Abstract
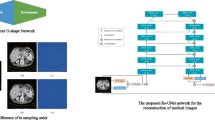
Digital subtraction angiography (DSA) is a powerful technique for visualizing blood vessels from X-ray images. However, the subtraction images obtained with this technique suffer from artifacts caused by patient motion. To avoid these artifacts, a new method called “Virtual DSA” is proposed, which generates DSA images directly from a single live image without using a mask image. The proposed Virtual DSA method was developed using the U-Net deep learning architecture. In the proposed method, a virtual DSA image only containing the extracted blood vessels was generated by inputting a single live image into U-Net. To extract the blood vessels more accurately, U-Net operates on each small area via a patch-based process. In addition, a different network was used for each zone to use the local information. The evaluation of the live images of the head confirmed accurate blood vessel extraction without artifacts in the virtual DSA image generated with the proposed method. In this study, the NMSE, PSNR, and SSIM indices were 8.58%, 33.86 dB, and 0.829, respectively. These results indicate that the proposed method can visualize blood vessels without motion artifacts from a single live image.







Similar content being viewed by others
References
Mistretta CA, Crummy AB, Strother CM (1981) Digital angiography: a perspective. Radiology 139(2):273–276. https://doi.org/10.1148/radiology.139.2.7012918
Brody WR (1982) Digital subtraction angiography. IEEE T Nucl Sci 29(3):1176–1180. https://doi.org/10.1109/TNS.1982.4336336
Takahashi M, Sato N, Fukui K, Kohrogi Y, Yamashita Y, Shinzato J, Higashida Y (1986) Hybrid digital subtraction angiography: Initial clinical experience. Computerized Radiology 10(4):147–154. https://doi.org/10.1016/0730-4862(86)90098-3
Yasuda M, Kato K, Sakiyama K, Uchiyama Y, Asanuma S, Fujimura K, Nakazawa Y (2010) Development of a prevention of body movement fixation appliance in leg digital subtraction angiography. Jpn J Radiol Technol 66(1):49–56. https://doi.org/10.6009/jjrt.66.49
Bale R, Lottersberger C, Vogele M, Prassl A, Czermak B, Dessl A, Jaschke W (2002) A novel vacuum device for extremity immobilisation during digital angiography: preliminary clinical experiences. Eur Radiol 12(12):2890–2894. https://doi.org/10.1007/s00330-002-1492-1
Zhang X, Zhang F, Li R (2010) DSA image registration based on 3D space-time detection. Procedia Eng 7:426–431. https://doi.org/10.1016/j.proeng.2010.11.070
Levin DC, Schapiro RM, Boxt LM, Dunham L, Harrington DP, Ergun DL (1984) Digital subtraction angiography: principles and pitfalls of image improvement techniques. AJR Am J Roentgenol 143(3):447–454. https://doi.org/10.2214/ajr.143.3.447
Pickens DR, Price RR, Erickson JJ, James AE Jr (1987) Digital image motion correction by spatial warp methods. Med Phys 14(1):56–61. https://doi.org/10.1118/1.596095
Oung H, Smith AM (1984) Real time motion detection in digital subtractive angiography. Proc SPIE 0515 Med Images Icons 515:336–339. https://doi.org/10.1117/12.964782
Meijering EH, Zuiderveld KJ, Viergever MA (1999) Image registration for digital subtraction angiography. Int J Comput Vision 31:227–246. https://doi.org/10.1023/A:1008074100927
Buzug TM, Weese J (1998) Image registration for DSA quality enhancement. Comput Med Imaging Graph 22(2):103–113. https://doi.org/10.1016/S0895-6111(98)00012-3
Liu B, Zhao Q, Dong J, Jia X, Yue Z (2013) A stretching transform-based automatic nonrigid registration system for cerebrovascular digital subtraction angiography images. Int J Imag Syst Tech 23(2):171–187. https://doi.org/10.1002/ima.22050
Lee S, Jeon CH, Sunwoo L, Oh DY, Lee KJ (2019) Phase-based nonrigid deformation for digital subtraction angiography. IEEE Access 7:32256–32265. https://doi.org/10.1109/ACCESS.2019.2902562
