Abstract
One of the markers of aging is lymphocyte telomere length (LTL), which is affected by genetic constitution of the organism and environmental conditions, such as development and diseases, including diabetes. The relationship of the later seems to be bilateral. The enzyme responsible for the maintenance of telomere length is a subunit of telomerase—telomerase reverse transcriptase (TERT). The aims of the present study were to (1) determine the influence of the TERT promoter sequence SNP variants on relative telomere length (RTL) in an elderly Polish population and (2) explore the potential associations of the SNPs with the type 2 diabetes mellitus (T2DM) in the obese individuals. Two highly homogenous subgroups of PolSenior participants were investigated, the first constituted 70 relatively healthy respondents and the second 70 individuals with T2DM. Telomere length ratio (T/S value) was measured; 1.5 kb part upstream of the transcription start site of the TERT promoter was sequenced, and the frequencies of polymorphisms were calculated and compared against analysed data. Low-frequency SNPs were evaluated but excluded from further comparative analyses to RTL and glucose metabolism markers. No significant difference in telomere length was found between the two studied subgroups. Univariate statistical analyses showed only a weak association of environmental or genetic factors altering this marker of aging. Approximate frequency of four SNPs in TERT promoter sequence was assessed in Polish population aged 65–95 years, but three of them (rs2735940, rs7712562 and rs2853669) were selected for further analyses. The SNP selection was based on their minor allele frequencies in general population and on published data. The univariate analysis has revealed that carriers of CC SNP (rs2853669) have had the shortest RTL in the T2DM group. Multivariate analysis has also revealed that the genetic effect of TERT promoter CC SNP was strengthened by the incidence of T2DM. The additional variation in RTL in paired groups indicates that in addition to T2DM and genetics, there are other factors contributing to development of the age-related diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The dynamic of telomeres’ length and their age-dependent shortening provide valuable insights into the pathogenesis of chronic inflammatory processes (Zhang et al. 2016) and consequently an aging-related disorders (Ishikawa et al. 2016), such as arterial hypertension (Ma et al. 2015) or cardiovascular disease (Nilsson et al. 2013). Although a growing body of evidence supports an association between short telomeres and type 2 diabetes mellitus (T2DM), most studies have been cross-sectional by nature, trying to answer whether the metabolic disturbances of T2DM cause telomere attrition or if the shorter telomeres lead to higher risk of T2DM. Biological hypotheses address both scenarios. Short telomeres may lead to premature cell senescence, resulting in the reduced cell mass and subsequently impaired insulin secretion and glucose tolerance (Elks and Scott 2014). Conversely, elevated blood glucose concentrations increase oxidative stress and potentially interfere with telomerase function leading to shorter telomeres. On the other hand, high concentration of blood glucose alone, as shown in cultured human fibroblasts, did not cause telomere shortening. However, it was significantly accelerated in cell cultures containing the pro-inflammatory cytokine, interleukin 1 beta (Salpea et al. 2013).
The normalized telomere to centromere signal ratio, determined for adipocytes by the method of in situ hybridization, was reduced in T2DM patients when compared to control ones. It, also, negatively correlated with blood glycated haemoglobin concentration (HbA1c) (Tamura et al. 2016a). Using autopsy samples, telomere length in pancreatic islet cells β and α were found shorter in patients with diabetes mellitus (DM) than in non-diabetic individuals (Tamura et al. 2016b). However, the relation between the relative lymphocyte telomere length (RTL) and the presence of diabetes was questioned in some reports. The results of the study on the US general population lead to the conclusion that RTL was not associated neither with diabetic status, its duration period, medical treatment, nor telomere attrition. Therefore, it was neither a cause nor a consequence of diabetes (Menke et al. 2015).
