Abstract
The analysis of cardiac activity is one of the most common elements for evaluating the state of a subject, either to control possible health risks, sports performance, stress levels, etc. This activity can be recorded using different techniques, with electrocardiogram and photoplethysmogram being the most common. Both techniques make significantly different waveforms, however the first derivative of the photoplethysmographic data produces a signal structurally similar to the electrocardiogram, so any technique focusing on detecting QRS complexes, and thus heartbeats in electrocardiogram, is potentially applicable to photoplethysmogram. In this paper, we develop a technique based on the wavelet transform and envelopes to detect heartbeats in both electrocardiogram and photoplethysmogram. The wavelet transform is used to enhance QRS complexes with respect to other signal elements, while the envelopes are used as an adaptive threshold to determine their temporal location. We compared our approach with three other techniques using electrocardiogram signals from the Physionet database and photoplethysmographic signals from the DEAP database. Our proposal showed better performances when compared to others. When the electrocardiographic signal was considered, the method had an accuracy greater than 99.94%, a true positive rate of 99.96%, and positive prediction value of 99.76%. When photoplethysmographic signals were investigated, an accuracy greater than 99.27%, a true positive rate of 99.98% and positive prediction value of 99.50% were obtained. These results indicate that our proposal can be adapted better to the recording technology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The analysis of cardiac activity is one of the most common elements for evaluating the state of a subject [1, 2], whether to control possible health risks [3], monitor sports performance [4], determine the level of stress [5], etc. There are different noninvasive methods for recording cardiac activity, of which two of the most common are the electrocardiogram (ECG) and the photoplethysmogram (PPG). The information about a subject’s state of health provided by the ECG and PPG is not comparable. While the ECG details the electrical activity of the heart, the PPG records variations in blood volume as a result of cardiac activity. Thus, the ECG provides greater diagnostic value than the PPG. In the ECG, the morphology of the QRS complex makes it possible to identify certain cardiac pathologies [6, 7], but above all, as it is an important element in the ECG signal, its location helps to determine the position of other waves and segments of the ECG, such as the ST, whose level above baseline is associated with cardiac ischemia and myocardial infarction [8, 9]. Also, the analysis of the variability of the heart rate (HR), better know as heart rate variability (HRV), has been successfully employed in a multitude of situations, such as revealing diabetic neuropathies [10], analyzing the growth and condition of a fetus [11], or evaluating the regulation of cardiac activity by the autonomic nervous system, which facilitates the detection of possible pathologies in the latter [12]. In [13], the PPG was compared with the ECG to analyze heart rate variability. The results showed that the data from both techniques were very similar, so that PPG may be a valid alternative. However, when determining which technology to use to analyze cardiac activity, one has to consider the purpose of the recording, whether the analysis can be performed with ECG and/or PPG, as well as possible limitations, such as sensitivity to movement of the PPG [14, 15], the use of electrodes for the ECG, and the comfort of the individual during data collection. On this last point, the PPG may be more comfortable with respect to the ECG, as it is found in a large number of commercial devices, such as smartwatches, with which subjects are more familiar [16].
The process for heartbeat detection in ECG data usually has a first preprocessing phase to eliminate artifacts, followed by a candidate selection process that is used in the last phase to determine the temporal location of the heartbeats. To reduce the artifacts, that make it difficult to process the ECG data, a bandpass filter or an adaptive multiresolution technique -like wavelet transform- is usually applied, selecting frequencies in the range between [5, 20] Hz, which are associated with the QRS complexes [17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36]. The second stage generates a new signal in which the QRS complexes are enhanced. These heartbeat improvements can be accomplished by applying two different procedures that we named as type-A or type-B processes. The type-A processes consist of a signal rectification (the absolute value or the square of the data) followed by a sliding window integrator [17, 18, 20, 24, 26, 32, 36,37,38], and sometimes the Shanon energy is set in an intermediate stage [18, 24, 26, 31, 38], where QRS is accented with respect to the remaining signal elements by concentrating the energy around them, which is calculated as \(-( d[n]^{2} log( d[n]^2 ) )\), where d[n] is the first derivative of the rescaled data between [0, 1]. As a result of the type-A process, the heartbeat appears as a kind of concave bell with a width approximately equal to the QRS-complex. The type-B processes are adaptive multiresolution techniques that decompose a signal into a set of signals with different frequency ranges associated with each one, so that temporal and frequency information is obtained simultaneously. Candidate selection is realized by choosing the levels to contain the frequency information associated with the QRS [20, 27, 39,40,41]. This way, type-A processes are simple and highly accurate, but sensitive to outliers, whereas type-B processes are more robust to outliers, but more complex. The final phase identifies the time points. This is typically accomplished by using fixed or adaptive thresholds. Hence, a heartbeat is associated with the time point at which these thresholds were exceeded [19, 20, 22, 25, 30, 33, 38, 42,43,44]. The accuracy of these techniques in determining the temporal location of the heartbeats is between 96.69% and 99.99%. Also, other authors use machine learning techniques on the preprocessed data to determine the location of the beats, eliminating the candidate selection phase [28, 45,46,47]. These studies have achieved an accuracy between 96.2% and 99.96% in locating heartbeats. Tables 1 and 2 summarize the main features of these studies. The databases ScopusFootnote 1 and IEEE XploreFootnote 2 were used for this purpose.
