Abstract
Correlations between the dimensions of a 2-D separation create trend lines that depend on structural or chemical characteristics of the compound class and thus facilitate classification of unknowns. This broadly applies to conventional ion mobility spectrometry (IMS)/mass spectrometry (MS), where the major biomolecular classes (e.g., lipids, peptides, nucleotides) occupy different trend line domains. However, strong correlation between the IMS and MS separations for ions of same charge has impeded finer distinctions. Differential IMS (or FAIMS) is generally less correlated to MS and thus could separate those domains better. We report the first observation of chemical class separation by trend lines using FAIMS, here for lipids. For lipids, FAIMS is indeed more independent of MS than conventional IMS, and subclasses (such as phospho-, glycero-, or sphingolipids) form distinct, often non-overlapping domains. Even finer categories with different functional groups or degrees of unsaturation are often separated. As expected, resolution improves in He-rich gases: at 70% He, glycerolipid isomers with different fatty acid positions can be resolved. These results open the door for application of FAIMS to lipids, particularly in shotgun lipidomics and targeted analyses of bioactive lipids.
Similar content being viewed by others
1 Introduction
Mass spectrometry (MS) has become the staple of biological analyses [1]. However, the complexity of most samples requires fractionation prior to MS, usually performed by electrophoresis (i.e., in liquids) or chromatography (i.e., on solid interfaces) [2]. Ion mobility spectrometry (IMS), where ions in gases are separated based on the properties of transport driven by electric field [3, 4], has emerged as an alternative or complement to condensed-phase methods. In conventional IMS, including the drift-tube (DT) [3] and traveling-wave (TW) [5] modes, ions are dispersed by the mobility (K) that controls motion at moderate electric field intensity (E). Fundamentally, this separation has limited orthogonality to MS because the mobility of an ion (with some charge state z) is related to its size and thus mass (m). The correlation increases for species of similar “density” (meaning close elemental composition) and especially chemical homologues [6–15].
A correlation between two separation dimensions (e.g., IMS and MS) is manifested as a “trend line,” which potentially permits separating different classes and classifying unknowns [6–15]. The classification power is proportional to the distance between the trend lines and the inverse width of distributions around the mean trends. Trend lines in the K versus m/z space were found for atomic clusters [16–19], organic molecules (e.g., amines, amino acids, and fatty acids) [15], and major biomolecular classes: glycans, nucleotides, carbohydrates, peptides, and lipids [6–14]. Lipids have little overlap with peptides and none with nucleotides and carbohydrates [12]. While subclasses form non-identical trend lines, the spacing between them is often within the scatter around each, precluding complete separation. For example, the mobilities in He for monophosphorylated 1+ peptides produced by matrix-assisted laser desorption ionization (MALDI) were 0%–10% (on average, 3%) above the trend for non-modified ones [8, 20], but the standard (σst) and maximum (σmax) deviations of the latter from the trend were ~2.5%–7.3% (depending on the source of data) and ~13% [9, 11, 20]. Hence, all phosphopeptides lied within the domain of non-modified peptides. The mass-mobility correlation for 1+ lipids (σst = 2.6%) is as tight as for peptides or tighter [11]. The σst value is also lower for lipids than for carbohydrates or nucleotides, perhaps reflecting the smaller size and less diverse structures of lipids [11]. Like for peptides, the trends for different subclasses (phospholipids, cerebrosides, sphingolipids) [12, 13, 21] and even finer categories (phospholipids with the ethanolamine and other head groups) [22] differ slightly, but the domains largely overlap, obstructing classification. As MS measures m/z, a set of similar species with unequal z exhibits multiple trend lines. In particular, tryptic peptide ions derived from electrospray ionization (ESI) have z = 1–5, and those with different z lie largely apart in the IMS/MS space [6]. The multiplicity of trends for a range of z substantially increases the orthogonality of IMS to MS, rendering IMS more useful for MS-based proteomics. However, most lipid ions generated by ESI, MALDI, or other ionization sources have z of 1 (in the negative mode, −1). This limits the utility of conventional IMS for lipid classification.
Differential or field asymmetric waveform IMS (FAIMS) [4, 23] exploits that K depends on E to separate species by the difference of mobility (∆K) at high and low E. In practice, a strong periodic electric field E(t), with the peak intensity called “dispersion field” (E D), is created in a gap between two electrodes using an asymmetric waveform (with the amplitude called “dispersion voltage,” DV) applied to one. Ions, carried through the gap by gas flow, oscillate and drift toward one of the electrodes where they are neutralized. A weak fixed “compensation field” (E C) superposed on E(t) may offset the drift for a species with certain K(E) function and permit it to pass the gap, while other ions remain unstable and are eliminated. Scanning E C reveals the spectrum of ions entering the gap.
