Abstract
The third SARS-CoV-2 pandemic wave causing Omicron variant has comparatively higher replication rate and transmissibility than the second wave-causing Delta variant. The exact mechanism behind the differential properties of Delta and Omicron in respect to infectivity and virulence is not properly understood yet. This study reports the analysis of different mutations within the receptor binding domain (RBD) of spike glycoprotein and non-structural protein (nsp) of Delta and Omicron strains. We have used computational studies to evaluate the properties of Delta and Omicron variants in this work. Q498R, Q493R and S375F mutations of RBD showed better docking scores for Omicron compared to Delta variant of SARS-CoV-2, whereas nsp3_L1266I with PARP15 (7OUX), nsp3_L1266I with PARP15 (7OUX), and nsp6_G107 with ISG15 (1Z2M) showed significantly higher docking score. The findings of the present study might be helpful to reveal the probable cause of relatively milder form of COVID-19 disease manifested by Omicron in comparison to Delta variant of SARS-CoV-2 virus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
SARS-CoV-2 has manifested itself in a variety of ways ranging from asymptomatic infection to severe pneumonia and death. The virulence and transmissibility might be regulated by the viral genomic constellation as well as host factors. The mutation rate of RNA viruses is quite higher than that of their host, causing rapid virus evolution [12]. SARS-CoV-2 is prone to accumulate rapid mutations while adapting to their new human host leading to the emergence of newer variants with time [30]. Presence of significant mutations across the genome of SARS-CoV-2 variants are key factor to the evolutionary dynamics of this rapidly mutating virus.
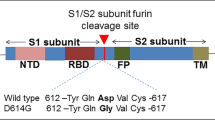
The SARS-CoV-2 Delta variant of concern (VoC) seemed to be one of the drivers of the detrimental COVID-19 ‘second wave’ across India, during early 2021 (Supplementary Table 1). The transmission rate of Delta was 40–60% higher in comparison to Alpha (B.1.1.7) VOC. The probability of Delta infection is higher in case of unvaccinated or partially vaccinated population compared to the vaccinated population [38]. Thereafter, the third COVID-19 wave was driven by the Omicron VoC, which outcompeted Delta VoC in context to transmissibility, which revealed 29% less disease severity than Delta along with high immune evasion potential [6, 20]. The R0 value (average basic reproduction number) of Omicron is as high as 10, whereas it is 5–9 in case of Delta [21]. The Delta variant carries 9 signature mutations within the structural spike glycoprotein, of which 2 are in the RBD, while the role of many of these mutations remains to be elucidated. There are approximately 30 signature mutations within the Omicron variant which have been reported across the spike protein of which 15 are within the receptor binding domain (RBD), including three deletions and one insertion [17] (Supplementary Table 2). The alignment of different RBD mutations in the Delta and Omicron variant has been depicted in Supplementary Figure 1.
The non-structural proteins (NSPs) are fundamentals in viral replication, transcription, assembly of mature virions as well as modulation of the host immune system [16, 29]. It has been found that the ORF1a (Open Reading Frame 1a) contributes to immune evasion potential of the virus and this positive selection drives the evolution of the NSPs [7, 15] (Supplementary Table 3). The ‘cytokine storm’ is a major cause of acute tissue injury among several patients with COVID-19 infection [33, 44]. To date, very few biologically relevant specific protein structures and mutant models of angiotensin peptide and Zn-bound ACE2 (Angiotensin Converting Enzyme 2) receptors have been proposed. In the current study, multiple point mutations within the RBD region of the spike (S) protein have been elucidated. Since the Omicron variant accumulated a lot of mutations specifically at RBD region of spike protein and since RBD plays a significant role in early event of viral infection, multiple point mutations within the RBD region have been examined in the current study. Moreover, we emphasized on the effect of the conformational alterations of these variants on their functional significance. In silico docking-based study has also been implicated on the immunological pathway related proteins responsible for evoking the ‘cytokine storm’ which might give some indication regarding the differential response between Delta and Omicron variants.
Materials and methods
Data retrieval
The FASTA sequence of the spike protein and non-structural proteins of SARS-CoV-2 of the Wuhan-Hu-1 (Wild type) was obtained from Uniport (https://www.uniprot.org/) (Accession no: P0DTC2) (The UniProt Consortium). The spike protein (GenBank Accession no: QWK65230.1) of the Delta variant was obtained from ViPR (Virus Pathogen Resource, https://www.viprbrc.org). The common mutations in the receptor binding domain (RBD) of the spike gene (S) within VoCs of SARS-CoV-2 were enlisted from GISAID (Global Initiative on Sharing Avian Influenza Data) (https://gisaid.org/). GISAID helps in rapid sharing of data from all influenza viruses and the coronavirus causing COVID-19. At the same time, it helps to understand how viruses evolve and spread during pandemics [40]. The corresponding nucleotide sequences of mutation was obtained from Next Clade (https://clades.nextstrain.org/). It is a tool that performs genetic sequence alignment, phylogenetic clade analysis, mutation study for SARS-CoV-2, Monkeypox, Influenza (Flu) and other clinically significant pathogens [1].