Although the hypothesis linking obesity and especially coexisting T2DM with accelerated telomere shortening is commonly accepted, the high inter-individual variability of RTL makes it difficult to confirm in a cohort studies. A significant positive association between shorter telomeres and T2DM risk was demonstrated in meta-analysis of 429 records (Willeit et al. 2014) but only in the quartile of participants with the shortest telomeres. In parallel study of a cohort of 2721 elderly subjects, it has been revealed that the shortening of RTL correlated with obesity. However, the correlation was significant with some obesity-related parameters, such as body fat percentage, subcutaneous fat and plasma leptin concentration but not with the others, such as BMI and visceral fat. Therefore, the mutual interplay between obesity and RTL is still uncertain (Njajou et al. 2012; Müezzinler et al. 2014). Only few studies investigating the association between lymphocyte telomere length (LTL) and obesity reported that weight gain, rather than the obese status per se, is the more important factor affecting telomere shortening rates (Buxton et al. 2011). Similarly, in the case of patients with T2DM (501 patients) diagnosed and medically treated, the LTL seemed to be correlated rather with diabetic complications than with the risk of T2DM itself (Testa et al. 2011). The shorter was the LTL baseline; the more pronounced and significant was the insulin resistance over the follow-up period. This effect was additive to that of BMI in parallel twins’ studies (Verhulst et al. 2016). Thus, telomere attrition could provide additive prognostic information on mortality risk in T2DM patients (Bonfigli et al. 2016).
Relatively large variability of RTL could be influenced not only by the above listed factors. The obesity is influenced not only by diet, physical activity, the type of diabetes, etc., but also by sex (Gardener et al. 2014), the type of analysed cells (Svenson et al. 2011; Meyer et al. 2016) and the cell proliferative activity in the analysed tissue. The RTL results depend also on the measurement method (Gutmajster et al. 2013; Rode et al. 2015; Mazidi et al. 2017), the DNA preparation (Raschenberger et al. 2016) and genetic constitution of an organism. RTL is a heritable trait with variability ranging from 34 to 82% and depends on a number of specific genetic variants associated with RTL, including variations in sequence of genes such as TERT, TERC, OBFC1, CTC1, CEP95 and SMURF2 (Codd et al. 2013; Lee et al. 2013; Zhou et al. 2016). The dynamics of RTL undergo the age-dependent shortening at remarkably rapid rates of attritions until the first 20 years of life. However, inter-individual variation in the initial length of telomeres was remarkable, in spite of its high heritability (Liu 2014). Telomere length was stable in the healthy old (range 61–75 years) and oldest old individuals (range 76–91 years) when compared with the younger ones (Houben et al. 2011; Franzke et al. 2015). There was also less of RTL variation between men and women. Slower telomere attrition rate in women resulted from the oestrogen protective function on the telomere length, which is not the case in post-menopausal women population (Gardener et al. 2014).
SNP analyses, always, are conducted on as big, as possible populations; therefore, in our work to avoid the low power of statistical tests, instead of enlarging the tested population, we screened the entire available to us population for participants that constituted group as homogenous as possible. The human telomerase reverse transcriptase gene sequence (hTERT), coding the catalytic subunit of telomerase holoenzyme, was the target in this study. The enzyme has been defined as the rate-limiting factor in regulating telomerase activity in maintaining the telomere length (Ozturk et al. 2017). No previously reported mutations leading to the telomeropathies (Opresko and Shay 2017) were expected. Also, both T2DM patients and controls with cancer histories were excluded from the study (Heidenreich et al. 2014). Additionally, 1100 bps DNA fragment upstream of the ATG start codon of the gene has been well characterized by others (Wick et al. 1999; Lewis and Tollefsbol 2016). The hTERT promoter sequence variants were reported as related to premature telomere shortening (Melicher et al. 2015), increased risk of cancer (Heidenreich et al. 2014) and cardiovascular diseases (CAD) (Bressler et al. 2015). Some polymorphisms were described as of no-clinical significance; however, three polymorphic changes (rs2853669, rs3215401, rs2735940) were found to influence telomerase expression (Matsubara et al. 2006a; Helbig et al. 2017). Nevertheless, there is yet no such data available for diabetes. Recently, additional functions of hTERT, beyond the maintenance of chromosome stability, have been explored. Telomerase/TERT may act as a transcription modulator through its interaction with transcription factors p65, β-catenin or Brahma-related gene-1 (BRG1) that might regulate transcription of some other genes. TERT is able to form different complexes in different cell contexts and regulate the gene expression in certain pathways (Zhou et al. 2014). TERT has also been shown to shuttle dynamically between different cellular compartments, under increased oxidative stress (Singhapol et al. 2013). Inhibition of TERT expression reduced basal 2-deoxyglucose uptake by 50% in human and mouse cell lines, while its overexpression upregulated glucose uptake by 3.25-fold. Therefore, loss of TERT expression (e.g. in diabetes or aging) may accompany insulin sensitivity and glucose uptake (Shaheen et al. 2014). It has been also postulated that TERT overexpression could induce cell survival and therefore to be applied to ease diabetes mellitus and its vascular complications (Qi Nan et al. 2015).
