ABSTRACT
The lipid droplet (LD) is a unique multi-functional organelle that contains a neutral lipid core covered with a phospholipid monolayer membrane. The LDs have been found in almost all organisms from bacteria to humans with similar shape. Several conserved functions of LDs have been revealed by recent studies, including lipid metabolism and trafficking, as well as nucleic acid binding and protection. We summarized these findings and proposed a hypothesis that the LD is a conserved organelle.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
The lipid droplet (LD) is a multi-functional organelle with unique structure that distinguishes it from other cellular organelles (Murphy and Vance, 1999; Martin and Parton, 2006; Farese and Walther, 2009; Welte, 2015). Since discovered in 1674 by Van Leeuwenhoek, the LD has been found to be an organelle necessary for many cellular functions that are essential for the organismic energy homeostasis, and more importantly for human health and aging. In addition to its role in lipid storage and metabolism (Cao et al., 2008; Cohen et al., 2011), recent studies have revealed that the LD is critical for membrane trafficking (Liu et al., 2004; Bartz et al., 2007), protein storage (Li et al., 2012) and degradation (Ploegh, 2007), and has a key role in hepatitis C virus (HCV) replication and assembly (Miyanari et al., 2007) and neurodegeneration (Liu et al., 2015). As important sites of neutral lipid storage and metabolism, the ectopic storage of lipids in LDs is a key cellular component in many diseases. On other hand, LDs in plants and oleaginous microorganisms provide not only food oil but also feedstock for biodiesel and industrial oil (Murphy, 2001; Alvarez and Steinbuchel, 2002; Murphy, 2012; Chen et al., 2014).
LIPID DROPLETS EXIST FROM BACTERIA TO HUMANS
LDs are found in almost all organisms from bacteria to humans (Murphy, 2012; Waltermann et al., 2005). So far, except for knowing that LDs exist in all eukaryotic cells, it is also reported that some actinobacteria and cyanobacteria contain LDs, such as the genera Micromonospora, Dietzia, Nocardia, Rhodococcus, Mycobacterium, Gordonia, some streptomycetes (Murphy, 2001; Murphy, 2012), Nostoc punctiforme (Peramuna and Summers, 2014), Synechococcus lividus (Edwards et al., 1968), Anabaena variabilis (Wolk, 1973), and Synechocystis sp. PCC 6803 (Van de Meene et al., 2006). In addition, in comparison with other bacterial microcompartments including protein-based and lipid-bilayer membrane-based (Cornejo et al., 2014; Bobik et al., 2015), the LD is an unique organelle due to its particular structure and composition: neutral lipid core, phospholipid monolayer membrane, and peripheral proteins (Martin and Parton, 2006; Ding et al., 2012). This unique property is conserved from bacteria to humans.
THE STRUCTURE AND COMPOSITION OF LIPID DROPLETS ARE CONSERVED
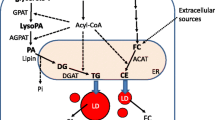
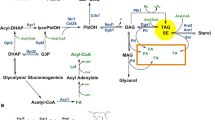
The core content of LDs in bacteria and eukaryotic cells is neutral lipid. Although some LDs contain retinyl ester (O’Mahony et al., 2015), polyhydroxyalkanoate or wax ester (Murphy, 2012), triacylglycerol (TAG) and cholesterol ester (CE) are the major neutral lipids of LDs in most cells (Waltermann and Steinbuchel, 2005; Barbosa and Siniossoglou, 2017). The neutral lipid core is coated by a phospholipid monolayer membrane in bacteria and eukaryotes (Martin and Parton, 2006; Farese and Walther, 2009; Waltermann and Steinbuchel, 2005), although the phospholipid composition may be different (Chitraju et al., 2012). In addition to the conserved lipid contents, the resident proteins of the organelle, including microorganism lipid droplet small (MLDS) and eukaryotic PERILIPIN (PLIN) family proteins (Kimmel et al., 2010), display conserved properties including the ability to target the phospholipid monolayer membrane and by the fact that they are all belong to apolipoprotein-like protein family (Yang et al., 2012).