Analysis of conserved residues and identification of mutation
NCBI Blastp Tool (National Centre for Biotechnology Information Basic Local Alignment Search Tool Protein; https://blast.ncbi.nlm.nih.gov/Blast.cgi?PAGE=Proteins) is a bioinformatics programme, which was used to align the Wuhan-Hu-1 (wild type) sequence with variants of Delta and Omicron sequences. The box shade application was used to create the alignment figure.
Structural modification
Amino acid FASTA sequences for Homo sapiens ACE2 protein (NCBI GenBank ID: BAB40370.1) and SARS-CoV-2 spike protein RBD domain (NCBI RefSeq ID: NC_045512.2) were obtained from NCBI. The RBD mutant sequence was prepared by manually incorporating substitution in the amino acid sequence. The corresponding protein structure for wild type and mutant RBD was generated using RaptorX (http://raptorx.uchicago.edu/) software (https://doi.org/10.1093/nar/gkw306). It performs multiple sequence alignment (MSA) based on a deep-learning method known as DeepCNF (Deep Convolutional Neural Fields) which deals with complex sequence-structure relationship for accurate prediction without homology or template-based modelling.
The PDB (Protein Data Bank) file of Homo sapiens ACE2 (PDB ID: 6VW1) have been modified by removing water and bound ligand. Hydrogen bonds have been added to the PDB file and RBDs by using autodock tools panel [28]. Computational docking and scoring techniques have revolutionized structural bioinformatics by providing unprecedented insights on key aspects of ligand-receptor interaction. Docking is used for optimizing known drugs and for identifying novel binders by predicting their binding mode and affinity. AutoDock and AutoDockTools (https://autodocksuite.scripps.edu/adt/) are free of charge techniques which are generally used for structure-based drug design. It is an automated procedure for predicting the interaction of ligands with biomacromolecular targets. AutoDock calculations are performed in several steps: (1) preparation of coordinate files using AutoDockTools, (2) precalculation of atomic affinities using AutoGrid, (3) docking of ligands using AutoDock, and (4) analysis of results using AutoDockTools. In this work, we used AutoDock for checking the pdb files (ACE2, RBD, various nsps and other proteins like STAT1) in search of missing amino acids and correct loops and turns.
The structure of the SARS-CoV2 non-structural proteins (nsps) has been predicted by homology modeling through the phyre2 webserver (http://www.sbg.bio.ic.ac.uk/phyre2) and human proteins have been retrieved from the protein data bank (Table 1). Phyre2 is an online tool available for predicting as well as analysing the protein structure, function, and mutations. It has an advanced remote homology detection method to build 3D models, analyse the effects of amino acid variants for the given protein sequence, and determine the ligand binding site. Depending on sequence length, number of homology sequences and frequency, Phyre2 server generally takes 30 min to several hours for a prediction to complete after pasting the protein amino acid sequence. Usually in Phyre 2, the submitted protein sequences are first scanned against a large sequence database using PSI-BLAST (https://www.ebi.ac.uk/Tools/sss/psiblast/). After that, generated PSI-BLAST is processed by the neural network secondary structure prediction programme PsiPred (PSI-blast based secondary structure PREDiction; http://bioinf.cs.ucl.ac.uk/psipred/) and the protein disorder predictor Disopred (https://bio.tools/disopred3). The colour coded confidence bar is used to predict the presence of alpha-helices, beta-strands and disordered regions. Here, we pasted the FASTA sequences to the phyre2 server, given job ids and email addresses, and opted for intensive modelling mode. After about 24 h, we got the homology models by email.
Molecular docking
Molecular dockings have been done by cluspro server (https://cluspro.org). It is a widely used tool for protein–protein docking study. The server only requires two files in PDB format and offers many advanced options to modify the search. It also removes the unstructured protein regions, helps in construction of homo-multimers, accounts for pairwise distance restraints, and locate the heparin-binding sites [11, 22, 23]. Cluspro delivers the scores coming from Piper (https://www.schrodinger.com/products/piper) which ranks models by cluster size. In this method, one of the proteins (receptor protein) is positioned at the origin of the coordinate system on a fixed grid, the second protein (ligand protein) is sited on a movable grid, and the interaction energy is documented as a correlation function or as a sum of a few correlation functions. Cluspro rotates the ligand with 70,000 rotations. For each rotation, it translates the ligand in x,y,z coordinates relative to the receptor on a grid. Then it selects the translation with the best score from each rotation. To find the ligand position with the most "neighbours" in 9 angstroms, and to become a cluster centre, clustering of ligand positions with a 9-angstrom C-alpha root-mean-square deviation radius is done. The docking score is calculated based on the following equation in cluspro: E = w1Erep + w2Eattr + w3Eelec + w4EDARS. Here, Erep, Eattr, and Eelect denote the repulsive and attractive contribution to the van der Waals interaction and electrostatic energy, respectively. EDARS represents the Decoys as Reference State (DARS) approach which is measured by the amount of free energy change following the removal of water molecules from the protein interface. This parameter takes the desolvation contribution into account. W1–4 coefficients are weighted which is calculated for different types of docking problems [45]. Here, we first submitted the pdb files of ACE2 and RBD in cluspro server and used default parameters for docking to observe the interactions between ACE2 and RBD and the various nsps with other proteins like STAT1. After that, we got our job id on cluspro servers in running job section. We have checked the job id section to see that the server has taken the correct pdb files in the user input section. After three hours, we got the docking results by email.
