Abstract
Hutchinson-Gilford progeria syndrome (HGPS) and Werner syndrome (WS) are two of the best characterized human progeroid syndromes. HGPS is caused by a point mutation in lamin A (LMNA) gene, resulting in the production of a truncated protein product—progerin. WS is caused by mutations in WRN gene, encoding a loss-of-function RecQ DNA helicase. Here, by gene editing we created isogenic human embryonic stem cells (ESCs) with heterozygous (G608G/+) or homozygous (G608G/G608G) LMNA mutation and biallelic WRN knockout, for modeling HGPS and WS pathogenesis, respectively. While ESCs and endothelial cells (ECs) did not present any features of premature senescence, HGPS- and WS-mesenchymal stem cells (MSCs) showed aging-associated phenotypes with different kinetics. WS-MSCs had early-onset mild premature aging phenotypes while HGPS-MSCs exhibited late-onset acute premature aging characterisitcs. Taken together, our study compares and contrasts the distinct pathologies underpinning the two premature aging disorders, and provides reliable stem-cell based models to identify new therapeutic strategies for pathological and physiological aging.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Progeroid syndromes are heritable human disorders characterized by progeroid features that recapitulate typical features of normal aging. Among all the progeroid syndromes, Hutchinson-Gilford progeria syndrome (HGPS) and Werner syndrome (WS) are best characterized (Kudlow et al., 2007). HGPS is a sporadic autosomal dominant syndrome, and most HGPS patients were heterozygous for LMNA mutation (p.G608G/+). LMNA encodes A-type lamins that belongs to the family of nuclear lamina proteins, and a point mutation (p.G608G) in LMNA creates an aberrant splicing site in exon 11, resulting in the production of a truncated protein, progerin (Chojnowski et al., 2015; DeBoy et al., 2017; Luo et al., 2014). Another commonly seen progeroid syndrome is WS, caused by mutations in WRN gene that encodes a RecQ DNA helicase (Yu et al., 1996) important to DNA replication and DNA damage repair. Loss-of-function WRN leads to genomic instability, heterochromatin alterations, and cell growth defects, which contribute to WS pathogenesis (Li et al., 2016; Murfuni et al., 2012; Ren et al., 2017a; Ren et al., 2011; Seki et al., 2008; Shamanna et al., 2017; Zhang et al., 2015).
Both HGPS and WS patients present a wide range of aging-associated syndromes such as alopecia, lipodystrophy, osteoporosis and atherosclerosis. Studies on fibroblasts from HGPS and WS patients reveal features of accelerated cellular senescence and decreased proliferation potential (Brunauer and Kennedy, 2015; Chen et al., 2017; Cheung et al., 2014; Cheung et al., 2015; Kudlow et al., 2007; Liu et al., 2011a). Despite these common features, differences exist between HGPS and WS in the scope, intensity and duration of symptoms. For example, most patients with HGPS show symptoms resembling aspects of aging at a very early age and die at a median age from 11 to 13. By comparison, WS patients usually develop normally in the childhood and can live up to their fifties (Cox and Faragher, 2007; Ding and Shen, 2008; Hennekam, 2006; Kudlow et al., 2007; Mazereeuw-Hautier et al., 2007; Muftuoglu et al., 2008; Oshima et al., 2017).
In recent years, technologies based on stem cells and gene editing have been widely used to model various human diseases (Atchison et al., 2017; Duan et al., 2015; Fu et al., 2016; Liu et al., 2011a; Liu et al., 2012; Liu et al., 2014; Liu et al., 2011b; Lo Cicero and Nissan, 2015; Miller et al., 2013; Pan et al., 2016; Ren et al., 2017b; Wang et al., 2017; Yang et al., 2017; Zhang et al., 2015). Of note, HGPS-specific induced pluripotent stem cells (iPSCs) and WS-specific iPSCs and embryonic stem cells (ESCs) have been separately generated. Based on the findings by us and other groups, although the iPSCs and ESCs do not have any premature aging defects, mesenchymal stem cells (MSCs) and vascular smooth muscle cells (VSMCs) derived from these pluripotent stem cells display premature aging, consistent with the observations in fibroblasts from HGPS and WS patients (Chen et al., 2017; Cheung et al., 2014; Liu et al., 2011a; Miller et al., 2013; Zhang et al., 2011). Both being typical cases of progeroid syndromes, comparative analysis on HGPS and WS is very limited. More information about the similarities and differences in the pathological processes and molecular mechanisms of HGPS and WS remains to be uncovered via comparative studies.
Here, we successfully developed a reliable and isogenic platform for side-by-side investigation of HGPS and WS. Taking advantage of gene editing, we generated human ESCs harboring heterozygous LMNA p.G608G mutation and WRN deficiency, mimicking HGPS and WS, respectively. Notably, a genetically enhanced HGPS-specific ESCs bearing biallelic LMNA p.G608G mutation were also created. We found that HGPS- and WS-MSCs, but not ESCs or ECs, exhibited typical aging-associated characteristics. Interestingly, distinct aging kinetics were detected between HGPS- and WS-MSCs. For the first time, we achieved a contemporaneous comparison between HGPS and WS under the same genetic background to unravel the molecular and cellular differences, opening a window into the understanding of the pathology of human aging and providing a platform for screening for therapeutic strategies against aging-associated disorders.
Results
Generation of LMNA-mutated and WRN-deficient human ESCs
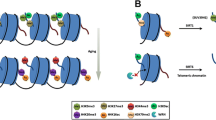
Using a genome-editing technique with a helper-dependent adenoviral vector (HDAdV), we generated heterozygous and homozygous LMNA-mutated human ESC lines (Fig. 1A). Combined with our previously reported WRN-deficient human ESCs (Zhang et al., 2015), we obtained ESCs with heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) LMNA mutation, and homozygous WRN deficiency (WRN−/−) under the same genetic background (Fig. 1B–D). All the three ESC lines displayed normal karyotypes and morphologies indistinguishable from those of WT-ESCs (Fig. 1B and 2A). All clones expressed the pluripotency markers OCT4, SOX2, NANOG, and were hypomethylated at the OCT4 promoter region (Fig. 1B and 2B). Each cell line was maintained for more than 30 passages without detectable growth abnormalities (data not shown) and was assessed for pluoripotency by differentiation into the three embryonic germ layers in vivo, by teratoma formation (Fig. 2C). Ki67 staining and cell cycle analysis also confirmed comparable proliferation potential of HGPS-ESCs and WS-ESCs with that of WT-ESCs (Fig. 2D and 2E). As expected, progerin was suppressed in both HGPS-ESCs and WS-ESCs (Fig. 1D). In addition, the levels of nuclear lamina component LAP2β, and heterochromatin markers H3K9me3 and HP1α were each normal in HGPS-ESCs and WS-ESCs compared to WT-ESCs (Fig. 2F and 2G). These data indicate that despite the progeroid-associated mutations, premature senescence phenotypes and chromosomal instability are well concealed in HGPS-ESCs and WS-ESCs at the pluripotent stage.
