). J. Biol. Chem. 262, 9412–9420 (1987). https://doi.org/10.1016/S0021-9258(18)48095-7
J. Conde-Vancells, E. Rodriguez-Suarez, N. Embade, D. Gil, R. Matthiesen, M. Valle, F. Elortza, S.C. Lu, J.M. Mato, J.M. Falcon-Perez, Characterization and comprehensive proteome profiling of exosomes secreted by hepatocytes. J. Proteome Res. 7, 5157–5166 (2008). https://doi.org/10.1021/pr8004887
E. Chargaff, R. West, The biological significance of the thromboplastic protein of blood. J. Biol. Chem. 166, 189–197 (1946). https://doi.org/10.1016/S0021-9258(17)34997-9
P. Wolf, The nature and significance of platelet products in human plasma. Br. J. Haematol. 13, 269–288 (1967). https://doi.org/10.1111/J.1365-2141.1967.TB08741.X
H.C. Anderson, Vesicles associated with calcification in the matrix of epiphyseal cartilage. J. Cell. Biol. 41, 72 (1969). https://doi.org/10.1083/JCB.41.1.59
Y. Couch, E.I. Buzàs, D.D. Vizio, Y.S. Gho, P. Harrison, A.F. Hill, J. Lötvall, G. Raposo, P.D. Stahl, C. Théry et al., A brief history of nearly EV-erything– the rise and rise of extracellular vesicles. J. Extracell. Vesicles 10, e12144 (2021). https://doi.org/10.1002/JEV2.12144
B. György, T.G. Szabó, M. Pásztói, Z. Pál, P. Misják, B. Aradi, V. László, É. Pállinger, E. Pap, Á. Kittel et al., Membrane vesicles, current state-of-the-art: emerging role of extracellular vesicles. Cell. Mol. Life Sci. 68, 2667–2688 (2011). https://doi.org/10.1007/S00018-011-0689-3
M. Colombo, G. Raposo, C.B. Théry, Secretion, and intercellular interactions of exosomes and other extracellular vesicles. Annu. Rev. Cell Dev. Biol. 30, 255–289 (2014). https://doi.org/10.1146/ANNUREV-CELLBIO-101512-122326
S. Araujo-Abad, A. Manresa-Manresa, E. Rodríguez-Cañas, M. Fuentes-Baile, P. García-Morales, R. Mallavia, M. Saceda, C. De, J. Romero, Glioblastoma-derived small extracellular vesicles: nanoparticles for glioma treatment. Int. J. Mol. Sci. 24, 5910 (2023). https://doi.org/10.3390/IJMS24065910
S. Araujo-Abad, A. Manresa-Manresa, E. Rodríguez-Cañas, M. Fuentes- Baile, P. García-Morales, R. Mallavia, M. Saceda, C. de Juan Romero, New therapy for pancreatic cancer based on extracellular vesicles. Biomed. Pharmacother. 162, 114657 (2023). https://doi.org/10.1016/J.BIOPHA.2023.114657
M. Mathieu, L. Martin-Jaular, G. Lavieu, C. Théry, Specificities of secretion and uptake of exosomes and other extracellular vesicles for cell-to-cell communication. Nat. Cell Biol. 21, 9–17 (2019). https://doi.org/10.1038/s41556-018-0250-9
J.H. Hurley, E. Boura, L.A. Carlson, B. Róycki, Membrane budding. Cell 143, 887 (2010). https://doi.org/10.1016/J.CELL.2010.11.030
Q.F. Han, W.J. Li, K.S. Hu, J. Gao, W.L. Zhai, J.H. Yang, S.J. Zhang, Exosome biogenesis: machinery, regulation, and therapeutic implications in cancer. Mol. Cancer 21, 207 (2022). https://doi.org/10.1186/S12943-022-01671-0
W.M. Henne, N.J. Buchkovich, S.D. Emr, The ESCRT pathway. Dev. Cell. 21, 77–91 (2011). https://doi.org/10.1016/J.DEVCEL.2011.05.015
M. Tschuschke, I. Kocherova, A. Bryja, P. Mozdziak, A. Angelova Volponi, K. Janowicz, R. Sibiak, H. Piotrowska-Kempisty, D. Iżycki, D. Bukowska et al., Inclusion biogenesis, methods of isolation and clinical application of human cellular exosomes. J. Clin. Med. 9, 436 (2020). https://doi.org/10.3390/jcm9020436
T. Skotland, N.P. Hessvik, K. Sandvig, A. Llorente, Exosomal lipid composition and the role of ether lipids and phosphoinositides in exosome biology. J. Lipid Res. 60, 18 (2019). https://doi.org/10.1194/JLR.R084343
G. Dawson, Isolation of lipid rafts (detergent-resistant microdomains) and comparison to extracellular vesicles (exosomes). Methods Mol. Biol. 2187, 99–112 (2021). https://doi.org/10.1007/978-1-0716-0814-2_6
L. Mashouri, H. Yousefi, A.R. Aref, A.M. Ahadi, F. Molaei, S.K. Alahari, Exosomes: composition, biogenesis, and mechanisms in cancer metastasis and drug resistance. Mol. Cancer 18, 1–14 (2019). https://doi.org/10.1186/S12943-019-0991-5
H.G. Zhang, W.E. Grizzle, Exosomes: a novel pathway of local and distant intercellular communication that facilitates the growth and metastasis of neoplastic lesions. Am. J. Pathol. 184, 28–41 (2014). https://doi.org/10.1016/J.AJPATH.2013.09.027
L. Cruz, J.A.A. Romero, R.P. Iglesia, M.H. Lopes, Extracellular vesicles: decoding a new language for cellular communication in early embryonic development. Front. Cell Dev. Biol. 6 (2018). https://doi.org/10.3389/FCELL.2018.00094
A. Zomer, C. Maynard, F.J. Verweij, A. Kamermans, R. Schäfer, E. Beerling, R.M. Schiffelers, E. De Wit, J. Berenguer, S.I.J. Ellenbroek et al., In vivo imaging reveals extracellular vesicle-mediated phenocopying of metastatic behavior. Cell 161, 1046–1057 (2015). https://doi.org/10.1016/j.cell.2015.04.042
M. Katoh, Therapeutics targeting angiogenesis: genetics and epigenetics, extracellular MiRNAs and signaling networks (review). Int. J. Mol. Med. 32, 763–767 (2013). https://doi.org/10.3892/IJMM.2013.1444
A. Monteforte, B. Lam, M.B. Sherman, K. Henderson, A.D. Sligar, A. Spencer, B. Tang, A.K. Dunn, A.B. Baker, Glioblastoma exosomes for therapeutic angiogenesis in peripheral ischemia. Tissue Eng. Part A 23, 1261 (2017). https://doi.org/10.1089/TEN.TEA.2016.0508
L.H. Schmidt, T. Spieker, S. Koschmieder, J. Humberg, D. Jungen, E. Bulk, A. Hascher, D. Wittmer, A. Marra, L. Hillejan et al., The long noncoding MALAT-1 RNA indicates a poor prognosis in non-small cell lung cancer and induces migration and tumor growth. J. Thorac. Oncol. 6, 1984–1992 (2011). https://doi.org/10.1097/JTO.0B013E3182307EAC
S. Pompili, A. Vetuschi, R. Sferra, A. Cappariello, Extracellular vesicles and resistance to anticancer drugs: a tumor skeleton key for unhinging chemotherapies. Front. Oncol. 12 (2022). https://doi.org/10.3389/FONC.2022.933675
C.H. Chang, S. Pauklin, Extracellular vesicles in pancreatic cancer progression and therapies. Cell Death Dis. 12, 1–12 (2021). https://doi.org/10.1038/s41419-021-04258-7
R. Safaei, B.J. Larson, T.C. Cheng, M.A. Gibson, S. Otani, W. Naerdemann, S.B. Howell, Abnormal lysosomal trafficking and enhanced exosomal export of cisplatin in drug-resistant human ovarian carcinoma cells. Mol. Cancer Ther. 4, 1595–1604 (2005). https://doi.org/10.1158/1535-7163.MCT-05-0102
A. Yoshida, T. Fujiwara, K. Uotani, T. Morita, M. Kiyono, S. Yokoo, J. Hasei, E. Nakata, T. Kunisada, T. Ozaki, Clinical and functional significance of intracellular and extracellular microRNA-25-3p in osteosarcoma. Acta Med. Okayama 72, 165–174 (2018). https://doi.org/10.18926/AMO/55857
T.L. Whiteside, Tumor-derived exosomes and their role in cancer progression. Adv. Clin. Chem. 74, 103 (2016). https://doi.org/10.1016/BS.ACC.2015.12.005
S. Araujo-Abad, M. Saceda, C. de Juan Romero, Biomedical application of small extracellular vesicles in cancer treatment. Adv. Drug Deliv. Rev. 182, 114117 (2022). https://doi.org/10.1016/J.ADDR.2022.114117
J. Javadi, A. Görgens, H. Vanky, D. Gupta, A. Hjerpe, S. El-andaloussi, D. Hagey, K. Dobra, Diagnostic and prognostic utility of the extracellular vesicles subpopulations present in pleural effusion. Biomolecules 11, 1606 (2021). https://doi.org/10.3390/BIOM11111606/S1
P. Luo, K. Mao, J. Xu, F. Wu, X. Wang, S. Wang, M. Zhou, L. Duan, Q. Tan, G. Ma et al., Metabolic characteristics of large and small extracellular vesicles from pleural effusion reveal biomarker candidates for the diagnosis of tuberculosis and malignancy. J. Extracell. Vesicles 9 (2020). https://doi.org/10.1080/20013078.2020.1790158