The correlation to the ion mass or m/z is generally weaker for ∆K than for K, thus FAIMS is more orthogonal to MS than conventional IMS [4, 24, 25]. Hence, FAIMS/MS should produce more disparate trend lines than conventional IMS/MS, yet their separation has not yet been reported for any compound classes. Also, no FAIMS studies of lipids have been undertaken. Here, we show that major lipid subclasses occupy distinct, often non-overlapping domains in the FAIMS/MS space that may aid the lipid fractionation and classification. We also demonstrate that high-resolution FAIMS can separate similar lipids, such as regioisomers and species differing by one degree of unsaturation.
2 Experimental
We employed a planar FAIMS unit coupled to a modified ion trap mass spectrometer (Thermo Fisher LTQ) [26, 27]. The carrier gas at ambient pressure and temperature was delivered to the unit at 2 L/min. We used DV = 4 kV (for E D = 21 kV/cm) or 5.4 kV (E D = 29 kV/cm), allowing He/N2 mixtures with up to 75% or 50% He (vol/vol), respectively [26, 27]. To enable comparison with the results from a commercial system, a cylindrical FAIMS device [23] (Ionalytics/Thermo Fisher) that permits a maximum E D = 21 kV/cm and 50% He, most analyses were performed in that regime [28]. To maximize the resolution and accuracy of peak positions (E C), the scan speed was moderate at 1–5 V/(cm × min).
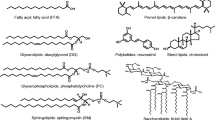
Lipid ions were generated by positive-mode ESI, with emitter at ~2 kV above the FAIMS inlet and liquid flow of 0.3 - 0.5 μL/min. The samples were 1–20 μM solutions of standards in 30:66.5:3.5 chloroform/methanol/water with 10 mM ammonium acetate. The standards, purchased from Avanti (Alabaster, AL, USA), Nu-Chek Prep (Elysian, MN, USA), or Larodan (Malmo, Sweden), included saturated and unsaturated lipids representing the key subclasses analyzed as cations: phospholipids and lysophospholipids with various head groups; glycerolipids, including mono-, di-, and triacylglycerols (MG, DG, and TG); galactolipids; and sphingolipids (ceramides and sphingomyelin, SM). To elicit the trends for each subclass, we sought several species preferably spanning a significant mass range. To gauge the capability for separations of isomers and near-isomers, we procured DG regioisomers and otherwise identical phospholipids with 0–3 double bonds. The galactolipid and some SM samples were mixtures of nonisobaric species readily distinguishable by MS. The lipids studied (62 total) are listed in Table 1 with the structures summarized in Scheme 1.
Structures of lipid classes studied in this work: (a) phospholipids with head groups as listed, called lysophospholipids when a hydrogen replaces the highlighted R chain; (b) TG, called DG or MG when H atom(s) replace(s) one or two of the R chains, respectively; (c) DGDG, called MGDG when an H atom replaces the terminal galactose; (d) ceramide, called sphingomyelin when the highlighted hydroxyl is replaced by phosphocholine. The R1, R2, and R3 are same or different fatty acids
These lipids produce protonated or ammoniated 1+ ions, depending on the chemistry (Table 1). In N2 and He/N2 mixtures, all species with m > ~300–400 Da examined by FAIMS (including all peptides) behaved as “type C” [29], where K decreased at higher E and thus E C > 0 for E D > 0. This has proven true for the lipids investigated here.
The E C scale drifts because of small variations of DV and gas composition, pressure, or temperature [30]. To offset these effects and obtain transferable absolute E C, the scale was anchored by internal calibration. The primary standard was TG 18:1/18:1/16:0, which generates a single ion (the NH +4 adduct) that has no mass overlap with other lipids studied and produces intense well-shaped FAIMS peaks. The accepted E C for this species was set by averaging numerous measurements made over time. Most lipids lie at higher E C values and, to avoid scanning excessive E C ranges and to improve the calibration accuracy, other lipids with greater E C were adopted as secondary standards in some runs. We linearly scaled the E C axis in all spectra to match the calibration, but re-measured any displaying a deviation over 2%. This procedure has greatly reduced the scatter of replicate E C measurements for any lipid.
3 Results
We first report the global mapping of lipids in the FAIMS/MS space, which was performed at {DV = 4 kV; 50% He} with E C values cataloged in Table 1. Then we move to targeted separations, optimizing the DV and He fraction for maximum resolution.