In this work, we hypothesize that (1) differences in RTL, previously seen by others, between controls and T2DM patients are results of inflammation and oxidative stress caused among others by obesity; therefore, if all participants are obese, no changes in RTL length between both groups should be detected. (2) In clinically homogenous groups, where the effects of genetic constitution of individuals are more pronounced, changes detected in the sequence of the TERT promoter affect telomerase activity in both telomeres’ length and glucose transport; thus, increasing the risk of T2DM in elder and obese individuals could serve as its prognostic marker.
Materials and methods
Participants
The analyses were carried out on a group of participants carefully selected from the cohort of the PolSenior study. Information on age, sex, socio-demographic characteristics, medical history, health status, family history and lifestyle were obtained based on detail questionnaires in a standardized manner (Bledowski et al. 2011).
In the group of 1842 subjects with assessed telomere length (data not shown), 277 participants were treated for diabetes and among them 140 individuals were obese, according to WHO criteria. The number of obese participants without T2DM was 411. From this cohort, participants with inflammatory conditions, namely rheumatoid diseases, acute and chronic infections, history of cancer, stroke, congestive heart failure, dementia or chronic obstructive pulmonary disease (except hypertension) were excluded. The group of T2DM included only patients previously diagnosed and already pharmacologically treated for diabetes with insulin. The inclusion criterion was the coexistence of insulin resistance (HOMA-IR values above 2.5). Selected subjects from the group without T2DM were matched with age, sex and in equal proportion of men and women.
The final group, strictly fulfilling the presented above criteria, consisted of 140 participants, of which 70 were obese with T2DM and 70 were control obese without T2DM. For this group, the power of statistical tests, for calculated variance, was satisfied. Specifically, 28 of age 65–69 years included 15 with T2DM, 28 of age 70–74 years included 14 with T2DM, 28 of age 75–79 years included 13 with T2DM, 28 of age 80–84 years included 14 with T2DM and 28 of age 85–89 and 90–95 years included 14 with T2DM. Detailed characteristics of both groups are presented in Table 1.
DNA isolation
Whole blood samples were obtained from all participants. Genomic DNA was extracted by salting-out method and stored at − 80 °C. The concentration and purity of the DNA were assessed by UV spectroscopy (Nanodrop, Thermo Fisher Scientific Inc., Wilmington, DE, USA).
The TL assay
In white blood cells, TL was measured using the real-time quantitative polymerase chain reaction (Q-PCR) as described previously (Gutmajster et al. 2013). Briefly, the amount of telomeric DNA (T) was divided by the amount of single-copy control gene DNA (S) which encodes acidic ribosomal phosphoprotein P0 (36B4, accession number NC_000012.12), producing a relative measurement of the telomere length (T/S ratio). Results were related to a control sample, used for standard curve generation. The quality of PCR products was assessed by the melting curve analysis. All samples were run in triplicates, and the control sample (Human Genomic DNA, ROCHE, Germany) was run in each experiment to ensure correct normalization among experiments.
Promoter genotyping
DNA amplification was performed using the mix OptiTaq™ PCR Master Mix (Eurix®) as follows: 5 min at 94 °C, 35 cycles at 94 °C for 30 s, 56 °C for 30 s, 72 °C for 1 min 30 s, with a final step at 72 °C for 7 min. Primer sequences were as follows: forward primer for TERT (TERT_F): 5′ATTCGACCTCTCTCCGCTGG3′; reverse primer (TERT_R): 5′CTGGAAGGTGAAGGGGCAG3′. PCR product was treated with exonuclease/alkalic phosphatase mix (Eurix®).
The sequencing PCRs were performed with the BigDye v.3.1 (Fisher Scientific, USA) with internal primers and sequences thanks to courtesy of Prof. Grzybowska (Varadi et al. 2009). Products were cleaned with the BigDye Terminator X (Fisher Scientific, USA). DNA sequencing was performed using Sanger’s technique and analyzed with the ABIPrism3130xl instrument (Fisher Scientific, USA). Results were analyzed with the Blast software (NCBI, USA).
Laboratory measurements
Serum total cholesterol, LDL cholesterol, HDL cholesterol, triglycerides, glucose, uric acid and C-reactive protein concentrations were assessed by an automated system (Modular PPE, Roche Diagnostics GmbH, Mannheim, Germany) in a single certified laboratory.