One of the most widely used algorithms to detect QRS complexes is the pan-tompkins (PT) [48]. It applies adaptive thresholds to the output resulting from integrating a 150 ms sliding window over the square of the derivative of a bandpass filter’s output. The accuracy of this technique is better than 99.3% [48]. Other techniques employ wavelet transform, envelopes and/or some classification technique, such as k-means, to detect QRS complexes [49, 50]. For example, in [49], envelope-based filtering is used to isolate QRS complexes, followed by a k-means classifier to differentiate between QRS and other spikes (hereafter, we refer to this technique as EK). The lower/upper envelopes are the result of a process that (1) locates the minima/maxima and (2) finds the interpolation curve passing through them [51]. This way, EK obtains the average of two lower envelopes twice to remove non-QRS-complex waves. In [50] is presented a modified version of PT, where the derivative of the bandpass filter output is replaced by the stationary wavelet transform, all other components being equal, so that it employs the same adaptive thresholding system (hereafter, we will refer to this technique with the acronym SWT). In this one, applying the wavelet transform (WT), data is decomposed into 2 levels using a Daubechies mother wavelet with vanishing moments of 3. Level 2 coefficients correspond to the frequency band of QRS complexes. In this way, both algorithms achieved an accuracy of better than 99.7%.
PPG signals are usually processed in 2 stages as shown in [52,53,54,55], where the accuracy was reported to be between 85% and 97.67%. In the first one, the signal noise is reduced by a 0.6–5 Hz bandpass or a median filter, while in the second one, the peaks from the smoothed PPG signal or its first derivative are located. Applying the first derivate to PPG (dPPG) [56] it is possible to obtain a signal with concave-bell-shaped waveform segments, similar to the QRS complex (Fig. 1), so techniques for detecting QRS in ECG signals are potentially applicable to PPG data. The techniques described above are mainly focused on ECG signals, so their performance with PPG signals has not been evaluated (no study from January 2017 to July 2022 has performed this analysis). In turn, the adaptive thresholds may present difficulties in adapting to an abrupt drop in amplitude, as occurs in the QRS or in the dPPG signal (Fig. 1), which may cause some QRS to go undetected. The abovementioned underlines the need to establish a process for determining the locations of heartbeats automatically and reliably in both recording techniques.
Recording using PPG measures in participant’s thumb (sampling rate of 128 Hz from DEAP dataset—Sect. 2.2). The signal of PPG is shown above, and its first derivative (dPPG) is shown below. The x-axes are set in samples
Methodology
In this paper, we present a technique to detect heartbeats in ECG and PPG signals, which is explained in detail in the next section, along with the databases and analysis performed.
Heartbeat detection using wavelet transform and envelopes
To locate heartbeats in ECG and dPPG signals, we propose a new technique based on maximal overlap discrete wavelet transform and upper envelops (WE) (Fig. 2). The maximal overlap discrete wavelet transform (MOWT) aims to select the peaks with the highest amplitude of the signal, while the QRS complex, and its position, is associated with the peak of the signal that exceeds the average of the envelopes. The envelopes result from locating all local maxima or minima and interpolating a signal passing through all of them [51]. If these peaks are maxima, the result is the upper envelope, while if they are minima, the lower envelope is obtained. Thus, the process is divided into 3 parts: (1) heartbeat enhancement using the MOWT, (2) calculation of the local maxima, (3) obtaining the envelopes and localization of the QRS complexes. The “Appedix A” shows the pseudocode of our proposal.
Summary of our proposal. Example of PPG signal processing with our proposal for a subject from DEAP dataset with sampling rate of 128 Hz and amplitude range of [− 262.144, + 262,144] mV. The x-axes are set in samples. a Schematic of the proposed technique. b The different phases of the algorithm: absolute value of the output of the MOWT ( \(\mid cQRS \mid\)), the selection of peaks and the resulting envelopes. On the left, the iterated process to obtain the envelopes, and on the right the envelopes and the QRS candidates. c The derivative of the PPG with the location of the QRS that provides the output of our proposal
Heartbeat enhancement
MOWT divides a signal into different components with temporal and frequency information, which makes it easy to detect from QRS complexes. Three fundamental aspects should be considered to select QRS complex candidates:
-
1.
The wavelet mother to be used. The more this resembles the QRS complex, the better the decomposition. For that reason, we selected the Symlets wavelet with 4 vanishing moments. This is a nearly symmetric variation of the Daubechies family of wavelet mother, and its resemblance to the QRS complex ensures a correct decomposition of the data.
-
2.
The number of components, N, into which the ECG signal is to be decomposed. Since the sampling rate may vary from one capture to another, the number of components is set as the value resulting from taking the integer part of the logarithm in base 2 of the sampling rate, that is, \(N=\lfloor log_2( F_s ) \rfloor\).
-
3.
The frequency range associated with QRS complexes is in the interval [5, 20] Hz. Thus, those levels of the MOWT decomposition that contain this interval will be chosen. The remaining ones are discarded.
After the previous steps, we proceed to calculate the modulus of the inverse transform of the MOWT, obtaining a signal with the possible QRS candidates, which we will call from now on \(\mid cQRS \mid\) (Fig. 2b).