3.1 Phospholipids (P)
These include (Scheme 1a) a glycerol core attaching straight-chain fatty acids (FA) with typically 14–24 carbons and differing number of double bonds in the first (sn1) and second (sn2) positions and a phosphate “head group” that controls the chemistry in the third position (sn3). Various head groups in combination with a large pool of FA, varying in both chain length and degree of saturation, create rich structural and biological diversity. Phosphatidylcholines (PC), with the choline (N,N,N-Trimethylethanolammonium) head group, are major constituents of mammalian cell membranes [31–33]. The E C values for “symmetric” PC featuring identical saturated FA with 14–23 carbons decrease with increasing mass linearly (r 2 = 0.990), Figure 1a. A negative mass-to-E C correlation for a compound class is common (for example, as seen for tryptic peptides of a given z), [26, 34, 35] but the linearity here is impressive. The two FA can differ, resulting in asymmetric PC such as 16:0/14:0 (with the 16- and 14-carbon FA), an isomer of the symmetric PC 15:0/15:0. The E C difference is negligible for these isomers, but appears larger for PC 16:0/16:0 and 14:0/18:0 with more unequal FA (Table 1). As the difference is still well within the peak width (~2 V/cm), these isomers are not presently resolved. To compare, the symmetric and asymmetric PC track the same trend line in conventional IMS [36].
Measurements for phospholipids (a) and lysophospholipids (b): data (symbols) and linear regressions for categories with at least two points (lines). Filled symbols are for saturated lipids, empty symbols are for unsaturated ones, and the number after lipid classification indicates the degree of unsaturation
Phosphatidylethanolamines (PE) and phosphatidylserines (PS) are parallel to PC, with the choline replaced by ethanolamine or serine. While PE are the second most abundant phospholipids in animals and the main lipids in microbial membranes, PS are found mainly in bacteria and mammalian nervous tissue [31, 37]. The E C values for saturated lipids of these categories decrease for heavier ions as for PC, but the offsets (and possibly, slopes) may differ slightly (Figure 1a). Analyses over broader mass ranges are needed to establish the trends for PE and PS more accurately.
Many lipids include unsaturated FA, with the number of double bonds (in nature, almost always cis-) influencing the membrane fluidity. PC with monounsaturated FA (such as 18:1) appear to form a trend line at E C values below those for saturated analogs (Figure 1a). The trend nears that for saturated PC for larger lipids, where a double bond should make a lesser difference. The examined PC with 3 or 4 double bonds in each FA have even lower E C, but no trend lines could be drawn as each type was represented by one species (Figure 1a).
Lysophospholipids (LP) have the same head groups as phospholipids, but include only one FA (normally in the sn1 position) [31] and thus tend to be smaller. The E C versus mass correlation for saturated lyso-PC (LPC) is as or even more linear (r 2 = 0.998) than that for PC (Figure 1b). Saturated lyso-PE (LPE) also align well and the difference between their and LPC trends is significant, exceeding that between PE and PC: the head group may contribute more to the mobility properties of smaller lipids. The trend for saturated lysophosphatidylglycerols (LPG) with the glycerol head group is also apart from that for LPC, though close to that for LPE. The E C value for the unsaturated LPC 18:1 is above that for the saturated analog (Figure 1b). This contrasts with the findings for PC, including those with the same FA. Whether this is a general distinction between PC and LPC remains to be seen. The P and LP domains are spaced apart much more than those for categories of either (Figure 2a).
Data for all lipids studied in this work, grouped by subclass (a) and in total (b). In (b), solid symbols are for E C values (regression in dash) and blank symbols are for peak widths (regression in dotted). The categories for saturated and unsaturated phospho- and lysophospholipids are aggregated; the nomenclature is per Figure 1
3.2 Glycerolipids
In this major subclass, the glycerol is esterified to one (MG), two (DG), or three (TG) same or different FA. Typical lipids naturally get heavier from MG to DG to TG, though the mass ranges overlap as the FA lengths vary. The trend lines for saturated MG and saturated symmetric DG have decreasing slopes similar to those for LP and P, while the offsets increase in the order: MG, LP, DG, P (Figure 2a). Unsaturated MG may have slightly lower E C than saturated ones, though more data are needed to conclude. Unsaturated DG lie at E C much below those for saturated ones, but the shift is not governed simply by the degree of unsaturation: DG 20:3/20:3 lies slightly above the line set by DG 18:1/18:1 and 24:1/24:1. This behavior follows that for phospholipids above. In the result, the domains of MG (saturated or not), LP, and saturated DG are fully separated in the mass overlap regions. The trend for unsaturated DG is close to that for LP extended, and those domains may mix at similar masses.
The trend for unsaturated TG (with each FA having 0 or 1 double bonds) continues that for saturated DG, and those categories coincide over at least some of the overlap region. The resulting domain appears fully separate from that for saturated phospholipids, but not from unsaturated ones. For the largest TG (m > 1 kDa), the trend line starts curving up (Figure 2a). This should happen for all lipids with increasing mass, as E C must stay positive (except for macro-ions with m > ~30 kDa where strong permanent ion dipoles may be aligned, causing E C < 0) [29, 38]. Perhaps this curving is observed here only for TG because they are among the heaviest lipids, and all others analyzed were under 1 kDa. The only saturated TG that we studied (with FA 19:0) has E C slightly above the trend for unsaturated TG, in line with the results for MG and DG. The results for asymmetric DG are described below.