Serum insulin concentration was assessed by electrochemiluminescence method (ECLIA) using commercially available kits and the Cobas E411 analyzer (Roche Diagnostics GmbH, Mannheim, Germany). Homeostatic model assessment of insulin resistance (HOMA-IR) was calculated with the standard formula: HOMA-IR = fasting serum insulin (μIU/ml) × fasting glucose (mg/dl)/405.
The plasma concentration of interleukin 6 (R&D Systems, Mineapolis, MN, USA) and adiponectin (B-Bridge International Inc., San Jose, CA, USA) was measured by the immunoenzymatic (ELISA) method.
Data analysis
Results from both the assays and tests and from the questionnaire-derived data on gender, age, body mass index (BMI), blood pressure and smoking status were analysed. Characteristics of the study population are presented as mean and standard deviation (SD) or median and lower-upper quartile (for non-normal distribution of the data). Categorical variables are reported as frequency and percentage. Pearson’s or Spearman’s correlation coefficients were calculated whenever necessary. Comparison between mean values in the two groups (patients and controls) according to RTL was evaluated using the Student’s t test or the Mann-Whitney U test when applicable. The association between TERT genotypes and variables of interest was initially assessed using multivariate logistic regression analysis, and then comparisons of groups were performed using chi-square test. The lowest analysed minor allele frequency (MAF) was accepted as 0.23, and the calculated power of the test was 0.81, which is the value statistically acceptable for the number of participants in this study (140). Univariate analysis of the association of RTL, anthropometric and biochemical findings between TERT genotypes of SNPs was conducted using Pearson correlation. For each SNP, Hardy-Weinberg equilibrium was assessed according to methods described elsewhere www.oege.org/software (Rodriguez et al. 2009). The association between TERT genotypes and T2DM risk was determined by calculating the odds ratio (OR) and 95% confidence interval (CI) using binary logistic regression. All the statistical analyses were performed using Statistica v.12 (StatSoft, DELL, USA), and the criterion for statistical significance was p < 0.05.
Results
Characteristics of the study groups
RTL level was higher in men (99.9 ± 92.4, log RTL 4.35 ± 0.84) than in women (77.9 ± 70.1, log RTL 4.06 ± 0.89), with relatively high variance and the p values not significant. Also, no statistically significant correlation between RTL and age was found as expected (Table 1).
T2DM and control groups differed significantly only in respect of the markers of glucose metabolism, such as fasting plasma glucose concentration, HOMA-IR (each p < 0.001), serum insulin (p < 0.05), total cholesterol and LDL cholesterol (p < 0.05) concentrations (Table 1).
RTL value was higher in T2DM patients group (99.5 ± 93.7, log RTL 4.36 ± 0.87) than in control group (78.1 ± 68.1, log RTL 4.04 ± 0.86) but without statistical significance (Table 1).
Distribution of genotypes within study subjects
Seven already described polymorphisms within the analysed TERT promoter fragment were detected: rs2735940, rs3215401, rs7712562, rs33958877, rs35161420, rs35226131 and rs2853669. Four of them, rs3215401, rs33958877, rs35161420 and rs35226131, were excluded from further analyses due to their low MAFs (Table S1).
The rs2735940 polymorphism showed distribution against the Hardy-Weinberg law in both the diabetic participants and in the control group. Additionally, distribution of MAF in the group of healthy control subjects differed significantly from those in general European populations (Table S1).
Biochemical characteristics according to genotypes
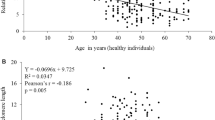
The rs2853669 genotypes negatively correlated with fasting plasma glucose concentrations in whole study group. However, TC and CC genotypes were accompanied by the lower plasma glucose concentrations (R2 = 0.03, p = 0.03) (Fig. 1). Also, the RTL was lower for TC and CC genotypes of this polymorphism in T2DM study subjects, as calculated from multiple regression analysis (R2 = 0.11, p = 0.005) (Fig. 2). Results presented in Table 2 indicated no elevated risk of T2DM due to the presence of any analysed SNPs. No other statistically significant relations with diabetes-related traits were found (data not shown).
Correlation of relative telomere length (RTL) with the genotypes of the rs2853669 polymorphism in T2DM group and compared to control group. The controls are black boxes, and the T2DM are filled with diagonal check pattern. The values are means + SD (error bars). The outer boxes mark 25–75%. Circles indicate Det. X that indicates Ext.