These apolipoprotein-like proteins have also the ability to target LDs in diverse organisms, for example, mammalian LD proteins (PLINs) are targeted to LDs in yeast (Rowe et al., 2016) and bacteria (Hanisch et al., 2006). The LD resident proteins in C. elegans, DHS-3 and MDT-28/PLIN1 (Chughtai et al., 2015) behave similarly to target mammalian LDs (Na et al. 2015). In addition, a Drosophila LD resident protein, LSD1/PLIN1 localizes to LDs in C. elegans (Liu et al., 2014). The LD resident proteins, human adipose differentiation-related protein (ADRP)/PLIN2, C. elegans MDT-28, and bacterial MLDS are all able to bind to adiposomes that contain a TAG core with a phospholipid (DOPC) monolayer to mimic LDs in vitro (Wang et al., 2016). The ability of these proteins to target LDs of other organisms indicates that this fundamental process is highly conserved.
THE LIPID DROPLET IS A FUNCTIONALLY CONSERVED ORGANELLE FROM BACTERIA TO HUMANS
Several functions of LDs are common through bacteria to humans, such as lipid storage and metabolism. However, the study of other functions of LDs, especially in bacteria, is insufficient. Recently, we found that the LDs in a bacterium, Rhodococcus jostii RHA1 (RHA1), bind to genomic DNA (Fig. 1) (Zhang et al., 2017) and protect it via their major protein, MLDS, which promotes bacterial survival under stress (Zhang et al., 2017). Furthermore, the study also reports that LDs are involved in transcriptional regulation via a LD-associated transcriptional regulator, MLDSR (Zhang et al., 2017). These two newly identified functions in bacteria suggest that LDs are unique endomembrane organelles involved in nucleic acid handling and facilitate bacterial survival in and adaptation to extreme environments (Zhang et al., 2017).
The conserved lipid droplet functions of binding and regulating nucleic acids from bacterial to human cells. In bacteria (left), LDs bind and protect genomic DNA via the major LD-associated protein, MLDS, which enhances the survival and adaptation of bacteria in extreme environments. Furthermore, a LD-associated transcriptional regulator, MLDSR, whose gene is in the same operon as mlds, induces or reduces the expression of MLDS when its cytosolic concentration is low or high, respectively. LDs have key role in transcriptional regulation by recruiting MLDSR to control its cytosolic concentration. Similar functions of LDs are also found in mammalian cells. In liver cells, hepatitis C virus is assembled around the LD surface and viral RNA is located to LDs through NS5A and core proteins. A hypothesis is proposed that after replication of viral RNA on the ER membrane, the newly synthesized RNA is moved by NS5A to the core protein on LDs, which triggers the initial viral assembly (right, part 1). In adipocytes, moreover, a transcriptional factor NFAT5 can be sequestered to LDs by Fsp27, which prevents its nuclear importation to initiate transcription (right, part 2). Several histones such as H2A, H2B, and H2Av are localized to LDs via the anchor protein Jabba in Drosophila (right, part 3). In addition, LDs are also present in the liver cell nucleus (right, part 4). The facts that both bacterial and mammalian LDs possess the function of nucleic acid handling indicate that LDs in living cells on earth are evolutionary conserved from prokaryotes to humans
In eukaryotic and prokaryotic cells, LD proteomic analysis has revealed that RNA-binding proteins, ribosomal subunits, and/or translation factors are present on LDs (Ding et al., 2012; Sato et al., 2006; Zhang et al., 2012). Ribosomes and RNA are also found on mammalian LDs (Dvorak et al., 2003; Dvorak, 2005; Wan et al., 2007). In addition, HCV localizes and assembles around the LD surface (Fig. 1) (Miyanari et al., 2007; Shi et al., 2002; Gentzsch et al., 2013; Fiches et al., 2016). Furthermore, a mammalian homologue of the most abundant LD resident protein in C. elegans, MDT-28, is a mediator of RNA polymerase II (Zhang et al., 2012; Li et al., 2015). LDs in Drosophila store histones via the Jabba protein (Fig. 1) (Li et al., 2012, 2014; Cermelli et al., 2006). Interestingly, several recent studies identified LDs in the nuclei of mammalian cells (Fig. 1) (Layerenza et al. 1831; Wang et al., 2013; Ohsaki et al., 2016). LDs inhibit the translocation of NFAT5 to the nucleus via the LD-associated protein FSP27 and reduce NFAT5 transcriptional activity (Fig. 1) (Ueno et al., 2012). Altogether, these reports suggest that eukaryotic LDs partially mimic some nuclear functions, which is similar to bacterial LDs.
According to these previous studies, both bacterial and eukaryotic LDs are involved in nucleic acid handling, suggesting that the LD is a functionally conserved organelle. In the evolution from prokaryotes to eukaryotes, the most important feature is the protection of hereditary material (nuclear emergence). Thus, the function of bacterial LDs to protect and regulate nucleic acids indicates that they are analogous to the eukaryotic nuclear membrane.