Why do the trend lines for unsaturated and saturated lipids differ? The mobilities of unsaturated PC exceed those of saturated analogs by ~0.5% per double bond in DTIMS [22] and by more (~5% for the first double bond and further ~1% for each beyond that) in TWIMS [36]. This was attributed to a double bond bending and rigidizing a FA, which reduces the collision cross section [36]. A larger effect in TWIMS was rationalized [36] as a consequence of unequal unfolding upon the field heating of ions by the traveling wave [5], where floppy saturated lipids expand more than rigid unsaturated ones. Though the molecular modeling was performed for PC [36], this effect related to the shape and flexibility of FA should transfer to other lipids. However, the effect of such structural compression on FAIMS analyses is unclear. For peptides or proteins, unfolding (including that induced by the field heating) decreases E C. Had this applied to lipids, the introduction of double bonds would increase the E C values for PC and, presumably, other classes. We have observed this for one LPC species only and the opposite in other cases, as discussed above. Hence the shifts between saturated and unsaturated lipids in FAIMS remain to be explained. This exemplifies the orthogonality between conventional IMS and FAIMS dimensions, which has previously been shown for amino acids and peptides [24, 25, 39]. Analytically, these shifts may allow identifying saturated or unsaturated lipids, and clarifying the degree of unsaturation. They also enable detecting small amounts of lipids in the presence of analogs with one more double bond, which is often difficult in MS because of isotopic distribution (see below). The E C values and thus trends for unsaturated lipids might depend on the double bond position: how sensitive is FAIMS to that remains to be researched.
3.3 Other Lipids
To sample the separation properties of lipids more comprehensively, we have measured species of three other classes. Glycolipids contain sugar(s) on the sn3 position, and FA on the sn1 and sn2 positions, of the glycerol. The simplest glycolipids are galactolipids where the sugar is galactose. Monogalactosyl diacylglycerols (MGDG) involve a single galactose and digalactosyl diacylglycerols (DGDG) have a chain of two; both are key components of plant membranes and photosynthetic tissues [40]. The saturated MGDG and DGDG fall on the same trend line lying close to and perhaps slightly above that for saturated phospholipids, but much above that for TG with the two domains apparently separated (Figure 2a).
Sphingolipids are built on a sphingoid base, typically sphingosine (an 18-carbon amino alcohol with monounsaturated straight chain) connected to FA via an amide linkage. Among other categories, ceramides are the simplest sphingolipids with no head group, and sphingomyelins (SM) have the phosphocholine group bound to sphingosine. SM are major components of animal tissues [31, 41]. Ceramides are rare in tissues (except skin), but are important intermediates in the biosynthesis of complex sphingolipids [31, 41]. The trend for ceramides with saturated FA emerges above those for saturated LP and unsaturated DG but below that for saturated DG or unsaturated TG (Figure 2a). The trend line for SM lies on the other side of that for saturated DG or unsaturated TG, below those for saturated phospho- or galactolipids, and passes through the domain for unsaturated phospholipids (Figure 2a). Thus FAIMS apparently can distinguish categories within the sphingolipid subclass.
For all lipids probed (Figure 2b), E C and m/z are substantially correlated (r 2 = 0.82 with σst = 7.4% and σmax = 22%), but the distribution about the mean trend is ~3 times wider than that for K and m/z. The peak widths vary within a limited range with a bias to increase for larger lipids (Figure 2b), reflecting the peak broadening for less mobile ions in planar FAIMS devices [4]. We have omitted many lipid subclasses such as cholesterol esters, glycosphingolipids (cerebro-, globo-, and gangliosides), sterol and prenol lipids, saccharolipids, and polyketides, some of which are structurally distant from the included subclasses. For the subclasses investigated, the ranges of molecular sizes and degrees of unsaturation were limited. Incorporation of missing classes and extension to larger/smaller and more unsaturated species would likely reduce the correlation between FAIMS and MS dimensions.