Generation of the heterozygous (LMNAG608G/+) and homozygous (LMNAG608G/G608G) ESCs. (A) Schematic representation of LMNA gene editing strategy by HDAdV-mediated homologous recombination. Blue triangles, FRT sites. (B) Morphology and immunofluorescence analysis of the pluripotency markers in WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− ESCs. Scale bar, 100 μm (left); 25 μm (right). (C) Confirmation of the heterozygous and homozygous mutation of LMNA by DNA sequencing. (D) Immunoblotting analysis of progerin and WRN expression in WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− ESCs. Progerin expression in homozygous (LMNAG608G/G608G) MSCs was carried out as a positive control
Characterization of HGPS-ESCs and WS-ESCs. (A) Karyotyping analysis of heterozygous (LMNAG608G/+) and homozygous (LMNAG608G/G608G) ESCs. (B) DNA methylation analysis of the OCT4 promoter region. (C) Immunostaining of representative markers of three germ layers in teratomas derived from heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− ESCs. Scale bar, 50 μm. (D) Ki67 immunostaining analysis of WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− ESCs. Scale bar, 25 μm. All cells were Ki67 positive. (E) Cell cycle analysis of ESCs. Data were presented as mean ± SEM, n = 3. (F) Representative immunofluorescence staining of LAP2β and HP1α in ESCs. Scale bar, 25 μm. All cells were LAP2β and HP1α positive. (G) Western blot analysis of LAP2β, HP1α and H3K9me3 expression in ESCs
HGPS-MSCs and WS-MSCs exhibit aging-associated phenotypes with different kinetics
Clinical observations in HGPS and WS patients indicate that premature aging disorders are often accompanied with defects in mesenchymal lineages, such as lipodystrophy, osteoporosis and atherosclerosis (Cox and Faragher, 2007). MSCs are adult stem cells originated from mesoderm and can be differentiated into osteocytes, chondrocytes, adipocytes and many other cell types (Lepperdinger, 2011; Marofi et al., 2017; Uccelli et al., 2008). We postulated that MSC exhaustion may play an important role in premature aging disorders. Here, HGPS-ESCs and WS-ESCs were differentiated into HGPS-MSCs and WS-MSCs. Both MSC lines expressed MSC-specific markers including CD90, CD73 and CD105 (Fig. 3A) and exhibited multiple-lineage differentiation potentials including adipogenesis, osteogenesis and chondrogenesis, though the differentiation ability of WS-MSCs towards adipocytes and osteoblasts was partly compromised (Fig. 3B–D).
Acquisition and characterization of HGPS-MSCs and WS-MSCs. (A) FACS analysis of MSC-specific markers (CD73, CD90, CD105) in WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− MSCs. (B) Left: characterization of adipogenesis potential of MSCs by Oil Red O staining. Right: Oil Red O positive areas were calculated by Image J. Data were presented as mean ± SEM, n = 3. *P < 0.05; ns, not significant. Scale bar, 100 μm. (C) Left: characterization of osteogenesis potential of MSCs by Von Kossa staining. Right: Von Kossa positive areas were calculated by Image J. Data were presented as mean ± SEM, n = 3. *P < 0.05; ns, not significant. Scale bar, 100 μm. (D) Left: characterization of chondrogenesis potential of MSCs by Toluidine Blue O staining. Right: the diameters of chondrocyte spheres were measured. Data were presented as mean ± SEM, n = 11. ns, not significant. Scale bar, 100 μm
Senescence-associated cellular changes were profiled in HGPS-MSCs and WS-MSCs at early and late passages. Population doubling curve indicated the early-onset senescence in WS-MSCs (Fig. 4A). By comparison, heterozygous (LMNAG608G/+) and homozygous (LMNAG608G/G608G) HGPS-MSCs grew at normal rate up to passage 6. Differences in cell cycle distribution were also observed between HGPS-MSCs and WS-MSCs (Fig. 4B). As previously described (Zhang et al., 2015), WS-MSCs exhibited cell cycle arrest at G2/M phase with decreased cell population at S phase as early as at passage 3, which later became more severe at passage 9 (Fig. 4B). By comparison, HGPS-MSCs did not show any defects until late passages, with even smaller cell population at S phase in homozygous MSCs than that in heterozygous MSCs (Fig. 4B). Consistent with the observations in growth curve and cell cycle analyses, the results of clonal expansion assay and SA-β-Gal staining also proved early-onset senescence in WS-MSCs (Fig. 4C and 4D). Interestingly, compared to the absence of progerin in HGPS-ESCs, differentiation into MSCs resulted in the re-expression of progerin at early passages and much more accumulation at late passages (Fig. 4E and 4F). With the accumulation of progerin, both heterozygous and homozygous MSCs displayed robust cell cycle arrest, proliferation defects and SA-β-Gal activity starting at passage 7, with more than 75% SA-β-Gal-positive MSCs at passage 11 (Fig. 4B–D). In addition, the doubled progerin levels (Fig. 4E–F) in homozygous HGPS-MSCs were correlated with faster kinetics of cellular senescence when compared to heterozygous HGPS-MSCs (Fig. 4A–F).
Phenotypic analyses of HGPS-MSCs and WS-MSCs indicate different kinetics between cell models of two different progeroid syndromes. (A) Growth curve showing the population doubling of MSCs, n = 3. (B) Cell cycle analysis of MSCs at passage 3 and passage 9. Data were presented as mean ± SEM, n = 3. (C) Analysis of clonal expansion abilities of WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− MSCs. Upper: representative images of crystal violet staining at passage 9. Lower: relative clonal expansion abilities at passage 3 and passage 9. Data were shown as mean ± SEM, n = 3. ***P < 0.001; ns, not significant. (D) Analysis of SA-β-Gal activity in WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− MSCs. Upper: representative images of SA-β-Gal staining at passage 9. Lower: frequency of SA-β-Gal positive cells. n = 3. (E) RT-qPCR analysis of progerin expression in WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− MSCs at passage 3 and passage 9. Data were shown as mean ± SEM, n = 3. ***P < 0.001; ns, not significant. (F) Western blot analysis of aging-related markers in WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− MSCs at passage 3 and passage 9. β-Tubulin were used as loading controls
Consistent with the defects in cell cycle progression and clonal expansion abilities, decrease in Ki67-positive cells was accompanied by misexpression of LAP2β and decreased expression of HP1α in WS-MSCs at passage 3 and further at passage 9 (Figs. 4F, 5A and 5B), indicative of impaired proliferation potential and heterochromatin disorganization since early passages. As for HGPS-MSCs, loss of Ki67-positive cells and misexpression of LAP2β were detected only at late passages in both homozygous and heterozygous HGPS-MSCs, with a even worse LAP2β defect in homozygous HGPS-MSCs (Figs. 4F and 5A).
Immunostaining of aging-related markers in HGPS-MSCs and WS-MSCs demonstrates different aging kinetics. (A) Left: representative immunostaining of LAP2β and Ki67 in WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− MSCs. Dashed lines indicate the nuclear boundaries and white arrows indicate abnormal nuclei. Scale bar, 10 μm. Right: percentages of LAP2β positive cells (upper) and Ki67 positive cells (lower) were shown as mean ± SEM, number of cells ≥ 300. ***P < 0.001; ns, not significant. (B) Left: representative immunostaining of progerin and HP1α in WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− MSCs. Dashed lines indicate the nuclear boundaries and white arrows indicate abnormal nuclei. Scale bar, 10 μm. Right: percentages of progerin positive cells (upper) and HP1α positive cells (lower) were shown as mean ± SEM, number of cells ≥ 300. ***P < 0.001; **P < 0.01; ns, not significant
Previous studies have reported that cells derived from HGPS and WS patients exhibit abnormal nuclear architecture (Adelfalk et al., 2005; Choi et al., 2011; De Sandre-Giovannoli et al., 2003; Eriksson et al., 2003; Goldman et al., 2004; Mallampalli et al., 2005; Saha et al., 2014; Scaffidi and Misteli, 2006; Toth et al., 2005; Verstraeten et al., 2008; Yang et al., 2005). Here, we also observed nuclear deformations in HGPS-MSCs and WS-MSCs (Figs. 5A, 5B, 6A and 6B). Increased number of cells with abnormal nuclear architecture was seen only in WS-MSCs at passage 3, but later in both WS-MSCs and HGPS-MSCs (Fig. 6A). In fact, there were even more cells with aberrant nuclear architecture in HGPS-MSCs, especially the homozygous ones, than WS-MSCs at passage 9, correlated with increased expression levels of progerin (Figs. 4E, 4F, 5B and 6A).