Based on the extensive distribution, as well as the conservation of structure, composition, and functions of LDs from almost all living organisms, we propose a hypothesis that the LD is a conserved organelle from bacteria to humans (Fig. 1).
References
Alvarez HM, Steinbuchel A (2002) Triacylglycerols in prokaryotic microorganisms. Appl Microbiol Biotechnol 60:367–376
Barbosa AD, Siniossoglou S (2017) Function of lipid droplet-organelle interactions in lipid homeostasis. Biochimica et Biophysica Acta.
Bartz R, Zehmer JK, Zhu M, Chen Y, Serrero G, Zhao Y, Liu P (2007) Dynamic activity of lipid droplets: protein phosphorylation and GTP-mediated protein translocation. J Proteome Res 6:3256–3265
Bobik TA, Lehman BP, Yeates TO (2015) Bacterial microcompartments: widespread prokaryotic organelles for isolation and optimization of metabolic pathways. Mol Microbiol 98:193–207
Cao H, Gerhold K, Mayers JR, Wiest MM, Watkins SM, Hotamisligil GS (2008) Identification of a lipokine, a lipid hormone linking adipose tissue to systemic metabolism. Cell 134:933–944
Cermelli S, Guo Y, Gross SP, Welte MA (2006) The lipid-droplet proteome reveals that droplets are a protein-storage depot. Curr Biol 16:1783–1795
Chen Y, Ding YF, Yang L, Yu JH, Liu GM, Wang XM, Zhang SY, Yu D, Song L, Zhang HX, Zhang CY, Huo LH, Huo CX, Wang Y, Du YL, Zhang HN, Zhang P, Na HM, Xu SM, Zhu YX, Xie ZS, He T, Zhang Y, Wang GL, Fan ZH, Yang FQ, Liu HL, Wang XW, Zhang XG, Zhang MQ, Li YD, Steinbuchel A, Fujimoto T, Cichello S, Yu J, Liu PS (2014) Integrated omics study delineates the dynamics of lipid droplets in Rhodococcus opacus PD630. Nucleic Acids Res 42:1052–1064
Chitraju C, Trotzmuller M, Hartler J, Wolinski H, Thallinger GG, Lass A, Zechner R, Zimmermann R, Kofeler HC, Spener F (2012) Lipidomic analysis of lipid droplets from murine hepatocytes reveals distinct signatures for nutritional stress. J Lipid Res 53:2141–2152
Chughtai AA, Kassak F, Kostrouchova M, Novotny JP, Krause MW, Saudek V, Kostrouch Z, Kostrouchova M (2015) Perilipin-related protein regulates lipid metabolism in C. elegans. PeerJ 3:e1213
Cohen JC, Horton JD, Hobbs HH (2011) Human fatty liver disease: old questions and new insights. Science 332:1519–1523
Cornejo E, Abreu N, Komeili A (2014) Compartmentalization and organelle formation in bacteria. Curr Opin Cell Biol 26:132–138
Ding Y, Yang L, Zhang S, Wang Y, Du Y, Pu J, Peng G, Chen Y, Zhang H, Yu J, Hang H, Wu P, Yang F, Yang H, Steinbuchel A, Liu P (2012) Identification of the major functional proteins of prokaryotic lipid droplets. J Lipid Res 53:399–411
Dvorak AM (2005) Mast cell secretory granules and lipid bodies contain the necessary machinery important for the in situ synthesis of proteins. Chem Immunol Allergy 85:252–315
Dvorak AM, Morgan ES, Weller PF (2003) RNA is closely associated with human mast cell lipid bodies. Histol Histopathol 18:943–968
Edwards MR, Berns DS, Ghiorse WC, Holt SC (1968) Ultrastructure of the thermophilic blue-green alga, synechococcus lividus copeland(1). J Phycol 4:283–298
Farese RV, Walther TC (2009) Lipid droplets finally get a little R-E-S-P-E-C-T. Cell 139:855–860
Fiches GN, Eyre NS, Aloia AL, Van Der Hoek K, Betz-Stablein B, Luciani F, Chopra A, Beard MR (2016) HCV RNA traffic and association with NS5A in living cells. Virology 493:60–74
Gentzsch J, Brohm C, Steinmann E, Friesland M, Menzel N, Vieyres G, Perin PM, Frentzen A, Kaderali L, Pietschmann T (2013) Hepatitis C virus p7 is critical for capsid assembly and envelopment. PLoS Pathogens 9:e1003355