C. Théry, S. Amigorena, G. Raposo, A. Clayton, Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr. Protoc. Cell. Biol. 30, 3.22.1–3.22.29 (2006). https://doi.org/10.1002/0471143030.CB0322S30
P. Tang, L. Tao, C. Yuan, L. Zhang, D. Xiu, Serum derived exosomes from pancreatic cancer patients promoted metastasis: an ITRAQ-based proteomic analysis. Onco. Targets Ther. 12, 9329–9339 (2019). https://doi.org/10.2147/OTT.S229494
H. Zhang, X. Zhang, X. Li, Intraocular exosomes in eye diseases. Curr. Mol. Med. 22, 540–548 (2021). https://doi.org/10.2174/1566524021666210901122948
A. González-Sarrías, C.E. Iglesias-Aguirre, A. Cortés-Martín, F. Vallejo, A. Cattivelli, L. Del Pozo-Acebo, A. Del Saz, M.C.L. De Las Hazas, A. Dávalos, J.C. Espín, Milk-derived exosomes as nanocarriers to deliver curcumin and resveratrol in breast tissue and enhance their anticancer activity. Int. J. Mol. Sci. 23 (2022). https://doi.org/10.3390/IJMS23052860
N. Yunusova, E. Dzhugashvili, A. Yalovaya, L. Kolomiets, A. Shefer, A. Grigor’eva, A. Tupikin, I. Kondakova, S. Tamkovich, Comparative analysis of tumor-associated microRNAs and tetraspanines from exosomes of plasma and ascitic fluids of ovarian cancer patients. Int. J. Mol. Sci. 24, 464 (2023). https://doi.org/10.3390/IJMS24010464/S1
A.S. Vickram, P.S. Srikumar, S. Srinivasan, P. Jeyanthi, K. Anbarasu, S. Thanigaivel, D. Nibedita, D. Jenila Rani, K. Rohini, Seminal exosomes– an important biological marker for various disorders and syndrome in human reproduction. Saudi J. Biol. Sci. 28, 3607–3615 (2021). https://doi.org/10.1016/J.SJBS.2021.03.038
I. Campoy, L. Lanau, T. Altadill, T. Sequeiros, S. Cabrera, M. Cubo-Abert, A. Pérez-Benavente, A. Garcia, S. Borrós, A. Santamaria et al., Exosome-like vesicles in uterine aspirates: a comparison of ultracentrifugation-based isolation protocols. J. Transl. Med. 14, 1–12 (2016). https://doi.org/10.1186/S12967-016-0935-4/FIGURES/4
G.O. Skryabin, A.V. Komelkov, K.I. Zhordania, D.V. Bagrov, S.V. Vinokurova, S.A. Galetsky, N.V. Elkina, D.A. Denisova, A.D. Enikeev, E.M. Tchevkina, Extracellular vesicles from uterine aspirates represent a promising source for screening markers of gynecologic cancers. Cells 11, 1064 (2022). https://doi.org/10.3390/CELLS11071064/S1
S. Nair, K.D. Tang, L. Kenny, C. Punyadeera, Salivary exosomes as potential biomarkers in cancer. Oral. Oncol. 84, 31–40 (2018). https://doi.org/10.1016/J.ORALONCOLOGY.2018.07.001
P. Kangas, T.A. Nyman, L. Metsähonkala, C. Burns, R. Tempest, T. Williams, J. Karttunen, T.S. Jokinen, Towards optimised extracellular vesicle proteomics from cerebrospinal fluid. Sci. Rep. 13, 1–13 (2023). https://doi.org/10.1038/s41598-023-36706-z
M. Tinè, T. Neri, D. Biondini, N. Bernardinello, A. Casara, M. Conti, M. Minniti, M.G. Cosio, M. Saetta, A. Celi et al., Do circulating extracellular vesicles strictly reflect bronchoalveolar lavage extracellular vesicles in COPD? Int. J. Mol. Sci. 24, 2966 (2023). https://doi.org/10.3390/IJMS24032966
A.S. Carvalho, M.C.S. Moraes, C.H. Na, I. Fierro-Monti, A. Henriques, S. Zahedi, C. Bodo, E.M. Tranfield, A.L. Sousa, A. Farinho et al., Is the proteome of bronchoalveolar lavage extracellular vesicles a marker of advanced lung cancer? Cancers 12, 3450 (2020). https://doi.org/10.3390/CANCERS12113450
J. Woo, S. Santasusagna, J. Banks, S. Pastor-Lopez, K. Yadav, M. Carceles-Cordon, A. Dominguez-Andres, R.B. Den, L.R. Languino, R. Pippa et al., Urine extracellular vesicle GATA2 MRNA discriminates biopsy result in men with suspicion of prostate cancer. J. Urol. 204, 691–700 (2020). https://doi.org/10.1097/JU.0000000000001066
B. Dhondt, E. Geeurickx, J. Tulkens, J. Van Deun, G. Vergauwen, L. Lippens, I. Miinalainen, P. Rappu, J. Heino, P. Ost et al., Unravelling the proteomic landscape of extracellular vesicles in prostate cancer by density-based fractionation of urine. J. Extracell. Vesicles 9 (2020). https://doi.org/10.1080/20013078.2020.1736935
R. Muraki, Y. Morita, S. Ida, R. Kitajima, S. Furuhashi, M. Takeda, H. Kikuchi, Y. Hiramatsu, Y. Takanashi, Y. Hamaya et al., Phosphatidylcholine in bile-derived small extracellular vesicles as a novel biomarker of cholangiocarcinoma. Cancer Med. 12, 13007–13018 (2023). https://doi.org/10.1002/CAM4.5973
S. Chaiyadet, J. Sotillo, M. Smout, C. Cantacessi, M.K. Jones, M.S. Johnson, L. Turnbull, C.B. Whitchurch, J. Potriquet, M. Laohaviroj et al., Carcinogenic liver fluke secretes extracellular vesicles that promote cholangiocytes to adopt a tumorigenic phenotype. J. Infect. Dis. 212, 1636–1645 (2015). https://doi.org/10.1093/INFDIS/JIV291
M. Nikanjam, S. Kato, R. Kurzrock, Liquid biopsy: current technology and clinical applications. J. Hematol. Oncol. 15, 131 (2022). https://doi.org/10.1186/S13045-022-01351-Y
P. Stejskal, H. Goodarzi, J. Srovnal, M. Hajdúch, L.J. van’t Veer, M.J.M. Magbanua, Circulating tumor nucleic acids: biology, release mechanisms, and clinical relevance. Mol. Cancer 22, 15 (2023). https://doi.org/10.1186/S12943-022-01710-W
S.N. Lone, S. Nisar, T. Masoodi, M. Singh, A. Rizwan, S. Hashem, W. El-Rifai, D. Bedognetti, S.K. Batra, M. Haris et al., Liquid biopsy: a step closer to transform diagnosis, prognosis and future of cancer treatments. Mol. Cancer 21, 1–22 (2022). https://doi.org/10.1186/S12943-022-01543-7
Z. Zhu, E. Hu, H. Shen, J. Tan, S. Zeng, The functional and clinical roles of liquid biopsy in patient-derived models. J. Hematol. Oncol. 16, 1–17 (2023). https://doi.org/10.1186/S13045-023-01433-5
E. Zhou, Y. Li, F. Wu, M. Guo, J. Xu, S. Wang, Q. Tan, P. Ma, S. Song, Y. Jin, Circulating extracellular vesicles are effective biomarkers for predicting response to cancer therapy. EBioMedicine 67 (2021). https://doi.org/10.1016/j.ebiom.2021.103365
G. Brock, E. Castellanos-Rizaldos, L. Hu, C. Coticchia, J. Skog, Liquid biopsy for cancer screening, patient stratification and monitoring. Transl. Cancer Res. 4, 280–290 (2015). https://doi.org/10.3978/J.ISSN.2218-676X.2015.06.05
T. Guo, X.H. Tang, X.Y. Gao, Y. Zhou, B. Jin, Z.Q. Deng, Y. Hu, X.F. Xing, Z.Y. Li, J.F. Ji, A liquid biopsy signature of circulating exosome-derived MRNAs, MiRNAs and LncRNAs predict therapeutic efficacy to neoadjuvant chemotherapy in patients with advanced gastric cancer. Mol. Cancer 21 (2022). https://doi.org/10.1186/S12943-022-01684-9
Li, H., Sui, T., Chen, X., Gu, Y., Luo, X., Liu, Y., He, Q., Screening and identification of serum exosomal protein ZNF587B in liquid biopsy for ovarian cancer diagnosis. Am. J. Cancer Res. 14, 1904–1913 (2024). https://doi.org/10.62347/RBTM1834
S.A. Melo, L.B. Luecke, C. Kahlert, A.F. Fernandez, S.T. Gammon, J. Kaye, V.S. LeBleu, E.A. Mittendorf, J. Weitz, N. Rahbari et al., Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature 523, 177–182 (2015). https://doi.org/10.1038/NATURE14581
P. Moutinho-Ribeiro, I.A. Batista, S.T. Quintas, B. Adem, M. Silva, R. Morais, A. Peixoto, R. Coelho, P. Costa-Moreira, R. Medas et al., Exosomal glypican-1 is elevated in pancreatic cancer precursors and can signal genetic predisposition in the absence of endoscopic ultrasound abnormalities. World J. Gastroenterol. 28, 4310–4327 (2022). https://doi.org/10.3748/WJG.V28.I31.4310
P. Moutinho-Ribeiro, B. Adem, I. Batista, M. Silva, S. Silva, C.F. Ruivo, R. Morais, A. Peixoto, R. Coelho, P. Costa-Moreira et al., Exosomal glypican-1 discriminates pancreatic ductal adenocarcinoma from chronic pancreatitis. Dig. Liver Dis. 54, 871–877 (2022). https://doi.org/10.1016/J.DLD.2021.10.012
T. Bardol, A.M. Dujon, V. Taly, C. Dunyach-Remy, J.P. Lavigne, B. Costa-Silva, K. Kurma, Z. Eslami-S, L. Cayrefourcq, C. Canivet et al., Early detection of pancreatic cancer by liquid biopsy “PANLIPSY”: a French nation-wide study project. BMC Cancer 24 (2024). https://doi.org/10.1186/S12885-024-12463-8