Local maximums
The search for local maxima is performed on \(\mid cQRS \mid\). This process is iterated, with the aim of obtaining a signal that dynamically adapts to the temporal evolution of the QRS complexes and allows them to be detected (Fig. 2b). The steps are as follows:
-
1.
Let \(P_A\) be the set of the magnitudes of all local maxima present in \(\mid cQRS \mid\) and \(L_A\) be the set containing their temporal locations in the signal.
-
2.
Let \(P_C\) and \(L_C\) be the sets containing the local maxima of \(P_A\) and their time positions that are more than 300 ms apart. This is done by applying an iterated procedure consisting of choosing the highest peak among all the candidates and eliminating all those that are less than 300 ms away from it. This is then repeated for the highest of the remaining peaks, and reiterated until there are no candidates left.
-
3.
Let \(aux_{Pc}\) and \(aux_{Lc}\) be a copy of \(P_C\) and \(L_C\) respectively. With those copies, we proceed to execute the following procedure 3 times (Fig. 2b, left):
-
(a)
All elements in \(aux_{Pc}\) and \(aux_{Lc}\) are removed from \(P_A\) and \(L_A\).
-
(b)
A new signal, midSig, of equal length to \(\mid cQRS \mid\), resulting from interpolating all remaining local maxima in \(P_A\) is generated. This interpolation is based on a piecewise cubic Hermite interpolating polynomial.
-
(c)
Let \(P_{ms}\) and \(L_{ms}\) be all local maxima of midSig that are at least 300 ms apart (as explained above), discarding those peaks that are in \(aux_{Lc}\).
-
(d)
The maximums resulting from the previous step are added, in temporal order, to the variables \(aux_{Pc}\) and \(aux_{Lc}\).
-
(a)
After finishing the previous procedure, the variables \(P_C\), \(L_C\), \(P_{ms}\) and \(L_{ms}\) will store the peaks that will be used to generate the envelopes.
Envelopes
From the above peaks, we compute two upper envelopes: one centered on the variables \(P_C\) and \(L_C\), and another based on \(P_{ms}\) and \(L_{ms}\) and passing through the positions stored in \(L_C\). These are obtained by finding the local maxima of the variables \(P_C\) and \(P_{ms}\), and interpolating a signal passing through these points using a piecewise Hermite cubic interpolation polynomial (Fig. 2b, right). From the envelopes, the average of both is calculated, so that all \(P_C\) peaks above this average will be classified as a heartbeat.
Online implementation
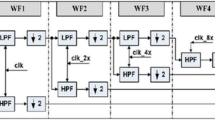
The implementation of our proposal to detect QRS complexes in real time requires the establishment of a data window as well as a shifting between windows (Fig. 3). To ensure that the window contains enough data, such that the envelopes match the variations of the QRS complex amplitudes, a length of 7.5 s and a shift of 750 ms have been set, so that if an individual has 40 beat per minutes (BPM), the window will contain at least 5 QRS complexes. The data buffer is divided into 3 areas: processed, active and shift. The process described above is applied to the entire window, so that only the QRS detected in the active zone will be taken into account. The processed and shift zones ensure that the envelopes are properly adjusted to the amplitudes of the QRS complexes. The length of the active and shift zones are identical, being set at 750 ms, ensuring that QRS is detected in subjects with a minimum of 40 BPM (1.5 s between beats), which minimizes the possibility of loss of complex detection.
Heartbeat database
The data used in this study were obtained from PhysioNet databases (PDB) [57,58,59,60,61] and DEAP dataset (DEAP) [62] (Table 3 shows the detail of the databases used). PDB contains signals from ECG whose selection was made based on [49]. In that article, the signals were randomly selected to analyze the effectiveness of the algorithm against different morphologies of QRS complexes. Specifically, three of the databases contained cardiographic recordings from subjects with various heart diseases, while the other two selected were from healthy subjects. In total, we analyzed 22 h and 11 min of data with expert annotations, which we used to compare the QRS detected by our proposal and the other techniques using an automated process. PDB provides the data with the power grid interference already removed. In DEAP, the cardiac activity was recorded using PPG, from which, using the dPPG, data resembling the ECG signals were obtained. A low-pass filter with a cutoff frequency of 30 Hz was applied to this signal, whose output was used as input to the different detectors. From the DEAP database, subjects 23 to 32 were discarded for having incorrect records. A total of 15 h and 24 min were analyzed manually, since the database lacked annotations with the locations of the heartbeats. Finally, it should be noted that no additional processing was applied to reduce noise, except for those already mentioned.
Evaluation and statistical analysis
Our proposal is compared with 3 techniques were described in Sect. “Introduction”: PT [48], EK [49], and SWT [50]. The first was selected because it is one of the most cited techniques; the second was used to select the ECG databases and uses envelope calculation as its main core; the third was selected because it uses the wavelet transform and is implemented in the py-ecg-detectorsFootnote 3 package for Python [63]. All the techniques were implemented, simulated and analyzed with Matlab version 9.12.0.1975300 (R2022a).