Discussion
Relations between RTL and T2DM
Short RTL has been postulated in some population studies as a risk factor for the development of T2DM (Tamura et al. 2016a). Results of the presented work, which was conducted on selected population of elderly obese participants with and without T2DM, did not reveal significant difference in RTL of the two groups. Also, the risk of T2DM reported previously by others was fairly attenuated in postmenopausal women (You et al. 2012). Additionally, no correlations were obtained when some diabetes related traits were analysed. It was hypothesized that telomere attrition would be a marker of insulin resistance (Verhulst et al. 2016); however, in this work, no significant correlation was found between RTL and HOMA-IR. According to other reports, RTL was already reduced in individuals with impaired glucose tolerance (Bethancourt et al. 2017), but high plasma glucose concentration alone did not result in faster telomere shortening (Salpea et al. 2013). Our data do not differ from previously reported findings. One explanation is that T2DM patients usually regularly take glucose-lowering medicines and other formulations; thus, the pathological effect of the disease could be, to some extent, distorted. Additionally, numerous studies have reported that the association between telomere attrition and both types of DM were correlated with the time of the disease duration (Willeit et al. 2014). Considering the fact, that weight gain during adulthood is associated with shortened telomeres, although this dependence is stronger at age under 60 years (Müezzinler et al. 2016), it seems that this phenomenon, but not T2DM itself, is responsible for RTL in the population analysed here.
Selected TERT genotypes in the participants with and without T2DM
In our study, the rs2853669 CC homozygotes had the shortest RTL when compared to other genotypes of this polymorphism. The difference, however, was significant only in T2DM subjects. This SNP is localized in the erythroblast transformation specific - 2 (ETS2) transcription factor consensus binding site. ETS2 is a positive regulator of TERT expression and the C allele was shown to reduce telomerase activity (Soerensen et al. 2012). The plasma glucose concentration was lowest in the CC homozygotes presumably due to reduced telomerase activity. It is in correlation with the previous observation that TERT regulates the pentose phosphate pathway; therefore, cells bearing TERT mutations promoting telomerase activation exhibited diminished glycogen accumulation (Ahmad et al. 2016).
For the other detected polymorphisms, the genotypes did not differ significantly, although for the SNP rs2735940, the 25% higher promoter activity for genotype TT and longer telomeres were previously reported (Matsubara et al. 2006b). In our studies, this polymorphism showed significant deviation from the Hardy-Weinberg distribution, with higher proportion of homozygotes in T2DM subjects. The observation was correlated with similar results obtained for a larger population (Montesanto et al. 2018). Therefore, we conclude that the presence of rs2735940-C allele could provide additive prognostic value on the presence of T2DM.
Also, the polymorphism rs7712562 showed deviation from H-W disequilibrium in both analysed groups. This observation, together with the statistically important difference of MAF distribution of polymorphism rs2735940 between our control group of old, relatively healthy participants and data presented for Central Europeans by NCBI, USA, gives an impact to further investigation in a larger study group on their influence on longevity. Polymorphisms under study are linked (data not shown); further study on variants combinations can set a new light on the TERT promoter functioning. Also, the observed variation in RTL that could not be linked directly neither to T2DM nor to the TERT promoter sequence variants indicates that there are also other contributing factors that need more attention.
General conclusions
Our study has some limitations. One of them is that the absolute Q-PCR results from different laboratories widely differ from each other. Next one is that the telomere length is likely to differ among blood cell types. Finally, the study was conducted on a relatively small sample size, which might have led or partially contributed to the weakness of the observed effects.
On the other hand, our study has a number of strengths. Firstly, the study participants were well selected in terms of morbidity: except for T2DM in the test group, they have had no other diagnosed diseases. In addition, characteristics of both groups were very similar as they differed only by the presence or absence of glucose and lipid disturbances. Secondly, a large fragment of the promoter was sequenced and analysed, including the core promoter region, where no other sequence variations were detected. Further analyses of the promoter/regulatory regions of TERT are necessary because the single base changes especially in DNA sequences of transcription factors and/or general transcription factors, recognition/binding sites might affect TERT transcription initiation and its rate.