A.K. Rupp, C. Rupp, S. Keller, J.C. Brase, R. Ehehalt, M. Fogel, G. Moldenhauer, F. Marmé, H. Sültmann, P. Altevogt, Loss of EpCAM expression in breast cancer derived serum exosomes: role of proteolytic cleavage. Gynecol. Oncol. 122, 437–446 (2011). https://doi.org/10.1016/j.ygyno.2011.04.035
C. Liu, X. Xu, B. Li, B. Situ, W. Pan, Y. Hu, T. An, S. Yao, L. Zheng, Single-exosome-counting immunoassays for cancer diagnostics. Nano Lett. 18, 4226–4232 (2018). https://doi.org/10.1021/ACS.NANOLETT.8B01184/SUPPL_FILE/NL8B01184_SI_001.PDF
G. Zhong, K. Wang, J. Li, S. Xiao, W. Wei, J. Liu, Determination of serum exosomal H19 as a noninvasive biomarker for breast cancer diagnosis. Onco Targets Ther. 13, 2563–2571 (2020). https://doi.org/10.2147/OTT.S243601
S. Keller, J. Ridinger, A.K. Rupp, J.W.G. Janssen, P. Altevogt, Body fluid derived exosomes as a novel template for clinical diagnostics. J. Transl. Med. 9, 1–9 (2011). https://doi.org/10.1186/1479-5876-9-86/FIGURES/4
Y. Ogawa, M. Kanai-Azuma, Y. Akimoto, H. Kawakami, R. Yanoshita, Exosome-like vesicles with dipeptidyl peptidase IV in human saliva. Biol. Pharm. Bull. 31, 1059–1062 (2008). https://doi.org/10.1248/BPB.31.1059
R.J. Berckmans, A. Sturk, L.M. Van Tienen, M.C.L. Schaap, R. Nieuwland, Cell-derived vesicles exposing coagulant tissue factor in saliva. Blood 117, 3172–3180 (2011). https://doi.org/10.1182/BLOOD-2010-06-290460
C. Lässer, V. Seyed Alikhani, K. Ekström, M. Eldh, P. Torregrosa Paredes, A. Bossios, M. Sjöstrand, S. Gabrielsson, J. Lötvall, H.H.S. Valadi, Plasma and breast milk exosomes contain RNA: uptake by macrophages. J. Transl. Med. 9, 1–8 (2011). https://doi.org/10.1186/1479-5876-9-9/FIGURES/6
P. Kumar, S. Gupta, B.C. Das, Saliva as a potential non-invasive liquid biopsy for early and easy diagnosis/prognosis of head and neck cancer. Transl. Oncol. 40, 101827 (2024). https://doi.org/10.1016/J.TRANON.2023.101827
L. He, F. Ping, Z. Fan, C. Zhang, M. Deng, B. Cheng, J. Xia, Salivary exosomal MiR-24-3p serves as a potential detective biomarker for oral squamous cell carcinoma screening. Biomed. Pharmacother. 121 (2020). https://doi.org/10.1016/J.BIOPHA.2019.109553
K. Barreiro, H. Holthofer, Urinary extracellular vesicles. A promising shortcut to novel biomarker discoveries. Cell Tissue Res. 369, 217–227 (2017). https://doi.org/10.1007/S00441-017-2621-0
A. Ranghino, V. Dimuccio, E. Papadimitriou, B. Bussolati, Extracellular vesicles in the urine: markers and mediators of tissue damage and regeneration. Clin. Kidney J. 8, 23–30 (2015). https://doi.org/10.1093/CKJ/SFU136
G. Cricrì, L. Bellucci, G. Montini, F. Collino, Urinary extracellular vesicles: uncovering the basis of the pathological processes in kidney-related diseases. Int. J. Mol. Sci. 22 (2021). https://doi.org/10.3390/IJMS22126507
P. Svenningsen, R. Sabaratnam, B.L. Jensen, Urinary extracellular vesicles: origin, role as intercellular messengers and biomarkers; efficient sorting and potential treatment options. Acta Physiol. (Oxf) 228 (2020). https://doi.org/10.1111/APHA.13346
U. Erdbrügger, C.J. Blijdorp, I.V. Bijnsdorp, F.E. Borràs, D. Burger, B. Bussolati, J.B. Byrd, A. Clayton, J.W. Dear, J.M. Falcón-Pérez et al., Urinary extracellular vesicles: a position paper by the urine task force of the international society for extracellular vesicles. J. Extracell. Vesicles 10 (2021). https://doi.org/10.1002/JEV2.12093
M. Wachalska, D. Koppers-Lalic, M. van Eijndhoven, M. Pegtel, A.A. Geldof, A.D. Lipinska, R.J. van Moorselaar, I.V. Bijnsdorp, Protein complexes in urine interfere with extracellular vesicle biomarker studies. J. Circ. Biomark. 5 (2016). https://doi.org/10.5772/62579
A. Kalani, A. Mohan, M.M. Godbole, E. Bhatia, A. Gupta, R.K. Sharma, S. Tiwari, Wilm’s tumor-1 protein levels in urinary exosomes from diabetic patients with or without proteinuria. PLoS One 8 (2013). https://doi.org/10.1371/JOURNAL.PONE.0060177
J. Skog, T. Würdinger, S. van Rijn, D.H. Meijer, L. Gainche, W.T. Curry, B.S. Carter, A.M. Krichevsky, X.O. Breakefield, Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat. Cell Biol. 10, 1470–1476 (2008). https://doi.org/10.1038/NCB1800
C. Liu, C. Hu, T. Chen, Y. Jiang, X. Zhang, H. Liu, Y. Wang, Z. Li, K. Hui, X. Jiang, The role of plasma exosomal Lnc-SNAPC5-3:4 in monitoring the efficacy of anlotinib in the treatment of advanced non-small cell lung cancer. J. Cancer Res. Clin. Oncol. 148, 2867–2879 (2022). https://doi.org/10.1007/S00432-022-04071-5
T.Y. Huang, C.Y. Wang, K.Y. Chen, L.T. Huang, Urinary exosomal thyroglobulin in thyroid cancer patients with post-ablative therapy: a new biomarker in thyroid cancer. Front. Endocrinol. (Lausanne) 11 (2020). https://doi.org/10.3389/FENDO.2020.00382
R. Tutrone, B. Lowentritt, B. Neuman, M.J. Donovan, E. Hallmark, T.J. Cole, Y. Yao, C. Biesecker, S. Kumar, V. Verma et al., ExoDx prostate test as a predictor of outcomes of high-grade prostate cancer - an interim analysis. Prostate Cancer Prostatic. Dis. 26, 596–601 (2023). https://doi.org/10.1038/S41391-023-00675-1
C. Admyre, S.M. Johansson, K.R. Qazi, -J.-J. Filén, R. Lahesmaa, M. Norman, E.P.A. Neve, A. Scheynius, S. Gabrielsson, Exosomes with immune modulatory features are present in human breast milk. J. Immunol. 179, 1969–1978 (2007). https://doi.org/10.4049/JIMMUNOL.179.3.1969
X. Li, L. Su, X. Zhang, Q. Chen, Y. Wang, Z. Shen, T. Zhong, L. Wang, Y. Xiao, X. Feng et al., Recent advances on the function and purification of milk exosomes: a review. Front. Nutrit. 9 (2022). https://doi.org/10.3389/FNUT.2022.871346
W. Qin, Y. Tsukasaki, S. Dasgupta, N. Mukhopadhyay, M. Ikebe, E.R. Sauter, Exosomes in human breast milk promote EMT. Clin. Cancer Res. 22, 4517–4524 (2016). https://doi.org/10.1158/1078-0432.CCR-16-0135
R. Upadhya, A.K. Shetty, Extracellular vesicles for the diagnosis and treatment of parkinson’s disease. Aging Dis. 12, 1438–1450 (2021). https://doi.org/10.14336/AD.2021.0516
D.M. Pegtel, L. Peferoen, S. Amor, Extracellular vesicles as modulators of cell-to-cell communication in the healthy and diseased brain. Philos. Trans. R. Soc. B. 369 (2014). https://doi.org/10.1098/RSTB.2013.0516
N. MacAulay, R.F. Keep, T. Zeuthen, Cerebrospinal fluid production by the choroid plexus: a century of barrier research revisited. Fluids Barriers CNS 19, 1–18 (2022). https://doi.org/10.1186/S12987-022-00323-1
K.Y. Lee, J.H. Im, W. Lin, H.S. Gwak, J.H. Kim, B.C. Yoo, T.H. Kim, J.B. Park, H.J. Park, H.J. Kim et al., Nanoparticles in 472 human cerebrospinal fluid: changes in extracellular vesicle concentration and MiR-21 expression as a biomarker for leptomeningeal metastasis. Cancers 12, 2745 (2020). https://doi.org/10.3390/CANCERS12102745
W.W. Chen, L. Balaj, L.M. Liau, M.L. Samuels, S.K. Kotsopoulos, C.A. Maguire, L. LoGuidice, H. Soto, M. Garrett, L.D. Zhu et al., BEAMing and droplet digital PCR analysis of mutant IDH1 MRNA in glioma patient serum and cerebrospinal fluid extracellular vesicles. Mol. Ther. Nucleic Acids 2, e109 (2013). https://doi.org/10.1038/MTNA.2013.28
J.M. Figueroa, J. Skog, J. Akers, H. Li, R. Komotar, R. Jensen, F. Ringel, I. Yang, S. Kalkanis, R. Thompson et al., Detection of wild-type EGFR amplification and EGFRvIII mutation in CSF-derived extracellular vesicles of glioblastoma patients. Neuro. Oncol. 19, 1494–1502 (2017). https://doi.org/10.1093/NEUONC/NOX085
