Abstract
The diversity of filamentous fungi that can grow at high ambient pH values (i.e., 8–11) remains largely understudied. Here we study 100 alkalitolerant and alkaliphilic isolates from the soils around the basin of soda lakes in Asia and Africa to assess the major evolutionary lineages and morphologies pertinent to the alkaliphilic trait in filamentous fungi. The Emericellopsis lineage (Hypocreales, Hypocreomycetidae), along with Plectosphaerellaceae (Hypocreomycetidae), Pleosporaceae (Dothideomycetes), Chaetomiaceae (Sordariomycetidae) families appeared to be overrepresented with strong alkalitolerants and effective alkaliphiles. In particular, Sodiomyces species (Plectosphaerellaceae), Acrostalagmus luteoalbus (Plectosphaerellaceae), Emericellopsis alkalina (Hypocreales), Thielavia sp. (Chaetomiaceae), and Alternaria sect. Soda (Pleosporaceae) grew best at high ambient pH. The pH tolerance of Chordomyces antarcticum, Acrostalagmus luteoalbus and some other species was largely affected by the presence of extra Na+ in the growth medium. Moderate alkalitolerants included Scopulariopsis members (Microascales), Fusarium, Cladosporium, and many asexual acremonium-like species from Bionectriaceae. Weak alkalitolerants were represented by sporadic isolates of Penicillium, Purpureocillium lilacinum, and Alternaria alternata species, with the growth optimum at neutral or acidic pH. Weak alkalitolerants develop loose dry chains of spores easily dispersed by air. Their presence at low frequency with the growth optimum at neutral or acidic pH leads us to treat them as transient species in the alkaline soils, as those are also ubiquitous saprobes in normal soils. Our phylogenetic analyses show that the alkaliphilic trait in filamentous fungi has evolved several times. Several lineages harboring strong alkalitolerants derived from the known marine-borne fungi (Emericellopsis, Alternaria sect. Phragmosporae), or fall within the fungi associated with halophytic grasses (Pleosporaceae). Soda soils contain a diversity of fungi that range from weak alkalitolerant to alkaliphilic, which in few cases is associated with darkly pigmented mycelium and formation of microsclerotia. The alkaliphilic trait is spread throughout the Ascomycota, and usually juxtaposes with slime-covered polyphyletic acremonium-, verticillium-, gliocladium-types of asexual morphology, hyphae aggregating in chords, and enclosed fruit bodies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Taxonomic novelties
New species –
Sodiomyces magadii Bondarenko, A.A. Grum-Grzhim., A.J.M. Debets & Bilanenko,
Sodiomyces tronii Bondarenko, A.A. Grum-Grzhim., A.J.M. Debets & Bilanenko,
Chordomyces antarcticum Bilanenko, Georgieva & A.A. Grum-Grzhim.,
Alternaria kulundii Bilanenko, Georgieva & A.A. Grum-Grzhim.,
Alternaria petuchovskii Bilanenko, Georgieva & A.A. Grum-Grzhim.,
Alternaria shukurtuzii Bilanenko, Georgieva & A.A. Grum-Grzhim.
New genera –
Chordomyces Bilanenko, Georgieva & A.A. Grum-Grzhim.
New sections –
Alternaria sect. Soda Bilanenko, Georgieva & A.A. Grum-Grzhim.
Introduction
Nature possesses ample variation in environmental niches with diverse abiotic conditions. In some locations, abiotic factors (such as temperature, ion content and pH) may deviate from those of most habitats, resulting in the formation of specific zones termed ‘extreme habitats’ restricting growth of most organisms. Soda soils (or alkaline soils) with high pH values (>8) represent an example of such extreme habitats. They usually develop in arid and semi-arid lands throughout the world, and may vary in salt concentrations from low to saturation. The driving force for the soda accumulation is the depletion of Ca2+ trapped by CO3 2− ion, leaving Na+ as the dominant cation (Jones et al. 1998). Soda soils form as a result of carbonate accumulation under poor Ca2+ and Mg2+ conditions. Water evaporation reinforces the process of soda accumulation. With few exceptions, soda soils usually are highly saline as both abiotic factors coincide.
Naturally, as high ambient pH and salts impose significant stress on a living organism, the overall biodiversity is expected to be compromised. Surprisingly, soda lakes, water basins which are often formed in the area of soda soils, have been shown to be highly productive, in fact, more productive than their freshwater counterparts. As pointed out by Zavarzin (1993), soda lakes should exhibit complete recycling of the nutrients. Studies on the biodiversity of microorganisms in soda lakes have revealed abundant bacterial communities, which also act as primary producers, usually dominated by cyanobacteria species (Antony et al. 2013). High densities of bacterial and archaeal blooms may even cause the coloration of water basins associated with the soda soils, for example, as in Magadi Lake (Kenya) due to massive expansion of red haloalkaliphilic Archaea (Jones et al. 1998). Studies of 16S rDNA gene sequences revealed a complex phylogenetically heterogeneous structure of the bacterial communities inhabiting soda lakes (Duckworth et al. 1996; Rees et al. 2004; Dong et al. 2006; Wani et al. 2006) often with the recovery of many new bacterial species (Sorokin and Muyzer 2010; Sorokin et al. 2011; Kompantseva et al. 2012; Antony et al. 2012). The accumulated data suggest that soda lakes in diverse geographical regions harbor alkaliphilic bacteria and Archaea from all major trophic groups (Rees et al. 2004). Biodiversity surveys showed the presence of eukaryotic organisms thriving in alkaline lakes too, like plankton diatoms, green algae, cryptophytes and haptophytes (Nagy et al. 2006; Keresztes et al. 2012). The vegetation in the soda soils area is usually dominated by halophytic grasses from Chenopodiaceae (Grum-Grzhimaylo et al. 2013a). Often few dominant eukaryotic species may be encountered in soda lakes. For example, diverse crustaceans, like brine shrimp Artemia salina, as well as many species of copepods, are characteristic of hyper-saline alkaline water environments (Hammer 1986; Schneider et al. 2012). Intriguingly, eukaryotic 18S rDNA sequences pool analysis seems to have recovered signatures of alveolata, stramenopiles, choanoflagellates, amoebozoans and cercozoans present in Lonar Lake (India) sediments (Antony et al. 2013). Recent studies by Xiong et al. (2012) and Lanzén et al. (2013) confirmed both prokaryotic and eukaryotic biodiversity richness and heterogeneity by investigating several soda lakes employing high-throughput sequencing techniques.
