Abstract
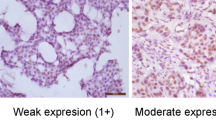
While p53 mutation is found in the majority of triple-negative breast cancer (TNBC) and despite recent developments in p53-targeting agents, their therapeutic application is still limited by the absence of standard biomarkers and ambiguousness of its essential biological role in cancer. Whole sections from 305 TNBC cases were stained for p53 to determine the correlation with lymph node metastasis and clinical outcomes in the whole cohort as well as in stratified patient groups according to AJCC stage and the use of adjuvant chemotherapy. Reduced immunohistochemical expression of p53 was an independent risk factor for lymph node metastasis. p53 overexpression was predictive of better clinical outcome in all patients (P = 0.012, disease-free survival and P = 0.008, overall survival) and the stratified cohorts of those who had early breast cancer and received adjuvant chemotherapy. Suppression of endogenous mutant p53 by siRNA and induction of wild-type p53 repressed TNBC cell invasion in vitro. In TNBC, increased immunohistochemical expression of p53 may reflect the accumulation of wild-type p53 rather than the mutant form. Strong p53 protein expression may serve as a favorable prognostic indicator and provide evidence for the use of specific agents targeting p53.



Similar content being viewed by others
References
Perou CM, Sorlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, et al. Molecular portraits of human breast tumours. Nature. 2000;406:747–52.
Kumar V, Abbas AK, Aster JC. Robbins and Cotran pathologic basis of disease. 9th ed. Philadelphia: Elsevier Saunders; 2014. p. 293–6.
Soussi T, Beroud C. Assessing TP53 status in human tumours to evaluate clinical outcome. Nat Rev Cancer. 2001;1:233–40.
Roger L, Jullien L, Gire V, Roux P. Gain of oncogenic function of p53 mutants regulates E-cadherin expression uncoupled from cell invasion in colon cancer cells. J Cell Sci. 2010;123:1295–305.
Muller PA, Vousden KH, Norman JC. p53 and its mutants in tumor cell migration and invasion. J Cell Biol. 2011;192:209–18.
Caulin C, Nguyen T, Lang GA, Goepfert TM, Brinkley BR, Cai WW, et al. An inducible mouse model for skin cancer reveals distinct roles for gain- and loss-of-function p53 mutations. J Clin Invest. 2007;117:1893–901.
Roger L, Gadea G, Roux P. Control of cell migration: a tumour suppressor function for p53? Biol Cell. 2006;98:141–52.
Muller PA, Caswell PT, Doyle B, Iwanicki MP, Tan EH, Karim S, et al. Mutant p53 drives invasion by promoting integrin recycling. Cell. 2009;139:1327–41.
Desmedt C, Voet T, Sotiriou C, Campbell PJ. Next-generation sequencing in breast cancer: first take home messages. Curr Opin Oncol. 2012;24:597–604.
Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature. 2012;490:61–70.
Duffy MJ, Synnott NC, McGowan PM, Crown J, O'Connor D, Gallagher WM. p53 as a target for the treatment of cancer. Cancer Treat Rev. 2014;40:1153–60.
Silwal-Pandit L, Vollan HK, Chin SF, Rueda OM, McKinney S, Osako T, et al. TP53 mutation spectrum in breast cancer is subtype specific and has distinct prognostic relevance. Clin Cancer Res. 2014;20:3569–80.
Hong B, van den Heuvel AP, Prabhu VV, Zhang S, El-Deiry WS. Targeting tumor suppressor p53 for cancer therapy: strategies, challenges and opportunities. Curr Drug Targets. 2014;15:80–9.
Maeda T, Nakanishi Y, Hirotani Y, Fuchinoue F, Enomoto K, Sakurai K, et al. Immunohistochemical co-expression status of cytokeratin 5/6, androgen receptor, and p53 as prognostic factors of adjuvant chemotherapy for triple negative breast cancer. Med Mol Morphol. 2015. doi:10.1007/s00795-015-0109-0.
Kashiwagi S, Yashiro M, Takashima T, Aomatsu N, Ikeda K, Ogawa Y, et al. Advantages of adjuvant chemotherapy for patients with triple-negative breast cancer at Stage II: usefulness of prognostic markers E-cadherin and Ki67. Breast Cancer Res. 2011;13:R122.
Piovan C, Palmieri D, Di Leva G, Braccioli L, Casalini P, Nuovo G, et al. Oncosuppressive role of p53-induced miR-205 in triple negative breast cancer. Mol Oncol. 2012;6:458–72.
Coates AS, Millar EK, O'Toole SA, Molloy TJ, Viale G, Goldhirsch A, et al. Prognostic interaction between expression of p53 and estrogen receptor in patients with node-negative breast cancer: results from IBCSG Trials VIII and IX. Breast Cancer Res. 2012;14:R143.
Lacroix M, Toillon RA, Leclercq G. p53 and breast cancer, an update. Endocr Relat Cancer. 2006;13:293–325.
Jasar D, Smichkoska S, Kubelka K, Filipovski V, Petrushevska G. Expression of p53 protein product in triple negative breast cancers and relation with clinical and histopathological parameters. Prilozi. 2015;36:69–79.
Neilsen PM, Noll JE, Mattiske S, Bracken CP, Gregory PA, Schulz RB, et al. Mutant p53 drives invasion in breast tumors through up-regulation of miR-155. Oncogene. 2013;32:2992–3000.