The analysis of our proposal assesses its effectiveness and efficiency, while comparing it with the three algorithms mentioned. To do so, we counted how many beats were correctly detected—true positive (TP), how many were incorrect—false positive (FP), and how many were not detected—false negative (FN). With this information, we calculated: (a) true positive rate (TPR), which measures the probability that a true QRS complex is correctly detected; (b) positive prediction value (PPV), which gives the probability that the positive results of the processing are true QRS complexes; (c) accuracy (ACC), which refers to the overall performance of the algorithm. The Eqs (1–3) show how these parameters are calculated. In addition, the Wilcoxon signed-rank test was applied to these three parameters to verify whether the differences of our proposal with respect to the others were due to random chance. For each technique, three vectors were obtained with the values of TPR, PPV and ACC. The p-value of the statistical test was obtained taking our proposal as a reference.
Results
The results obtained after applying the techniques to the databases are shown in the Table 4. For ECG data one can see that all the algorithms present very similar values, with PT, and our proposal, being the ones with the best results according to the three evaluated indicators (TPR, PPV and ACC are greater than \(>99.93\)%), although without significant statistical differences between all techniques, except with EK. The PT algorithm obtained a slightly higher TPR but lower PPV and ACC with respect to our proposal. On the other hand, the analysis of the data recorded by PPG shows that our technique was superior to the others, both in terms of accuracy, ability to detect QRS complexes and reliability (TPR, PPV and ACC are greater than \(>99.27\)%), with this difference being significant except for PPV with respect to the data from PT and SWT. Furthermore, statistical analysis of the combined PPG and ECG results also shows that our technique performs significantly better than the others.
Discussion
The proposed technique demonstrated a high level of accuracy in the detection of heartbeats for ECG and PPG signals, which implies more independence with respect to the method used for data recording. Both the overall results yielded from the offline version, as well as from its real-time implementation, were high (\(>99.66\)%). These values were higher than those obtained by the other algorithms with which they were compared. These differences were mainly due to the PPG, whose signal had first to be derived, and then have a 30 Hz low-pass filter applied, to obtain waves similar to the QRS complexes of the ECG. In this way, we were able to apply the same algorithms as for the localization of the QRS in ECG. However, the amplitudes of these complexes in the PPG showed greater variability with respect to the ECG data. This variability makes the EK technique performs worse performance in comparison with our proposal, since it uses lower envelopes to isolate the QRS from the remaining components of the ECG signal and other noises. Thus, if a QRS has variations in its peak, so that it presents local minima, the lower envelope process will cause the QRS amplitude to decrease or be eliminated. On the other hand, the PT and SWT algorithms employ the same adaptive thresholding process. This results in certain QRS complexes with a much smaller amplitude not being detected. Finally, in the SWT technique, in addition to the aforementioned thresholding problem, a higher occurrence of FP is added due to the oscillations resulting from applying the stationary wavelet transform. Thus, our proposal, which uses envelopes as adaptive thresholds to detect QRS complexes, shows greater accuracy by adapting appropriately to the temporal evolution of QRS amplitudes.
The TPR of the online version of our proposal, applied to the ECG, was slightly higher with respect to the offline version, while its PPV was slightly lower. These small differences were due to the fact that in the offline version, the envelopes were globally adjusted to the totality of the QRS, while in the online version the adjustment was by smaller sections which included a few complexes. This caused some QRS misclassified in the offline version to be detected in the online version, but in turn this caused the online version to generate more FP. In both implementations, some of these errors occurred at the beginning and end of the data vector, since the fit at these ends was not correct. A possible solution could be to check whether the amplitude of the first and last QRS complexes differ by 50% in amplitude with respect to the posterior and anterior, respectively, and discard them if the difference is greater than 50%. However, we believe that this is not a significant problem because it is located at the beginning and end of the data.
On the other hand, the offline version requires 2 parameters which are: (a) the sampling rate and (b) the distance in seconds between local maxima. The online version needs 2 additional parameters: (a) the buffer length and (b) the shifting length. Focusing on the sampling rate, the MOWT is used to select the frequency range of [5, 20] Hz. This assumes that the minimum sampling rate must be 40 Hz, and the minimum data time to have a spectral resolution of 5 Hz at this sampling rate is 200 ms. For there to be at least one heartbeat in this time, the HR must be at least 300 BPM, which implies ventricular fibrillation. Therefore, in general, the minimum data length needs to be greater than 200 ms, which ensures sufficient spectral resolution to select the frequency interval associated with the QRS complexes. With respect to the second parameter, the temporal distance of 300 ms between consecutive QRS candidate peaks is a factor that limits the applicability of our technique, since if the HR is greater than 200 BPM, some QRS complexes may not be detected. However, HR above this value are infrequent, around 35 per 100,000 patients per year [64], so we consider this limitation to be uncritical. Regarding the parameters of the online version, the proposed buffer length can be reduced if memory requirements so require. Taking as a reference the 200 BPM, a value well above the normal resting range [65], the shift, active and processed zones must have a duration of at least 300 ms for the buffer to contain at least 2 QRS complexes, so the window would be 900 ms. Likewise, to detect a minimum HR of 40 BPM, the buffer should be at least 3 s and the shifting 750 ms. Another factor affected by the buffer length is system delay in providing the location of the QRS complexes. For the values set in the Sect. 2.1.4, the time it takes to give the first data is 7.5 s, and then 1.5 s, which can be reduced if necessary as indicated above. Additionally, the PPG signal has an intrinsic delay of up to 250 ms due to its dependence on blood pressure [66], resulting in a higher variability that must be taken into account when used for monitoring individual. These values are acceptable in a multitude of situations such as day-to-day activities, physical exercise, hospital monitoring, etc.