Despite the general preference of fungi to grow at neutral or slightly acidic pH, some have been shown to grow at high pH too (Di Menna 1959; Goto et al. 1981; Aono 1990; Steiman et al. 2004). In many cases, alkaline soils were surveyed using neutral or acidic isolation media, thereby lowering the chances of recovering alkaliphilic strains. Lisichkina et al. (2003) reported a first recovery of alkalitolerant yeast species from soda soils using alkaline selective medium. In 1993, Okada and collaborators isolated the first alkaliphilic acremonium-like hyphomycete from manure, and named it Acremonium alcalophilum, as there was no other known alkaliphilic Acremonium species at that time (Okada et al. 1993).
Later, extensive studies by Nagai et al. (1995; 1998) focused on the taxonomic distribution of fungi in alkaline limestone caves in Japan as well as from Indonesian grassland. Using an alkaline medium, members of Acremonium, Fusarium, Gliocladium, Mucor and Plectosporium genera were isolated frequently. Many other species were termed alkalitolerant as they are capable of growing to some extent at elevated pH, but with the growth optimum still lying at neutral or acidic pH values. In particular, Acremonium species had shown wide alkalitolerance and this genus sensu lato was proposed to have properties to tolerate high ambient pH values. The abundance of alkalitolerant strains from alkaline habitats led to assumption that these strains are physiologically active in this environment. Interestingly, however, according to Nagai et al. (1995), acidic soils harbored few alkaliphiles as well, which is in line with similar observations in the studies of alkaliphilic bacterial communities (Horikoshi and Akiba 1982; Horikoshi 1999). Generally speaking, the frequency of alkaliphilic fungi is low, while alkalitolerants seem to be far more widespread, as alkalitolerants may often be encountered in many neutral types of soil.
The vast majority of studies on the alkaliphilic and alkalitolerant fungi conducted so far lack DNA sequence data to robustly identify the strains. As mentioned above, many alkalitolerant fungi possess acremonium-type asexual development as one of the simplest structures found in asexual fungi known to be highly polyphyletic throughout Ascomycota (Summerbell et al. 2011), imposing substantial difficulties for proper identification. It is therefore important to use DNA-based information to be able to pinpoint the taxonomy of a studied isolate. The sequencing data are also needed to understand whether the alkaliphilic trait in fungi has evolved once or several times throughout evolutionary history. Evidence for the latter scenario has been provided by our recent studies, in which we described few new taxonomically diverse alkaliphilic and alkalitolerant filamentous fungi inhabiting soda soils (Grum-Grzhimaylo et al. 2013a, b). Being polyphyletic, halophilic filamentous fungi have been shown to utilize different strategies of dealing with extreme ambient salt stress (Kis-Papo et al. 2014).
In the present study, we undertook a comprehensive approach to better understand the evolution of alkaliphily in fungi. To address this, on a buffered selective medium (pH 10) with an antibiotic, as well as on neutral common media, we isolated over 100 strains of alkalitolerant and alkaliphilic fungi sampled from the alkaline soils with different degree of salinity collected in Russia, Mongolia, Kazakhstan, Kenya, Tanzania, and Armenia. Essentially all isolated strains were tested for their pH preference to provide insight into the frequency and distribution of alkaliphiles in natural populations. The pH effect on the growth of the recovered isolates could discriminate between the physiologically active species and accidental transient species in alkaline habitats. Next, we sequenced several loci and reconstructed phylogenies to determine the positions of the alkaliphilic fungal lineages. Our strains were genetically heterogeneous and displayed various taxonomic affinities. We hence selected phylogenetically informative loci, which would better resolve the taxonomic group of interest according to the recent published studies. We studied representative strains using light microscopy as well as with a scanning electron microscope, and made photographic plates highlighting morphological features along with some drawings to better depict the key morphologies.
To make the present work more complete, we combined our new data with selected data from our previous studies (Grum-Grzhimaylo et al. 2013a, b). Noteworthy, our isolation method recovered only ascomycetous fungi. Overall, we found the alkaliphilic/strong alkalitolerant phenotype in about 2/3 of our recovered strains from soda soils. In addition, our data show a strong bias towards fungi having simple reproductive structures, which is in line with the previous studies. The results indicate that asexual sporulation with conidia likely glued together by slime (for example, in Sodiomyces, Acremonium, Acrostalagmus, Verticillium) forming compact heads correlates with the ability to tolerate high ambient pH. The systematic approach involving growth, microscopy and phylogenetic multi-gene analyses helped us to evaluate characteristic morphological characters of alkalitolerant and alkaliphilic fungi that evolved independently in several lineages (genus Emericellopsis and families Plectosphaerellaceae, Pleosporaceae, Chaetomiaceae) across Ascomycota.
Materials and methods
Locations and soda soil samples
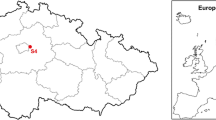
We collected samples of the soda soils at the locations shown on the map in Fig. 1. The pH of the samples varied from 7.6 to 11 with the range of total soluble salts of 1.4–310 g/kg. The details on the isolation places are provided in Table 1.
Strains and media
For initial isolation of alkalitolerant/alkaliphilic strains we used alkaline agar (AA) selective medium (ca. pH 10) supplemented with antibiotic rifampicin (2 g/l), as in Grum-Grzhimaylo et al. (2013a). Rifampicin was selected as the most effective antibiotic for the suppression of bacterial growth at alkaline conditions without hindering the fungal growth, among tested: ampicillin, amoxicillin (clavulanic acid), amikacin, bacitracin, chloramphenicol, clindamycin, doxycycline, erythromycin, gentamicin, kanamycin, lincomycin, levofloxacin, neomycin, novobiocin, ofloxacin, oxytetracycline, penicillin, tetracycline, tobramycin, vancomycin and rifampicin. In parallel, we implemented MYA, CZ and WA media (Mueller et al. 2004) for the possible recovery of weak alkalitolerant fungi. Emerged fungal isolates were purified and pre-identified by morphological characters. To avoid redundancy, in few cases of joint isolations from both media, we discarded the co-isolated strains down to a minimum number of morphotypes from a particular soil sample. The recovered fungal isolates were kept at 6 °C for further characterization. Newly acquired strains were deposited at the CBS-KNAW Fungal Biodiversity Centre (Utrecht, The Netherlands) and partly in the All-Russian Collection of Microorganisms (Pushchino, Russia). Colony morphology was characterized by growing the strains on several types of media: WA, CZ, MYA, PDA, OA, and AA (malt/yeast extract based) (Mueller et al. 2004; Grum-Grzhimaylo et al. 2013a) at 28 °C in the dark. Additionally, newly acquired Alternaria strains were grown on SNA (synthetic nutrient-poor agar) medium (Nirenberg 1976) at room temperature to induce sporulation as described in Woudenberg et al. (2013). For pH-optimum elucidation, we implemented race tubes (Perkins and Pollard 1986) with MYA-based medium buffered at pH from 4 to 11.2, as follows. The complete medium consisted of two components: the salt/buffer component (1) and the nutrient component (2). 500 ml of (1) had 0.2 M concentration of buffer supplemented with 5 g NaCl (Merck), 1 g KNO3 (Merck), 1 g K2HPO4 (Merck). 500 ml of (2) contained 17 g malt extract (Merck), 1 g yeast extract (BBL), 20 g agar (Duchefa Biochemie). (1) and (2) were autoclaved separately at 120 °C for 20 min, allowed to cool down to 55 °C and then mixed in 1:1 ratio, yielding final concentrations of the complete medium: 0.1 M buffer, 5 g/l NaCl, 1 g/l KNO3, 1 g/l K2HPO4, 17 g/l malt extract, 1 g/l yeast extract, 20 g/l agar. Buffer choices for generating different final pH values were: Na3C6H5O7/citric acid system for pH 4, 5.2, Na2HPO4/NaH2PO4 system for pH 5.9, 7, 7.8, Na2CO3/NaHCO3 carbonate system for pH 8.7, 9.8, Na2HPO4/NaOH system for pH 11.2. The above listed pH values are measured final pH of the complete media, which in few cases differed from the initial pH of the (1) component alone. Race tubes were inoculated in duplicate with agar plugs (1x1 mm) from the parental strain grown on Petri dish. Race tubes were put at 28 °C in the dark, and depending on the strain, the growth expansion was followed for 1–2 months.