J.C. Akers, V. Ramakrishnan, R. Kim, S. Phillips, V. Kaimal, Y. Mao, W. Hua, I. Yang, C.C. Fu, J. Nolan et al., MiRNA contents of cerebrospinal fluid extracellular vesicles in glioblastoma patients. J. Neurooncol. 123, 205–216 (2015). https://doi.org/10.1007/S11060-015-1784-3
L. Geng, J. Xu, Y. Zhu, X. Hu, Y. Liu, K. Yang, H. Xiao, Y. Zou, H. Liu, J. Ji et al., Targeting MiR-9 in glioma stem cell-derived extracellular vesicles: a novel diagnostic and therapeutic biomarker. Transl. Oncol. 22 (2022). https://doi.org/10.1016/J.TRANON.2022.101451
Y. Qi, C. Jin, W. Qiu, R. Zhao, S. Wang, B. Li, Z. Zhang, Q. Guo, S. Zhang, Z. Gao et al., The dual role of glioma exosomal microRNAs: glioma eliminates tumor suppressor MiR-1298-5p via exosomes to promote immunosuppressive effects of MDSCs. Cell Death Dis. 13 (2022). https://doi.org/10.1038/S41419-022-04872-Z
Y. Tian, Z. Wang, Y. Wang, B. Yin, J. Yuan, B. Qiang, W. Han, X. Peng, Glioma-derived endothelial cells promote glioma cells migration via extracellular vesicles-mediated transfer of MYO1C. Biochem. Biophys. Res. Commun. 525, 155–161 (2020). https://doi.org/10.1016/J.BBRC.2020.02.017
J. Yin, X. Ge, Z. Shi, C. Yu, C. Lu, Y. Wei, A. Zeng, X. Wang, W. Yan, J. Zhang et al., Extracellular vesicles derived from hypoxic glioma stem-like cells confer temozolomide resistance on glioblastoma by delivering MiR-30b-3p. Theranostics 11, 1763–1779 (2021). https://doi.org/10.7150/THNO.47057
J. Rezaie, M. Feghhi, T. Etemadi, A review on exosomes application in clinical trials: perspective, questions, and challenges. Cell Commun. Signaling 20, 1–13 (2022). https://doi.org/10.1186/S12964-022-00959-4/TABLES/7
K.W. Witwer, E.I. Buzás, L.T. Bemis, A. Bora, C. Lässer, J. Lötvall, E.N. Nolte-’t Hoen, M.G. Piper, S. Sivaraman, J. Skog et al., Standardization of sample collection, isolation and analysis methods in extracellular vesicle research. J. Extracell. Vesicles 2 (2013). https://doi.org/10.3402/JEV.V2I0.20360
X.X. Yang, C. Sun, L. Wang, X.L. Guo, New insight into isolation, identification techniques and medical applications of exosomes. J. Control Release 308, 119–129 (2019). https://doi.org/10.1016/J.JCONREL.2019.07.021
L. Zhu, H.T. Sun, S. Wang, S.L. Huang, Y. Zheng, C.Q. Wang, B.Y. Hu, W. Qin, T.T. Zou, Y. Fu et al., Isolation and characterization of exosomes for cancer research. J. Hematol. Oncol. 13, 1–24 (2020). https://doi.org/10.1186/S13045-020-00987-Y
F. Momen-Heravi, Isolation of extracellular vesicles by ultracentrifugation. Methods Mol. Biol. 1660, 25–32 (2017). https://doi.org/10.1007/978-1-4939-7253-1_3
S. Gupta, S. Rawat, V. Arora, S.K. Kottarath, A.K. Dinda, P.K. Vaishnav, B. Nayak, S. Mohanty, An improvised one-step sucrose cushion ultracentrifugation method for exosome isolation from culture supernatants of mesenchymal stem cells. Stem Cell Res. Ther. 9, 1–11 (2018). https://doi.org/10.1186/S13287-018-0923-0/FIGURES/6
A. Cheruvanky, H. Zhou, T. Pisitkun, J.B. Kopp, M.A. Knepper, P.S.T. Yuen, R.A. Star, Rapid isolation of urinary exosomal biomarkers using a nanomembrane ultrafiltration concentrator. Am. J. Physiol. Renal. Physiol. 292, F1657 (2007). https://doi.org/10.1152/AJPRENAL.00434.2006
A.N. Böing, E.V.D. Pol, A.E. Grootemaat, F.A.W. Coumans, A. Sturk, R. Nieuwland, Single-step isolation of extracellular vesicles by size-exclusion chromatography. J. Extracell. Vesicles 3 (2014). https://doi.org/10.3402/JEV.V3.23430
D.D. Taylor, W. Zacharias, C. Gercel-Taylor, Exosome isolation for proteomic analyses and RNA profiling. Methods Mol. Biol. 728, 235–246 (2011). https://doi.org/10.1007/978-1-61779-068-3_15
D.W. Greening, R. Xu, H. Ji, B.J. Tauro, R.J. Simpson, A protocol for exosome isolation and characterization: evaluation of ultracentrifugation, density-gradient separation, and immunoaffinity capture methods. Methods Mol. Biol. 1295, 179–209 (2015). https://doi.org/10.1007/978-1-4939-2550-6_15
B.J. Tauro, D.W. Greening, R.A. Mathias, H. Ji, S. Mathivanan, A.M. Scott, R.J. Simpson, Comparison of ultracentrifugation, density gradient separation, and immunoaffinity capture methods for isolating human colon cancer cell line LIM1863-derived exosomes. Methods 56, 293–304 (2012). https://doi.org/10.1016/J.YMETH.2012.01.002
M.A. Rider, S.N. Hurwitz, D.G. Meckes, ExtraPEG: a polyethylene glycol-based method for enrichment of extracellular vesicles. Sci. Rep. 6, 1–14 (2016). https://doi.org/10.1038/srep23978
A. Gámez-Valero, M. Monguió-Tortajada, L. Carreras-Planella, M. Franquesa, K. Beyer, F.E. Borràs, Size-exclusion chromatography-based isolation minimally alters extracellular vesicles’ characteristics compared to precipitating agents. Sci. Rep. 6 (2016). https://doi.org/10.1038/SREP33641
M. He, J. Crow, M. Roth, Y. Zeng, A.K. Godwin, Integrated immunoisolation and protein analysis of circulating exosomes using microfluidic technology. Lab. Chip. 14, 3780 (2014). https://doi.org/10.1039/C4LC00662C
L. Saludas, E. Garbayo, A. Ruiz-Villalba, S. Hernández, P. Vader, F. Prósper, M.J. Blanco-Prieto, Isolation methods of large and small extracellular vesicles derived from cardiovascular progenitors: a comparative study. Eur. J. Pharm. Biopharm. 170, 187–196 (2022). https://doi.org/10.1016/J.EJPB.2021.12.012
W.Z. Liu, Z.J. Ma, X.W. Kang, Current status and outlook of advances in exosome isolation. Anal. Bioanal. Chem. 414, 7141 (2022). https://doi.org/10.1007/S00216-022-04253-7
K. Brennan, K. Martin, S.P. FitzGerald, J. O’Sullivan, Y. Wu, A. Blanco, C. Richardson, M.M. Mc Gee, A comparison of methods for the isolation and separation of extracellular vesicles from protein and lipid particles in human serum. Sci. Rep. 10, 1–13 (2020). https://doi.org/10.1038/s41598-020-57497-7
J. Fitzgerald, P. Leonard, E. Darcy, S. Sharma, R. O’Kennedy, Immunoaffinity chromatography: concepts and applications. Methods Mol. Biol. 1485, 27–51 (2017). https://doi.org/10.1007/978-1-4939-6412-3_3
A. Liga, A.D.B. Vliegenthart, W. Oosthuyzen, J.W. Dear, M. Kersaudy-Kerhoas, Exosome isolation: a microfluidic road-map. Lab. Chip. 15, 2388–2394 (2015). https://doi.org/10.1039/C5LC00240K
S.G. Antimisiaris, S. Mourtas, A. Marazioti, Exosomes and exosome-inspired vesicles for targeted drug delivery. Pharmaceutics 10, 218 (2018). https://doi.org/10.3390/PHARMACEUTICS10040218
S.A.A. Kooijmans, L.A.L. Fliervoet, R. Van Der Meel, M.H.A.M. Fens, H.F.G. Heijnen, P.M.P. Van Bergen En Henegouwen, P. Vader, R.M. Schiffelers, PEGylated and targeted extracellular vesicles display enhanced cell specificity and circulation time. J. Control Release 224, 77–85 (2016). https://doi.org/10.1016/J.JCONREL.2016.01.009
S.V. Chitti, C. Nedeva, R. Manickam, P. Fonseka, S. Mathivanan, Extracellular vesicles as drug targets and delivery vehicles for cancer therapy. Pharmaceutics 14 (2022). https://doi.org/10.3390/PHARMACEUTICS14122822
M.A.C. Pomatto, B. Bussolati, S. D’Antico, S. Ghiotto, C. Tetta, M.F. Brizzi, G. Camussi, Improved loading of plasma-derived extracellular vesicles to encapsulate antitumor MiRNAs. Mol. Ther. Methods Clin. Dev. 13, 133–144 (2019). https://doi.org/10.1016/J.OMTM.2019.01.001
L.H. Lv, Y.L. Wan, Y. Lin, W. Zhang, M. Yang, G.N. Li, H.M. Lin, C.Z. Shang, Y.J. Chen, J. Min, Anticancer drugs cause release of exosomes with heat shock proteins from human hepatocellular carcinoma cells that elicit effective natural killer cell antitumor responses in vitro. J. Biol. Chem. 287, 15874–15885 (2012). https://doi.org/10.1074/JBC.M112.340588
E. Torreggiani, L. Roncuzzi, F. Perut, N. Zini, N. Baldini, Multimodal transfer of MDR by exosomes in human osteosarcoma. Int. J. Oncol. 49, 189–196 (2016). https://doi.org/10.3892/IJO.2016.3509