Immunofluorescence analysis of DNA damage response and nucleolar changes in HGPS-MSCs and WS-MSCs. (A) Left: representative immunostaining of γ-H2AX and 53BP1 in WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− MSCs. Dashed lines indicate the nuclear boundaries and white arrows indicate abnormal nuclei. Scale bar, 10 μm. Right: percentages of cells with aberrant nuclear architecture (upper) and γ-H2AX/53BP1 double-positive cells (lower) were shown as mean ± SEM, number of cells ≥ 300. ***P < 0.001; **P < 0.01; *P < 0.05; ns, not significant. (B) Left: representative immunostaining of Ki67 and nucleolin in WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− MSCs. Dashed lines indicate the nuclear boundaries and white arrows indicate abnormal nuclei. Scale bar, 10 μm. Right: percentages of cells with different numbers of nucleoli were shown as mean ± SEM, number of cells ≥ 300. Upper, passage 3; lower, passage 9
Having shown the distinct senescence-associated kinetics in HGPS-MSCs and WS-MSCs, we continued to evaluate other aging-related phenotypes. Increased DNA damage response is an important feature of aging (Brunauer and Kennedy, 2015; Burtner and Kennedy, 2010; Liu et al., 2005; Lopez-Otin et al., 2013; Mostoslavsky et al., 2006; Musich and Zou, 2011; Saha et al., 2014; Wang et al., 2009; Zhang et al., 2015). Here, increase in γ-H2AX and 53BP1 double-positive cells, indicative of increased DNA damage response, was observed only in WS-MSCs at passage 3 (Fig. 6A). At passage 9, both WS-MSCs and HGPS-MSCs exhibited increased DNA damage response, with the most observed in homozygous HGPS-MSCs (Fig. 6A). Increased size and decreased number of nucleoli can also serve as aging biomarkers (Buchwalter and Hetzer, 2017; Tiku et al., 2016). We observed that only WS-MSCs had fewer but larger nucleoli at early passages, and both WS-MSCs and HGPS-MSCs exhibited increased size and decreased numbers of nucleoli at late passages (Fig. 6B).
Taken together, these results suggest that HGPS-MSCs and WS-MSCs exhibit aging-associated phenotypes with different kinetics, and progerin exerts a dose-dependent effect on cellular senescence of HGPS-MSCs.
HGPS-ECs and WS-ECs do not exhibit phenotypes of accelerated senescence
Arterosclerosis have been observed in HGPS and WS patients, and progerin is widely present in the vascular cells including endothelial cells (Lo et al., 2014; McClintock et al., 2006; Miyamoto et al., 2014; Olive et al., 2010). As the inner layer of blood vessels, endothelial cells have unique functions in vascular biology, including barrier effect, vascular tone control, blood clotting regulation and inflammatory response (Bochenek et al., 2016; Hansen et al., 2017; Sturtzel, 2017). To explore whether LMNA mutation or WRN deficiency may cause aging-associated defects in endothelial cells (ECs), HGPS-ESCs and WS-ESCs were differentiated into HGPS-ECs and WS-ECs, respectively. CD31 and CD144 double-positive cells were sorted (Fig. 7A). All EC lines had typical endothelial morphology (Fig. 7B) and expressed endothelial-specific markers (Fig. 7C). Despite the expression of progerin in HGPS-ECs and the loss of WRN in WS-ECs (Fig. 7D), HGPS-ECs and WS-ECs were still able to form lattice-like vessel structures on matrigel and maintain normal lipid uptake capacities, nitric oxide (NO) synthesis abilities (Fig. 7F, 7G and 7H), proliferation potentials (Fig. 7E and 8A), as well as genomic stability (Fig. 8B and 8C). Therefore, LMNA mutation and WRN deficiency does not facilitate EC senescence, suggesting that the premature aging caused by progeria-associated mutations are cell-type-specific.
Acquisition and characterization of HGPS-ECs and WS-ECs. (A) CD31/CD144 positive cells were sorted as ECs by FACS. (B) Representative morphology of WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− ECs. Scale bar, 50 μm. (C) Immunostaining of EC-specific markers (CD31/vWF/CD144/eNOS) in WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− ECs. Scale bar, 50 μm. (D) Western blot analysis of WRN and progerin expression in WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− ECs. Actin was used as loading control. (E) Growth curve analysis showing the population doubling of ECs, n = 3. (F) The abilities of in vitro tube formation in WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− ECs. Cells were stained by Calcein-AM. (G) The uptake abilities of Dil-Ac-LDL in WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− ECs. Scale bar, 50 μm.(H) Measurement of Dil-Ac-LDL and nitric oxide (NO) by FACS in WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− ECs
HGPS-ECs and WS-ECs do not exhibit phenotypes of accelerated senescence. (A) Left: representative immunostaining of Lamin B1 and Ki67 in WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− ECs. Scale bar, 10 μm. Right: percentages of Ki67 positive cells and abnormal nuclei were shown as mean ± SEM, number of cells ≥ 300. ns, not significant. (B) Left: representative immunostaining of LAP2β and HP1α in WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− ECs. Scale bar, 10 μm. Right: percentages of LAP2β positive cells and HP1α positive cells were shown as mean ± SEM, number of cells ≥ 300. ns, not significant. (C) Left: representative immunostaining of γ-H2AX and 53BP1 in WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− ECs. Dashed lines indicate the nuclear boundaries. Scale bar, 10 μm. Right: percentages of γ-H2AX/53BP1 double-positive cells were shown as mean ± SEM, number of cells ≥ 300. ns, not significant
To be noted, both HGPS-ECs and WS-ECs were more apoptotic compared to WT-ECs at baseline, indicating impaired EC homeostasis (Fig. 9). Additionally, WS-ECs were more sensitive to TNF-α-induced apoptosis (Fig. 9). Thus, despite the absence of premature senescence, ECs bearing HGPS or WS-associated mutations demonstrated increased susceptibility to apoptosis.
Cellular apoptosis analysis in HGPS-ECs and WS-ECs. Left: cellular apoptosis analysis by FACS after treatment with vehicle or TNFα in WT, heterozygous (LMNAG608G/+), homozygous (LMNAG608G/G608G) and WRN−/− ECs. Right: percentages of apoptotic cells were presented as mean ± SEM, n = 3. ***P < 0.001; **P < 0.01; ns, not significant
Discussion
Human progeroid syndromes are characterized by typical features resembling normal aging, and therefore studies on progeroid disorders have provided important clues to understanding the molecular mechanisms underlying premature and normal aging (Burtner and Kennedy, 2010; Cao et al., 2011; Ding and Shen, 2008; Dreesen and Stewart, 2011; Kudlow et al., 2007; Miyamoto et al., 2014; Scaffidi and Misteli, 2006). As two of the best characterized progeroid syndromes, HGPS and WS have attracted a lot of attention during the last decade; related studies have been greatly conducive to our understanding of the pathology of these two disorders (Atchison et al., 2017; Chen et al., 2017; Cheung et al., 2014; De Sandre-Giovannoli et al., 2003; Ding and Shen, 2008; Egesipe et al., 2016; Kubben et al., 2016; Kudlow et al., 2007; Li et al., 2016; Liu et al., 2011a; Liu et al., 2011b; Lo Cicero and Nissan, 2015; Scaffidi and Misteli, 2006; Zhang et al., 2011; Zhang et al., 2015). However, there are no effective treatments so far and more information about the molecular pathology of these two premature aging syndromes are to be unveiled.