Morphology
We used light microscopy (LM), scanning electron microcopy (SEM), and low-temperature scanning electron microscopy (cryoSEM) to morphologically characterize the isolated strains. For microscopy studies, fungal strains were taken from the medium, where the most developed morphology was observed. Specimens for SEM and cryoSEM were prepared as described previously (Grum-Grzhimaylo et al. 2013a). Alternaria specimens for LM were prepared as in Woudenberg et al. (2013). Characteristic structures were recorded digitally. However, we drew the reproductive structures of a few representative strains to clarify the morphology seen in the photomicrographs. Taxonomic novelties were deposited in MycoBank (www.MycoBank.org; Crous et al. 2004).
DNA extraction, PCR, and sequencing
For total genomic DNA (gDNA) extraction, mycelium was grown on a cellophane membrane placed on top of the AA medium. The collected mycelium was stored in Eppendorf tubes at −80 °C until used. For gDNA extraction, we used approximately 0.1 g of wet mycelium for each sample preparation with DNeasy Plant Mini kit (Qiagen, Chatsworth, CA), according to the manufacturer’s instructions. The quality and quantity of the isolated gDNA was verified on a NanoDrop 2000. We amplified various loci of our newly isolated strains, however, not all of them were used for phylogenetic reconstructions. A total set of the studied genes include the following: large and small subunit rDNA (LSU and SSU), internal transcribed spacers 1 and 2 including 5.8S rDNA (ITS), second largest subunit of RNA polymerase II (RPB2), transcriptional elongation factor 1 subunit α (TEF1-α), actin (Act), glyceraldehyde-3-phosphate dehydrogenase (GAPDH), and tryptophan synthase (TS). PCR primer sets for above listed loci were the following: LR0R/LR9 (LSU), NS1/NS8 (SSU), ITS1f/ITS4r (ITS), fRPB2-5F/fRPB2-7cR (RPB2), EF1α-983f/EF1α-2218r (TEF1-α). For verticillia strains, we used VActf/VActr (Act), VGPDf2/VGPDr (GAPDH), and VTs3f/VTs3r (TS) as in (Inderbitzin et al. 2011a). For Alternaria isolates in section Soda, we additionally amplified and sequenced the GAPDH marker with gpd1/gpd2 primer pair, as in (Berbee et al. 1999). PCR mixes, PCR products purification, and sequencing procedures of the remainder loci were performed as described previously (Grum-Grzhimaylo et al. 2013a). For the details on the selection of the annealing temperatures in PCR, see Table 2. Raw chromatograms of the newly generated sequences were viewed, edited and assembled with CodonCodeAligner v. 3. 7. 1 (CodonCode Corporation, Dedham, MA) and DNAStar Lasergene EditSeq v. 7. 1. 0 (DNASTAR Inc., Madison, WI). The resultant sequences were deposited in GenBank (Online Resource 1–8).
Phylogenetic analyses
To analyze the taxonomy of our strains, we performed several phylogenetic reconstructions with the various sets of loci resulting in a different degree of resolution. Appropriate reference sequences were obtained from GenBank. Multiple sequence alignments were created with online MAFFT v. 7 service (Katoh and Standley 2013) using automatic alignment strategy. The resulted alignments were reviewed and corrected manually. Ambiguously aligned regions, long gaps and introns were removed from the alignments using BioEdit v. 7. 1. 3. 0 (Hall 1999). The first dataset focused on the class-wide taxonomy of Ascomycota using single SSU locus to accommodate our all studied strains from soda soils. Next, for most of the strains, we produced separate trees on levels of families or genera. The set of trees include the taxonomic reconstructions of the studied strains within: Plectrosphaerellaceae (LSU, ITS), Verticillium s. str. (Act, GAPDH, TS, ITS), Sodiomyces (ITS), Scopulariopsis (LSU, TEF1-α), Lasiosphaeriaceae/Chaetomiaceae (LSU, ITS), Pleosporaceae (SSU, LSU, RPB2, ITS), and Alternaria sect. Soda (GAPDH). For the details on the above mentioned phylogenetic analyses, see Table 2. In the phylogenetic analyses, which involved more than one gene, matrixes were concatenated using Mesquite v. 2. 75 (Maddison and Maddison 2011). All multi-gene matrixes were divided into unlinked partitions corresponding to each gene. The best fit model for DNA substitution was estimated with jModelTest v. 2.1.1 (Guindon and Gascuel 2003; Darriba et al. 2012). Phylogenetic reconstructions were made with both Bayesian and maximum likelihood analyses. For Bayesian estimation of phylogeny, we used MrBayes v. 3. 1. 2. (Huelsenbeck and Ronquist 2001) with 4 Metropolis Coupled Markov Chain Monte Carlo (MCMCMC) chains running for 10 M generations starting with a random tree topology. Each 100th tree was sampled and their likelihood scores were checked in TRACER v. 1. 5 (Rambaut and Drummond 2007) to verify the convergence of the runs. The burn-in factor was set to 50 % in all our phylogenetic inferences, which accepted stationary stable set of trees. The consensus tree was generated employing the 50 % majority rule with the recovery of posterior probabilities (BI PP) at each node. Maximum likelihood bootstrap analysis (ML BP) was performed with GARLI v. 2. 0 (Zwickl 2006) with 200 bootstrap replicates including five searches for each. A 50 % majority rule consensus tree was constructed with SumTrees v. 3. 3. 1. script within DendroPy v. 3. 11. 0. library (Sukumaran and Holder 2010) running in Python v. 2. 6. environment. For visualization of the consensus trees, we used TreeGraph v. 2. 0. 47–206 beta (Stöver and Müller 2010) and Adobe Illustrator CS6 (Adobe Systems, San Jose, CA). The node supports were considered strong and displayed as thickened lines, if they gained joint supports of ML > 90 and PP > 0.94. Phylogenetic analyses and matrixes were deposited in TreeBase (submission #15631).