S. Salarpour, H. Forootanfar, M. Pournamdari, M. Ahmadi-Zeidabadi, M. Esmaeeli, A. Pardakhty, Paclitaxel incorporated exosomes derived from glioblastoma cells: comparative study of two loading techniques. Daru 27, 533–539 (2019). https://doi.org/10.1007/S40199-019-00280-5
S.A.A. Kooijmans, S. Stremersch, K. Braeckmans, S.C. De Smedt, A. Hendrix, M.J.A. Wood, R.M. Schiffelers, K. Raemdonck, P. Vader, Electroporation-induced SiRNA precipitation obscures the efficiency of SiRNA loading into extracellular vesicles. J. Control. Release 172, 229–238 (2013). https://doi.org/10.1016/J.JCONREL.2013.08.014
D. Zhang, H. Lee, Z. Zhu, J.K. Minhas, Y. Jin, Enrichment of selective MiRNAs in exosomes and delivery of exosomal MiRNAs in vitro and in vivo. Am. J. Physiol. Lung Cell. Mol. Physiol. 312, L110–L121 (2017). https://doi.org/10.1152/AJPLUNG.00423.2016
S. Ramanathan, S.R. Douglas, G.M. Alexander, B.B. Shenoda, J.E. Barrett, E. Aradillas, A. Sacan, S.K. Ajit, Exosome microRNA signatures in patients with complex regional pain syndrome undergoing plasma exchange. J. Transl. Med. 17, 1–12 (2019). https://doi.org/10.1186/S12967-019-1833-3/FIGURES/6
X.-M. Xi, S.-J. Xia, R. Lu, Drug loading techniques for exosome-based drug delivery systems. Pharmazie 76, 61–67 (2021). https://doi.org/10.1691/ph.2021.0128
S. Bosch, L. De Beaurepaire, M. Allard, M. Mosser, C. Heichette, D. Chrétien, D. Jegou, J.M. Bach, Trehalose prevents aggregation of exosomes and cryodamage. Sci. Rep. 6, 1–11 (2016). https://doi.org/10.1038/srep36162
G. Won Lee, M. Thangavelu, M. Joung Choi, E. Yeong Shin, H. Sol Kim, J. Seon Baek, Y. Woon Jeong, J. Eun Song, C. Carlomagno, J. Miguel Oliveira et al., Exosome mediated transfer of MiRNA-140 promotes enhanced chondrogenic differentiation of bone marrow stem cells for enhanced cartilage repair and regeneration. J. Cell. Biochem. 121, 3642–3652 (2020). https://doi.org/10.1002/JCB.29657
M. Sancho-Albero, M.D.M. Encabo-Berzosa, M. Beltrán-Visiedo, L. Fernández-Messina, V. Sebastián, F. Sánchez-Madrid, M. Arruebo, J. Santamaría, P. Martín-Duque, Efficient encapsulation of theranostic nanoparticles in cell-derived exosomes: leveraging the exosomal biogenesis pathway to obtain hollow gold nanoparticle-hybrids. Nanoscale 11, 18825–18836 (2019). https://doi.org/10.1039/C9NR06183E
F. Lin, R. Wang, Hemolytic mechanism of dioscin proposed by molecular dynamics simulations. J. Mol. Model. 16, 107–118 (2010). https://doi.org/10.1007/S00894-009-0523-0
A. Jeyaram, T.N. Lamichhane, S. Wang, L. Zou, E. Dahal, S.M. Kronstadt, D. Levy, B. Parajuli, D.R. Knudsen, W. Chao et al., Enhanced loading of functional MiRNA cargo via PH gradient modification of extracellular vesicles. Mol. Ther. 28, 975–985 (2020). https://doi.org/10.1016/J.YMTHE.2019.12.007
G. Fuhrmann, A. Serio, M. Mazo, R. Nair, M.M. Stevens, Active loading into extracellular vesicles significantly improves the cellular uptake and photodynamic effect of porphyrins. J. Control. Release 205, 35–44 (2015). https://doi.org/10.1016/J.JCONREL.2014.11.029
L. Alvarez-Erviti, Y. Seow, H. Yin, C. Betts, S. Lakhal, M.J.A. Wood, Delivery of SiRNA to the mouse brain by systemic injection of targeted exosomes. Nat. Biotechnol. 29, 341–345 (2011). https://doi.org/10.1038/nbt.1807
M.J. Haney, N.L. Klyachko, Y. Zhao, R. Gupta, E.G. Plotnikova, Z. He, T. Patel, A. Piroyan, M. Sokolsky, A.V. Kabanov et al., Exosomes as drug delivery vehicles for parkinson’s disease therapy. J. Control. Release 207, 18–30 (2015). https://doi.org/10.1016/J.JCONREL.2015.03.033
O.A. Ojo, T.R. Adeyemo, D. Rotimi, G.E.S. Batiha, G. Mostafa-Hedeab, M.E. Iyobhebhe, T.C. Elebiyo, B. Atunwa, A.B. Ojo, C.M.G. Lima et al., Anticancer properties of curcumin against colorectal cancer: a review. Front. Oncol. 12 (2022). https://doi.org/10.3389/FONC.2022.881641
G.E. Melling, E. Carollo, R. Conlon, J.C. Simpson, D.R.F. Carter, The challenges and possibilities of extracellular vesicles as therapeutic vehicles. Eur. J. Pharm. Biopharm. 144, 50–56 (2019). https://doi.org/10.1016/J.EJPB.2019.08.009
K. O’Brien, S. Ughetto, S. Mahjoum, A.V. Nair, X.O.U. Breakefield, Functionality, and re-release of extracellular vesicle-encapsulated cargo. Cell. Rep. 39, 110651 (2022). https://doi.org/10.1016/J.CELREP.2022.110651
H. Hirose, Y. Hirai, M. Sasaki, H. Sawa, S. Futaki, Quantitative analysis of extracellular vesicle uptake and fusion with recipient cells. Bioconjug. Chem. 33, 1852–1859 (2022). https://doi.org/10.1021/ACS.BIOCONJCHEM.2C00307/ASSET/IMAGES/LARGE/BC2C00307_0003.JPEG
S. Rani, K. O’Brien, F.C. Kelleher, C. Corcoran, S. Germano, M.W. Radomski, J. Crown, L. O’Driscoll, Isolation of exosomes for subsequent MRNA, microRNA, and protein profiling. Methods Mol. Biol. 784, 181–195 (2011). https://doi.org/10.1007/978-1-61779-289-2_13
T. Skotland, K. Sandvig, A. Llorente, Lipids in exosomes: current knowledge and the way forward. Prog. Lipid Res. 66, 30–41 (2017). https://doi.org/10.1016/j.plipres.2017.03.001
R. Wubbolts, R.S. Leckie, P.T.M. Veenhuizen, G. Schwarzmann, W. Möbius, J. Hoernschemeyer, J.W. Slot, H.J. Geuze, W. Stoorvogel, Proteomic and biochemical analyses of human B cell-derived exosomes: potential implications for their function and multivesicular body formation. J. Biol. Chem. 278, 10963–10972 (2003). https://doi.org/10.1074/jbc.M207550200
Y. Huang, Z. Liu, N. Li, C. Tian, H. Yang, Y. Huo, Y. Li, J. Zhang, Z. Yu, Parkinson’s disease derived exosomes aggravate neuropathology in SNCA*A53T mice. Ann. Neurol. 92, 230–245 (2022). https://doi.org/10.1002/ANA.26421
S. Biagiotti, F. Abbas, M. Montanari, C. Barattini, L. Rossi, M. Magnani, S. Papa, B. Canonico, Extracellular vesicles as new players in drug delivery: a focus on red blood cells-derived EVs. Pharmaceutics 15 (2023). https://doi.org/10.3390/PHARMACEUTICS15020365
I.K. Herrmann, M.J.A. Wood, G. Fuhrmann, Extracellular vesicles as a next-generation drug delivery platform. Nat. Nanotechnol. 16, 748–759 (2021). https://doi.org/10.1038/S41565-021-00931-2
Y. Zhang, J. Bi, J. Huang, Y. Tang, S. Du, P. Li, Exosome: a review of its classification, isolation techniques, storage, diagnostic and targeted therapy applications. Int. J. Nanomed. 15, 6917–6934 (2020). https://doi.org/10.2147/IJN.S264498
S.M. Kim, Y. Yang, S.J. Oh, Y. Hong, M. Seo, M. Jang, Cancer-derived exosomes as a delivery platform of CRISPR/Cas9 confer cancer cell tropism-dependent targeting. J. Control. Release 266, 8–16 (2017). https://doi.org/10.1016/J.JCONREL.2017.09.013
Y. Lin, J. Wu, W. Gu, Y. Huang, Z. Tong, L. Huang, J. Tan, Exosome-liposome hybrid nanoparticles deliver CRISPR/Cas9 system in MSCs. Adv. Sci. (Weinh) 5 (2018). https://doi.org/10.1002/ADVS.201700611
P. Gee, M.S.Y. Lung, Y. Okuzaki, N. Sasakawa, T. Iguchi, Y. Makita, H. Hozumi, Y. Miura, L.F. Yang, M. Iwasaki et al., Extracellular nanovesicles for packaging of CRISPR-Cas9 protein and SgRNA to induce therapeutic exon skipping. Nat. Commun. 11 (2020). https://doi.org/10.1038/S41467-020-14957-Y
X. Zhang, H. Zhang, J. Gu, J. Zhang, H. Shi, H. Qian, D. Wang, W. Xu, J. Pan, H.A. Santos, Engineered extracellular vesicles for cancer therapy. Adv. Mater. 33 (2021). https://doi.org/10.1002/ADMA.202005709
A. Mizrak, M.F. Bolukbasi, G.B. Ozdener, G.J. Brenner, S. Madlener, E.P. Erkan, T. Ströbel, X.O. Breakefield, O. Saydam, Genetically engineered microvesicles carrying suicide MRNA/protein inhibit schwannoma tumor growth. Mol. Ther. 21, 101–108 (2013). https://doi.org/10.1038/MT.2012.161