As mentioned above, the PPG can be recorded on various areas of the body [67], so in certain situations where subject comfort is sought, and where noise due to movement is minimal, the PPG may be preferred over the ECG. Also, people tend to be more familiar with the idea of wearing a watch-like device [16], which could be a key factor in selecting which technology to use. On the other hand, the reuse of modules and code across manufacturers reduces the expense of product development [68]. As our proposal is highly reliable in both ECG and PPG signals, it enables the reuse of code and/or modules at manufacturers, reducing development cost. At the same time, it facilitates decision-making at the level of scientific and/or medical research, since the analysis of the variability of the HR is carried out with guarantees with both recording technologies.
Finally, our proposal presents a first stage for highlighting QRS complexes followed by an adaptive threshold, like most of the techniques contained in Sect. 1. However, the first stage does not seek to generate a concave bell that covers the QRS after rectification of the MOWT, but rather to obtain peaks that allow the threshold to be adjusted to the variation in QRS amplitudes and thus improve the localization of the QRS. Thus, the results obtained by our proposal are superior to the majority of the articles included in the Sect. 1 when used in ECG signals (Tables 1 and 2). They are located in the first quartile, so that only 4 of them have a higher accuracy [23, 30, 32, 45]. However, none of these reflect their effectiveness on data derived from the PPG signal, so we cannot establish a comparison with our proposal.
Conclusion
In this work we have described an algorithm applicable to ECG and PPG signals, with a high accuracy in detecting heartbeats due to the use of envelopes as an adaptive threshold, as these are better adjusted to the variability of the amplitude of QRS complexes. For offline version, the accuracy in detecting such QRS was higher than 99.94% for ECG and 99.42% for PPG, with a high probability of detecting true QRS complexes (TPR \(>99.8\)% for PPG and \(>99.96\)% for ECG) and a high reliability that the processing outputs were real QRS complexes (PPV \(>99.50\)% for PPG and \(>99.98\)% for ECG). On the other hand, the online version was slightly better at detecting QRS in ECG signals, but at the cost of a higher number of FP.
The analysis of cardiac activity is used in multiple applications: processes to detect pathologies, in the calculation of caloric expenditure, analysis of possible health risks during sports activities, in systems that seek to determine the emotional state of a subject, stress, cognitive load, etc. Therefore, the applicability of our proposal is evident, as it is flexible with respect to the data recording technique and highly accurate in the localization of QRS complexes.
References
Molina-Cantero AJ, Merino-Monge M, Castro-García JA, Pousada-García T, Valenzuela-Muñoz D, Gutiérrez-Párraga J, López-Álvarez S, Gómez-González IM (2021) A study on physical exercise and general mobility in people with cerebral palsy: Health through costless routines. Int J Environ Res Public Health. https://doi.org/10.3390/ijerph18179179
Castro-García JA, Molina-Cantero AJ, Gómez-González IM, Lafuente-Arroyo S, Merino-Monge M (2022) Towards human stress and activity recognition: a review and a first approach based on low-cost wearables. Electronics. https://doi.org/10.3390/electronics11010155
Carrarini C, Di Stefano V, Russo M, Dono F, Di Pietro M, Furia N, Onofrj M, Bonanni L, Faustino M, De Angelis MV (2022) ECG monitoring of post-stroke occurring arrhythmias: an observational study using 7-day Holter ECG. Sci Rep. https://doi.org/10.1038/s41598-021-04285-6
Marutani Y, Konda S, Ogasawara I, Yamasaki K, Yokoyama T, Maeshima E, Nakata K (2022) An experimental feasibility study evaluating the adequacy of a sportswear-type wearable for recording exercise intensity. Sensors 22(7):2577. https://doi.org/10.3390/s22072577
Hjortskov N, Rissén D, Blangsted AK, Fallentin N, Lundberg U, Søgaard K (2004) The effect of mental stress on heart rate variability and blood pressure during computer work. Euro J Appl Physiol 92(1–2):84–89. https://doi.org/10.1007/s00421-004-1055-z
Parra JC, López A, Casado MT, Ramírez RM (2002) Wolff-Parkinson-White: a propósito de un caso. SEMERGEN Med Fam 28(1):44–46. https://doi.org/10.1016/s1138-3593(02)74403-1
Wesley K (2017) Huszar’s ECG and 12-lead interpretation. Elsevier
Järvenpää J, Oikarinen L, Korhonen P, Väänänen H, Toivonen L, Viitasalo M (2007) Changing capacity of electrocardiographic ventricular repolarization in post-myocardial infarction patients with and without nonfatal cardiac arrest. Am J Cardiol 99(3):295–299. https://doi.org/10.1016/j.amjcard.2006.08.027