PyMol structural modeling
The PyMol molecular graphics system (version 2.4.2) from Schrödinger, Inc. was used for downloading structure files from the PDB database for further analysis and image generation. PyMOL (https://pymol.org/2/) is a widely used open-source Python-based software for visualization, editing and analysis of structure of biomolecules like proteins, nucleic acids, and other small molecules (https://doi.org/10.1002/wcms.1298). Structural images were cropped via Adobe Photoshop (https://adobe-photoshop-7-0-1-update.en.softonic.com/) for further representation through proper illustration.
Results
Docking result and docking position analysis
After completing the docking simulation, we got the top 30 docking poses between ACE2 and RBD. The docking poses were sorted based on cluspro lowest docking score and largest cluster size. The docking score of ACE2 and RBD ranged from (− 720) to 1265.9 (Table 2). The comprehensive perception of SARS-CoV-2 (Omicron Variant) spike protein RBD and human ACE2 after docking has been described in Fig. 1. 723649_model_5 of S_Q498R showed a docking score of (− 1100.6) and cluster size of 174. The docking score of 723647_model_2 (Q493R) had been (− 1265.9) with a cluster size of 96. 723366_model_5 (S375F) had been (− 1141.1) with a cluster size of 183. When these best three models were compared to the crystal structure of SARS-CoV-2 RBD bound human ACE2, it was found that 723649_model_5 of Q498R bound was closer than the other two models (Fig. 2). The SARS-CoV-2 RBD and human ACE2 complex formed 16 hydrogen bonds in the range of 2.67 Å to 3.17 Å. Moreover, the complex formed 16 salt bridges between them too.
Comprehensive perception of SARS-CoV-2 (Omicron Variant) spike protein RBD and human ACE2 after docking. A The secondary structure of ACE2 has been shown by ribbon, and RBD by ribbon and transparent surface structure. B Interaction between the amino acids ACE2, and RBD, where chain A denotes ACE2 and chain B denotes RBD respectively
The docking score of nsps ranged from (− 763.7) to (− 1479.1) (Table 3). The best docking score (− 1479.1) was exhibited by nsp6_G107 (W) with a cluster size of 69. The highest docking score and experimental evidence helped us to make this strain as most docked nsps in comparison to other nsps [5, 49]. The details of the docking analysis of NSP6 (G107w) (Omicron variant) and human STAT1 has been shown in Fig. 3.
Comprehensive perception of SARS-CoV-2 NSP6 (G107w) and human STAT1 after docking. A The secondary structure of STAT1 has been shown by ribbon, and NSP6 by ribbon and transparent surface structure. B Interaction between the amino acids of STAT1 and NSP6, where chain A denotes NSP6 and chain B denotes STAT1 respectively
Chemical interactions
The number of non-bonded contacts between SARS-CoV-2 RBD and human ACE2 was 159. The 16 hydrogen bonds and their binding distance that were less than four angstroms supported the tight binding of the SARS-CoV-2 RBD and human ACE2 complex. The salt bridges contributed to the stability of the entropically unfavourable folded conformation of the complex. The abundance of non-bonded contacts between SARS-CoV-2 RBD and human ACE2 were also in favour of strong binding. Non-bonded contacts between ACE2 (Chain A) and RBD (Chain B), hydrogen bonds between ACE2 (Chain A) and RBD (Chain B), as well as salt bridges between ACE2 (Chain A) and RBD (Chain B) have been described in Tables 4, 5, and Supplementary Table 4 respectively. The ribbon structures of different RBD mutations of Delta and Omicron strains of SARS-CoV-2 has been described in Fig. 4.
The SARS-CoV-2 NSP6 (G107w) and human STAT1 complex formed 15 hydrogen bonds in the range of 2.54 Å to 3.30 Å. The number of non-bonded contacts between SARS-CoV-2 NSP6 (G107w) and human STAT1 was 260. The 16 hydrogen bonds and their binding distance that were less than four angstroms supported the tight binding of the SARS-CoV-2 NSP6 (G107w) and human STAT1 complex. The abundance of non-bonded contacts between SARS-CoV-2 NSP6 (G107w) and human STAT1 were also in favour of strong binding. Hydrogen bonds between nsp6 (Chain A) and STAT1 (Chain B) and non-bonded contacts have been represented in Table 6 and Supplementary Table 5 respectively.
Discussion
We attempted to identify the probable host and SARS-CoV-2 interacting protein determinants, encompassing variants of concern (VoC) like Delta and Omicron. The spike glycoprotein (S) facilitates the viral entry into host cells by binding with the receptor binding domain (RBD) via S1 subunit and then fusing the viral and host membrane with the help of S2 subunit [31]. The RBD domain of SARS-CoV-2 Spike is optimal for binding to human ACE-2 through additional hydrogen bond formation and hydrophobic force in comparison to other species [48]. Mutations in RBD increase the stability of the spike virus structure and reduce the binding efficiency of vaccine-induced antibodies [26]. At the initial stage of the pandemic, S-protein mutations were very rare specially in the RBD region. Gradually, considerable number of new mutations have been accumulated for adapting new host species by increasing the binding affinity for the host receptor (Supplementary Table 6). It is important to address RBD in the S protein of SARS-CoV-2 as it is the most likely target for the invention of virus attachment inhibitors, vaccines, and neutralizing antibodies [43]. Till now, there are only few reports available on the mutation spectrum of RBD and its effects for binding with hACE-2. So, in our present study the RBD region has been focussed instead of the whole S-protein.