In this study, we generated LMNA-mutated and WRN-deficient human ESC lines with the same genetic background, making it possible to compare and contrast the cellular consequences of the genetic defects underlying HGPS and WS side-by-side. Similar to the iPSCs derived from the fibroblasts of HGPS and WS patients, HGPS- and WS-ESCs did not show any premature aging defects, indicating that pluripotent stem cells are able to conceal aging defects caused by LMNA mutation or WRN deficiency (Liu et al., 2012; Zhang et al., 2013). Upon mesenchymal differentiation, however, HGPS- and WS-MSCs exhibited aging-associated phenotypes that recapitulate those reported in fibroblasts and iPSC-derived MSCs from HGPS and WS patients (Cheung et al., 2014; Cheung et al., 2015; Compagnucci and Bertini, 2017; Zhang et al., 2011), with different kinetics. By measuring proliferation potential, SA-β-gal positivity, cell cycle, DNA damage response, and nuclear architecture, we showed that WS-MSCs had early-onset mild premature aging phenotypes while HGPS-MSCs exhibited late-onset acute premature aging characterisitics. To some extent, these dynamic features may mimic the patterns of disease progression of these two premature aging disorders (Fig. 10). To our knowledge, this is the first study evaluating the similarities and differences of HGPS- and WS-stem cells side by side. Our platform provides powerful tools to study aging by mimicking human genetic diseases in a petridish, facilitating the understanding of the pathology of different types of progeroid disorders and more importantly, making it possible for targeted high-throughput drug screening in human genetic background.
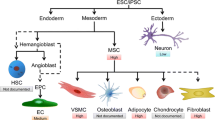
Schematic drawing of the major cellular phenotypes observed in HGPS- and WS-specific ESCs, ECs and MSCs. HGPS-ESCs and WS-ESCs were generated by gene editing under the same genetic background, and further differentiated to ECs and MSCs. LMNA-mutant and WRN-deficient ESCs or ECs show no accelerated senescence related defects, while HGPS-MSCs and WS-MSCs exhibited aging-associated phenotypes with different kinetics, including self-renewal ability, DNA damage response, nucleolar expansion, as well as nuclear architecture and heterochromatin alterations. WS-MSCs had early-onset mild premature aging phenotypes while HGPS-MSCs exhibited late-onset acute premature aging characterisitics
In addition, we observed that the homozygous HGPS-MSCs exhibited more severe aging phenotypes with a higher level of progerin than the heterozygous MSCs. Thus, the MSCs with homozygous or heterozygous LMNA mutation generated in our study also provide opportunities to investigate the role of progerin in a dose-dependent manner. Given the propriety of higher homogenicity in MSCs bearing homozygous LMNA mutation (e.g., expression of progerin), these cells may be particularly amenable to mechanistic studies using multi-omics techniques.
Different from HGPS-MSCs, HGPS-ECs did not display any premature senescence phenotypes, consistent with previous observations in HGPS-iPSC-derived ECs (Zhang et al., 2011). Similarly, WS-ECs did not show aging defects, either. These results indicate that the senescence-associated defects caused by LMNA mutation or WRN deficiency are cell-type-specific (Fig. 10). However, further analyses show that these cells were not otherwise normal; HGPS-ECs and WS-ECs were more apoptotic at baseline than WT-ECs. Moreover, WS-ECs, but not HGPS-ECs, exhibited a more pronounced response to inflammatory factor TNF-α, again indicating different molecular pathologies between the two progeroid syndromes.
MSCs and ECs as the outer and inner layers of blood vessels, respectively, play important roles in maintaining vascular homeostasis (Bochenek et al., 2016; Fang et al., 2010; Hansen et al., 2017; Hoshino et al., 2008; Kramann et al., 2016; Pasquinelli et al., 2007; Sturtzel, 2017; Wang et al., 2018). VSMCs, a cellular component of tunica media, have been proved defective in HGPS patients (Atchison et al., 2017; Chen et al., 2017; Compagnucci and Bertini, 2017; Gonzalo and Kreienkamp, 2015; Harhouri et al., 2017; Kinoshita et al., 2017; Liu et al., 2011a; Olive et al., 2010; Ragnauth et al., 2010; Vidak and Foisner, 2016; Zhang et al., 2011). Based on our data, it is reasonable to postulate that the exhaustion of MSC components in tunica adventitia may also be a common cause of accelerated aging defects in HGPS and WS patients. In addition, increased apoptosis of WS-ECs under inflammatory condition (e.g., TNF-α) may contribute to the vascular pathology in WS.
Therefore, we have generated in vitro models to compare and contrast the pathogenesis of HGPS and WS for the first time, providing high-throughput platforms to efficiently screen for effective treatments for both progeria syndromes and normal aging. In the future, it would be interesting to employ multi-omics technologies, including genomics, epigenomics, transcriptomics, proteomics and metabonomics, to unravel the molecular patterns of HGPS and WS under the same human genetic background, shedding light on the complex mechanisms underlying premature and normal aging and providing new evidence for the prevention and treatment of age-associated disorders.
Materials and Methods
Cell culture
WT-ESCs (Human H9 ESCs, WiCell Research) and three genetically edited ESCs were maintained on mitomycin C-inactivated mouse embryonic fibroblast (MEF) feeder in human ESC culture medium: 80% DMEM/F12 (Gibco), 20% Knockout Serum Replacement (Gibco), 0.1 mmol/L non-essential amino acids (NEAA, Gibco), 2 mmol/L GlutaMAX (Gibco), 55 μmol/L β-mercaptoethanol (Invitrogen), and 10 ng/mL FGF2 (Joint Protein Central); ESCs were also cultured on Matrigel (BD Biosciences) with mTeSR medium (STEMCELL Techonologies). All MSCs were cultured in MSC culture medium: 90% α-MEM + Glutmax (Gibco), 10% fetal bovine serum (FBS, Gemcell, Lot A77E01F), 1% penicillin/streptomycin (Gibco) and 1 ng/mL FGF2 (Joint Protein Central). ECs were cultured in EGM2 medium (Lonza).
Generation of LMNA G608 mutation knock-in ESCs
Helper-dependent adenoviral vector (HDAdV) for LMNA G608G knock-in was generated same as previous report with some modifications (Yang et al., 2017). In brief, exon 11 of LMNA gene was PCR-amplified from LNMA gene correction vector (LMNA-c-HDAdV) (Liu et al., 2011b) and subcloned into the pCR2.1-TOPO vector (Invitrogen). The G608G mutation at exon 11 was introduced using the GeneTailor Site-Directed Mutagenesis System (Invitrogen). The mutated exon 11 was replaced into LMNA-c-HDAdV (Liu et al., 2011b), and generated LMNA G608G knock-in HDAdV plasmid. The generated plasmids was packaged into HDAdV following previous report (Yang et al., 2017). To generate heterozygous LMNA G608G mutation knock-in ESCs, ESCs were infected with LMNA G608G knock-in HDAdV at MOI of 0.3–3 btu/cell and followed previous report (Yang et al., 2017). To generate homozygous LMNA G608G knock-in ESCs, we repeated 2nd round of mutation knock-in using the generated heterozygous LMNA G608G knock-in clones. Successful targeted knock-in events were verified by PCR amplification and DNA sequencing with the following primers: LMNA exon 11-F, 5′-TTGGGCCTGAGTGGTCAGTC-3′; LMNA exon 11-R, 5′-GACCCGCCTGCAGGATTTGG-3′.