References
Ahmad F, Dixit D, Sharma V et al (2016) Nrf2-driven TERT regulates pentose phosphate pathway in glioblastoma. Cell Death Dis 7:e2213. https://doi.org/10.1038/cddis.2016.117
Bethancourt HJ, Kratz M, Beresford SAA et al (2017) No association between blood telomere length and longitudinally assessed diet or adiposity in a young adult Filipino population. Eur J Nutr 56:295–308. https://doi.org/10.1007/s00394-015-1080-1
Bledowski P, Mossakowska M, Chudek J et al (2011) Medical, psychological and socioeconomic aspects of aging in Poland: assumptions and objectives of the PolSenior project. Exp Gerontol 46:1003–1009. https://doi.org/10.1016/j.exger.2011.09.006
Bonfigli AR, Spazzafumo L, Prattichizzo F et al (2016) Leukocyte telomere length and mortality risk in patients with type 2 diabetes. Oncotarget 7:50835–50844. https://doi.org/10.18632/oncotarget.10615
Bressler J, Franceschini N, Demerath EW et al (2015) Sequence variation in telomerase reverse transcriptase (TERT) as a determinant of risk of cardiovascular disease: the Atherosclerosis Risk in Communities (ARIC) study. BMC Med Genet 16:52. https://doi.org/10.1186/s12881-015-0194-x
Buxton JL, Walters RG, Visvikis-Siest S et al (2011) Childhood obesity is associated with shorter leukocyte telomere length. J Clin Endocrinol Metab 96:1500–1505. https://doi.org/10.1210/jc.2010-2924
Codd V, Nelson CP, Albrecht E et al (2013) Identification of seven loci affecting mean telomere length and their association with disease. Nat Genet 45:422–427. https://doi.org/10.1038/ng.2528
Elks CE, Scott RA (2014) The long and short of telomere length and diabetes. Diabetes 63:65–67. https://doi.org/10.2337/db13-1469
Franzke B, Neubauer O, Wagner K-H (2015) Super DNAging—new insights into DNA integrity, genome stability and telomeres in the oldest old. Mutat Res 766:48–57
Gardener M et al (2014) Gender and telomere length: systematic review and meta- analysis. ☆ HHS Public Access. Exp Gerontol 51:15–27. https://doi.org/10.1016/j.exger.2013.12.004
Gutmajster E, Witecka J, Wyskida M et al (2013) Telomere length in elderly Caucasians weakly correlates with blood cell counts. Sci World J 2013:153608. https://doi.org/10.1155/2013/153608
Heidenreich B, Rachakonda PS, Hemminki K, Kumar R (2014) TERT promoter mutations in cancer development. Curr Opin Genet Dev 24:30–37
Helbig S, Wockner L, Bouendeu A et al (2017) Functional dissection of breast cancer risk-associated TERT promoter variants. Oncotarget 8:67203–67217. https://doi.org/10.18632/oncotarget.18226
Houben JMJ, Giltay EJ, Rius-Ottenheim N et al (2011) Telomere length and mortality in elderly men: the Zutphen Elderly Study. J Gerontol A Biol Sci Med Sci 66A:38–44. https://doi.org/10.1093/gerona/glq164
Ishikawa N, Nakamura K-I, Izumiyama-Shimomura N et al (2016) Changes of telomere status with aging: an update. Geriatr Gerontol Int 16:30–42. https://doi.org/10.1111/ggi.12772
Lee JH, Cheng R, Honig LS et al (2013) Genome wide association and linkage analyses identified three loci-4q25, 17q23.2, and 10q11.21-associated with variation in leukocyte telomere length: the long life family study. Front Genet 4. https://doi.org/10.3389/fgene.2013.00310
Lewis KA, Tollefsbol TO (2016) Regulation of the telomerase reverse transcriptase subunit through epigenetic mechanisms. Front Genet 7:83. https://doi.org/10.3389/fgene.2016.00083
Liu J-P (2014) Molecular mechanisms of ageing and related diseases. Clin Exp Pharmacol Physiol 41:445–458. https://doi.org/10.1111/1440-1681.12247
Ma L, Li Y, Wang J (2015) Telomeres and essential hypertension. Clin Biochem 48:1195–1199. https://doi.org/10.1016/j.clinbiochem.2015.07.013
Matsubara Y, Murata M, Watanabe K et al (2006a) Coronary artery disease and a functional polymorphism of hTERT. Biochem Biophys Res Commun 348:669–672. https://doi.org/10.1016/j.bbrc.2006.07.103
Matsubara Y, Murata M, Yoshida T et al (2006b) Telomere length of normal leukocytes is affected by a functional polymorphism of hTERT. Biochem Biophys Res Commun 341:128–131. https://doi.org/10.1016/j.bbrc.2005.12.163