Mutations are the fundamental basis of most of the evolution and they can bring about different types of variations upon which natural selection can act. Although most of the mutations are not beneficial for the organisms and are deleterious in nature. The percentage of mutations which are harmful versus beneficial may alter in different organisms over time, deleterious mutations are assumed to always outnumber the beneficial mutations. In case of RNA viruses, the mutation rate is much higher and small increase in mutation rate can cause RNA viruses to become extinct [12]. Therefore, researchers are now focussed upon the impacts of mutations within the viruses. As time passes, a considerable number of new mutations have been accumulated in the SARS-CoV-2 variants, altering the binding affinity for the host receptor (Supplementary Table 6). There are scanty reports available so far on the mutation spectrum of RBD and its effects for binding with hACE-2. Among the different mutations reported in the receptor binding domain (RBD) of S protein, most of them are of synonymous mutations (an alteration in the DNA sequence that codes for amino acids in a protein sequence, but does not modify the encoded amino acid), and only few were reported to affect the functions of the S protein [8, 42] (Supplementary Table 7).
According to our findings, alterations in the RBD region of the Omicron variant might account for the high binding specificity with hACE2, which might contribute to the higher transmissibility and decreased virulence, when compared to the Delta variant. The 3 mutations in RBD (Q498R, Q493R and S375F) of Omicron exhibited better affinity and binding efficiency with hACE2. The Delta variant shared two RBD mutations with Omicron, including K417N, and T478K. It has been linked to S protein structural alterations which may facilitate immune evasion [32]. The T478K substitution helped in efficient RBD binding and enhanced immunological escape [10]. Delta L452R mutation facilitated the affinity for ACE2 receptors found in various kinds of human cells including the lungs [37]. In addition to these two common mutations, there are many amino acid alterations found within the RBD of the Spike protein including N440K, G446S, S477N, E484A, Q493K, G496S, Q498R, N501Y, and Y505H in the Omicron strain. It is evident from previous studies that few mutations within the RBD are pathologically significant. For example, K417N and N501Y mutations have contributions in immune escape and enhanced infectivity [32]. The combination of mutations Q498R and N501Y significantly increased the binding capacity of ACE2 [51]. The functional attributions of many other mutations still require further investigation. Amongst the different RBD mutations of these two strains, Q498R mutation of Omicron gave best docking score which indicated the higher binding efficiency with ACE2. In addition, Q493R and S375F mutations of Omicron also gave better docking scores. The influence of different amino acid changes also triggered the differential infection rate of SARS-CoV-2. It explains the possible reason behind the high rate of infectivity in case of Omicron in comparison to Delta.
One of the earlier studies showed docking based assay to examine the differential properties of Delta and Omicron variants and found that the Omicron variant had a higher affinity for binding with hACE2 than the Delta variant due to the presence of different mutations in the RBD which was responsible for higher transmission [34]. Based on computational analysis, Q493R, N501Y, S371L, S373P, S375F, Q498R, and T478K mutations played significant role in binding with hACE2 [24]. Similarly in the present study, we have also found Q498R, Q493R and S375F mutations of Omicron showed more affinity with hACE2. More recently, a study on SARS-CoV-2 lineages have been investigated to find out multiple point mutations on spike RBD [18]. The structural impacts of selected S1 mutations within B.1.617.2 sub-lineage (Delta) have been discussed previously [36]. Therefore, the findings obtained in our study support the previous studies.
Mutations within the non-structural proteins further promote the weaker response of the immune system in defending the infection, resulting in delayed hyper inflammation with a weakened interferon (IFN) response. Delayed immune activation, extends infection, and endorses viral replication. The substantial production of IL-6, IL-1, TNF-α, and interferon may lead to form ‘cytokine storm’ which is a major cause of acute tissue injury during Covid infection [27]. It has been found that the adaptive evolution in ORF1a contribute to immune evasion due to selection pressure and the positive selection drives the evolution of nsps [13, 47]. Interferons (IFN) were key players behind the cytokine storm, which is a major complication of SARS-CoV-2 and can lead to respiratory distress syndrome (ARDS) and death. Here, in this present study we have selected nsp3, 5, 6 and 13 due to their potential role in generating immune evasion [3, 25, 50]. Nsp3 (PLpro) participates actively in innate immune response and it has indirect (deISGylation) as well as direct (cleavage) effects on the interferon regulatory factor 3 (IRF3) pathways. Moreover, it effectively inhibits IRF3 activity and leads to complete suppression of the type-I IFN response [52]. SARS-CoV-2 nsp1, PLpro, and nsp13 oppose to type I IFN signaling by mediating the inhibition of STAT1 and STAT2 phosphorylation, the deISGylation of IRF3, and the inhibition of STAT2 phosphorylation, correspondingly. Likewise, SARS-CoV-2 PLpro, nsp 6, and nsp 13 reduce TBK1 (TANK-binding kinase) phosphorylation, leading to the blockade of the RIG-1 pathway. Nsp3 (PLpro) can also interfere with RIG-1 (retinoic acid-inducible gene I) signaling through the deISGylate of MdeISGylate of MAD5 (melanoma differentiation-associated protein 5) [9]. Previous studies revealed that there are several mutations in nsps including nsp3 (K38R, V1069I, Δ1265, L1266I, A1892T), nsp5 (P132H), nsp6 (Δ105-107, A189V), nsp12 (P323L), and nsp14 (I42V) [39]. Another study showed mutations in NSP3, NSP6, NSP13, M protein, ORF7b, and ORF9b might have several important roles like higher transmission rate, lower infectivity rate, host immune evasion through natural killer cell inactivation, disruption of host protein synthesis and autophagosome-lysosome fusion prevention [19]. Therefore, we have selected few nsps for docking study by following the importance obtained from previous works.