Generation of ECs
Briefly, WT-ESCs and three genetically edited ESCs were plated on Matrigel in EC differentiation medium I (EC basal medium with 25 ng/mL BMP4, 3 μmol/L CHIR99021, 3 μmol/L IWP2 and 4 ng/mL FGF2) for 3 days. Differentiation medium II (EC basal medium with 50 ng/mL VEGF, 20 ng/mL FGF2,10 ng/mL IL-6) was used for another 3 days then CD31/CD144 double-positive cells were sorted by FACS.
Generation of MSCs
MSCs were differentiated as previously described (Duan et al., 2015; Fu et al., 2016; Wang et al., 2018; Zhang et al., 2015). Briefly, hESCs were dissociated into EBs and then were plated on Matrigel coated plates in MSC differentiation medium (α-MEM + GlutaMAX (Gibco), 10% FBS (Gemcell, Lot A77E01F), 1% penicillin/streptomycin (Gibco), 10 ng/mL FGF2 (Joint Protein Central) and 5 ng/mL TGFβ (HumanZyme)). About 10 days later, the confluent MSC-like cells were passaged on gelatin coated plate and cultured in MSC culture medium: 90% α-MEM + Glutmax (Gibco), 10% FBS (Gemcell, Lot A77E01F), 1% penicillin/streptomycin (Gibco) and 1 ng/mL FGF2 (Joint Protein Central). Then CD73/CD90/CD105 tripositive cells were sorted by FACS. MSCs were further differentiated towards adipocytes, osteoblasts, and chondrocytes to verify their multiple-lineage differentiation capacities (Pan et al., 2016; Zhang et al., 2015). Oil red O (adiopogenesis), Von Kossa (osteogenesis), and Toluidine blue (chondrogenesis) staining was performed respectively.
Bisulfite sequencing of the OCT4 promoter
Bisulfite treatment of DNA was carried out by using EZ DNA Methylation Kit (Zymo Research) following the manufacturer’s instructions. About 1 μg of genomic DNA was used. A genomic fragment of the OCT4 promoter was amplified using LA Taq Hot Start Version (TAKARA) as previously described. In brief, PCR products were purified by using gel extraction kit (Qiagen), and subsequently cloned into the pMD20 T vector (Transgen). 7 clones from each sample were sequenced with the universal primer M13.
Primers used for PCR: meF-OCT4, 5′-ATTTGTTTTTTGGGTAGTTAAAGGT-3′; meR-OCT4, 5′-CCAACTATCTTCATCTTAATAACATCC-3′.
Teratoma analysis
Teratoma assay was performed as described (Duan et al., 2015; Fu et al., 2016; Zhang et al., 2015). Briefly, 5 × 106 ESCs were administrated subcutaneously into NOD/SCID mice (male, 6–8 weeks). 6–12 weeks after injection, mice were killed and teratomas were analyzed by immunostaining. All animal experiments were conducted with approval of the Institute of Biophysics, Chinese Academy of Science.
Fluorescence-activated cell sorting (FACS)
MSCs or ECs were collected by using TrypLE Express (Gibco) and washed by PBS twice. Cells were incubated with primary antibody diluted with 10% FBS in PBS for 1 h at room temperature and then sorted by using a flow cytometer (BD FACSAria IIIu).
Antibodies used for cell sorting: anti-CD73 (550741), anti-CD90 (555595), anti-CD31 (555445), anti-CD144 (560410) antibodies were from BD Biosciences; anti-CD105 (17–1057) antibody was from eBioscience.
Analysis of cell cycle distribution
For cell cycle analysis, about 1 × 106 cells were processed with the Click-iT EdU Flow Cytometry Assay Kits (Invitrogen) according to the manufacturer’s instructions. In brief, the cells were harvested after 2 h incubation with EdU and stained with Alexa Fluor 647 dye azide and propidium iodide. Cells were examined by fluorescence-activated cell sorting (FACS) using a flow cytometer (BD LSRFortesa).
Clonal expansion assay
The single-cell clonal expansion assay was carried out as described (Duan et al., 2015). Briefly, 2,000 MSCs were seeded in a gelatin-coated 12-well plate. The relative cell density was then determined by Image J after crystal violet staining.
SA-β-Gal staining
SA-β-Gal staining was performed as described previously (Duan et al., 2015; Zhang et al., 2015). Briefly, cultured cells were washed in PBS and fixed at room temperature for 5 min in 2% formaldehyde and 0.2% glutaraldehyde. Fixed cells were stained with SA-β-Gal staining solution at 37°C overnight, percentage of SA-β-Gal positive cells were then calculated.
Measurement of cell apoptosis and nitric oxide (NO)
For cellular apoptosis analysis, cells were collected freshly and stained with Annexin V-EGFP Apoptosis Detection Kit (Vigorous Biotechnology), and then apoptotic cells were quantified by FACS. For NO detection, cells were treated with DAF-FM (Molecular Probes) for 30 min and quantified by FACS.
Dil-Ac-LDL uptake assay
In brief, ECs were collected after 6 h incubation with Dil-Ac-LDL (Molecular Probes) in EC culture medium. For FACS analysis, cells were collected by using TrypLE Express (Gibco) and measured by a flow cytometer (BD LSRFortesa). For immunofluorescence detection, cells were processed following the immunofluorescence microscopy protocol.
In vitro tube formation assay
Briefly, 5 × 104 cells were suspended in 500 μL EC medium and then seeded on Matrigel coated plate. After 6–8 h, lattice-like vessel structures formed and then cells were stained with Calcein-AM (Invitrogen) and examined by using fluorescence microscope (Olympus).
Western blotting
1 × 106 cells were lysed in 100 μL RIPA buffer [50 mmol/L Tris-HCl (pH = 7.5), 150 mmol/L NaCl, 1% NP-40, 0.5% sodium deoxycholate, 0.1% SDS] supplemented with NaF, NaVO4 and a protease-inhibitor mixture (Roche). Typically 20 μg of proteins were separated by SDS-PAGE, transferred to a PVDF membrane (Millipore), and blotted with one of the following primary antibodies and then HRP-conjugated secondary antibodies. The quantification of western blot was performed with Image Lab software for ChemiDoc XRS system (Bio-Rad).
Primary antibodies for western blotting include anti-WRN (Santa Cruz Biotechnology, Inc.), anti-Progerin (Santa Cruz Biotechnology, Inc.), anti-P21 (Cell Signaling Technology, Inc.), anti-LAP2β (BD Bioscience, Inc.), anti-HP1α (Cell Signaling Technology, Inc.), anti-Actin (Santa Cruz, Inc.), anti-Lamin B1 (Abcam, Inc.), anti-P16 (BD Bioscience, Inc.), anti-H3K9me3 (Abcam, Inc.), anti-β-Tubulin (Santa Cruz, Inc.).
RT-PCR
Total RNA was extracted by using TRIzol reagent (Invitrogen). 2 μg of RNA was converted to cDNA by using GoScript Reverse Transcription System (Promega), and 1/50 volume of the cDNA reaction was applied to PCR using primers for human GAPDH, Progerin. RT-qPCR was performed by using iTaq Universal SYBR Green Supermix (Bio-Rad).
Primers used for RT-PCR: GAPDH-F, 5′-TCGGAGTCAACGGATTTGGT-3′; GAPDH-R, 5′-TTGCCATGGGTGGAATCATA-3′; Progerin-F, 5′-ACTGCAGCAGCTCGGGG-3′; Progerin-R, 5′-TCTGGGGGCTCTGGGC-3′.