Results
Soda soil samples
Initially we screened our soda soil samples using soil suspensions diluted in water, which did not recover any fungal isolates, but rather revealed massive bacterial expansion. This observation led us to use soil clumps put directly on the selective medium for the recovery of fungi. Additionally, we boosted up the concentration of antibiotic rifampicin to a final concentration of 2 g/l.
Strains
The frequency of fungi recovery from soda soils was substantially lower in comparison to neutral habitats even with salinification. Nonetheless, we recovered 48 new strains within the current framework, however, we shall take into account a total set of strains derived from our three studies (Grum-Grzhimaylo et al. 2013a, b) to draw a broader picture of the distribution and features of the fungi from soda soils. All combined, from soda soils at the edge of the lake basins, we isolated 100 strains of fungi with various degrees of alkalitolerance and taxonomic affinity within Ascomycota. The majority of the recovered strains (>90) readily produced asexual spores, whereas only members of Sodiomyces and two strains of Emericellopsis developed the sexual stage. The rest of the strains remained sterile during our investigation, although they often formed resting structures, like dark thick-walled mycelium or, sometimes, chlamydospores and microsclerotia (observed in Pleosporaceae). Our isolation method employing the MYA- and CZ-based medium buffered at pH 10 did not yield yeasts or basidiomycetous, zygomycetous, chytridiomycetous fungi as well as lower fungi-like organisms, like Oomycota.
Growth experiments
The growth experiments using plates with various media (WA, CZ, MYA, PDA, OA, AA) and race tubes with MYA based medium (containing 5 g/l NaCl) ranging in pH values, showed different effect on growth of the studied isolates. The interpretation of growth experiments is often an ambiguous task, resulting in a high degree of subjectivity. Therefore we first shall set a few anchoring definitions, to orient across the growth pattern data we obtained. In Fig. 2 we plotted three distinct growth classes with relation to pH: acidophiles, neutrophiles, and alkaliphiles. Acidomyces acidophilum species grows best at acidic pH (from 3 to 5), while Sodiomyces species display the opposite – their growth optimum lie at pH between 8.5 and 11 with no ability to grow at pH 4. The neutrophilic physiology is the most abundant, and here is represented by Aspergillus nidulans, which growth optimum is at pH around 5.5–6. These clearly distinct physiological boundaries are rarely observed, as there are many intermediate growth types.
In the current framework, we accept several categories of growth types with relation to pH, adopted from the existing bacterial studies. The alkaliphiles, with the growth optimum at pH above 8 are subdivided into obligate and facultative. Obligate alkaliphiles cannot grow at pH below 4–5, while facultative can. The next group is alkalitolerants, which can cope with high pH to different extent, however, the growth optimum lies at pH below 8 (at neutral, or even acidic pH). We further divide alkalitolerants into three classes by the severity at which the growth is hindered as the pH increases. These classes are: strong, moderate, and weak alkalitolerants. Weak alkalitolerants are often neutrophiles, which can barely cope with high ambient pH showing highly reduced growth. Moderate alkalitolerants are the isolates which grow at about half the growth rate at high pH, as compared to neutral pH. Strong alkalitolerants show little or no reduction in growth at neutral pH with often wide pH growth optimum. In some cases, our growth patterns in Petri dishes on commonly used media at neutral pH and MYA-based media with various pHs are inconsistent. In such cases, the pH tolerance is greatly affected by the presence of extra Na+ (in the form of NaCl) in the media. The terminology used in the current study is summarized in Fig. 3. It should be pointed out, that studying the pH effect alone is challenging, as this factor interweaves with osmotic pressure, water potential and ion concentrations effect, making the pH factor hard to disentangle.
In general, our results show that the Plectosphaerellaceae (Zare et al. 2007) harbours a large number of strong alkalitolerants and alkaliphilic isolates (Figs. 4, 5). Genera Chordomyces, Acrostalagmus, and Sodiomyces of Plectosphaerellaceae showed alkaliphilic phenotype, as seen in Figs. 6, 7, 8 and 9. For Sodiomyces, comparisons of growth in race tubes and on plates suggest that the presence of extra Na+ does not have a large impact on the growth pattern. From the graphs and plate images, it is seen that the neutral pH media show reduced growth, while at high pH the growth substantially improved. These results led us to assume that Sodiomyces is more affected by the pH factor than the extra Na+ supplementation, displaying truly alkaliphilic phenotype. Chordomyces antarcticum and Verticillium zaregamsianum isolates, on the other hand, seem to be more affected by the presence of extra Na+. As seen in Figs. 6 and 11, the growth in tubes at neutral pH supplemented with NaCl was almost as good as at high pH. However, on plates with commonly used neutral media like MYA, PDA, and to a lesser extent on OA, the growth was greatly inhibited. The growth on AA medium was best for these species. Interestingly, CBS 987.87 isolate of Chordomyces antarcticum (identified as Acremonium antarcticum before) seems to possess less pH tolerance capacity compared to our isolates of Chordomyces from soda soils, as the growth was best at pH 6–7 with extra Na+ in the medium. Facultative alkaliphile Acrostalagmus luteoalbus displays intermediate effect of extra Na+ − its growth was indeed improved with the addition of NaCl, but not as dramatic as for Chordomyces antarcticum or Verticillium zaregamsianum (see Fig. 7).