Hanisch J, Waltermann M, Robenek H, Steinbuchel A (2006) Eukaryotic lipid body proteins in oleogenous actinomycetes and their targeting to intracellular triacylglycerol inclusions: Impact on models of lipid body biogenesis. Appl Environ Microbiol 72:6743–6750
Kimmel AR, Brasaemle DL, McAndrews-Hill M, Sztalryd C, Londos C (2010) Adoption of PERILIPIN as a unifying nomenclature for the mammalian PAT-family of intracellular lipid storage droplet proteins. J Lipid Res 51:468–471
Layerenza JP, Gonzalez P, Garcia de Bravo MM, Polo MP, Sisti MS, Ves-Losada A (1831) Nuclear lipid droplets: a novel nuclear domain. Biochem Biophys Acta 2013:327–340
Li Z, Thiel K, Thul PJ, Beller M, Kuhnlein RP, Welte MA (2012) Lipid droplets control the maternal histone supply of Drosophila embryos. Curr Biol 22:2104–2113
Li Z, Johnson MR, Ke Z, Chen L, Welte MA (2014) Drosophila lipid droplets buffer the H2Av supply to protect early embryonic development. Curr Biol 24:1485–1491
Li L, Walsh RM, Wagh V, James MF, Beauchamp RL, Chang YS, Gusella JF, Hochedlinger K, Ramesh V (2015) Mediator subunit Med28 is essential for mouse peri-implantation development and pluripotency. PLoS ONE 10:e0140192
Liu P, Ying Y, Zhao Y, Mundy DI, Zhu M, Anderson RG (2004) Chinese hamster ovary K2 cell lipid droplets appear to be metabolic organelles involved in membrane traffic. J Biol Chem 279:3787–3792
Liu Z, Li X, Ge Q, Ding M, Huang X (2014) A lipid droplet-associated GFP reporter-based screen identifies new fat storage regulators in C. elegans. J Genet Genomics 41:305–313
Liu L, Zhang K, Sandoval H, Yamamoto S, Jaiswal M, Sanz E, Li Z, Hui J, Graham BH, Quintana A, Bellen HJ (2015) Glial lipid droplets and ROS induced by mitochondrial defects promote neurodegeneration. Cell 160:177–190
Martin S, Parton RG (2006) Lipid droplets: a unified view of a dynamic organelle. Nat Rev Mol Cell Biol 7:373–378
Miyanari Y, Atsuzawa K, Usuda N, Watashi K, Hishiki T, Zayas M, Bartenschlager R, Wakita T, Hijikata M, Shimotohno K (2007) The lipid droplet is an important organelle for hepatitis C virus production. Nat Cell Biol 9:1089–1097
Murphy DJ (2001) The biogenesis and functions of lipid bodies in animals, plants and microorganisms. Prog Lipid Res 40:325–438
Murphy DJ (2012) The dynamic roles of intracellular lipid droplets: from archaea to mammals. Protoplasma 249:541–585
Murphy DJ, Vance J (1999) Mechanisms of lipid-body formation. Trends Biochem Sci 24:109–115
Na H, Zhang P, Chen Y, Zhu X, Liu Y, Liu Y, Xie K, Xu N, Yang F, Yu Y, Cichello S, Mak HY, Wang MC, Zhang H, Liu P (2015) Identification of lipid droplet structure-like/resident proteins in Caenorhabditis elegans. Biochem Biophys Acta 1853:2481–2491
Ohsaki Y, Kawai T, Yoshikawa Y, Cheng J, Jokitalo E, Fujimoto T (2016) PML isoform II plays a critical role in nuclear lipid droplet formation. J Cell Biol 212:29–38
O’Mahony F, Wroblewski K, O’Byrne SM, Jiang H, Clerkin K, Benhammou J, Blaner WS, Beaven SW (2015) Liver X receptors balance lipid stores in hepatic stellate cells through Rab18, a retinoid responsive lipid droplet protein. Hepatology 62:615–626
Peramuna A, Summers ML (2014) Composition and occurrence of lipid droplets in the cyanobacterium Nostoc punctiforme. Arch Microbiol 196:881–890
Ploegh HL (2007) A lipid-based model for the creation of an escape hatch from the endoplasmic reticulum. Nature 448:435–438
Rowe ER, Mimmack ML, Barbosa AD, Haider A, Isaac I, Ouberai MM, Thiam AR, Patel S, Saudek V, Siniossoglou S, Savage DB (2016) Conserved amphipathic helices mediate lipid droplet targeting of perilipins 1-3. J Biol Chem 291:6664–6678