3.4 Targeted Separations of Isomers and Near-Isomers
Dispersion of data around the mean trend for a compound type in FAIMS/MS (or IMS/MS) space is detrimental to classification, but enables separation of isomers or isobars within a class. All lipid classes permit numerous isomers, and many are encountered in biology and differ in activity [42]. The isomeric diversity of lipids was underestimated, and ostensibly pure compounds now appear to be mixtures [42]. In particular, glycerolipids (except TG with identical FA) have regioisomers resulting from the FA permutation between the glycerol sites. The sn1 and sn3 positions are equivalent, but a DG with different FA can have FA1, FA2, or none in the sn2 slot, creating three isomers, such as 16:0 (sn1)/12:0 (sn2), 16:0 (sn1)/12:0 (sn3), and 12:0 (sn1)/16:0 (sn2). We have analyzed the first two (Figure 3), and their separation at 50% He and DV = 4 kV is incomplete. As with amino acids and peptides [26], raising the He fraction increases E C values while narrowing the peaks. By ~75% He, the resolution doubles and these DG are baseline-separated. The spectrum for each isomer features a peak at E C of the other with ~10% abundance (Figure 3). This is likely due to migration of acyl chains with time and, thus, isomeric mixing [43]: the isomeric abundances of 1,2- and 1,3-DGs in solution can often shift by ~10% upon 2 mo of room-temperature storage [44]. Hence FAIMS can be useful to separate lipid regioisomers and control their isomeric purity. The inability to distinguish isomeric lipids is viewed as the major “limitation of existing lipidomics weaponry that seriously impedes the understanding of lipid biochemistry” [42].
FAIMS spectra for a mixture of DG 16:0 (sn1)/12:0 (sn2) and 16:0 (sn1)/12:0 (sn3) at DV = 4 kV and He fractions as marked (solid lines). On top are the widths (V/cm) for major peaks with the resolving powers underlined. Values below are the distances between two peaks with the resolution (r) given. Features for isomers with FA 12:0 in the sn2 and sn3 slots at 50% He are shown as dashed lines
All FA can be unsaturated to various degrees, creating lipids that differ by one double bond, such as LPC 18:0 and 18:1. Such species differ by 2 Da, but the isotopic distributions obstruct the MS detection of the more saturated one in the presence of the other. For example, the protonated LPC 18:1 has isotopologues with cumulative ~6% abundance at m/z = 524.360–524.366, which differ from the base peak of 18:0 (m/z = 524.372) by ~10–20 ppm and thus require MS resolving power >105 to be separated. As the ionization yields of saturated lipids tend to be below those of unsaturated analogs, saturated lipids are more challenging to detect and quantify by MS in a mixture with unsaturated lipids than the isotopic distributions suggest. This problem may be addressed by FAIMS, which provides baseline separation of LPC 18:0 and 18:1 already at {DV = 4 kV; 50% He} (Figure 4). The difficulty caused by isotopic overlaps in MS escalates for larger lipids, as the relative mass differences between isotopologues decrease and heavier ones grow in abundance. For example, protonated PC 23:1/23:0 has isotopologues with cumulative ~20% abundance at only ~6–12 ppm from the base peak for PC 23:0/23:0 (m/z = 930.8). The FAIMS resolution of species with different degrees of unsaturation may well diminish for larger lipids, but perhaps increasing the DV and/or He content could offset that.
4 Conclusions
We have explored lipids by FAIMS, focusing on major subclasses analyzed as cations: phospho-, lysophospho-, glycero-, galacto-, and sphingolipids. The protonated and ammoniated ions generated by ESI exhibit (in 1:1 He/N2) the expected C-type behavior with an inverse correlation between ion mass and compensation field within each subclass. However, the subclasses follow distinct trend lines and are completely or partly separated in the FAIMS/MS space, and the classification power substantially exceeds that of conventional IMS. The trend lines also differ for finer categories, such as lipids with different head groups or degree of unsaturation, and the sign and magnitude of difference vary between subclasses. The dispersion of data points within a category allows FAIMS to separate isomers with different fatty acid attachments, such as 1,2- and 1,3-diacylglycerols. Lipids differing by one double bond and, thus, mixed in MS because of isotopic distributions such as saturated and monounsaturated LPC, can also be separated. As for peptides [26–28, 45], the resolution improves at higher He content, and species not fully separated at 50% He often can be at 75% He. The highest possible dispersion field (E D) and maximum He content permitted at that field normally provide the best resolution, making optimization of those variables rarely needed in practice.
At this point, the utility of FAIMS in lipid analyses can be contemplated in provisional terms only. First, FAIMS could increase the specificity of shotgun lipidomics, where unseparated crude lipid extracts are subjected to MS/MS, and product, precursor, or neutral loss scans specific to lipid classes are performed [46]. For example, a precursor scan for m/z = 184 shows all lipids containing phosphocholine: PC, LPC, and SM [46]. These could be pre-separated using FAIMS, with linked FAIMS and MS scans to track the desired trend line. One can also classify lipids exploiting the Kendrick mass defect defined by the elemental composition, however, (1) a resolving power of ~200,000 and sub-ppm mass accuracy exclusive to FTICR MS are required, and (2) isomers and classes with identical compositions, such as PC and PE, are fundamentally indistinguishable [47, 48]. In targeted analyses, FAIMS could filter specific bioactive lipids such as ceramides or other sphingolipids that are often not abundant and lost in the “chemical noise.” As implemented for proteomics [34, 35, 49], FAIMS can be inserted between liquid chromatography (LC) and MS stages to raise the overall peak capacity. Recent introduction of lipidomic analyses by 2-D LC, comprising normal-phase NPLC for subclass fractionation and reverse-phase RPLC for separations based on other factors such as the degree of unsaturation [50], speaks to the insufficiency of single-stage LC for modern lipid research [48]. The 2-D chromatography approach is powerful and well-established in proteomics [51], but inherently suffers from very low throughput (~1–5 samples/d) [50]. A 1-D LC/FAIMS hybrid would compete with 2-D LC in separation power, but offer much higher speed. Similar considerations apply to isomer separations, the traditional focus of IMS application. Structural isomers of lipids, including positional isomers of DG or TG, can often be resolved by LC, but the needed analysis times are commonly ~2–10 h [52–54] compared with seconds with FAIMS. We continue exploring the utility of FAIMS for other lipids and lipid classes and evaluate it in new challenging tasks, such as separation of isomers differing in the position of double bond [42] or branching point on the fatty acid [54].