Immunofluorescence microscopy
Cells seeded on microscope coverslips were fixed with 4% formaldehyde in PBS for 30 min, permeabilized with 0.4% Triton X-100 in PBS for 25 min, and blocked with 10% donkey serum in PBS for 1 h. The coverslips were incubated with primary antibody (diluted with 1% donkey serum in PBS) overnight at 4°C and then incubated with fluorescence-labeled secondary antibody (diluted with 1% donkey serum in PBS at 1:500) at room temperature for 1 h. Hoechst 33342 (Invitrogen) was used to stain nuclear DNA.
Antibodies for immunofluorescence were purchased from the following companies. Abcam: anti-NANOG (ab21624), anti-Nucleolin (ab22758); ZSGB-Bio: anti-hSMA (ZM-0003); Sigma: anti-TUJ1 (T2200); Santa Cruz Biotechnology: anti-OCT4 (sc-5279), anti-SOX2 (sc-17320), anti-Progerin (sc-81611), anti-Lamin B (sc-6217); Cell Signaling Technology: anti-HP1α (2616), anti-FOXA2 (8186S), anti-CD144 (2158); Bethyl Laboratories: anti-53BP1 (A300-273A); Millipore: anti-γ-H2AX (05-636); BD Bioscience: anti-LAP2β (611000), anti-eNOS (610296); Vector: anti-Ki67 (VP-RM04); Dako: anti-vWF (A082).
Statistical analysis
Student’s t-test was used to analyse differences between different cell lines. Results were presented as mean ± SEM. P values < 0.05, P values < 0.01 and P values < 0.001 were considered statistically significant (*, **, ***).
References
Adelfalk C, Scherthan H, Hirsch-Kauffmann M, Schweiger M (2005) Nuclear deformation characterizes Werner syndrome cells. Cell Bbiol Int 29:1032–1037
Atchison L, Zhang H, Cao K, Truskey GA (2017) A tissue engineered blood vessel model of Hutchinson-Gilford progeria syndrome using human iPSC-derived smooth muscle cells. Sci Rep 7:8168
Bochenek ML, Schutz E, Schafer K (2016) Endothelial cell senescence and thrombosis: ageing clots. Thromb Res 147:36–45
Brunauer R, Kennedy BK (2015) Medicine. Progeria accelerates adult stem cell aging. Science 348:1093–1094
Buchwalter A, Hetzer MW (2017) Nucleolar expansion and elevated protein translation in premature aging. Nat Commun 8:328
Burtner CR, Kennedy BK (2010) Progeria syndromes and ageing: what is the connection? Nat Rev Mol Cell Biol 11:567–578
Cao K, Blair CD, Faddah DA, Kieckhaefer JE, Olive M, Erdos MR, Nabel EG, Collins FS (2011) Progerin and telomere dysfunction collaborate to trigger cellular senescence in normal human fibroblasts. J Clin Investig 121:2833–2844
Chen Z, Chang WY, Etheridge A, Strickfaden H, Jin Z, Palidwor G, Cho JH, Wang K, Kwon SY, Dore C et al (2017) Reprogramming progeria fibroblasts re-establishes a normal epigenetic landscape. Aging Cell 16:870–887
Cheung HH, Liu X, Canterel-Thouennon L, Li L, Edmonson C, Rennert OM (2014) Telomerase protects werner syndrome lineage-specific stem cells from premature aging. Stem Cell Rep 2:534–546
Cheung HH, Pei D, Chan WY (2015) Stem cell aging in adult progeria. Cell Regener 4:6
Choi S, Wang W, Ribeiro AJ, Kalinowski A, Gregg SQ, Opresko PL, Niedernhofer LJ, Rohde GK, Dahl KN (2011) Computational image analysis of nuclear morphology associated with various nuclear-specific aging disorders. Nucleus 2:570–579
Chojnowski A, Ong PF, Dreesen O (2015) Nuclear lamina remodelling and its implications for human disease. Cell Tissue Res 360:621–631
Compagnucci C, Bertini E (2017) The Potential of iPSCs for the treatment of premature aging disorders. Int J Mol Sci 18:2350
Cox LS, Faragher RG (2007) From old organisms to new molecules: integrative biology and therapeutic targets in accelerated human ageing. Cell Mol Life Sci: CMLS 64:2620–2641
De Sandre-Giovannoli A, Bernard R, Cau P, Navarro C, Amiel J, Boccaccio I, Lyonnet S, Stewart CL, Munnich A, Le Merrer M et al (2003) Lamin a truncation in Hutchinson-Gilford progeria. Science 300:2055
DeBoy E, Puttaraju M, Jailwala P (2017) Identification of novel RNA isoforms of LMNA. Nucleus 8:573–582
Ding SL, Shen CY (2008) Model of human aging: recent findings on Werner’s and Hutchinson-Gilford progeria syndromes. Clin Interv Aging 3:431–444
Dreesen O, Stewart CL (2011) Accelerated aging syndromes, are they relevant to normal human aging? Aging 3:889–895
Duan S, Yuan G, Liu X, Ren R, Li J, Zhang W, Wu J (2015) PTEN deficiency reprogrammes human neural stem cells towards a glioblastoma stem cell-like phenotype. Nat Commun 6:10068
Egesipe AL, Blondel S, Cicero AL, Jaskowiak AL, Navarro C, Sandre-Giovannoli A, Levy N, Peschanski M, Nissan X (2016) Metformin decreases progerin expression and alleviates pathological defects of Hutchinson-Gilford progeria syndrome cells. npj Aging Mech Dis 2:16026
Eriksson M, Brown WT, Gordon LB, Glynn MW, Singer J, Scott L, Erdos MR, Robbins CM, Moses TY, Berglund P et al (2003) Recurrent de novo point mutations in lamin A cause Hutchinson-Gilford progeria syndrome. Nature 423:293–298
Fang B, Li Y, Song Y, Li N (2010) Isolation and characterization of multipotent progenitor cells from the human fetal aorta wall. Exp Biol Med 235:130–138
Fu L, Xu X, Ren R, Wu J, Zhang W, Yang J, Ren X, Wang S, Zhao Y, Sun L et al (2016) Modeling xeroderma pigmentosum associated neurological pathologies with patients-derived iPSCs. Protein Cell 7:210–221
Goldman RD, Shumaker DK, Erdos MR, Eriksson M, Goldman AE, Gordon LB, Gruenbaum Y, Khuon S, Mendez M, Varga R et al (2004) Accumulation of mutant lamin A causes progressive changes in nuclear architecture in Hutchinson-Gilford progeria syndrome. Proc Natl Acad Sci USA 101:8963–8968
Gonzalo S, Kreienkamp R (2015) DNA repair defects and genome instability in Hutchinson-Gilford progeria syndrome. Curr Opin Cell Biol 34:75–83
Hansen NW, Hansen AJ, Sams A (2017) The endothelial border to health: mechanistic evidence of the hyperglycemic culprit of inflammatory disease acceleration. IUBMB Life 69:148–161
Harhouri K, Navarro C, Depetris D, Mattei MG, Nissan X, Cau P, De Sandre-Giovannoli A, Levy N (2017) MG132-induced progerin clearance is mediated by autophagy activation and splicing regulation. EMBO Mol Med 9:1294–1313
Hennekam RC (2006) Hutchinson-Gilford progeria syndrome: review of the phenotype. Am J Med Genet Part A 140:2603–2624