In the present study, we did not observe significant changes in the docking scores between wild and mutant type nsps in case of Omicron variant. The results reflected that there was no such impact of mutations upon the activity of nsps in the case of Omicron. As described earlier, selected nsps play a crucial role in the inflammatory pathway leading to generate ‘cytokine storm’. So, the pathogenicity of Omicron was not so severe due to the presence of mutations in nsps. On the other hand, mutation in nsp13 was present in the Delta strain and the docking score between the wild (− 967.6) and mutant (− 940.3) types differs significantly which indicated that mutation in nsp13 played an important role in regulating the virulence of the disease in case of Delta variant (Table 3).
Considerable number of studies have been done on the evolutionary pattern of SARS-CoV-2. The RBD of the spike proteins and the region of the nucleocapsid protein associated with nuclear localisation signal (NLS) contain positively selected amino acid replacements. These replacements play important role in SARS-CoV-2 phylogeny. Genome sequence analysis of different strains of SARS-CoV-2 helps in deciphering the adaptive evolution of virus. Selection appears to act on combinations of mutations in spike domain as well as nucleocapsid domain [35]. One more study have been done on the recurrent mutations of SARS-CoV-2 Spike protein at residues K417, L452, E484, N501 and P681 emerging across various strains of SARS-CoV-2 including Alpha, Beta, Gamma, and Delta. With the advent of Omicron and its other sub lineages, additional group of mutations have been observed at different amino acid residues, namely R346, K444, N450, N460, F486, F490, Q493, and S494 [14]. The Spike (S) protein of SARS-CoV-2 exhibits both purifying selection and ancestral recombination events which lead to make S-protein capable of infecting human and other mammalian cells. The additional mutational variability leads to enhance the opportunities for future recombination [41]. Although there are substantial number of experiments have been done on spike proteins (S) of SARS-CoV-2, scanty reports are available on RBD of S-protein. Hence, in this present study, we have emphasized on the mutations within the RBD of both Delta and Omicron in a comparative pattern. With best of our knowledge, it is the pilot study on non-structural proteins (nsps) in searching the key for the differential virulence properties of Delta and Omicron strains of SARS-CoV-2. Moreover, docking based computational study on different mutations within RBD helped in gaining knowledge regarding the pathogenic properties of Delta and Omicron, the two different strains of SARS-CoV-2 which led to evoke waves of pandemic throughout the world.
Our present study relied on computer-based strategies to investigate multiple aspects of the virus and was supported by the previous findings [2, 4, 46]. Computational structural biology is the rapidly emerging integral part of applied immunology which continuously aid in the proper understanding in the structural basis of protein [36]. Computational study is very useful as there is very limited experimental and literature results available on the epidemiological distribution, mutational fitness, structure, and function of SARS-CoV-2. Therefore, scientists are now relied on simulation-based techniques to investigate differential properties of the virus. Computational tools of immunoinformatic are very convenient to reveal different aspects of infectious pathogens. Our present study is followed by these considerations. The present computational technique is a cost-effective approach for the prediction of newly emerged SARS-CoV-2 variants at molecular level. The acquired knowledge might be helpful for the scientific communities for further investigations. The findings of the present study may give probable clues for the differential clinical presentation of Delta and Omicron in terms of virulence and infectivity. Although, the present study has not included any in vitro experimental approach which gives opportunity for future researchers to reveal the mystery behind the differential viral properties.
References
Aksamentov I, Roemer C, Hodcroft EB, Neher RA. Nextclade: clade assignment, mutation calling and quality control for viral genomes. J Open Source Softw. 2021;6(67):3773. https://doi.org/10.21105/joss.03773.
Al-Karmalawy AA, Dahab MA, Metwaly AM, Elhady SS, Elkaeed EB, Eissa IH, Darwish KM. Molecular docking and dynamics simulation revealed the potential inhibitory activity of ACEIs against SARS-CoV-2 targeting the hACE2 receptor. Front Chem. 2021;9:6612. https://doi.org/10.3389/fchem.2021.661230.