Korhonen P, Husa T, Konttila T, Tierala I, Mäkijärvi M, Väänänen H, Toivonen L (2009) Complex T-wave morphology in body surface potential mapping in prediction of arrhythmic events in patients with acute myocardial infarction and cardiac dysfunction. Europace 11(4):514–520. https://doi.org/10.1093/europace/eup051
Ceriello A, Prattichizzo F (2021) Variability of risk factors and diabetes complications Cardiovasc Diabetol. https://doi.org/10.1186/s12933-021-01289-4
Cerritelli F, Frasch MG, Antonelli MC, Viglione C, Vecchi S, Chiera M, Manzotti A (2021) A review on the vagus nerve and autonomic nervous system during fetal development: searching for critical windows. Front Neurosci. https://doi.org/10.3389/fnins.2021.721605
Tulen JHM, Manin’t Veld AJ, van Steenis HG, Mechelse K (1991) Sleep patterns and blood pressure variability in patients with pure autonomic failure. Clin Auton Res 1(4):309–315. https://doi.org/10.1007/BF01819837
Lu G, Yang F, Taylor JA, Stein JF (2009) A comparison of photoplethysmography and ECG recording to analyse heart rate variability in healthy subjects. J Med Eng Technol 33(8):634–641. https://doi.org/10.3109/03091900903150998
Allen J (2007) Photoplethysmography and its application in clinical physiological measurement. Physiol Meas. https://doi.org/10.1088/0967-3334/28/3/R01
Fine J, Branan KL, Rodriguez AJ, Boonya-Ananta T, Ajmal Ramella-Roman JC, McShane MJ, Coté GL (2021) Sources of inaccuracy in photoplethysmography for continuous cardiovascular monitoring. Biosensors (Basel). https://doi.org/10.3390/bios11040126
Jeng MY, Yeh TM, Pai FY (2020) Analyzing older adults’ perceived values of using smart bracelets by means-end chain. Healthcare (Switzerland). https://doi.org/10.3390/healthcare8040494
Khadirnaikar S, Aparna P (2017) A feasible QRS detection algorithm for arrhythmia diagnosis. 2016 International conference on advances in electrical, electronic and systems engineering. ICAEES, pp 32–37
Nayak C, Saha SK, Kar R, Mandal D (2018) Automated QRS complex detection using MFO-based DFOD. IET Signal Process 12(9):1172–1184. https://doi.org/10.1049/iet-spr.2018.5230
Burguera A (2019) Fast QRS detection and ECG compression based on signal structural analysis. IEEE J Biomed Health Info 23(1):123–131. https://doi.org/10.1109/JBHI.2018.2792404
Razzaq Hussein EA, Hassooni AS, Al-Libawy H (2019) Detection of electrocardiogram QRS complex based on modified adaptive threshold. Int J Electric Comput Eng 9(5):3512–3521. https://doi.org/10.11591/ijece.v9i5.pp3512-3521
Kurniawan A, Yuniarno EM, Setijadi E, Yusuf M, Ketut Eddy Purnama I (2020) QVAT: QRS complex detection based on variance analysis and adaptive threshold for electrocardiogram signal. In: Proceedings - 2020 international seminar on intelligent technology and its application: humanification of reliable intelligent systems, ISITIA 2020, pp. 175–179. https://doi.org/10.1109/ISITIA49792.2020.9163784
Chen A, Zhang Y, Zhang M, Liu W, Chang S, Wang H, He J, Huang Q (2020) A real time QRS detection algorithm based on ET and PD controlled threshold strategy. Sensors (Switzerland) 20(14):1–15. https://doi.org/10.3390/s20144003
Avishek Paul SP, Nantu Das Mitra M (2022) Automated detection of cardinal points of ECG signal for feature extraction using a single median filter. J Inst Eng (India) Serie B. https://doi.org/10.1007/s40031-022-00754-2
Xu W, Du F (2022) A robust qrs complex detection method based on shannon energy envelope and hilbert transform. J Mech Med Biol https://doi.org/10.1142/S0219519422400139
Rahul J, Sora M, Sharma LD (2021) A novel and lightweight P, QRS, and T peaks detector using adaptive thresholding and template waveform. Comput Biol Med. https://doi.org/10.1016/j.compbiomed.2021.104307
Ma J, Wang X, Wu X, Zhao T, Li Q (2019) Design and implementation of a novel r-peak detection algorithm. In: Proceedings - 10th International conference on information technology in medicine and education, ITME 2019, pp. 110–114. https://doi.org/10.1109/ITME.2019.00035
Singh N, Deora P, Pradhan PM (2019) Simultaneously concentrated PSWF-based Synchrosqueezing S-transform and its application to R peak detection in ECG signal. In: 2019 28th IEEE international conference on robot and human interactive communication, RO-MAN 2019. https://doi.org/10.1109/RO-MAN46459.2019.8956391
Chen H, Maharatna K (2020) An Automatic R and T Peak detection method based on the combination of hierarchical clustering and discrete wavelet transform. IEEE J Biomed Health Info 24(10):2825–2832. https://doi.org/10.1109/JBHI.2020.2973982
Hadji S (2021) R wave localization from transformed electrocardiogram signal by EMD. Proc Int Conf Electric Comput Energy Technol (ICECET). https://doi.org/10.1109/ICECET52533.2021.9698708