Sato S, Fukasawa M, Yamakawa Y, Natsume T, Suzuki T, Shoji I, Aizaki H, Miyamura T, Nishijima M (2006) Proteomic profiling of lipid droplet proteins in hepatoma cell lines expressing hepatitis C virus core protein. J Biochem 139:921–930
Shi ST, Polyak SJ, Tu H, Taylor DR, Gretch DR, Lai MMC (2002) Hepatitis C virus NS5A colocalizes with the core protein on lipid droplets and interacts with apolipoproteins. Virology 292:198–210
Ueno M, Shen WJ, Patel S, Greenberg AS, Azhar S, Kraemer FB (2012) Fat-specific protein 27 modulates nuclear factor of activated T cells 5 and the cellular response to stress. J Lipid Res 54:734–743
Van de Meene AM, Hohmann-Marriott MF, Vermaas WF, Roberson RW (2006) The three-dimensional structure of the cyanobacterium Synechocystis sp. PCC 6803. Arch Microbiol 184:259–270
Waltermann M, Steinbuchel A (2005) Neutral lipid bodies in prokaryotes: recent insights into structure, formation, and relationship to eukaryotic lipid depots. J Bacteriol 187:3607–3619
Waltermann M, Hinz A, Robenek H, Troyer D, Reichelt R, Malkus U, Galla HJ, Kalscheuer R, Stoveken T, von Landenberg P, Steinbuchel A (2005) Mechanism of lipid-body formation in prokaryotes: how bacteria fatten up. Mol Microbiol 55:750–763
Wan HC, Melo RC, Jin Z, Dvorak AM, Weller PF (2007) Roles and origins of leukocyte lipid bodies: proteomic and ultrastructural studies. FASEB J 21:167–178
Wang L, Wang Y, Liang Y, Li J, Liu Y, Zhang J, Zhang A, Fu J, Jiang G (2013) Specific accumulation of lipid droplets in hepatocyte nuclei of PFOA-exposed BALB/c mice. Sci Rep 3:2174
Wang Y, Zhou XM, Ma X, Du Y, Zheng L, Liu P (2016) Construction of nano-droplet/adiposome and artificial lipid droplets. ACS Nano 10:3312–3322
Welte MA (2015) Expanding roles for lipid droplets. Curr Biol 25:R470–481
Wolk CP (1973) Physiology and cytological chemistry blue-green algae. Bacteriol Rev 37:32–101
Yang L, Ding YF, Chen Y, Zhang SY, Huo CX, Wang Y, Yu JH, Zhang P, Na HM, Zhang HN, Ma YB, Liu PS (2012) The proteomics of lipid droplets: structure, dynamics, and functions of the organelle conserved from bacteria to humans. J Lipid Res 53:1245–1253
Zhang P, Na H, Liu Z, Zhang S, Xue P, Chen Y, Pu J, Peng G, Huang X, Yang F, Xie Z, Xu T, Xu P, Ou G, Zhang SO, Liu P (2012) Proteomic study and marker protein identification of Caenorhabditis elegans lipid droplets. Mol Cell Proteomics 11:317–328
Zhang C, Yang L, Ding Y, Wang Y, Lan L, Ma Q, Chi X, Wei P, Zhao Y, Steinbuchel A, Zhang H, Liu P (2017) Bacterial lipid droplets bind to DNA via an intermediary protein that enhances survival under stress. Nat Commun 8:15979
ACKNOWLEDGEMENTS
The authors thank Dr. Mark Christian for his critical reading and useful suggestions. The authors also thank Ms. Libing Mu for the graphical summary. This work was supported by the Ministry of Science and Technology of China (Grant No. 2016YFA0500100), National Natural Science Foundation of China (Grant Nos. U1402225, 31571388, 61273228, and 81270932), Chinese Academy of Sciences (Grant No. XDA12030201).
ABBREVIATIONS
CE, cholesterol ester; HCV, hepatitis C virus; LD, lipid droplet; MLDS, microorganism lipid droplet small; TAG, triacylglycerol
COMPLIANCE WITH ETHICS GUIDELINES
Congyan Zhang declares that he has no conflict of interest. Pingsheng Liu declares that he has no conflict of interest. This article does not contain any studies with human or animal subjects performed by the any of the authors.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Zhang, C., Liu, P. The lipid droplet: A conserved cellular organelle. Protein Cell 8, 796–800 (2017). https://doi.org/10.1007/s13238-017-0467-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13238-017-0467-6