Hoshino A, Chiba H, Nagai K, Ishii G, Ochiai A (2008) Human vascular adventitial fibroblasts contain mesenchymal stem/progenitor cells. Biochem Biophys Res Commun 368:305–310
Kinoshita D, Nagasawa A, Shimizu I, Ito TK, Yoshida Y, Tsuchida M, Iwama A, Hayano T, Minamino T (2017) Progerin impairs vascular smooth muscle cell growth via the DNA damage response pathway. Oncotarget 8:34045–34056
Kramann R, Goettsch C, Wongboonsin J, Iwata H, Schneider RK, Kuppe C, Kaesler N, Chang-Panesso M, Machado FG, Gratwohl S et al (2016) Adventitial MSC-like cells are progenitors of vascular smooth muscle cells and drive vascular calcification in chronic kidney disease. Cell Stem Cell 19:628–642
Kubben N, Zhang W, Wang L, Voss TC, Yang J, Qu J, Liu GH, Misteli T (2016) Repression of the antioxidant NRF2 pathway in premature aging. Cell 165:1361–1374
Kudlow BA, Kennedy BK, Monnat RJ Jr (2007) Werner and Hutchinson-Gilford progeria syndromes: mechanistic basis of human progeroid diseases. Nat Rev Mol Cell Biol 8:394–404
Lepperdinger G (2011) Inflammation and mesenchymal stem cell aging. Curr Opin Immunol 23:518–524
Li Y, Zhang W, Chang L, Han Y, Sun L, Gong X, Tang H, Liu Z, Deng H, Ye Y et al (2016) Vitamin C alleviates aging defects in a stem cell model for Werner syndrome. Protein Cell 7:478–488
Liu B, Wang J, Chan KM, Tjia WM, Deng W, Guan X, Huang JD, Li KM, Chau PY, Chen DJ et al (2005) Genomic instability in laminopathy-based premature aging. Nat Med 11:780–785
Liu GH, Barkho BZ, Ruiz S, Diep D, Qu J, Yang SL, Panopoulos AD, Suzuki K, Kurian L, Walsh C et al (2011a) Recapitulation of premature ageing with iPSCs from Hutchinson-Gilford progeria syndrome. Nature 472:221–225
Liu GH, Ding Z, Izpisua Belmonte JC (2012) iPSC technology to study human aging and aging-related disorders. Curr Opin Cell Biol 24:765–774
Liu GH, Suzuki K, Li M, Qu J, Montserrat N, Tarantino C, Gu Y, Yi F, Xu X, Zhang W et al (2014) Modelling Fanconi anemia pathogenesis and therapeutics using integration-free patient-derived iPSCs. Nat Commun 5:4330
Liu GH, Suzuki K, Qu J, Sancho-Martinez I, Yi F, Li M, Kumar S, Nivet E, Kim J, Soligalla RD et al (2011b) Targeted gene correction of laminopathy-associated LMNA mutations in patient-specific iPSCs. Cell Stem Cell 8:688–694
Lo Cicero A, Nissan X (2015) Pluripotent stem cells to model Hutchinson-Gilford progeria syndrome (HGPS): current trends and future perspectives for drug discovery. Ageing Res Rev 24:343–348
Lo CY, Tjong YW, Ho JC, Siu CW, Cheung SY, Tang NL, Yu S, Tse HF, Yao X (2014) An upregulation in the expression of vanilloid transient potential channels 2 enhances hypotonicity-induced cytosolic Ca(2)(+) rise in human induced pluripotent stem cell model of Hutchinson-Gillford progeria. PLoS ONE 9:e87273
Lopez-Otin C, Blasco MA, Partridge L, Serrano M, Kroemer G (2013) The hallmarks of aging. Cell 153:1194–1217
Luo YB, Mastaglia FL, Wilton SD (2014) Normal and aberrant splicing of LMNA. J Med Genet 51:215–223
Mallampalli MP, Huyer G, Bendale P, Gelb MH, Michaelis S (2005) Inhibiting farnesylation reverses the nuclear morphology defect in a HeLa cell model for Hutchinson-Gilford progeria syndrome. Proc Natl Acad Sci USA 102:14416–14421
Marofi F, Vahedi G, Biglari A, Esmaeilzadeh A, Athari SS (2017) Mesenchymal stromal/stem cells: a new era in the cell-based targeted gene therapy of cancer. Front Immunol 8:1770
Mazereeuw-Hautier J, Wilson LC, Mohammed S, Smallwood D, Shackleton S, Atherton DJ, Harper JI (2007) Hutchinson-Gilford progeria syndrome: clinical findings in three patients carrying the G608G mutation in LMNA and review of the literature. Br J Dermatol 156:1308–1314
McClintock D, Gordon LB, Djabali K (2006) Hutchinson-Gilford progeria mutant lamin A primarily targets human vascular cells as detected by an anti-Lamin A G608G antibody. Proc Natl Acad Sci USA 103:2154–2159
Miller JD, Ganat YM, Kishinevsky S, Bowman RL, Liu B, Tu EY, Mandal PK, Vera E, Shim JW, Kriks S et al (2013) Human iPSC-based modeling of late-onset disease via progerin-induced aging. Cell Stem Cell 13:691–705
Miyamoto MI, Djabali K, Gordon LB (2014) Atherosclerosis in ancient humans, accelerated aging syndromes and normal aging: is lamin a protein a common link? Global Heart 9:211–218
Mostoslavsky R, Chua KF, Lombard DB, Pang WW, Fischer MR, Gellon L, Liu P, Mostoslavsky G, Franco S, Murphy MM et al (2006) Genomic instability and aging-like phenotype in the absence of mammalian SIRT6. Cell 124:315–329
Muftuoglu M, Oshima J, von Kobbe C, Cheng WH, Leistritz DF, Bohr VA (2008) The clinical characteristics of Werner syndrome: molecular and biochemical diagnosis. Hum Genet 124:369–377
Murfuni I, De Santis A, Federico M, Bignami M, Pichierri P, Franchitto A (2012) Perturbed replication induced genome wide or at common fragile sites is differently managed in the absence of WRN. Carcinogenesis 33:1655–1663
Musich PR, Zou Y (2011) DNA-damage accumulation and replicative arrest in Hutchinson-Gilford progeria syndrome. Biochem Soc Trans 39:1764–1769
Olive M, Harten I, Mitchell R, Beers JK, Djabali K, Cao K, Erdos MR, Blair C, Funke B, Smoot L et al (2010) Cardiovascular pathology in Hutchinson-Gilford progeria: correlation with the vascular pathology of aging. Arterioscler Thromb Vasc Biol 30:2301–2309
Oshima J, Sidorova JM, Monnat RJ Jr (2017) Werner syndrome: Clinical features, pathogenesis and potential therapeutic interventions. Ageing Res Rev 33:105–114
Pan H, Guan D, Liu X, Li J, Wang L, Wu J, Zhou J, Zhang W, Ren R, Zhang W et al (2016) SIRT6 safeguards human mesenchymal stem cells from oxidative stress by coactivating NRF2. Cell Res 26:190–205
Pasquinelli G, Tazzari PL, Vaselli C, Foroni L, Buzzi M, Storci G, Alviano F, Ricci F, Bonafe M, Orrico C et al (2007) Thoracic aortas from multiorgan donors are suitable for obtaining resident angiogenic mesenchymal stromal cells. Stem Cells 25:1627–1634
Ragnauth CD, Warren DT, Liu Y, McNair R, Tajsic T, Figg N, Shroff R, Skepper J, Shanahan CM (2010) Prelamin A acts to accelerate smooth muscle cell senescence and is a novel biomarker of human vascular aging. Circulation 121:2200–2210