References
Siuzdak, G.: The Expanding Role of Mass Spectrometry in Biotechnology. MCC Press (2006)
Niessen, W.M.A.: Liquid Chromatography-Mass Spectrometry. CRC Press, Boca Raton (2006)
Eiceman, G.A., Karpas, Z.: Ion Mobility Spectrometry. CRC Press, Boca Raton (2005)
Shvartsburg, A.A.: Differential ion mobility spectrometry. CRC Press, Boca Raton (2008)
Shvartsburg, A.A., Smith, R.D.: Fundamentals of traveling wave ion mobility spectrometry. Anal. Chem. 80, 9689–9699 (2008)
Srebalus, C.A., Li, J., Marshall, W.S., Clemmer, D.E.: Gas-phase separations of electrosprayed peptide libraries. Anal. Chem. 71, 3918–3927 (1999)
Merenbloom, S.I., Koeniger, S.L., Valentine, S.J., Plasencia, M.D., Clemmer, D.E.: IMS-IMS and IMS-IMS-IMS/MS for separating peptide and protein fragment ions. Anal. Chem. 78, 2802–2809 (2006)
Ruotolo, B.T., Verbeck, G.F., Thompson, L.M., Woods, A.S., Gillig, K.J., Russell, D.H.: Distinguishing between phosphorylated and nonphosphorylated peptides with ion mobility-mass spectrometry. J. Proteome Res. 1, 303–306 (2002)
Ruotolo, B.T., McLean, J.A., Gillig, K.J., Russell, D.H.: Peak capacity of ion mobility mass spectrometry: the utility of varying drift gas polarizability for the separation of tryptic peptides. J. Mass Spectrom. 39, 361–367 (2004)
Woods, A.S., Ugarov, M., Egan, T., Koomen, J., Gillig, K.J., Fuhrer, K., Gonin, M., Schultz, J.A.: Lipid/Peptide/Nucleotide separation with MALDI-ion mobility-TOFMS. Anal. Chem. 76, 2187–2195 (2004)
Fenn, L.S., Kliman, M., Mahsut, A., Zhao, S.R., McLean, J.A.: Characterizing ion mobility-mass spectrometry conformation space for the analysis of complex biological samples. Anal. Bioanal. Chem. 394, 235–244 (2009)
McLean, J.A.: The mass-mobility correlation redux: the conformational landscape of anhydrous biomolecules. J. Am. Soc. Mass Spectrom. 20, 1775–1781 (2009)
Ridenour, W.B., Kliman, M., McLean, J.A., Caprioli, R.M.: Structural characterization of phospholipids and peptides directly from tissue sections by MALDI traveling-wave ion mobility-mass spectrometry. Anal. Chem. 82, 1881–1889 (2010)
Fenn, L.S., McLean, J.A.: Simultaneous glycoproteomic strategies utilizing ion mobility-mass spectrometry. Mol. Biosyst. 5, 1298–1302 (2009)
Dwivedi, P., Schultz, A.J., Hill, H.H.: Metabolic profiling of human blood by high-resolution ion mobility mass spectrometry (IM-MS). Int. J. Mass Spectrom. 298, 78–90 (2010)
Bowers, M.T., Kemper, P.R., von Helden, G., van Koppen, P.A.M.: Gas-phase ion chromatography: transition metal state selection and carbon cluster formation. Science 260, 1446–1451 (1993)
Shvartsburg, A.A., Schatz, G.C., Jarrold, M.F.: Mobilities of carbon cluster ions: critical importance of the molecular attractive potential. J. Chem. Phys. 108, 2416–2423 (1998)
Shvartsburg, A.A., Hudgins, R.R., Dugourd, P., Gutierrez, R., Frauenheim, T., Jarrold, M.F.: Observation of “Stick” and “Handle” intermediates along the fullerene road. Phys. Rev. Lett. 84, 2421–2424 (2000)
Shvartsburg, A.A., Hudgins, R.R., Dugourd, P., Jarrold, M.F.: Structural information from ion mobility measurements: applications to semiconductor clusters. Chem. Soc. Rev. 30, 26–35 (2001)
Ruotolo, B.T., Gillig, K.J., Woods, A.S., Egan, T.F., Ugarov, M.V., Schultz, J.A., Russell, D.H.: Analysis of phosphorylated peptides by ion mobility-mass spectrometry. Anal. Chem. 76, 6727–6733 (2004)
Trimpin, S., Tan, B., Bohrer, B.C., O’Dell, D.K., Merenbloom, S.I., Pazos, M.X., Clemmer, D.E., Walker, J.M.: Profiling of phospholipids and related lipid structures using multidimensional ion mobility spectrometry-mass spectrometry. Int. J. Mass Spectrom. 287, 58–69 (2009)