Mazidi M, Penson P, Banach M (2017) Association between telomere length and complete blood count in US adults. Arch Med Sci 13:601–605. https://doi.org/10.5114/aoms.2017.67281
Melicher D, Buzas EI, Falus A (2015) Genetic and epigenetic trends in telomere research: a novel way in immunoepigenetics. Cell Mol Life Sci 72:4095–4109. https://doi.org/10.1007/s00018-015-1991-2
Menke A, Casagrande S, Cowie CC (2015) Leukocyte telomere length and diabetes status, duration, and control: the 1999-2002 National Health and Nutrition Examination Survey. BMC Endocr Disord 15:52. https://doi.org/10.1186/s12902-015-0050-1
Meyer A, Salewsky B, Buchmann N et al (2016) Relative leukocyte telomere length, hematological parameters and anemia—data from the Berlin Aging Study II (BASE-II). Gerontology 62:330–336. https://doi.org/10.1159/000430950
Montesanto A, Bonfigli AR, Crocco P et al (2018) Genes associated with type 2 diabetes and vascular complications. Aging 10:178–196. https://doi.org/10.18632/aging.101375
Müezzinler A, Zaineddin AK, Brenner H (2014) Body mass index and leukocyte telomere length in adults: a systematic review and meta-analysis. Obes Rev 15:192–201. https://doi.org/10.1111/obr.12126
Müezzinler A, Mons U, Dieffenbach AK et al (2016) Body mass index and leukocyte telomere length dynamics among older adults: results from the ESTHER cohort. Exp Gerontol 74:1–8. https://doi.org/10.1016/j.exger.2015.11.019
Nilsson PM, Tufvesson H, Leosdottir E et al (2013) Telomeres and cardiovascular disease risk: an update 2013. Transl Res 162:371–380. https://doi.org/10.1016/j.trsl.2013.05.004
Njajou OT, Cawthon RM, Blackburn EH et al (2012) Shorter telomeres are associated with obesity and weight gain in the elderly. Int J Obes 36:1176–1179. https://doi.org/10.1038/ijo.2011.196
Opresko PL, Shay JW (2017) Telomere-associated aging disorders. Ageing Res Rev 33:52–66
Ozturk M, Li Y, Tergaonkar V (2017) Current insights to regulation and role of telomerase in human diseases. Antioxidants 6:17. https://doi.org/10.3390/antiox6010017
Qi Nan W, Ling Z, Bing C (2015) The influence of the telomere-telomerase system on diabetes mellitus and its vascular complications. Expert Opin Ther Targets 19:849–864. https://doi.org/10.1517/14728222.2015.1016500
Raschenberger J, Lamina C, Haun M et al (2016) Influence of DNA extraction methods on relative telomere length measurements and its impact on epidemiological studies. Sci Rep 6. https://doi.org/10.1038/srep25398
Rode L, Nordestgaard BG, Bojesen SE (2015) Peripheral blood leukocyte telomere length and mortality among 64 637 individuals from the general population. J Natl Cancer Inst 107. https://doi.org/10.1093/jnci/djv074
Rodriguez S, Gaunt TR, Day INM (2009) Hardy-Weinberg equilibrium testing of biological ascertainment for Mendelian randomization studies. Am J Epidemiol 169:505–514. https://doi.org/10.1093/aje/kwn359
Salpea KD, Maubaret CG, Kathagen A et al (2013) The effect of pro-inflammatory conditioning and/or high glucose on telomere shortening of aging fibroblasts. PLoS One 8:e73756. https://doi.org/10.1371/journal.pone.0073756
Shaheen F, Grammatopoulos DK, Müller J et al (2014) Extra-nuclear telomerase reverse transcriptase (TERT) regulates glucose transport in skeletal muscle cells. Biochim Biophys Acta Mol Basis Dis 1842:1762–1769. https://doi.org/10.1016/j.bbadis.2014.06.018
Singhapol C, Pal D, Czapiewski R et al (2013) Mitochondrial telomerase protects cancer cells from nuclear DNA damage and apoptosis. PLoS One 8:e52989. https://doi.org/10.1371/journal.pone.0052989
Soerensen M, Thinggaard M, Nygaard M et al (2012) Genetic variation in TERT and TERC and human leukocyte telomere length and longevity: a cross-sectional and longitudinal analysis. Aging Cell 11:223–227. https://doi.org/10.1111/j.1474-9726.2011.00775.x
Svenson U, Nordfjäll K, Baird D et al (2011) Blood cell telomere length is a dynamic feature. PLoS One 6:e21485. https://doi.org/10.1371/journal.pone.0021485