Ali A, Shah AS, Ahmad A. Gain-of-function of mutant p53: mutant p53 enhances cancer progression by inhibiting KLF17 expression in invasive breast carcinoma cells. Cancer Lett. 2014;354:87–96.
Arjonen A, Kaukonen R, Mattila E, Rouhi P, Hognas G, Sihto H, et al. Mutant p53-associated myosin-X upregulation promotes breast cancer invasion and metastasis. J Clin Invest. 2014;124:1069–82.
Noll JE, Jeffery J, Al-Ejeh F, Kumar R, Khanna KK, Callen DF, et al. Mutant p53 drives multinucleation and invasion through a process that is suppressed by ANKRD11. Oncogene. 2012;31:2836–48.
Adorno M, Cordenonsi M, Montagner M, Dupont S, Wong C, Hann B, et al. A Mutant-p53/Smad complex opposes p63 to empower TGFbeta-induced metastasis. Cell. 2009;137:87–98.
Boudreau HE, Casterline BW, Burke DJ, Leto TL. Wild-type and mutant p53 differentially regulate NADPH oxidase 4 in TGF-beta-mediated migration of human lung and breast epithelial cells. Br J Cancer. 2014;110:2569–82.
Romond EH, Perez EA, Bryant J, Suman VJ, Geyer Jr CE, Davidson NE, et al. Trastuzumab plus adjuvant chemotherapy for operable HER2-positive breast cancer. N Engl J Med. 2005;353:1673–84.
Osborne CK. Tamoxifen in the treatment of breast cancer. N Engl J Med. 1998;339:1609–18.
Elston CW, Ellis IO. Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. Histopathology. 1991;19:403–10.
Goldhirsch A, Wood WC, Coates AS, Gelber RD, Thurlimann B, Senn HJ. Strategies for subtypes—dealing with the diversity of breast cancer: highlights of the St. Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2011. Ann Oncol. 2011;22:1736–47.
Zlobec I, Steele R, Michel RP, Compton CC, Lugli A, Jass JR. Scoring of p53, VEGF, Bcl-2 and APAF-1 immunohistochemistry and interobserver reliability in colorectal cancer. Mod Pathol. 2006;19:1236–42.
Yamashita H, Toyama T, Nishio M, Ando Y, Hamaguchi M, Zhang Z, et al. p53 protein accumulation predicts resistance to endocrine therapy and decreased post-relapse survival in metastatic breast cancer. Breast Cancer Res. 2006;8:R48.
Kurshumliu F, Gashi-Luci L, Kadare S, Alimehmeti M, Gozalan U. Classification of patients with breast cancer according to Nottingham prognostic index highlights significant differences in immunohistochemical marker expression. World J Surg Oncol. 2014;12:243.
Mehta SA, Christopherson KW, Bhat-Nakshatri P, Goulet Jr RJ, Broxmeyer HE, Kopelovich L, et al. Negative regulation of chemokine receptor CXCR4 by tumor suppressor p53 in breast cancer cells: implications of p53 mutation or isoform expression on breast cancer cell invasion. Oncogene. 2007;26:3329–37.
Heinlein C, Krepulat F, Lohler J, Speidel D, Deppert W, Tolstonog GV. Mutant p53(R270H) gain of function phenotype in a mouse model for oncogene-induced mammary carcinogenesis. Int J Cancer. 2008;122:1701–9.
Gajdos C, Tartter PI, Bleiweiss IJ. Lymphatic invasion, tumor size, and age are independent predictors of axillary lymph node metastases in women with T1 breast cancers. Ann Surg. 1999;230:692–6.
Wu M, Wei W, Xiao X, Guo J, Xie X, Li L, et al. Expression of SIRT1 is associated with lymph node metastasis and poor prognosis in both operable triple-negative and non-triple-negative breast cancer. Med Oncol. 2012;29:3240–9.
Zhang J, Wang Y, Yin Q, Zhang W, Zhang T, Niu Y. An associated classification of triple negative breast cancer: the risk of relapse and the response to chemotherapy. Int J Clin Exp Pathol. 2013;6:1380–91.
Biganzoli E, Coradini D, Ambrogi F, Garibaldi JM, Lisboa P, Soria D, et al. p53 status identifies two subgroups of triple-negative breast cancers with distinct biological features. Jpn J Clin Oncol. 2011;41:172–9.
Wang J, Zhang C, Chen K, Tang H, Tang J, Song C, et al. ERbeta1 inversely correlates with PTEN/PI3K/AKT pathway and predicts a favorable prognosis in triple-negative breast cancer. Breast Cancer Res Treat. 2015;152:255–69.
Mahamodhossen YA, Liu W, Rong-Rong Z. Triple-negative breast cancer: new perspectives for novel therapies. Med Oncol. 2013;30:653.
Gossner W. A brief history of the Society for Histochemistry: its founders, its mission and the first 50 years. Histochem Cell Biol. 2002;118:91–4.
Maier T, Guell M, Serrano L. Correlation of mRNA and protein in complex biological samples. FEBS Lett. 2009;583:3966–73.
Vogel C, Marcotte EM. Insights into the regulation of protein abundance from proteomic and transcriptomic analyses. Nat Rev Genet. 2012;13:227–32.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
None
Rights and permissions
About this article
Cite this article
Jin, MS., Park, I.A., Kim, J.Y. et al. New insight on the biological role of p53 protein as a tumor suppressor: re-evaluation of its clinical significance in triple-negative breast cancer. Tumor Biol. 37, 11017–11024 (2016). https://doi.org/10.1007/s13277-016-4990-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-016-4990-5