Patrick CBG, Mercier P, Töreyin Hakan (2022) A stochastic resonance electrocardiogram enhancement algorithm for robust QRS detection. IEEE J Biomed Health Info. https://doi.org/10.1109/JBHI.2022.3178109
Rakshit M, Das S (2017) An efficient wavelet-based automated R-peaks detection method using Hilbert transform. Biocybern Biomed Eng 37(3):566–577. https://doi.org/10.1016/j.bbe.2017.02.002
Rodriguez VH, Medrano C, Plaza I (2018) A Real-Time QRS complex detector based on discrete wavelet transform and adaptive threshold as standalone application on ARM microcontrollers. In: 2018 international conference on biomedical engineering applications, ICBEA 2018 - Proceedings. https://doi.org/10.1109/ICBEA.2018.8471741
Hossain MB, Bashar SK, Walkey AJ, McManus DD, Chon KH (2019) An accurate QRS complex and P wave detection in ECG signals using complete ensemble empirical mode decomposition with adaptive noise approach. IEEE Access 7:128869–128880. https://doi.org/10.1109/ACCESS.2019.2939943
Liu W, Wang X, Gao H, Yang C, Li J, Liu C (2020) An octave convolution neural network-based QRS detector. In: International conference on sensing, measurement and data analytics in the era of artificial intelligence, ICSMD 2020 - Proceedings, pp. 413–418. https://doi.org/10.1109/ICSMD50554.2020.9261658
Banerjee S (2019) A first derivative based R-peak detection and DWT based beat delineation approach of single lead electrocardiogram signal. In: Proceedings of 2019 IEEE region 10 symposium, TENSYMP 2019, pp. 565–570. https://doi.org/10.1109/TENSYMP46218.2019.8971094
Modak S, Taha LY, Abdel-Raheem E (2020) Single channel QRS detection using wavelet and median denoising with adaptive multilevel thresholding. In: 2020 IEEE international symposium on signal processing and information technology, ISSPIT 2020. https://doi.org/10.1109/ISSPIT51521.2020.9408699
Jain S, Kumar A, Bajaj V (2018) QRS complex detection using cuckoo search optimization algorithm. In: Proceedings of the 2017 IEEE international conference on communication and signal processing, ICCSP, pp. 91–95. https://doi.org/10.1109/ICCSP.2017.8286606
Lee M, Park D, Dong SY, Youn I (2018) A novel R peak detection method for mobile environments. IEEE Access 6:51227–51237. https://doi.org/10.1109/ACCESS.2018.2867329
Rekik S, Ellouze N (2017) Enhanced and optimal algorithm for QRS detection. IRBM 38(1):56–61. https://doi.org/10.1016/j.irbm.2016.11.004
Guyot P, Voiriot P, Djermoune EH, Papelier S, Lessard C, Felices M, Bastogne T (2018) R-peak detection in holter ECG signals using non-negative matrix factorization. Comput Cardiol. https://doi.org/10.22489/CinC.2018.123
Bashar SK, Noh Y, Walkey AJ, Mcmanus DD, Chon KH (2019) VERB: VFCDM-based electrocardiogram reconstruction and beat detection algorithm. IEEE Access 7:13856–13866. https://doi.org/10.1109/ACCESS.2019.2894092
Sahoo S, Mohanty M, Behera S, Sabut SK (2017) ECG beat classification using empirical mode decomposition and mixture of features. J Med Eng Technol 41(8):652–661. https://doi.org/10.1080/03091902.2017.1394386
Bae TW, Kwon KK (2021) ECG PQRST complex detector and heart rate variability analysis using temporal characteristics of fiducial points. Biomed Signal Process Control. https://doi.org/10.1016/j.bspc.2020.102291
Modak S, Taha LY, Abdel-Raheem E (2021) A novel method of QRS detection using time and amplitude thresholds with statistical false peak elimination. IEEE Access 9:46079–46092. https://doi.org/10.1109/ACCESS.2021.3067179
Cai W, Hu D (2020) QRS complex detection using novel deep learning neural networks. IEEE Access 8:97082–97089. https://doi.org/10.1109/ACCESS.2020.2997473
Belkadi MA, Daamouche A, Melgani F (2021) A deep neural network approach to QRS detection using autoencoders. Exp Syst Appl. https://doi.org/10.1016/j.eswa.2021.115528
Liang X, Li L, Liu Y, Chen D, Wang X, Hu S, Wang J, Zhang H, Sun C, Liu C (2022) Ecg\_segnet: an ecg delineation model based on the encoder-decoder structure. Comput Biol Med 145:105445
Pan J, Tompkins WJ (1985) A real-time QRS detection algorithm. IEEE Trans Biomed Eng 32(3):230–236
Merino M, Gómez IM, Molina AJ (2015) Envelopment filter and k-means for the detection of qrs waveforms in electrocardiogram. Med Eng Phys 37(6):605–609. https://doi.org/10.1016/j.medengphy.2015.03.019
Kalidas V, Tamil L (2017) Real-time QRS detector using stationary wavelet transform for automated ECG analysis. In: Proceedings - 2017 IEEE 17th international conference on bioinformatics and bioengineering, BIBE, pp. 457–461. https://doi.org/10.1109/BIBE.2017.00-12
Huang NE, Shen SS (2014) Hilbert-Huang transform and its applications. World Sci 2014:1–10
Neha Kanawade R, Tewary S, Sardana HK (2019) Photoplethysmography based arrhythmia detection and classification. In: 2019 6th international conference on signal processing and integrated networks, SPIN 2019, pp. 944–948. https://doi.org/10.1109/SPIN.2019.8711737