Jackson, S.N., Ugarov, M., Post, J.D., Egan, T., Langlais, D., Schultz, J.A., Woods, A.S.: A study of phospholipids by ion mobility TOFMS. J. Am. Soc. Mass Spectrom. 19, 1655–1662 (2008)
Guevremont, R.: High-field asymmetric waveform ion mobility spectrometry: a new tool for mass spectrometry. J. Chromatogr. A 1058, 3–19 (2004)
Guevremont, R., Barnett, D.A., Purves, R.W., Vandermey, J.: Analysis of a tryptic digest of pig hemoglobin using ESI-FAIMS-MS. Anal. Chem. 72, 4577–4584 (2000)
Shvartsburg, A.A., Mashkevich, S.V., Smith, R.D.: Feasibility of higher-order differential ion mobility separations using new asymmetric waveforms. J. Phys. Chem. A 110, 2663–2673 (2006)
Shvartsburg, A.A., Danielson, W.F., Smith, R.D.: High-resolution differential ion mobility separations using helium-rich gases. Anal. Chem. 82, 2456–2462 (2010)
Shvartsburg, A.A., Prior, D.C., Tang, K., Smith, R.D.: High-resolution differential ion mobility separations using planar analyzers at elevated dispersion fields. Anal. Chem. 82, 7649–7655 (2010)
Shvartsburg, A.A., Tang, K., Smith, R.D.: Differential ion mobility separations of peptides with resolving power exceeding 50. Anal. Chem. 82, 32–35 (2010)
Shvartsburg, A.A., Bryskiewicz, T., Purves, R., Tang, K., Guevremont, R., Smith, R.D.: Field asymmetric waveform ion mobility spectrometry studies of proteins: dipole alignment in ion mobility spectrometry? J. Phys. Chem. B 110, 21966–21980 (2006)
Schneider, B.B., Covey, T.R., Coy, S.L., Krylov, E.V., Nazarov, E.G.: Chemical effects in the separation process of a differential mobility/mass spectrometry system. Anal. Chem. 82, 1867–1880 (2010)
Christie, W.W.: Lipid Analysis. The Oily Press, Dundee (2003)
Han, X., Gross, R.W.: Shotgun lipidomics: electrospray ionization mass spectrometric analysis and quantitation of the cellular lipidomes directly from crude extracts of biological samples. Mass Spectrom. Rev. 24, 367–412 (2005)
Pulfer, M., Murphy, R.C.: Electrospray mass spectrometry of phospholipids. Mass Spectrom. Rev. 22, 332–364 (2003)
Venne, K., Bonneil, E., Eng, K., Thibault, P.: Improvement in peptide detection for proteomics analysis using NanoLC-MS and high-field asymmetry waveform ion mobility mass spectrometry. Anal. Chem. 77, 2176–2186 (2005)
Saba, J., Bonneil, E., Pomies, C., Eng, K., Thibault, P.: Enhanced sensitivity in proteomics experiments using FAIMS coupled to a hybrid linear ion trap/orbitrap mass spectrometer. J. Proteome Res. 8, 3355–3366 (2009)
Kim, H.I., Kim, H., Pang, E.S., Ryu, E.K., Beegle, L.W., Loo, J.A., Goddard, W.A., Kanik, I.: Structural characterization of unsaturated phosphatidylcholines using traveling wave ion mobility spectrometry. Anal. Chem. 81, 8289–8297 (2009)
Wenk, M.R.: The emerging field of lipidomics. Nat. Rev. 4, 594–610 (2005)
Shvartsburg, A.A., Noskov, S.Y., Purves, R., Smith, R.D.: Pendular proteins in gases and new avenues for characterization of macromolecules by ion mobility spectrometry. Proc. Natl. Acad. Sci. U. S. A. 106, 6495–6500 (2009)
Tang, K., Li, F., Shvartsburg, A.A., Strittmatter, E.F., Smith, R.D.: Two-dimensional gas-phase separations coupled to mass spectrometry for analyses of complex mixtures. Anal. Chem. 77, 6381–6388 (2005)
Murata, N., Higashi, S., Fujimura, Y.: Glycerolipids in various preparations of photosystem II from spinach chloroplasts. Biochem. Biophys. Acta 1019, 261–268 (1990)
Wennekes, T., van den Berg, R.J.B.N.H., Boot, R.G., van der Marel, G.A., Overkleeft, H.S., Aerts, J.M.F.G.: Glycosphingolipids—nature, function, and pharmacological modulation. Angew. Chem. Int. Ed. 48, 8848–8869 (2009)
Mitchell, T.W., Pham, H., Thomas, M.C., Blanksby, S.J.: Identification of double bond position in lipids: from GC to OzID. J. Chromatogr. B 877, 2722–2735 (2009)
Private communication from Avanti Lipids, 2009.