Ren R, Deng L, Xue Y, Suzuki K, Zhang W, Yu Y, Wu J, Sun L, Gong X, Luan H et al (2017a) Visualization of aging-associated chromatin alterations with an engineered TALE system. Cell Res 27:483–504
Ren R, Ocampo A, Liu GH, Izpisua Belmonte JC (2017b) Regulation of stem cell aging by metabolism and epigenetics. Cell Metab 26:460–474
Ren X, Lim S, Ji Z, Yuh J, Peng V, Smith MT, Zhang L (2011) Comparison of proliferation and genomic instability responses to WRN silencing in hematopoietic HL60 and TK6 cells. PLoS ONE 6:e14546
Saha B, Cypro A, Martin GM, Oshima J (2014) Rapamycin decreases DNA damage accumulation and enhances cell growth of WRN-deficient human fibroblasts. Aging Cell 13:573–575
Scaffidi P, Misteli T (2006) Lamin A-dependent nuclear defects in human aging. Science 312:1059–1063
Seki M, Otsuki M, Ishii Y, Tada S, Enomoto T (2008) RecQ family helicases in genome stability: lessons from gene disruption studies in DT40 cells. Cell Cycle 7:2472–2478
Shamanna RA, Croteau DL, Lee JH, Bohr VA (2017) Recent advances in understanding werner syndrome. F1000Research 6:1779
Sturtzel C (2017) Endothelial cells. Advances in experimental medicine and biology. Can Med Assoc J 1003:71–91
Tiku V, Jain C, Raz Y (2016) Small nucleoli are a cellular hallmark of longevity. Nat Commun 8:16083
Toth JI, Yang SH, Qiao X, Beigneux AP, Gelb MH, Moulson CL, Miner JH, Young SG, Fong LG (2005) Blocking protein farnesyltransferase improves nuclear shape in fibroblasts from humans with progeroid syndromes. Proc Natl Acad Sci USA 102:12873–12878
Uccelli A, Moretta L, Pistoia V (2008) Mesenchymal stem cells in health and disease. Nat Rev Immunol 8:726–736
Verstraeten VL, Ji JY, Cummings KS, Lee RT, Lammerding J (2008) Increased mechanosensitivity and nuclear stiffness in Hutchinson-Gilford progeria cells: effects of farnesyltransferase inhibitors. Aging Cell 7:383–393
Vidak S, Foisner R (2016) Molecular insights into the premature aging disease progeria. Histochem Cell Biol 145:401–417
Wang C, Jurk D, Maddick M, Nelson G, Martin-Ruiz C, von Zglinicki T (2009) DNA damage response and cellular senescence in tissues of aging mice. Aging Cell 8:311–323
Wang L, Yi F, Fu L, Yang J, Wang S, Wang Z, Suzuki K, Sun L, Xu X, Yu Y et al (2017) CRISPR/Cas9-mediated targeted gene correction in amyotrophic lateral sclerosis patient iPSCs. Protein Cell 8:365–378
Wang S, Hu B, Ding Z, Dang Y, Wu J, Li D, Liu X, Xiao B, Zhang W, Ren R et al (2018) ATF6 safeguards organelle homeostasis and cellular aging in human mesenchymal stem cells. Cell Discov 4:2
Yang J, Li J, Suzuki K, Liu X, Wu J, Zhang W, Ren R, Zhang W, Chan P, Izpisua Belmonte JC et al (2017) Genetic enhancement in cultured human adult stem cells conferred by a single nucleotide recoding. Cell Res 27:1178–1181
Yang SH, Bergo MO, Toth JI, Qiao X, Hu Y, Sandoval S, Meta M, Bendale P, Gelb MH, Young SG et al (2005) Blocking protein farnesyltransferase improves nuclear blebbing in mouse fibroblasts with a targeted Hutchinson-Gilford progeria syndrome mutation. Proc Natl Acad Sci USA 102:10291–10296
Yu CE, Oshima J, Fu YH, Wijsman EM, Hisama F, Alisch R, Matthews S, Nakura J, Miki T, Ouais S et al (1996) Positional cloning of the Werner’s syndrome gene. Science 272:258–262
Zhang J, Lian Q, Zhu G, Zhou F, Sui L, Tan C, Mutalif RA, Navasankari R, Zhang Y, Tse HF et al (2011) A human iPSC model of Hutchinson Gilford progeria reveals vascular smooth muscle and mesenchymal stem cell defects. Cell Stem Cell 8:31–45
Zhang W, Li J, Suzuki K, Qu J, Wang P, Zhou J, Liu X, Ren R, Xu X, Ocampo A et al (2015) Aging stem cells. A Werner syndrome stem cell model unveils heterochromatin alterations as a driver of human aging. Science 348:1160–1163
Zhang W, Qu J, Suzuki K, Liu GH, Izpisua Belmonte JC (2013) Concealing cellular defects in pluripotent stem cells. Trends in cell biology 23:587–592
Acknowledgements
We are grateful to Lei Bai, Ruijun Bai, Mingming Wei and Shikun Ma for administrative assistance, to Hongyu Li, Xing Zhang, Zehua Wang and Pengze Yan for their technical assistance and to Junying Jia (IBP, CAS) and Shuang Sun (IBP, CAS) for their help in FACS. We thank Shuo Guo (IBP, CAS), Xinyi Wu (IBP, CAS) for breeding and management of laboratory animals and Lei Zhou (IBP, CAS) for providing veterinary care. This work was supported by the Strategic Priority Research Program of the Chinese Academy of Sciences (XDA16010100), the National Key R&D Program of China (2015CB964800, 2017YFA0103304, 2017YFA0102802, 2014CB910503), the National High Technology Research and Development Program of China (2015AA020307), CAS (KJZDEW-TZ-L05, and CXJJ-16M271), the National Natural Science Foundation of China (Grant Nos. 81625009, 81330008, 91749202, 91749123, 81371342, 81471414, 81422017, 81401159, 31671429, 81601233, 81671377, 31601109, 31601158, 81771515 and 81701388), Program of Beijing Municipal Science and Technology Commission (Z151100003915072), the Thousand Young Talents Program of China, and the State Key Laboratory of Stem Cell and Reproductive Biology (2016SRLab-KF13). Work in the laboratory of J.C.I.B. was supported by The Glenn Foundation, The Progeria Research Foundation, Universidad Catolica San Antonio de Murcia (UCAM) and Fundacion Dr. Pedro Guillen.
ABBREVIATIONS
ECs, endothelial cells; ESCs, embryonic stem cells; HGPS, Hutchinson-Gilford progeria syndrome; iPSCs, induced pluripotent stem cells; MSCs, mesenchymal stem cells; NO, nitric oxide; VSMCs, vascular smooth muscle cells; WS, Werner syndrome
COMPLIANCE WITH ETHICS GUIDELINES
Zeming Wu, Weiqi Zhang, Moshi Song, Wei Wang, Gang Wei, Wei Li, Jinghui Lei, Yu Huang, Yanmei Sang, Piu Chan, Chang Chen, Jing Qu, Keiichiro Suzuki, Juan Carlos Izpisua Belmonte and Guang-Hui Liu declare that they have no conflict of interest. This article does not contain any studies with human or animal subjects performed by the any of the authors.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Wu, Z., Zhang, W., Song, M. et al. Differential stem cell aging kinetics in Hutchinson-Gilford progeria syndrome and Werner syndrome. Protein Cell 9, 333–350 (2018). https://doi.org/10.1007/s13238-018-0517-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13238-018-0517-8