Sriharini K, Kumanan R, Sivagaminathan S, Koushik V, L V (2020) Analysis of inter-beat intervals from wrist pulse signals. In: 2020 IEEE 17th india council international conference, INDICON 2020 . https://doi.org/10.1109/INDICON49873.2020.9342333
Tum HM (2021) Photoplethysmography (PPG) scheming system based on finite impulse response (FIR) filter design in biomedical applications. Int J Electric Electron Eng Telecommun 10(4):272–282. https://doi.org/10.18178/ijeetc.10.4.272-282
Blok S, Piek MA, Tulevski II, Somsen GA, Winter MM (2021) The accuracy of heartbeat detection using photoplethysmography technology in cardiac patients. J Electrocardiol 67:148–157. https://doi.org/10.1016/j.jelectrocard.2021.06.009
Elgendi M, Liang Y, Ward R (2018) Toward generating more diagnostic features from photoplethysmogram waveforms. Diseases 6(1):20. https://doi.org/10.3390/diseases6010020
Goldberger AL, Amaral LAN, Glass L, Hausdorff JM, Ivanov PC, Mark RG, Mietus JE, Moody GB, Peng CK, Stanley HE (2000) PhysioBank, PhysioToolkit, and PhysioNet: components of a new research resource for complex physiologic signals. Circulation 101(23):215–220. https://doi.org/10.1161/01.CIR.101.23.e215
Greenwald SD (1990) Improved detection and classification of arrhythmias in noise-corrupted electrocardiograms using contextual information. PhD thesis, Harvard University–MIT Division of Health Sciences and Technology (1990)
Taddei A, Distante G, Emdin M, Pisani P, Moody GB, Zeelenberg C, Marchesi C (1992) The European ST-T database: Standard for evaluating systems for the analysis of ST-T changes in ambulatory electrocardiography. European Heart Journal 13(9):1164–1172. https://doi.org/10.1093/oxfordjournals.eurheartj.a060332
Iyengar N, Peng CK, Morin R, Goldberger AL, Lipsitz LA (1996) Age-related alterations in the fractal scaling of cardiac interbeat interval dynamics. Am J Physiol Regul Integr Comp Physiol. https://doi.org/10.1152/ajpregu.1996.271.4.r1078
Moody GB, Mark RG (2001) The impact of the mit-bih arrhythmia database. IEEE Eng Med Biol Mag 20(3):45–50. https://doi.org/10.1109/51.932724
Koelstra S, Mühl C, Soleymani M, Lee JS, Yazdani A, Ebrahimi T, Pun T, Nijholt A, Patras I (2012) DEAP: a database for emotion analysis; using physiological signals. IEEE Trans Affective Comput 3(1):18–31. https://doi.org/10.1109/T-AFFC.2011.15
Howell L, Porr B (2018) High precision ECG Database with annotated R peaks, recorded and filmed under realistic conditions
Orejarena LA, Vidaillet H, Destefano F, Nordstrom DL, Vierkant RA, Smith PN, Hayes JJ (1998) Paroxysmal supraventricular tachycardia in the general population. J Am Coll Cardiol 31(1):150–157. https://doi.org/10.1016/S0735-1097(97)00422-1
Hernandez JFT (2010) Fisiología humana, 4e
Bennis FC, van Pul C, van den Bogaart JJL, Andriessen P, Kramer BW, Delhaas T (2019) Artifacts in pulse transit time measurements using standard patient monitoring equipment. PLoS ONE. https://doi.org/10.1371/journal.pone.0218784
Tamura T, Maeda Y, Sekine M, Yoshida M (2014) Wearable photoplethysmographic sensors-past and present. Electronics. https://doi.org/10.3390/electronics3020282
Hardung B, Kölzow T, Krüger A (2004) Reuse of software in distributed embedded automotive systems. In: EMSOFT 2004 - Fourth ACM international conference on embedded software, pp. 203–210. https://doi.org/10.1145/1017753.1017787
Acknowledgements
We want to thank Patrick Partridge, for the revision of the manuscript.
Funding
Funding for open access publishing: Universidad de Sevilla/CBUA. This research was funded by project AAI, Grant PID2019-104323RB-C32 funded by MCIN/AEI/10.13039/501100011033.
Author information
Authors and Affiliations
Contributions
All the authors contributed to the conception and design of the study. The material preparation, data collection, and analysis were performed by MMM. The first draft of the manuscript was written by MMM, and all the authors commented on the previous versions of the manuscript. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
This article does not include any studies with human participants or animals performed by any of the authors.
Appendix A
Appendix A
Pseudocode of our offline proposal
To improve the understanding of the technique developed in this paper, the pseudocode that implements the offline version of our proposal is shown below.

Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Merino-Monge, M., Castro-García, J.A., Lebrato-Vázquez, C. et al. Heartbeat detector from ECG and PPG signals based on wavelet transform and upper envelopes. Phys Eng Sci Med 46, 597–608 (2023). https://doi.org/10.1007/s13246-023-01235-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13246-023-01235-6