Overview of the wide scale Ascomycota tree indicating the phylogenetic positions of the alkalitolerant and alkaliphilic isolates derived from the current study as well as from Grum-Grzhimaylo et al. 2013a, 2013b. Bayesian 50 % majority rule consensus tree as inferred from the phylogenetic analysis of the single locus (SSU). Thickened branches indicate strong combined support (ML > 90, PP > 0.94). The colour-coded pH growth preference is indicated
Position of the alkalitolerant and alkaliphilic isolates within Plectosphaerellaceae. Bayesian 50 % majority rule consensus tree as inferred from the phylogenetic analysis of two loci (LSU, ITS). Thickened branches indicate strong combined support (ML > 90, PP > 0.94). T – type/ex-type strains; designation holds for the remainder of the figures throughout the study
Facultative alkaliphilic Chordomyces antarcticum (Plectosphaerellaceae) colony morphology on various around-neutral pH (except for AA with pH 10) media (top) and linear growth rate patterns at different pH values in race tubes (below). Media in race tubes contained extra Na+ as compared to neutral WA, CZ, MYA, PDA, and OA media used for growth on plates. Extra Na+ boosts growth at neutral pH for Chordomyces
Obligate alkaliphilic Sodiomyces tronii and Sodiomyces magadii (Plectosphaerellaceae) species colony morphology on various media. The phylogenetic position of Sodiomyces species is shown on the 50 % majority rule Bayesian consensus tree, which is constructed based on single ITS locus. Thickened branch indicate strong combined support (ML > 90, PP > 0.94). The growth pattern at various pH values is given also for Acremonium alcalophilum CBS 114.92 ex-type isolate. Note that for Sodiomyces, extra Na+ present in linear growth experiment has little effect on alkaliphilic phenotype, compared to Chordomyces antarcticum
Moderate alkalitolerant isolates of Scopulariopsis (Microascales) colony morphology on various media. The tree is a 50 % majority rule Bayesian consensus as inferred from the analysis of two loci (LSU, TEF1-α). Thickened branches indicate strong combined support (ML > 90, PP > 0.94). Growth patterns at different pH values are displayed
Verticillium zaregamsianum colony morphology on various media. The position of the strong alkalitolerant (strains V201, V202, V204) and facultative alkaliphilic (strain V203) Verticillium zaregamsianum isolates are displayed on the tree. The tree is a 50 % majority rule Bayesian consensus as inferred from the phylogenetic analysis of the four loci (Act, GAPDH, TS, ITS). Thickened branches indicate strong combined support (ML > 90, PP > 0.94). The growth patterns at different pH values are given
Verticillium zaregamsianum V204 (= CBS 137624): a, b Conidiophores (LM). c Resting mycelium (LM). d–f Microsclerotia (LM). g Chlamydospore (LM). h Conidia (LM). i Conidiophore with conidia (cryoSEM). j Clump of conidia (cryoSEM). k Conidiophore, conidia, chlamydospores, resting mycelium and microsclerotia. l, m Microsclerotia (cryoSEM). Scale bars: a–h, k, l = 10 μm; i, m = 5 μm; j = 1 μm
Scopulariopsis fusca G414 (= CBS 138116): a–d Conidiophores and conidia (LM). e, f Conidiophores and conidia (cryoSEM). g Chain of conidia (cryoSEM). h Annellidic conidiogenesis (cryoSEM). i Conidia junction (cryoSEM). j Conidia junction as seen from beneath (cryoSEM). k Conidia surface element (cryoSEM). Scale bars: a–f = 10 μm; g = 5 μm; h = 4 μm; i = 2 μm; j = 1 μm; k = 0.5 μm
Scopulariopsis brevicaulis G415 (= CBS 137632): a–c Conidiophores and conidia (LM). d, e, g Conidiophores and conidia (cryoSEM). f Conidium within a conidia chain (cryoSEM). h Conidia junction as seen from beneath (cryoSEM). i Conidia junction as seen atop (cryoSEM). Scale bars: a–e, g = 10 μm; f = 3 μm; h, i = 1 μm
Position of the two obligate alkaliphilic isolates of Thielavia sp. within Chaetomiaceae. Ambiguous Lasiosphaeriaceae sp. M314 strain fall within sister family. Bayesian 50 % majority rule consensus tree as inferred from the phylogenetic analysis of two loci (LSU, ITS). Thickened branches indicate strong combined support (ML > 90, PP > 0.94)
Two Thielavia sp. isolates M316 and M317, Chaetomiaceae (de Hoog et al. 2013; Kruys et al. 2014; Wang et al. 2014), displayed an obligate alkaliphilic phenotype, as seen in Fig. 16. Isolate Lasiosphaeriaceae sp. M314 showed different results on plate and in race tubes. In tubes, it displayed alkaliphily, while on plates the growth was better at neutral pH values; these observations did not allow us to establish the relation to pH for this strain in our study. Three Scopulariopsis isolates (two S. brevicaulis and one S. fusca) showed moderate alkalitolerant abilities confirmed by the growth experiments both in tubes and on plates. As seen in Fig. 10, the growth optima for Scopulariopsis isolates are at neutral pH, and OA medium in particular supported a better growth over the rest of neutral pH media (MYA, PDA). Growth on CZ medium was the fastest, but the sporulation was reduced. Scopulariopsis fusca displayed somewhat better alkalitolerance as compared to S. brevicaulis isolates. Single case isolations of Cladosporium sphaerospermum, Purpureocillium lilacinum, Fusarium sp., and Penicillium sp. show weak to moderate alkalitolerance, as seen in Fig. 22.
Growth patterns at various pH values for Alternaria isolates. a Weak alkalitolerant Alternaria alternata G408 (= CBS 137513). b Strong alkalitolerant Alternaria molesta M312 (= CBS 137524) and moderate alkalitolerant Alternaria sp. M301 (= CBS 137514) (sect. Phragmosporae). c Strong alkalitolerants and facultative alkaliphiles of Alternaria sect. Soda isolates
Asexual low-frequent weak and moderate alkalitolerant isolates of Cladosporium sphaerospermum G402 (= CBS 137612), Purpureocillium lilacinum G406 (= CBS 137613), Fusarium sp. G412 (= CBS 137614), and Penicillium sp. G418 (= CBS 137615) colony morphology on various media. The growth patterns at various pH values are given
Asexual low-frequent isolates from soda soils. a Moderate alkalitolerant Cladosporium sphaerospermum G402 (= CBS 137612). b Weak alkalitolerant Purpureocillium lilacinum G406 (= CBS 137613). c Moderate alkalitolerant Fusarium sp. G412 (= CBS 137614). All images were taken with cryoSEM. Scale bars: a–c = 5 μm
Acremonium-like morphology of alkalitolerant isolates. a, b, f Acremonium sclerotigenum A101 (= CBS 138752): hyphal chords, conidiophores, conidial heads (cryoSEM) and a drawing of the morphology observed. c–e, g Sarocladium sp. A131 (= CBS 138121): hyphal chords, conidiophores, conidial heads (cryoSEM) and a drawing of the morphology observed. Scale bars: a–d, f, g = 10 μm, e = 5 μm
Strong alkalitolerant Acremonium roseolum A109 (= CBS 138118). a–c 11-d-old colony morphology on PDA, OA, and alkaline agar (AA), respectively (9 cm Petri dish). d Catenate conidia in native culture (LM). e, f Catenate conidia (cryoSEM). g–j Conidia and conidiophores (LM). k Conidium surface (cryoSEM). l Conidia being formed on a conidiophore emerging from a hyphal chord (cryoSEM). Scale bars: d = 50 μm, e, l = 5 μm, f, k = 2 μm, g–j = 10 μm
Sodiomyces magadii MAG2 (= CBS 137619). a Agar plug with fruiting bodies as seen through a binocular microscope. b Intact fruiting body with ascospores inside (LM). c Opened ascoma releasing ascospores. Note the peridium surface structure (LM). d Fruiting body (SEM). e Chlamydospores (SEM). f Ascospores (SEM). g Ascospores (note the septa) (LM). h Broken fruiting body (SEM). i, j, k Fruiting body, ascospores and chlamydospore, respectively. l Cracked ascoma with ascospores (SEM). Scale bars: b, c, i = 20 μm, d, h = 50 μm, e, g = 5 μm, f, j, k, l = 10 μm