Tamura Y, Takubo K, Aida J et al (2016a) Telomere attrition and diabetes mellitus. Geriatr Gerontol Int 16:66–74
Tamura Y, Izumiyama-Shimomura N, Kimbara Y et al (2016b) Telomere attrition in beta and alpha cells with age. Age (Dordr) 38:61. https://doi.org/10.1007/s11357-016-9923-0
Testa R, Olivieri F, Sirolla C et al (2011) Leukocyte telomere length is associated with complications of type 2 diabetes mellitus. Diabet Med 28:1388–1394. https://doi.org/10.1111/j.1464-5491.2011.03370.x
Varadi V, Brendle A, Grzybowska E et al (2009) A functional promoter polymorphism in the TERT gene does not affect inherited susceptibility to breast cancer. Cancer Genet Cytogenet 190:71–74. https://doi.org/10.1016/j.cancergencyto.2008.12.006
Verhulst S, Dalgård C, Labat C et al (2016) A short leucocyte telomere length is associated with development of insulin resistance. Diabetologia 59:1258–1265. https://doi.org/10.1007/s00125-016-3915-6
Wick M, Zubov D, Hagen G (1999) Genomic organization and promoter characterization of the gene encoding the human telomerase reverse transcriptase (hTERT ). Gene 232:97–106
Willeit P, Raschenberger J, Heydon EE et al (2014) Leucocyte telomere length and risk of type 2 diabetes mellitus: new prospective cohort study and literature-based meta-analysis. PLoS One 9:e112483. https://doi.org/10.1371/journal.pone.0112483
You N, Chen B, Song Y et al (2012) A prospective study of leukocyte telomere length and risk of type 2 diabetes in postmenopausal women. Diabetes 61:2998–3004. https://doi.org/10.2337/db12-0241
Zhang J, Rane G, Dai X et al (2016) Ageing and the telomere connection: an intimate relationship with inflammation. Ageing Res Rev 25:55–69
Zhou J, Ding D, Wang M et al (2014) Telomerase reverse transcriptase in the regulation of gene expression. BMP Rep 47(1):8–14. https://doi.org/10.5483/BMBRep.2014.47.1.284
Zhou Y, Ning Z, Lee Y et al (2016) Shortened leukocyte telomere length in type 2 diabetes mellitus: genetic polymorphisms in mitochondrial uncoupling proteins and telomeric pathways. Clin Transl Med 5:8. https://doi.org/10.1186/s40169-016-0089-2
Acknowledgements
The authors would like to acknowledge Profs. Jan E. Zejda, Andrzej Franek, Malgorzata Kowalska and Edward Błaszczak for discussing the statistical methods used in this work.
Funding
The work used the material collected within PolSenior study, which was implemented under publicly funded project no. PBZ-MEIN-9/2/2006, Ministry of Science and Higher Education. The work was in part financially supported by Institutional grants KNW-1-010/P/1/0, KNW-1-007/P/2/0 and KNW-1-041/K/3/0 (all awarded to ALS). The equipment for molecular analyses used in this laboratory work was purchased using the EU funds (European Regional Development Fund) within the Sectoral Operational Program “Increase of Economic Competitiveness”.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The work complied with ethical standards that were met by the PolSenior programme project.
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The study was approved by the Ethics Committee of the Medical University of Silesia and was carried out in accordance with The Code of Ethics of the World Medical Association (Declaration of Helsinki).
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Communicated by: Michal Witt
Electronic supplementary material
Table S1
Distribution of the TERT alleles in healthy obese controls and obese diabetic (T2DM) elderly subjects. (DOCX 23 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Gutmajster, E., Chudek, J., Augusciak-Duma, A. et al. Possible association of the TERT promoter polymorphisms rs2735940, rs7712562 and rs2853669 with diabetes mellitus in obese elderly Polish population: results from the national PolSenior study. J Appl Genetics 59, 291–299 (2018). https://doi.org/10.1007/s13353-018-0450-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-018-0450-9