E.P. Erkan, D. Senfter, S. Madlener, G. Jungwirth, T. Ströbel, N. Saydam, O. Saydam, Extracellular vesicle-mediated suicide MRNA/protein delivery inhibits glioblastoma tumor growth in vivo. Cancer Gene Ther. 24, 38–44 (2017). https://doi.org/10.1038/CGT.2016.78
M. Kanada, B.D. Kim, J.W. Hardy, J.A. Ronald, M.H. Bachmann, M.P. Bernard, G.I. Perez, A.A. Zarea, T.J. Ge, A. Withrow et al., Microvesicle-mediated delivery of minicircle DNA results in effective gene-directed enzyme prodrug cancer therapy. Mol. Cancer Ther. 18, 2331–2342 (2019). https://doi.org/10.1158/1535-7163.MCT-19-0299
A. Ursula, J. Jana, B. Katarina, P. Petra, P. Martin, P. Pavel, T. Ondrej, K. Juraj, Z. Martina, R. Vanda et al., Prodrug suicide gene therapy for cancer targeted intracellular by mesenchymal stem cell exosomes. Int. J. Cancer 144, 897–908 (2019). https://doi.org/10.1002/IJC.31792
S. Kamerkar, V.S. Lebleu, H. Sugimoto, S. Yang, C.F. Ruivo, S.A. Melo, J.J. Lee, R. Kalluri, Exosomes facilitate therapeutic targeting of oncogenic KRAS in pancreatic cancer. Nature 546, 498–503 (2017). https://doi.org/10.1038/NATURE22341
F. Pi, D.W. Binzel, T.J. Lee, Z. Li, M. Sun, P. Rychahou, H. Li, F. Haque, S. Wang, C.M. Croce et al., Nanoparticle orientation to control RNA loading and ligand display on extracellular vesicles for cancer regression. Nat. Nanotechnol. 13, 82–89 (2018). https://doi.org/10.1038/S41565-017-0012-Z
Z. Zheng, Z. Li, C. Xu, B. Guo, P. Guo, Folate-displaying exosome mediated cytosolic delivery of SiRNA avoiding endosome trapping. J. Control. Release 311–312, 43–49 (2019). https://doi.org/10.1016/J.JCONREL.2019.08.021
M. Katakowski, B. Buller, X. Zheng, Y. Lu, T. Rogers, O. Osobamiro, W. Shu, F. Jiang, M. Chopp, Exosomes from marrow stromal cells expressing MiR-146b inhibit glioma growth. Cancer Lett. 335, 201–204 (2013). https://doi.org/10.1016/J.CANLET.2013.02.019
F.M. Lang, A. Hossain, J. Gumin, E.N. Momin, Y. Shimizu, D. Ledbetter, T. Shahar, S. Yamashita, B. Parker Kerrigan, J. Fueyo et al., Mesenchymal stem cells as natural biofactories for exosomes carrying MiR-124a in the treatment of gliomas. Neuro. Oncol. 20, 380–390 (2018). https://doi.org/10.1093/NEUONC/NOX152
H. Zhang, J. Wang, T. Ren, Y. Huang, X. Liang, Y. Yu, W. Wang, J. Niu, W. Guo, Bone marrow mesenchymal stem cell-derived exosomal MiR-206 inhibits osteosarcoma progression by targeting TRA2B. Cancer Lett. 490, 54–65 (2020). https://doi.org/10.1016/J.CANLET.2020.07.008
G. Liang, S. Kan, Y. Zhu, S. Feng, W. Feng, S. Gao, Engineered exosome-mediated delivery of functionally active MiR-26a and its enhanced suppression effect in HepG2 cells. Int. J. Nanomed. 13, 585–599 (2018). https://doi.org/10.2147/IJN.S154458
G. Lou, X. Song, F. Yang, S. Wu, J. Wang, Z. Chen, Y. Liu, Exosomes derived from MiR-122-modified adipose tissue-derived MSCs increase chemosensitivity of hepatocellular carcinoma. J. Hematol. Oncol. 8 (2015). https://doi.org/10.1186/S13045-015-0220-7
H. Nie, X. Xie, D. Zhang, Y. Zhou, B. Li, F. Li, F. Li, Y. Cheng, H. Mei, H. Meng et al., Use of lung-specific exosomes for MiRNA-126 delivery in non-small cell lung cancer. Nanoscale 12, 877–887 (2020). https://doi.org/10.1039/C9NR09011H
S.I. Ohno, M. Takanashi, K. Sudo, S. Ueda, A. Ishikawa, N. Matsuyama, K. Fujita, T. Mizutani, T. Ohgi, T. Ochiya et al., Systemically injected exosomes targeted to EGFR deliver antitumor microRNA to breast cancer cells. Mol. Ther. 21, 185–191 (2013). https://doi.org/10.1038/MT.2012.180
J.L. Munoz, S.A. Bliss, S.J. Greco, S.H. Ramkissoon, K.L. Ligon, P. Rameshwar, Delivery of functional anti-MiR-9 by mesenchymal stem cell-derived exosomes to glioblastoma multiforme cells conferred chemosensitivity. Mol. Ther. Nucleic Acids 2 (2013). https://doi.org/10.1038/MTNA.2013.60
X. Wang, H. Zhang, M. Bai, T. Ning, S. Ge, T. Deng, R. Liu, L. Zhang, G. Ying, Y. Ba, Exosomes serve as nanoparticles to deliver anti-MiR-214 to reverse chemoresistance to cisplatin in gastric cancer. Mol. Ther. 26, 774–783 (2018). https://doi.org/10.1016/J.YMTHE.2018.01.001
R. Ji, X. Zhang, H. Gu, J. Ma, X. Wen, J. Zhou, H. Qian, W. Xu, J. Qian, J. Lin, MiR-374a-5p: a new target for diagnosis and drug resistance therapy in gastric cancer. Mol. Ther. Nucleic Acids 18, 320–331 (2019). https://doi.org/10.1016/J.OMTN.2019.07.025
N. Yim, S.W. Ryu, K. Choi, K.R. Lee, S. Lee, H. Choi, J. Kim, M.R. Shaker, W. Sun, J.H. Park et al., Exosome engineering for efficient intracellular delivery of soluble proteins using optically reversible protein–protein interaction module. Nat. Commun. 7 (2016). https://doi.org/10.1038/NCOMMS12277
H. Choi, H. Yim, C. Park, S.H. Ahn, Y. Ahn, A. Lee, H. Yang, C. Choi, Targeted delivery of exosomes armed with anti-cancer therapeutics. Membranes 12, 85 (2022). https://doi.org/10.3390/MEMBRANES12010085
M.Y. Koo, J. Park, J.M. Lim, S.Y. Joo, S.P. Shin, H.B. Shim, J. Chung, D. Kang, H.A. Woo, S.G. Rhee, Selective inhibition of the function of tyrosine-phosphorylated STAT3 with a phosphorylation site-specific intrabody. Proc. Natl. Acad. Sci. U. S. A. 111, 6269–6274 (2014). https://doi.org/10.1073/PNAS.1316815111/-/DCSUPPLEMENTAL/PNAS.201316815SI.PDF
J.R. Aspe, C.J.D. Osterman, J.M.S. Jutzy, S. Deshields, S. Whang, N.R. Wall, Enhancement of gemcitabine sensitivity in pancreatic adenocarcinoma by novel exosome-mediated delivery of the survivin-T34A mutant. J. Extracell. Vesicles 3, 1–9 (2014). https://doi.org/10.3402/JEV.V3.23244
L. Tao, J. Zhou, C. Yuan, L. Zhang, D. Li, D. Si, D. Xiu, L. Zhong, Metabolomics identifies serum and exosomes metabolite markers of pancreatic cancer. Metabolomics 15, 1–11 (2019). https://doi.org/10.1007/S11306-019-1550-1/METRICS
K.C. Vallabhaneni, P. Penfornis, S. Dhule, F. Guillonneau, K.V. Adams, Y.Y. Mo, R. Xu, Y. Liu, K. Watabe, M.C. Vemuri et al., Extracellular vesicles from bone marrow mesenchymal stem/stromal cells transport tumor regulatory microRNA, proteins, and metabolites. Oncotarget 6, 4953 (2015). https://doi.org/10.18632/ONCOTARGET.3211
J.L. Palacios-Ferrer, M.B. García-Ortega, M. Gallardo-Gómez, M.Á. García, C. Díaz, H. Boulaiz, J. Valdivia, J.M. Jurado, F.M. Almazan-Fernandez, S. Arias-Santiago et al., Metabolomic profile of cancer stem cell-derived exosomes from patients with malignant melanoma. Mol. Oncol. 15, 407 (2021). https://doi.org/10.1002/1878-0261.12823
E. Yang, X. Wang, Z. Gong, M. Yu, H. Wu, D. Zhang, Exosome-mediated metabolic reprogramming: the emerging role in tumor microenvironment remodeling and its influence on cancer progression. Signal Transd. Targeted Ther. 5, 1–13 (2020). https://doi.org/10.1038/s41392-020-00359-5
H. Qi, C. Liu, L. Long, Y. Ren, S. Zhang, X. Chang, X. Qian, H. Jia, J. Zhao, J. Sun et al., Blood exosomes endowed with magnetic and targeting properties for cancer therapy. ACS Nano 10, 3323–3333 (2016). https://doi.org/10.1021/ACSNANO.5B06939
L. Pascucci, V. Coccè, A. Bonomi, D. Ami, P. Ceccarelli, E. Ciusani, L. Viganò, A. Locatelli, F. Sisto, S.M. Doglia et al., Paclitaxel is incorporated by mesenchymal stromal cells and released in exosomes that inhibit in vitro tumor growth: a new approach for drug delivery. J. Control. Release 192, 262–270 (2014). https://doi.org/10.1016/J.JCONREL.2014.07.042
S. Araujo-Abad, A. Manresa-Manresa, E. Rodríguez-Cañas, M. Fuentes-Baile, P. García-Morales, R. Mallavia, M. Saceda, C. de Juan Romero, Glioblastoma-derived small extracellular vesicles: nanoparticles for glioma treatment. Int. J. Mol. Sci. 24 (2023). https://doi.org/10.3390/IJMS24065910
A.K. Agrawal, F. Aqil, J. Jeyabalan, W.A. Spencer, J. Beck, B.W. Gachuki, S.S. Alhakeem, K. Oben, R. Munagala, S. Bondada et al., Milk-derived exosomes for oral delivery of paclitaxel. Nanomedicine 13, 1627–1636 (2017). https://doi.org/10.1016/J.NANO.2017.03.001