Sodiomyces tronii MAG1 (= CBS 137618). a, b Conidiophore development (LM). c Mature conidiophore with conidia (LM). d–h Early and late stages of conidiophore development (cryoSEM). i Early stage of fruiting body development. Strings of thick-walled brown cells are involved in exoperidium formation (LM). j Opened ascoma with an empty ascus (arrowed) (LM). k Fruiting body (SEM). l Peridium structure as seen on a cross-section (SEM). m Broken fruiting body (SEM). n A drawing of the asexual structures observed. o Fruiting body development stages. Strings of thick-walled brown cells are involved in exoperidium formation. p Mature fruiting body (no ascospores have been observed). q Chains of thick-walled cells involved in exoperidium formation. Scale bars: a–h = 5 μm, i, k, o–q = 20 μm, j, l, n = 10 μm, m = 50 μm
Isolates falling in the Pleosporaceae (Hyde et al. 2013) showed different capabilities for the pH tolerance (Figs. 18, 19 and 21). Alternaria isolates displayed various degrees of alkalitolerance: weak alkalitolerant (single A. alternata sect. Alternata isolate G408), moderate alkalitolerant Alternaria sp. sect. Phragmosporae M301 and strong alkalitolerant A. molesta sect. Phragmosporae M312, and five strong alkalitolerant/facultative alkaliphilic isolates (M304, M307, M309, M310, M313) of new Alternaria species fall in a new section Soda described below. Outside Alternaria, allied to Pleospora, we obtained six isolates, which have been sterile. Four of them (M302, M303, M306, and M315) show strong alkalitolerance. Finally, sterile Pleosporaceae spp. M305 and M311 isolates display facultative alkaliphilic phenotype.
Phylogenetic analyses
Ascomycota (SSU)
We reconstructed a high-level Ascomycota tree to map all our newly isolated strains from soda soils onto the ascomycetous lineages (Fig. 4). We used the SSU locus as a single partition with 66 taxa sampled throughout several phyla within Ascomycota. The basidiomycetous Armillaria mellea PBM2470 and Saccharomyces castellii CBS 4309 yeast served as an outgroup for this phylogenetic analysis. The dataset contained 1629 characters of which 428 (26.3 %) were phylogenetically informative. The Bayesian MCMC runs converged after 5 M generations and remained stationary afterwards with the deviation of approximately 0.006 between the runs. Our single-locus tree provided enough resolution to identify the major lineages of Ascomycota. The clade delineation followed from the Spatafora et al. (2006). On this tree, we often used a single sequence reference to serve as a point for depicting the position of our multiple strains isolated from soda soils. To show the distribution of the alkalitolerant trait across our isolates, we included the colour-coded pH growth preference, as inferred from our growth experiments. Our alkalitolerant and alkaliphilic isolates fall within many taxonomic lineages across Ascomycota. However, our results indicate that subphylum Hypocreomycetidae seems to be particularly enriched with alkaliphilic and alkalitolerant fungi, as 80 % of our isolates belong there. The presented phylogeny based on the single SSU gene analysis approach is not meant for high phylogenetic accuracy, as it is used only to illustrate the approximate positions of our alkalitolerant fungi throughout the Ascomycota. Later in this study, we will treat most of the alkalitolerant and alkaliphilic fungal lineages in a greater detail. Noteworthy, for drawing a broader picture of the biodiversity of fungi and their frequencies in soda lakes, we include strains from our recent studies (Grum-Grzhimaylo et al. 2013a, b).
Plectosphaerellaceae (LSU, ITS)
The dataset for the Plectosphaerellaceae (Hypocreomycetidae) contained 65 taxa and included two loci concatenated in a single alignment (LSU and ITS). The sequences for the alignment varied little and thus were easily aligned. The alignment comprised 1286 characters with 283 (22 %) of phylogenetically informative characters, which resolved major genera within the Plectosphaerellaceae (Fig. 5). Sister members of Glomerellaceae, Glomerella cingulata FAU 553 and Colletotrichum boninense CMT74 were used to root the tree. Members of Plectosphaerella fall into 78/0.97 supported clade. Recently described isolates of P. oligotrophica (Liu et al. 2013) sister to P. ramiseptata and P. citrullae ex-types were not included in our analysis. Another recently introduced Plectosphaerellaceae genus Lectera, harboring plant-associated species (Cannon et al. 2012) clearly stand out in a long-branched clade with maximal support values. Our new twelve facultative alkaliphilic isolates robustly (100/1.0) group with the known isolate identified as Acremonium antarcticum CBS 987.87, which currently has no nomenclatural status. We designated a type isolate (strain M27) for this clade and accommodated a new genus, Chordomyces, with a single species – Chordomyces antarcticum. Morphology of the ex-type culture is provided in Fig. 30. Several other isolates of Chordomyces antarcticum were included in morphological analysis with low-temperature SEM (see Fig. 31). Also, we obtained five facultative alkaliphilic isolates, which were identified as Acrostalagmus luteoalbus (Fig. 8), as they unambiguously (100/1.0) group within known CBS 194.87 isolate, a combination introduced by Zare et al. (2004). Obligate alkaliphilic Sodiomyces isolates group in a well-supported clade (75/0.98), few of them display different morphological features and diverge molecularly, therefore, they were described here as new species, S. tronii and S. magadii. The morphology and ITS-based phylogeny of the Sodiomyces species is treated below.
Chordomyces antarcticum M27 (= CBS 120045). a 20-d-old colony on AA agar (9 cm Petri dish). b Hyphal chord with branching (LM). c Hyphal chord with unbranched conidiopohres (LM). d Sterilic hyphal chord tip (LM). e–h Various types of branching conidiophores (LM). i, j Hyphal chords with conidiation (SEM). k Conidiophore collarette (arrowed) (LM). l–o Various types of conidiophores (SEM). p A drawing of the morphology observed. Scale bars: b–i, k–p = 10 μm; j = 50 μm
Sodiomyces (ITS)
We isolated two new obligate alkaliphilic species of Sodiomyces from Magadi Lake (Kenya), which is supported by both morphological and phylogenetic analyses. The ITS phylogeny of Sodiomyces (Plectosphaerellaceae) isolates is depicted in Fig. 9, which is rooted to a close alkaliphilic asexual Acremonium alcalophilum CBS 114.92 ex-type strain. The matrix for this analysis had 509 characters, only 13 of which were phylogenetically informative (2.5 %). Such a small variation within the ITS locus was enough to reconstruct the phylogenetic relationships within the Sodiomyces. Two isolates of S. tronii (MAG1, MAG3) form a well-supported clade (86/0.97) within a group of S. alkalinus isolates, while S. magadii (MAG2) is placed more basally to the S. alkalinus group, closer to Acremonium alcalophilum CBS 114.92 ex-type isolate. Sodiomyces therefore has been sized up to 21 isolates grouped in 3 species. The morphology of new species is shown in Figs. 27, 28, and we also made a comparative morphological chart with three species of Sodiomyces showing differentiating criteria between the species (Fig. 29).