Hariharan SG, Kaethner C, Strobel N, Kowarschik M, DiNitto J, Fahrig R, Navab N (2019) Model-based motion artifact correction in digital subtraction angiography using optical-flow. Bildverarbeitung für die Medizin 2019:146–151. https://doi.org/10.1007/978-3-658-25326-4_31
Lo SB, Lou SA, Lin J, Freedman MT, Chien MV, Mun SK (1995) Artificial convolutional neural network techniques and applications for lung nodule detection. IEEE Trans Med Imaging 14(4):711–718. https://doi.org/10.1109/42.476112
Teramoto A, Fujita H, Yamamuro O, Tamaki T (2016) Automated detection of pulmonary nodules in PET/CT images: ensemble false-positive reduction using a convolutional neural network technique. Med Phys 43(6):2821–2827. https://doi.org/10.1118/1.4948498
Cha KH, Hadjiiski L, Samala R, Chan H, Caoili EM, Cohan RH (2016) Urinary bladder segmentation in CT urography using deep-learning convolutional neural network and level sets. Med Phys 43(4):1882–1896. https://doi.org/10.1118/1.4944498
Anthimopoulos M, Christodoulidis S, Ebner L, Christe A, Mougiakakou S (2016) Lung pattern classification for interstitial lung diseases using a deep convolutional neural network. IEEE Trans Med Imaging 35(5):1207–1216. https://doi.org/10.1109/TMI.2016.2535865
Pereira S, Pinto A, Alves V, Silva CA (2016) Brain tumor segmentation using convolutional neural networks in MRI images. IEEE Trans Med Imaging 35(5):1240–1251. https://doi.org/10.1109/TMI.2016.2538465
Dou Q, Chen H, Yu L, Zhao L, Qin J, Wang D, Mok VCT, Shi L, Heng P (2016) Automatic detection of cerebral microbleeds from MR images via 3D convolutional neural networks. IEEE Trans Med Imaging 35(5):1182–1195. https://doi.org/10.1109/TMI.2016.2528129
Samala RK, Chen H, Hadjiiski L, Helvie MA, Wei J, Cha K (2016) Mass detection in digital breast tomosynthesis: deep convolutional neural network with transfer learning from mammography. Med Phys 43(12):6654–6666. https://doi.org/10.1118/1.4967345
Miki Y, Muramatsu C, Hayashi T, Zhou X, Hara T, Katsumata A, Fujita H (2017) Classification of teeth in cone-beam CT using deep convolutional neural network. Comput Biol Med 80:24–29. https://doi.org/10.1016/j.compbiomed.2016.11.003
Ghafoorian M, Karssemeijier N, Heskes T, Bergkamp M, Wissink J, Obels J, Keizer K, de Leeuw F, van Ginneken B, Marchiori E, Platel B (2017) Deep multi-scale location-aware 3D convolutional neural networks for automated detection of lacunes of presumed vascular origin. Neuroimage-Clin 14:391–399. https://doi.org/10.1016/j.nicl.2017.01.033
Lakhani P, Sundaram B (2017) Deep learning at chest radiography: automated classification of pulmonary tuberculosis by using convolutional neural networks. Radiology 284(2):574–582. https://doi.org/10.1148/radiol.2017162326
Han X (2107) MR-based synthetic CT generation using a deep convolutional neural network method. Med Phys 44(4):1408–1419. Doi: https://doi.org/10.1002/mp.12155
Matsubara N, Teramoto A, Saito K, Fujita H (2020) Bone suppression for chest X-ray image using a convolutional neural filter. Phys Eng Sci Med 43:97–108. https://doi.org/10.1007/s13246-019-00822-w
Onishi Y, Teramoto A, Tsujimoto M, Tsukamoto T, Saito K, Toyama H, Fujita H (2019) Automated pulmonary nodule classification in computed tomography images using a deep convolutional neural network trained by generative adversarial networks. Biomed Res Int. https://doi.org/10.1155/2019/6051939
Ueda D, Yamamoto A, Nishimori M, Shimono T, Doishita S, Shimazaki A, Miki Y (2019) Deep learning for MR angiography: automated detection of cerebral aneurysms. Radiology 290(1):187–194. https://doi.org/10.1148/radiol.2018180901
Teramoto A, Yamada A, Kiriyama Y, Tsukamoto T, Yan K, Zhang L, Imaizumi K, Saito K, Fujita H (2019) Automated classification of benign and malignant cells from lung cytological images using deep convolutional neural network. Inf Med Unlocked 16:100205. https://doi.org/10.1016/j.imu.2019.100205