M.S. Kim, M.J. Haney, Y. Zhao, V. Mahajan, I. Deygen, N.L. Klyachko, E. Inskoe, A. Piroyan, M. Sokolsky, O. Okolie et al., Development of exosome-encapsulated paclitaxel to overcome MDR in cancer cells. Nanomedicine 12, 655–664 (2016). https://doi.org/10.1016/J.NANO.2015.10.012
M.S. Kim, M.J. Haney, Y. Zhao, D. Yuan, I. Deygen, N.L. Klyachko, A.V. Kabanov, E.V. Batrakova, Engineering macrophage-derived exosomes for targeted paclitaxel delivery to pulmonary metastases: in vitro and in vivo evaluations. Nanomedicine 14, 195–204 (2018). https://doi.org/10.1016/J.NANO.2017.09.011
F. Susa, T. Limongi, B. Dumontel, V. Vighetto, V. Cauda, Engineered extracellular vesicles as a reliable tool in cancer nanomedicine. Cancers 11, 1979 (2019). https://doi.org/10.3390/CANCERS11121979
A.K.A. Silva, J. Kolosnjaj-Tabi, S. Bonneau, I. Marangon, N. Boggetto, K. Aubertin, O. Clément, M.F. Bureau, N. Luciani, F. Gazeau et al., Magnetic and photoresponsive theranosomes: translating cell-released vesicles into smart nanovectors for cancer therapy. ACS Nano 7, 4954–4966 (2013). https://doi.org/10.1021/NN40026
A.K.A. Silva, N. Luciani, F. Gazeau, K. Aubertin, S. Bonneau, C. Chauvierre, D. Letourneur, C. Wilhelm, Combining magnetic nanoparticles with cell derived microvesicles for drug loading and targeting. Nanomedicine 11, 645–655 (2015). https://doi.org/10.1016/J.NANO.2014.11.009
Y. Liu, L. Bai, K. Guo, Y. Jia, K. Zhang, Q. Liu, P. Wang, X. Wang, Focused ultrasound-augmented targeting delivery of nanosonosensitizers from homogenous exosomes for enhanced sonodynamic cancer therapy. Theranostics 9, 5261–5281 (2019). https://doi.org/10.7150/THNO.33183
B. Dumontel, F. Susa, T. Limongi, V. Vighetto, D. Debellis, M. Canta, V. Cauda, Nanotechnological engineering of extracellular vesicles for the development of actively targeted hybrid nanodevices. Cell Biosci. 12 (2022). https://doi.org/10.1186/S13578-022-00784-9
B. Dumontel, C. Jiménez-Jiménez, M. Vallet-Regí, M. Manzano, Bioinspired extracellular vesicle-coated silica nanoparticles as selective delivery systems. Mater. Today Bio. 23, 100850 (2023). https://doi.org/10.1016/J.MTBIO.2023.100850
M. Haris, S.K. Yadav, A. Rizwan, A. Singh, E. Wang, H. Hariharan, R. Reddy, F.M. Marincola, Molecular magnetic resonance imaging in cancer. J. Transl. Med. 13, 313 (2015). https://doi.org/10.1186/S12967-015-0659-X
S. Rayamajhi, R. Marasini, T.D.T. Nguyen, B.L. Plattner, D. Biller, S. Aryal, Strategic reconstruction of macrophage-derived extracellular vesicles as a magnetic resonance imaging contrast agent. Biomater. Sci. 8, 2887–2904 (2020). https://doi.org/10.1039/D0BM00128G
Z. Varga, I. Gyurkó, K. Pálóczi, E.I. Buzás, I. Horváth, N. Hegedus, D. Máthé, K. Szigeti, Radiolabeling of extracellular vesicles with (99m)Tc for quantitative in vivo imaging studies. Cancer Biother Radiopharm. 31, 168–173 (2016). https://doi.org/10.1089/CBR.2016.2009
S. Bai, Y. Wei, R. Liu, R. Xu, L. Xiang, J. Du, Role of tumour-derived exosomes in metastasis. Biomed. Pharmacother. 147, 112657 (2022). https://doi.org/10.1016/J.BIOPHA.2022.112657
F. Huang, J. Wan, S. Hao, X. Deng, L. Chen, L. Ma, TGF-Β1-silenced leukemia cell-derived exosomes target dendritic cells to induce potent anti-leukemic immunity in a mouse model. Cancer Immunol. Immunother. 66, 1321–1331 (2017). https://doi.org/10.1007/S00262-017-2028-5/FIGURES/4
R. Hosseini, H. Sarvnaz, M. Arabpour, S.M. Ramshe, L. Asef-Kabiri, H. Yousefi, M.E. Akbari, N. Eskandari, Cancer exosomes and natural killer cells dysfunction: biological roles, clinical significance and implications for immunotherapy. Mol. Cancer 21, 1–18 (2022). https://doi.org/10.1186/S12943-021-01492-7
Z.H. Wang, Z.A. Li, X.Y. Gu, X.F. Bai, Review of research pertaining to the effect of tumor-derived exosomes on natural killer cells. J. Physiol. Pharmacol. 74, 481–487 (2023). https://doi.org/10.26402/JPP.2023.5.01
K.H. Han, C.H. Kim, S.H. Kim, C.H. Lee, M. Park, V.D. Bui, V.H. Duong, S. Kwon, M. Ha, H. Kang et al., Immunogenic extracellular vesicles derived from endoplasmic reticulum-stressed tumor cells: implications as the therapeutic cancer vaccine. ACS Nano 18, 199–209 (2024). https://doi.org/10.1021/ACSNANO.3C05645/SUPPL_FILE/NN3C05645_SI_001.PDF
A.M. Marleau, C.S. Chen, J.A. Joyce, R.H. Tullis, Exosome removal as a therapeutic adjuvant in cancer. J. Transl. Med. 10, 134 (2012). https://doi.org/10.1186/1479-5876-10-134
F. Huang, Z. Li, W. Zhang, J. Li, S. Hao, Enhancing the anti-leukemia immunity of acute lymphocytic leukemia-derived exosome-based vaccine by downregulation of PD-L1 expression. Cancer Immunol. Immunother. 71, 2197–2212 (2022). https://doi.org/10.1007/S00262-021-03138-5/TABLES/2
W. Zhou, X. Chen, Y. Zhou, S. Shi, C. Liang, X. Yu, H. Chen, Q. Guo, Y. Zhang, P. Liu et al., Exosomes derived from immunogenically dying tumor cells as a versatile tool for vaccination against pancreatic cancer. Biomaterials 280, 121306 (2022). https://doi.org/10.1016/J.BIOMATERIALS.2021.121306
Q. Tong, K. Li, F. Huang, Y. Dai, T. Zhang, M. Muaibati, A. Abuduyilimu, X. Huang, Extracellular vesicles hybrid plasmid-loaded lipid nanovesicles for synergistic cancer immunotherapy. Mater. Today Bio. 23, 100845 (2023). https://doi.org/10.1016/J.MTBIO.2023.100845
L. Huang, Y. Rong, X. Tang, K. Yi, P. Qi, J. Hou, W. Liu, Y. He, X. Gao, C. Yuan et al., Engineered exosomes as an in situ DC-primed vaccine to boost antitumor immunity in breast cancer. Mol. Cancer 21 (2022). https://doi.org/10.1186/S12943-022-01515-X
B. Zuo, Y. Zhang, K. Zhao, L. Wu, H. Qi, R. Yang, X. Gao, M. Geng, Y. Wu, R. Jing et al., Universal immunotherapeutic strategy for hepatocellular carcinoma with exosome vaccines that engage adaptive and innate immune responses. J. Hematol. Oncol. 15 (2022). https://doi.org/10.1186/S13045-022-01266-8
J. Li, J. Li, Y. Peng, Y. Du, Z. Yang, X. Qi, Dendritic cell derived exosomes loaded neoantigens for personalized cancer immunotherapies. J. Control. Release 353, 423–433 (2023). https://doi.org/10.1016/J.JCONREL.2022.11.053
I. Radziejewska, Tumor-associated carbohydrate antigens of MUC1 - implication in cancer development. Biomed. Pharmacother. 174 (2024). https://doi.org/10.1016/J.BIOPHA.2024.116619
H. Zhu, K. Wang, Z. Wang, D. Wang, X. Yin, Y. Liu, F. Yu, W. Zhao, An efficient and safe MUC1-dendritic cell-derived exosome conjugate vaccine elicits potent cellular and humoral immunity and tumor inhibition in vivo. Acta Biomater. 138, 491–504 (2022). https://doi.org/10.1016/J.ACTBIO.2021.10.041
A. Salmaninejad, S.M. Layeghi, Z. Falakian, S. Golestani, S. Kobravi, S. Talebi, M. Yousefi, An update to experimental and clinical aspects of tumor-associated macrophages in cancer development: hopes and pitfalls. Clin. Exp. Med. 24 (2024). https://doi.org/10.1007/S10238-024-01417-W
S. Kamerkar, C. Leng, O. Burenkova, S.C. Jang, C. McCoy, K. Zhang, K. Dooley, S. Kasera, T. Zi, S. Sisó et al., Exosome-mediated genetic reprogramming of tumor-associated macrophages by ExoASO-STAT6 leads to potent monotherapy antitumor activity. Sci. Adv. 8 (2022). https://doi.org/10.1126/SCIADV.ABJ7002
K.E. Sivick, A.L. Desbien, L.H. Glickman, G.L. Reiner, L. Corrales, N.H. Surh, T.E. Hudson, U.T. Vu, B.J. Francica, T. Banda et al., Magnitude of therapeutic STING activation determines CD8+ T cell-mediated anti-tumor immunity. Cell. Rep. 25, 3074–3085.e5 (2018). https://doi.org/10.1016/J.CELREP.2018.11.047