Verticillium sensu stricto (Act, GAPDH, TS, ITS)
Robust identification of four Verticillium strains (V201, V202, V203, V204) recovered from soda soil at different depths at Orongoyskoe Lake required the analysis of extra loci, other than ITS and LSU alone. For this, we followed the Inderbitzin et al. (2011b) and Hyde et al. (2014) studies utilizing Act, GAPDH, TS, along with ITS sequences. The phylogenetic analysis of these loci for Verticillium s. str. clade (Plectosphaerellaceae) is shown in Fig. 11. The final concatenated matrix contained 1840 characters with 31 % of them being phylogenetically informative. Members of the sister genus Gibellulopsis was used to root the tree. Our isolates group within the Verticillium zaregamsianum clade of reference isolates falling into Flavexundans-clade with good support values (80/0.99), and therefore identified as such. Verticillium species of Flavexundans-clade are characterized by the presence of yellow pigment, which is also present in our isolates (see Fig. 11) adding extra evidence for proper identification of our strains. Morphological features of the V. zaregamsianum V204 isolate are depicted in Fig. 12. Interestingly, although four isolates of V. zaregamsianum, sampled at different depths, were identical across four studied loci, but the growth patterns though seem to differ. V201 and V202 isolates recovered from 30 to 70 cm depth, seem to be less alkalitolerant than V203 and V204 recovered at greater depth at about 60 to 95 cm.
Scopulariopsis (LSU, TEF1-α)
Three Scopulariopsis isolates (Microascales) with moderate alkalitolerance were identified as S. brevicaulis and S. fusca based on both morphological data and two-locus phylogenetic analysis (see Figs. 10, 13 and 14). Double-loci, LSU and TEF1-α, concatenated phylogenetic matrix contained 1429 characters, which include 13 % of phylogenetically informative characters (mostly in TEF1-α gene). The reference species designation was taken from Sandoval-Denis et al. (2013). Two isolates, G413 and G415, fall within the well-supported S. brevicaulis-clade (85/1.0), whereas isolate G414 falls within the sister group – S. candida complex. The identification of the G414 isolate to the species level was largely based on the morphological features.
Lasiosphaeriaceae/Chaetomiaceae (LSU, ITS)
Two obligate alkaliphilic strains recovered from Kunkur Steppe (Russia) and Ararat Valley (Armenia) – M316 and M317 fall into the Chaetomiaceae grouping with Thielavia australiensis ATCC 28236 ex-type strain with good support values (83/1.0), as seen in Fig. 15. Unfortunately, M316 and M317 isolates have not formed morphological structures based on which we could describe these strains in detail and they are very closely related phylogenetically. We found two synonymous mutations in RPB2 gene and one insertion mutation in ITS region between M316 and M317 isolates. Interestingly, the eye-catching feature of M316 is that it contains a group 1 intron in the SSU gene spanning for 416 nucleotides as opposed to M317 isolate. In spite of phylogenetic identity of LSU and ITS to Thielavia australiensis ATCC 28236 ex-type, we therefore shall identify them as members of Thielavia, even though Thielavia species display a large degree of polyphyly across Chaetomiaceae. One ambiguous isolate M314 group within sister Lasiosphaeriaceae and have been sterile during our investigation. This isolate falls into the polyphyletic area of genera Zopfiella, Cercophora, Cladorrhinum, which makes proper identification based on sequencing data impossible. We keep the Lasiosphaeriaceae sp. designation for the M314 strain (Fig. 16).
Pleosporaceae (SSU, LSU, RPB2, ITS)
Fourteen isolates of various alkalitolerance are members of Pleosporaceae (see Fig. 17). Eight grouped within Alternaria and spread across three sections (Alternata, Phragmosporae, and Soda). Based largely on morphology, isolate G408 was identified as A. alternata, which is grouped within Alternata section. The other strains (M301 and M312) fall into Phragmosporae section. One of them, M312, produced conidia and was identified as A. molesta, the other one has been sterile and we keep the Alternaria sp. designation for the strain M301. Five isolates of Alternaria (M304, M307, M309, M310, and M313) are clearly diverging from the rest sister sections of Alternaria and we accommodate those isolates into a new section Soda, which is treated below. Six isolates are placed basal to the Alternaria, having grouped with Pleospora-allied genera. Unfortunately, as these six isolates have been sterile, we were not able to study them thoroughly. Interestingly, some of these isolates might represent new genera as, for example M306, M305, M311, and M302, distinctly diverged from the known reference species (Figs. 17, 18 and 19).
Alternaria sect. Soda (GAPDH)
In order to follow the evolutionary relationship between the isolates in a new section Soda and allied sections we constructed a separate phylogenetic tree based on the partial sequence of glyceraldehyde-3-phosphate dehydrogenase (GAPDH) gene containing introns which contribute well to the clade differentiation within the section rank (Fig. 20). The matrix contained 23 % of phylogenetically informative characters and was rooted to Pleospora tarda CBS 714.68 type. Our analysis shows that our five new isolates clearly diverge from the sister sections of Alternaria – sect. Infectoriae, Embellisia and Chalastospora, confirming similar observation based on a more conserved dataset as seen before in Fig. 17. GAPDH data provides evidence that five isolates split into two or three species. Here, we designate three A. kulundii sp. nov. (M309, M310, and M313), A. petuchovskii sp. nov. (M304), and A. shukurtuzii sp. nov. (M307). Although the statistical support for A. kulundii clade is extremely low (57/0.54), we shall designate it as a separate species from the M304 isolate, in spite the fact that the clade of A. kulundii together with M304 received maximal supports. Our decision to split this four-isolate cluster strongly relied on the growth patterns. The growth experiments provide the evidence for the species differentiation, as in Figs. 19, 21 it is seen that isolates of A. kulundii M309, M310, M313 behave somewhat similar on various media, however, A. petuchovskii M304 and A. shukurtuzii M307 perform differently with a highly reduced growth rate on neutral media and overall colony morphology without Na+ supplementation. The addition of extra Na+ strongly reinforced the growth at neutral pH for those species. A. shukurtuzii M307 is placed outside A. kulundii and A. petuchovskii isolates. Unfortunately, we were not able to amplify GAPDH gene in M301 and M312 isolates, which fall into Phragmosporae section, thus M312 isolate as one producing conidia was identified by morphology as A. molesta (Fig. 26e–j), and M301 being sterile was left as Alternaria sp. (Figs. 22 and 23).