Angeletti S, Benvenuto D, Bianchi M, Giovanetti M, Pascarella S, Ciccozzi M. COVID-2019: the role of the nsp2 and nsp3 in its pathogenesis. J Med Virol. 2020;92(6):584–8. https://doi.org/10.1002/jmv.25719.
Basu A, Sarkar A, Maulik U. Molecular docking study of potential phytochemicals and their effects on the complex of SARS-CoV2 spike protein and human ACE2. Sci Rep. 2020;10(1):17699. https://doi.org/10.1038/s41598-020-74715-4.
Benvenuto D, Angeletti S, Giovanetti M, Bianchi M, Pascarella S, Cauda R, Ciccozzi M, Cassone A. Evolutionary analysis of SARS-CoV-2: how mutation of non-structural protein 6 (NSP6) could affect viral autophagy. J Infect. 2020;81(1):e24–7. https://doi.org/10.1016/j.jinf.2020.03.058.
Burki TK. Omicron variant and booster COVID-19 vaccines. Lancet Respir Med. 2022;10(2):e17. https://doi.org/10.1016/S2213-2600(21)00559-2.
Cagliani R, Forni D, Clerici M, Sironi M. Computational inference of selection underlying the evolution of the novel coronavirus, severe acute respiratory syndrome coronavirus 2. J Virol. 2020;94(12):e00411-e420. https://doi.org/10.1128/JVI.00411-20.
Callaway E. The mutation that helps Delta spread like wildfire. Nature. 2021;596(7873):472–3. https://doi.org/10.1038/d41586-021-02275-2.
Chen Y, Liu Q, Guo D. Emerging coronaviruses: genome structure, replication, and pathogenesis. J Med Virol. 2020;92(10):2249. https://doi.org/10.1002/jmv.26234.
Cherian S, Potdar V, Jadhav S, Yadav P, Gupta N, Das M, Rakshit P, Singh S, Abraham P, Panda S, Team N. SARS-CoV-2 spike mutations, L452R, T478K, E484Q and P681R, in the second wave of COVID-19 in Maharashtra, India. Microorganisms. 2021;9(7):1542. https://doi.org/10.3390/microorganisms9071542.
Desta IT, Porter KA, Xia B, Kozakov D, Vajda S. Performance and Its limits in rigid body protein-protein docking. Structure. 2020;28(9):1071-1081.e3. https://doi.org/10.1016/j.str.2020.06.006.
Duffy S. Why are RNA virus mutation rates so damn high? PLoS Biol. 2018;16(8):e3000003. https://doi.org/10.1371/journal.pbio.3000003.
Emam M, Oweda M, Antunes A, El-Hadidi M. Positive selection as a key player for SARS-CoV-2 pathogenicity: insights into ORF1ab, S and E genes. Virus Res. 2021;302:198472. https://doi.org/10.1016/j.virusres.2021.
Focosi D, Quiroga R, McConnell S, Johnson MC, Casadevall A. Convergent evolution in SARS-CoV-2 spike creates a variant soup from which new COVID-19 waves emerge. Int J Mol Sci. 2023;24(3):2264. https://doi.org/10.3390/ijms24032264.
Forni D, Cagliani R, Mozzi A, Pozzoli U, Al-Daghri N, Clerici M, Sironi M. Extensive positive selection drives the evolution of nonstructural proteins in lineage C betacoronaviruses. J Virol. 2016;90(7):3627–39. https://doi.org/10.1128/JVI.02988-15.
Gordon DE, Jang GM, Bouhaddou M, Xu J, Obernier K, White KM, O’Meara MJ, Rezelj VV, et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature. 2020;583(7816):459–68. https://doi.org/10.1038/s41586-020-2286-9.
Han P, Li L, Liu S, Wang Q, Zhang D, Xu Z, Han P, Li X, Peng Q, Su C, Huang B, Li D, Zhang R, Tian M, Fu L, Gao Y, Zhao X, Liu K, Qi J, Gao GF, Wang P. Receptor binding and complex structures of human ACE2 to spike RBD from omicron and delta SARS-CoV-2. Cell. 2022;185(4):630-640.e10. https://doi.org/10.1016/j.cell.2022.01.001.
Harvey WT, Carabelli AM, Jackson B, Gupta RK, Thomson EC, Harrison EM, Ludden C, Reeve R, Rambaut A; COVID-19 Genomics UK (COG-UK) Consortium; Peacock SJ, Robertson DL. SARS-CoV-2 variants, spike mutations and immune escape. Nat Rev Microbiol. 2021;19(7):409–424. https://doi.org/10.1038/s41579-021-00573-0
Hossain A, Akter S, Rashid AA, Khair S, Alam ASMRU. Unique mutations in SARS-CoV-2 Omicron subvariants’ non-spike proteins: Potential impacts on viral pathogenesis and host immune evasion. Microb Pathog. 2022;170:105699. https://doi.org/10.1016/j.micpath.2022.105699.
Ito K, Piantham C, Nishiura H. Relative instantaneous reproduction number of Omicron SARS-CoV-2 variant with respect to the Delta variant in Denmark. J Med Virol. 2022;94(5):2265–8. https://doi.org/10.1002/jmv.27560.