Cossignani, L., Luneia, R., Damiani, P., Simonetti, M.S., Riccieri, R., Tiscornia, E.: Analysis of isomeric diacylglycerolic classes to evaluate the quality of olive oil in relation to storage conditions. Eur. Food Res. Technol. 224, 379–383 (2007)
Shvartsburg, A.A., Creese, A., Smith, R.D., Cooper, H.J.: Separation of peptide isomers with variant modified sites by high-resolution differential ion mobility spectrometry. Anal. Chem. 82, 8327–8334 (2010)
Hu, C., van der Heijden, R., Wang, M., van der Greef, J., Hankemeier, T., Xu, G.: Analytical strategies in lipidomics and applications in disease biomarker discovery. J. Chromatogr. B 877, 2836–2846 (2009)
He, H., Conrad, C.A., Nilsson, C.L., Ji, Y., Schaub, T.M., Marshall, A.G., Emmett, M.R.: A novel method for lipidomic analysis: p53 expression modulates sulfatide, ganglioside and phospholipid composition of U87 MG glioblastoma cells. Anal. Chem. 79, 8423–8430 (2007)
Lerno, L.A., German, J.B., Lebrilla, C.B.: Method for the identification of lipid classes based on referenced kendrick mass analysis. Anal. Chem. 82, 4236–4245 (2010)
Canterbury, J.D., Yi, X., Hoopmann, M.R., MacCoss, M.J.: Assessing the dynamic range and peak capacity of nanoflow LC-FAIMS-MS on an ion trap mass spectrometer for proteomics applications. Anal. Chem. 80, 6888–6897 (2008)
Nie, H., Liu, R., Yang, Y., Bai, Y., Guan, Y., Qian, D., Wang, T., Liu, H.: Lipid profiling of rat peritoneal surface layers by online normal- and reversed-phase 2D LC QTOF-MS. J. Lipid Res. 51, 2833–2844 (2010)
Washburn, M.P., Wolters, D., Yates, J.R.: Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nat. Biotechnol. 19, 242–247 (2001)
Momchilova, S., Itabashi, Y., Nikolova-Damyanova, B., Kuksis, A.: Regioselective separation of isomeric triacylglycerols by reversed phase high-performance liquid chromatography: stationary phase and mobile phase effects. J. Sep. Sci. 29, 2578–2583 (2006)
Leveque, N.L., Heron, S., Tchapla, A.: Regioisomer characterization of triacylglycerols by nonaqueous reversed-phase liquid chromatography/electrospray ionization mass spectrometry using silver nitrate as a postcolumn reagent. J. Mass Spectrom. 45, 284–296 (2010)
Schreiberova, O., Krulikovska, T., Sigler, K., Cejkova, A., Rezanka, T.: RP-HPLC/MS-APCI analysis of branched chain tag prepared by precursor-directed biosynthesis with Rhodococcus erythropolis. Lipids 45, 743–756 (2010)
Acknowledgments
The authors thank Rui Zhao and Dr. Keqi Tang for experimental help, and Professor John A. McLean for discussions. The authors acknowledge that parts of this research were supported by NIH NCRR (RR 18522), NIH grant DK071283, and NIAID Award U54AI081680. The content of this paper is solely the responsibility of the authors and does not necessarily represent the official views of the NIAID or NIH. The work was performed in the Environmental Molecular Sciences Laboratory, a scientific user facility at PNNL supported by the US DoE OBER.
Author information
Authors and Affiliations
Corresponding author
Additional information
An erratum to this article can be found at http://dx.doi.org/10.1007/s13361-011-0156-2
Rights and permissions
About this article
Cite this article
Shvartsburg, A.A., Isaac, G., Leveque, N. et al. Separation and Classification of Lipids Using Differential Ion Mobility Spectrometry. J. Am. Soc. Mass Spectrom. 22, 1146–1155 (2011). https://doi.org/10.1007/s13361-011-0114-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-011-0114-z