S.C. Jang, K.D. Economides, R.J. Moniz, C.L. Sia, N. Lewis, C. McCoy, T. Zi, K. Zhang, R.A. Harrison, J. Lim et al., ExoSTING, an extracellular vesicle loaded with STING agonists, promotes tumor immune surveillance. Commun. Biol. 4 (2021). https://doi.org/10.1038/S42003-021-02004-5
S.A.A. Kooijmans, P. Vader, S.M. van Dommelen, W.W. van Solinge, R.M. Schiffelers, Exosome mimetics: a novel class of drug delivery systems. Int. J. Nanomed. 7, 1525 (2012). https://doi.org/10.2147/IJN.S29661
J. Wolfram, M. Ferrari, Clinical cancer nanomedicine. Nano Today 25, 85–98 (2019). https://doi.org/10.1016/j.nantod.2019.02.005
K.W. Witwer, J. Wolfram, Extracellular vesicles versus synthetic nanoparticles for drug delivery. Nat. Rev. Mater. 6, 106 (2021). https://doi.org/10.1038/S41578-020-00277-6
C.K. Das, B.C. Jena, I. Banerjee, S. Das, A. Parekh, S.K. Bhutia, M. Mandal, Exosome as a novel shuttle for delivery of therapeutics across biological barriers. Mol. Pharm. 16, 24–40 (2018). https://doi.org/10.1021/ACS.MOLPHARMACEUT.8B00901
S. EL Andaloussi, S. Lakhal, I. Mäger, M.J.A. Wood, Exosomes for targeted SiRNA delivery across biological barriers. Adv. Drug Deliv. Rev. 65, 391–397 (2013). https://doi.org/10.1016/J.ADDR.2012.08.008
G. Kibria, E.K. Ramos, Y. Wan, D.R. Gius, H. Liu, Exosomes as a drug delivery system in cancer therapy: potential and challenges. Mol. Pharm. 15, 3633 (2018). https://doi.org/10.1021/ACS.MOLPHARMACEUT.8B00277
F. Soltani, H. Parhiz, A. Mokhtarzadeh, M. Ramezani, Synthetic and biological vesicular nano-carriers designed for gene delivery. Curr. Pharm. Des. 21, 6214–6235 (2015). https://doi.org/10.2174/1381612821666151027153410
T.L. Whiteside, Immune modulation of T-cell and NK (natural killer) cell activities by TEXs (tumour-derived exosomes). Biochem. Soc. Trans. 41, 251 (2013). https://doi.org/10.1042/BST20120265
J.L. Hood, S.A. Wickline, A systematic approach to exosome-based translational nanomedicine. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 4, 458–467 (2012). https://doi.org/10.1002/WNAN.1174
G. Van Niel, G. D’Angelo, G. Raposo, Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Biol. 19, 213–228 (2018). https://doi.org/10.1038/nrm.2017.125
C.Y. Ng, L.T. Kee, M.E. Al-Masawa, Q.H. Lee, T. Subramaniam, D. Kok, M.H. Ng, J.X. Law, Scalable production of extracellular vesicles and its therapeutic values: a review. Int. J. Mol. Sci. 23 (2022). https://doi.org/10.3390/IJMS23147986
Z. Wu, D. He, H. Li, Bioglass enhances the production of exosomes and improves their capability of promoting vascularization. Bioact. Mater. 6, 823–835 (2021). https://doi.org/10.1016/J.BIOACTMAT.2020.09.011
D. Jafari, S. Shajari, R. Jafari, N. Mardi, H. Gomari, F. Ganji, M. Forouzandeh Moghadam, A. Samadikuchaksaraei, Designer exosomes: a new platform for biotechnology therapeutics. BioDrugs 34, 567–586 (2020). https://doi.org/10.1007/S40259-020-00434-X
F. Li, J. Wu, D. Li, L. Hao, Y. Li, D. Yi, K.W.K. Yeung, D. Chen, W.W. Lu, H. Pan et al., Engineering stem cells to produce exosomes with enhanced bone regeneration effects: an alternative strategy for gene therapy. J. Nanobiotechnol. 20, 1–23 (2022). https://doi.org/10.1186/S12951-022-01347-3/TABLES/1
J. Wang, E.E. Bonacquisti, A.D. Brown, J. Nguyen, Boosting the biogenesis and secretion of mesenchymal stem cell-derived exosomes. Cells 9, 660 (2020). https://doi.org/10.3390/CELLS9030660
M.N.F.B. Hassan, M.D. Yazid, M.H.M. Yunus, S.R. Chowdhury, Y. Lokanathan, R.B.H. Idrus, A.M.H. Ng, J.X. Law, Large-scale expansion of human mesenchymal stem cells. Stem Cells Int. 2020 (2020). https://doi.org/10.1155/2020/9529465
D. Zha, S. Rayamajhi, J. Sipes, A. Russo, H.B. Pathak, K. Li, M.E. Sardiu, L.E. Bantis, A. Mitra, R.V. Puri et al., Proteomic profiling of fallopian tube-derived extracellular vesicles using a microfluidic tissue-on-chip system. Bioengineering 10 (2023). https://doi.org/10.3390/BIOENGINEERING10040423
H. Kang, Y.H. Bae, Y. Kwon, S. Kim, J. Park, Extracellular vesicles generated using bioreactors and their therapeutic effect on the acute kidney injury model. Adv. Healthc. Mater. 11, 2101606 (2022). https://doi.org/10.1002/ADHM.202101606
D.C. Watson, D. Bayik, A. Srivatsan, C. Bergamaschi, A. Valentin, G. Niu, J. Bear, M. Monninger, M. Sun, A. Morales-Kastresana et al., Efficient production and enhanced tumor delivery of engineered extracellular vesicles. Biomaterials 105, 195 (2016). https://doi.org/10.1016/J.BIOMATERIALS.2016.07.003
J. Cao, B. Wang, T. Tang, L. Lv, Z. Ding, Z. Li, R. Hu, Q. Wei, A. Shen, Y. Fu et al., Three-dimensional culture of MSCs produces exosomes with improved yield and enhanced therapeutic efficacy for cisplatin-induced acute kidney injury. Stem Cell Res. Ther. 11 (2020). https://doi.org/10.1186/S13287-020-01719-2
X. Feng, X. Chen, X. Zheng, H. Zhu, Q. Qi, S. Liu, H. Zhang, J. Che, Latest trend of milk derived exosomes: cargos, functions, and applications. Front. Nutrit. 8, 747294 (2021). https://doi.org/10.3389/FNUT.2021.747294/BIBTEX
J.L. Betker, B.M. Angle, M.W. Graner, T.J. Anchordoquy, The potential of exosomes from cow milk for oral delivery. J. Pharm. Sci. 108, 1496 (2019). https://doi.org/10.1016/J.XPHS.2018.11.022
M.A. Babaker, F.A. Aljoud, F. Alkhilaiwi, A. Algarni, A. Ahmed, M.I. Khan, I.M. Saadeldin, F.A. Alzahrani, The therapeutic potential of milk extracellular vesicles on colorectal cancer. Int. J. Mol. Sci. 23, 6812 (2022). https://doi.org/10.3390/IJMS23126812
C. Saura, C. Ortiz, J. Matito, E.J. Arenas, A. Suñol, Á. Martín, O. Córdoba, A. Martínez-Sabadell, I. García-Ruiz, I. Miranda et al., Early-stage breast cancer detection in breast milk. Cancer Discov. 13, 2191 (2023). https://doi.org/10.1158/2159-8290.CD-22-1340
Z.L. Tan, J.F. Li, H.M. Luo, Y.Y. Liu, Y. Jin, Plant extracellular vesicles: a novel bioactive nanoparticle for tumor therapy. Front. Pharmacol. 13, 1006299 (2022). https://doi.org/10.3389/FPHAR.2022.1006299/BIBTEX
J.F. Creeden, J. Sevier, J.T. Zhang, Y. Lapitsky, F.C. Brunicardi, G. Jin, J. Nemunaitis, J.Y. Liu, A. Kalinoski, D. Rao et al., Smart exosomes enhance PDAC targeted therapy. J. Control Release 368, 413–429 (2024). https://doi.org/10.1016/J.JCONREL.2024.02.037
S. Mu, H. Chang, F. Qu, Fabrication and characterization of microneedle patches for loading and delivery of exosomes. J. Visualized Exp. (2024). https://doi.org/10.3791/67109
Acknowledgements
The authors apologize to many colleagues whose work could not be cited owing to space restrictions. The authors thank Pilar García Morales and Víctor Manuel Barberá for frequent discussions.
Funding
Work in Dr. de Juan’s lab was supported by Instituto de Salud Carlos III (ISCIII) and co-funded by European Social Fund “Investing in your future” [MS19/00095 to CdJ] and by Instituto de Salud Carlos III (ISCIII) and co-funded by the European Union [PI22/00824 to MS and CdJ]; by Consellería de Innovación, Universidades, Ciencia y Sociedad Digital (Generalitat Valenciana) [CIAICO/2021/135 and CIAPOT/2022/3 to CdJ, CIAICO/2022/81 and CIAPOT/2022/7 to MS]. SAA was supported by Universidad de Las Americas funding [525.A.XV.24].
Author information
Authors and Affiliations
Contributions
C.dJ. and S.A.A. wrote the main manuscript text and J.M.B and E.L.L contributed to some subsections. A.L.C prepared the figures and C.dJ., S.A.A and J.M.B the Tables. M.S revised the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Araujo-Abad, S., Berna, J., Lloret-Lopez, E. et al. Exosomes: from basic research to clinical diagnostic and therapeutic applications in cancer. Cell Oncol. (2024). https://doi.org/10.1007/s13402-024-00990-2
Accepted:
Published:
DOI: https://doi.org/10.1007/s13402-024-00990-2