Khandia R, Singhal S, Alqahtani T, Kamal MA, El-Shall NA, Nainu F, Desingu PA, Dhama K. Emergence of SARS-CoV-2 Omicron (B.1.1.529) variant, salient features, high global health concerns and strategies to counter it amid ongoing COVID-19 pandemic. Environ Res. 2022;209:112816. https://doi.org/10.1016/j.envres.2022.112816.
Kozakov D, Beglov D, Bohnuud T, Mottarella SE, Xia B, Hall DR, Vajda S. How good is automated protein docking? Proteins. 2013;81(12):2159–66. https://doi.org/10.1002/prot.24403.
Kozakov D, Hall DR, Xia B, Porter KA, Padhorny D, Yueh C, Beglov D, Vajda S. The ClusPro web server for protein-protein docking. Nat Protoc. 2017;12(2):255–78. https://doi.org/10.1038/nprot.2016.169.
Kumar S, Thambiraja TS, Karuppanan K, Subramaniam G. Omicron and Delta variant of SARS-CoV-2: a comparative computational study of spike protein. J Med Virol. 2022;94(4):1641–9. https://doi.org/10.1002/jmv.27526.
Low ZY, Zabidi NZ, Yip AJW, Puniyamurti A, Chow VTK, Lal SK. SARS-CoV-2 Non-structural proteins and their roles in host immune evasion. Viruses. 2022;14(9):1991. https://doi.org/10.3390/v14091991.
Mengist HM, Kombe AJK, Mekonnen D, Abebaw A, Getachew M, Jin T. Mutations of SARS-CoV-2 spike protein: Implications on immune evasion and vaccine-induced immunity. Semin Immunol. 2021;55:101533. https://doi.org/10.1016/j.smim.2021.101533.
Montazersaheb S, Hosseiniyan Khatibi SM, Hejazi MS, Tarhriz V, Farjami A, Ghasemian Sorbeni F, Farahzadi R, Ghasemnejad T. COVID-19 infection: an overview on cytokine storm and related interventions. Virol J. 2022;19(1):92. https://doi.org/10.1186/s12985-022-01814-1.
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ. AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem. 2009;30(16):2785–91. https://doi.org/10.1002/jcc.21256.
Mousavizadeh L, Ghasemi S. Genotype and phenotype of COVID-19: their roles in pathogenesis. J Microbiol Immunol Infect. 2021;54(2):159–63. https://doi.org/10.1016/j.jmii.2020.03.022.
Pachetti M, Marini B, Benedetti F, Giudici F, Mauro E, Storici P, Masciovecchio C, Angeletti S, Ciccozzi M, Gallo RC, Zella D, Ippodrino R. Emerging SARS-CoV-2 mutation hot spots include a novel RNA-dependent-RNA polymerase variant. J Transl Med. 2020;18(1):179. https://doi.org/10.1186/s12967-020-02344-6.
Piplani S, Singh PK, Winkler DA, Petrovsky N. In silico comparison of SARS-CoV-2 spike protein-ACE2 binding affinities across species and implications for virus origin. Sci Rep. 2021;11(1):13063. https://doi.org/10.1038/s41598-021-92388-5.
Pondé RAA. Physicochemical effect of the N501Y, E484K/Q, K417N/T, L452R and T478K mutations on the SARS-CoV-2 spike protein RBD and its influence on agent fitness and on attributes developed by emerging variants of concern. Virology. 2022;572:44–54. https://doi.org/10.1016/j.virol.2022.05.003.
Ragab D, Salah Eldin H, Taeimah M, Khattab R, Salem R. The COVID-19 cytokine storm; what we know so far. Front Immunol. 2020;11:1446. https://doi.org/10.3389/fimmu.2020.01446.
Rashid PMA, Salih GF. Molecular and computational analysis of spike protein of newly emerged omicron variant in comparison to the delta variant of SARS-CoV-2 in Iraq. Mol Biol Rep. 2022;49(8):7437–45. https://doi.org/10.1007/s11033-022-07545-4.
Rochman ND, Wolf YI, Faure G, Mutz P, Zhang F, Koonin EV. Ongoing global and regional adaptive evolution of SARS-CoV-2. Proc Natl Acad Sci USA. 2021;118(29):e2104241118. https://doi.org/10.1073/pnas.2104241118.
Roy U. Comparative structural analyses of selected spike protein-RBD mutations in SARS-CoV-2 lineages. Immunol Res. 2022;70(2):143–51. https://doi.org/10.1007/s12026-021-09250-z.
Sanches PRS, Charlie-Silva I, Braz HLB, Bittar C, Freitas Calmon M, Rahal P, Cilli EM. Recent advances in SARS-CoV-2 Spike protein and RBD mutations comparison between new variants Alpha (B.1.1.7, United Kingdom), Beta (B.1.351, South Africa), Gamma (P.1, Brazil) and Delta (B.1.617.2, India). J Virus Erad. 2021;7(3):100054. https://doi.org/10.1016/j.jve.2021.100054.