Onishi Y, Teramoto A, Tsujimoto M, Tsukamoto T, Saito K, Toyama H, Fujita H (2020) Multiplanar analysis for pulmonary nodule classification in CT images using deep convolutional neural network and generative adversarial networks. Int J Comput Assist Radiol Surg 15(1):173–178. https://doi.org/10.1007/s11548-019-02092-z
Teramoto A, Tsukamoto T, Yamada A, Kiriyama Y, Imaizumi K, Saito K, Fujita H (2020) Deep learning approach to classification of lung cytological images: Two-step training using actual and synthesized images by progressive growing of generative adversarial networks. PLoS One 15(3):e0229951. https://doi.org/10.1371/journal.pone.0229951
Zreik M, van Hamersvelt RW, Wolterink JM, Leiner T, Viergever MA, Išgum I (2018) A recurrent CNN for automatic detection and classification of coronary artery plaque and stenosis in coronary CT angiography. IEEE Trans Med Imaging 38(7):1588–1598. https://doi.org/10.1109/TMI.2018.2883807
Wolterink JM, van Hamersvelt RW, Viergever MA, Leiner T, Išgum I (2019) Coronary artery centerline extraction in cardiac CT angiography using a CNN-based orientation classifier. Med Image Anal 51:46–60. https://doi.org/10.1016/j.media.2018.10.005
Ronneberger O, Fischer P, Brox T (2015) U-net: Convolutional networks for biomedical image segmentation. Int Conf Med Image Comput Comput Assist Interv 9351:234–241. https://doi.org/10.1007/978-3-319-24574-4_28
Alom MZ, Hasan M, Yakopcic C, Taha TM, Asari VK (2018) Recurrent residual convolutional neural network based on u-net (r2u-net) for medical image segmentation. arXiv:1802.06955.
Huang Q, Sun J, Ding H, Wang X, Wang G (2018) Robust liver vessel extraction using 3D U-Net with variant dice loss function. Comput Biol Med 101:153–162. https://doi.org/10.1016/j.compbiomed.2018.08.018
Gao Y, Song Y, Yin X, Wu W, Zhang L, Chen Y, Shi W (2019) Deep learning-based digital subtraction angiography image generation. Int J Comput Assist Radiol Surg 14(10):1775–1784. https://doi.org/10.1007/s11548-019-02040-x
Goodfellow I, Pouget-Abadie J, Mirza M, Xu B, Warde-Farley D, Ozair S, Courville A, Bengio Y (2014) Generative adversarial nets. NIPS’14: Proceedings of the 27th International Conference on Neural Information Processing Systems 2:2672–2680.
Song T, Song Y, Wang Y, Huang X (2018) Residual network with dense block. J Electron Imaging 27(5):053036
Szegedy C, Ioffe S, Vanhoucke V, Alemi AA (2017) Inception-v4, inception-resnet and the impact of residual connections on learning. Proceedings of the Thirty-First AAAI Conference on Artificial Intelligence 4278–4284.
Iandola F, Moskewicz M, Karayev S, Girshick R, Darrell T, Keutzer K (2014) Densenet: implementing efficient convnet descriptor pyramids. arXiv:1404.1869.
Nair V, Hinton GE (2010) Rectified linear units improve restricted boltzmann machines. Proceedings of the 27th International Conference on Machine Learning (ICML-10) 807–814.
Ioffe S, Szegedy C (2015) Batch normalization: Accelerating deep network training by reducing internal covariate shift. Proceedings of the 32nd International Conference on Machine Learning 37:448–456. arXiv:1502.03167.
Wang Z, Bovik AC, Sheikh HR, Simoncelli EP (2004) Image quality assessment: from error visibility to structural similarity. IEEE Trans Image Process 13(4):600–612. https://doi.org/10.1109/TIP.2003.819861
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures in the studies involving human participants were performed in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. This article does not involve any studies performed on animals by any of the authors.
Informed consent
Consent from the patients was obtained with the condition that all data are anonymized.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kimura, R., Teramoto, A., Ohno, T. et al. Virtual digital subtraction angiography using multizone patch-based U-Net. Phys Eng Sci Med 43, 1305–1315 (2020). https://doi.org/10.1007/s13246-020-00933-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13246-020-00933-9