Seppälä E, Veneti L, Starrfelt J, Danielsen AS, Bragstad K, Hungnes O, Taxt AM, Watle SV, Meijerink H. Vaccine effectiveness against infection with the Delta (B.1.617.2) variant, Norway, April to August 2021. Euro Surveill. 2021;26(35):2100793. https://doi.org/10.2807/1560-7917.ES.2021.26.35.2100793.
Shah M, Woo HG. Omicron: a heavily mutated SARS-CoV-2 variant exhibits stronger binding to ACE2 and potently escapes approved COVID-19 therapeutic antibodies. Front Immunol. 2022;12:830527. https://doi.org/10.3389/fimmu.2021.830527.
Shu Y, McCauley J. GISAID: global initiative on sharing all influenza data—from vision to reality. Euro Surveill. 2017;22(13):30494. https://doi.org/10.2807/1560-7917.ES.2017.22.13.30494.
Singh D, Yi SV. On the origin and evolution of SARS-CoV-2. Exp Mol Med. 2021;53(4):537–47. https://doi.org/10.1038/s12276-021-00604-z.
Starr TN, Greaney AJ, Hilton SK, Ellis D, Crawford KHD, Dingens AS, Navarro MJ, Bowen JE, Tortorici MA, Walls AC, King NP, Veesler D, Bloom JD. Deep mutational scanning of SARS-CoV-2 receptor binding domain reveals constraints on folding and ACE2 binding. Cell. 2020;182(5):1295-1310.e20. https://doi.org/10.1016/j.cell.2020.08.012.
Tai W, He L, Zhang X, Pu J, Voronin D, Jiang S, Zhou Y, Du L. Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell Mol Immunol. 2020;17(6):613–20. https://doi.org/10.1038/s41423-020-0400-4.
Tang Y, Liu J, Zhang D, Xu Z, Ji J, Wen C. Cytokine storm in COVID-19: the current evidence and treatment strategies. Front Immunol. 2020;11:1708. https://doi.org/10.3389/fimmu.2020.01708.
Vajda S, Yueh C, Beglov D, Bohnuud T, Mottarella SE, Xia B, Hall DR, Kozakov D. New additions to the ClusPro server motivated by CAPRI. Proteins. 2017;85(3):435–44. https://doi.org/10.1002/prot.25219.
Vardhan S, Sahoo SK. In silico ADMET and molecular docking study on searching potential inhibitors from limonoids and triterpenoids for COVID-19. Comput Biol Med. 2020;124:103936. https://doi.org/10.1016/j.compbiomed.2020.103936.
Velazquez-Salinas L, Zarate S, Eberl S, Gladue DP, Novella I, Borca MV. Positive selection of ORF1ab, ORF3a, and ORF8 genes drives the early evolutionary trends of SARS-CoV-2 during the 2020 COVID-19 pandemic. Front Microbiol. 2020;11:550674. https://doi.org/10.3389/fmicb.2020.550674.
Wang Y, Liu M, Gao J. Enhanced receptor binding of SARS-CoV-2 through networks of hydrogen-bonding and hydrophobic interactions. Proc Natl Acad Sci USA. 2020;117(25):13967–74. https://doi.org/10.1073/pnas.2008209117.
Yadav R, Chaudhary JK, Jain N, Chaudhary PK, Khanra S, Dhamija P, Sharma A, Kumar A, Handu S. Role of structural and non-structural proteins and therapeutic targets of SARS-CoV-2 for COVID-19. Cells. 2021;10(4):821. https://doi.org/10.3390/cells10040821.
Yan W, Zheng Y, Zeng X, He B, Cheng W. Structural biology of SARS-CoV-2: open the door for novel therapies. Signal Transduct Target Ther. 2022;7(1):26. https://doi.org/10.1038/s41392-022-00884-5.
Zahradník J, Marciano S, Shemesh M, Zoler E, Harari D, Chiaravalli J, Meyer B, Rudich Y, Li C, Marton I, Dym O, Elad N, Lewis MG, Andersen H, Gagne M, Seder RA, Douek DC, Schreiber G. SARS-CoV-2 variant prediction and antiviral drug design are enabled by RBD in vitro evolution. Nat Microbiol. 2021;6(9):1188–98. https://doi.org/10.1038/s41564-021-00954-4.
Zhang L, Jackson CB, Mou H, Ojha A, Peng H, Quinlan BD, Rangarajan ES, Pan A, Vanderheiden A, Suthar MS, Li W, Izard T, Rader C, Farzan M, Choe H. SARS-CoV-2 spike-protein D614G mutation increases virion spike density and infectivity. Nat Commun. 2020;11(1):6013. https://doi.org/10.1038/s41467-020-19808-4.
Acknowledgements
We gratefully acknowledge Dr. Anindita Banerjee, NIBMG for sharing the research idea and valuable suggestions throughout the work.
Funding
JR would like to acknowledge University grant commission (UGC), Govt. of India for fellowship and AM would like to acknowledge Department of Science and Technology (DST-SERB), Govt. of India for start-up research grant (SRG/2021/001000).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Panja, A., Roy, J., Mazumder, A. et al. Divergent mutations of Delta and Omicron variants: key players behind differential viral attributes across the COVID-19 waves. VirusDis. 34, 307–320 (2023). https://doi.org/10.1007/s13337-023-00823-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13337-023-00823-0