By a combination of morphology and DNA information, we identified isolates G402, G406, G412, and G418 as Cladosporium sphaerospermum, Purpureocillium lilacinum, Fusarium sp., and Penicillium sp., respectively (Figs. 22 and 23).
Morphology
Simply-structured acremonium-like morphotypes dominate across our collection of alkalitolerant and alkaliphilic fungi. Sodiomyces alkalinus, Acrostalagmus luteoalbus, Emericellopsis alkalina, Chordomyces antarcticum, Acremonium sp. are the most represented taxa in our study, which produce such type of asexual morphology (Figs. 8, 30 and 31). For the Sodiomyces alkalinus and Emericellopsis alkalina morphology details, see Grum-Grzhimaylo et al. (2013a, b). Sister to Emericellopsis lineage, we observed moderate alkalitolerants of Sarocladium affinity and Acremonium sclerotigenum, which produce similar conidiation, as seen in Fig. 24. Above mentioned taxa represent a large part of the observed biodiversity from soda soils within our framework and are polyphyletic. Despite being polyphyletic, they produce similar conidiospores clumped up in compact heads, likely to be glued together by slime. Sister to Emericellopsis, we recovered a strong alkalitolerant isolate identified as Acremonium roseolum (strain A109) forming atypical catenate conidia (Fig. 25), which falls into the inflatum-clade, as outlined by Summerbell et al. (2011). At substantially lower frequencies we observed chains of dry air-borne conidia in the species of Alternaria, Cladosporium, Purpureocillium, Fusarium, Penicillium, and Scopulariopsis (Figs. 13, 14, 23, 26 and 32). Unfortunately, a number of isolates have not developed reproductive stages during our investigation. Those include obligate alkaliphiles of Chaetomiaceae – Thielavia sp., single Lasiosphaeriaceae sp. isolate and Pleospora-related (Pleosporaceae) isolates of different affinity to pH. Ascomata formation has been observed only in Sodiomyces species, and two isolates of Emericellopsis alkalina. One of them, E101, was reported previously in Grum-Grzhimaylo et al. (2013b), however, later we observed the ascoma formation in another E. alkalina isolate – A117 (data not shown). Interestingly, we did not record ascospores inside the ascomata of Sodiomyces tronii. In Alternaria sect. Soda and Verticillium isolates we observed darkly pigmented thick-wall mycelium along with the formation of microsclerotia (Figs. 12, 32 and 33), which might be survival structures in harsh environments. We have not detected resting structures in facultative alkaliphiles like Acrostalagmus, Chordomyces. Obligate alkaliphilic Sodiomyces tronii and S. magadii have been able to produce single or short-chained chlamydospores (Fig. 29), as opposed to S. alkalinus, which did not form chlamydospores during our investigation. Curiously, obligate and facultative alkaliphilic isolates of our collection, Thielavia sp., Sodiomyces species, Acrostalagmus luteoalbus, Chordomyces antarcticum, Emericellopsis alkalina display brightly-coloured (white, salmon, pinkish, orange) colonies as seen on plates with the studied media, which is counterintuitive, as the stress-related and defensive morphology often encompasses darkly-coloured mycelium.
Taxonomy
Sodiomyces A.A. Grum-Grzhim. et al., Persoonia 31:154, 2013.
A genus of the Plectosphaerellaceae (Glomerellales, Hypocreomycetidae).
Colonies on alkaline agar (AA, pH 10.0–10.2) rather fast-growing, on MEA (pH 6.0–6.5) growing more slowly. Young colonies white; later, darkening zones appearing as a result of formation of abundant ascomata, velvety to woolly. Reverse colourless. Odour pleasant. Exudate absent. Mycelium superficial or partly immersed, consisting of hyaline, smooth-walled septate hyphae, often fasciculate. Decumbent vegetative hyphae thin-walled, hyaline.
Asexual morph acremonium- and gliocladium-like. Conidiation nematogenous and plectonematogenous. Conidiophores are characterized by considerable variation in the branching patterns including solitary phialides early formed laterally from simple and fasciculate vegetative hyphae, basitonously verticillate, penicillate (monoverticillate and biverticillate) arrangement. Phialides variable, gradually tapering towards the apex, rather thin-walled. Conidia aseptate, hyaline, subglobose, oval, ellipsoidal, cylindrical, with rounded apices, smooth as observed by SEM, aggregated in slimy masses. Chlamydospores absent or present, 1–2-multi-celled, intercalary, terminal, in chains, darkening with age, forming bands and microsclerotia.
Ascomata cleistothecial, dark-brown, globose, superficial. Peridium multi-layered, pseudoparenchymatous. Paraphyses absent. Asci thin-walled, without apical apparatus, saccate, unitunicate, scattered irregularly in the ascocarp, embedded in a gelatinous matrix. Ascospores ellipsoidal or ovoid, medially 1-septate, not constricted at the septum, thick-walled, pale brown, smooth. Ascospores released by dissolution of the ascus wall before maturity, accumulating within the ascocarp, released in a slimy mass, liberated by pressure within the ascocarp.
Type species: Sodiomyces alkalinus (Bilanenko & M. Ivanova) A.A. Grum-Grzhim. et al.
Grum-Grzhimaylo et al., Persoonia 31:157. 2013.
= Heleococcum alkalinum Bilanenko & M. Ivanova, Mycotaxon 91:501. 2005.
Key to Sodiomyces species
1. Ascospores present on alkaline agar, asexual morph acremonium-like or absent, conidia subglobose, L/B < 2, chlamydospores not in chains if present (2)
Ascospores absent on alkaline agar, asexual morph gliocladium-like, conidia cylindrical, L/B > 2, chlamydospores in chains (S. tronii)
2. Ascospores length more than 10 μm (S. alkalinus)
Ascospores length less than 10 μm (S. magadii)
Sodiomyces magadii Bondarenko et al., sp. nov. MycoBank MB 811263 Fig. 27.
Etymology: Name refers to the Magadi Lake in Kenya (Africa), where the fungus was isolated.
Ascomata dark brown, superficial on the substratum, globose, 90–220 (on average 160) μm diam, non-ostiolate, cleistothecial, not changing colour in 3 % KOH and lactic acid. Peridium multi-layered, pseudoparenchymatous, exoperidium on average 9 μm thick, composed of 3–5 layers of flattened cells. Paraphyses absent. Asci not observed. Ascospores ovoid, 6–8 × 4–7 μm, medially 1-septate at maturity, not constricted at the septum, thick-walled, pale brown, smooth.
Asexual morph unknown.
Culture characteristics — Colonies on alkaline agar (AA, pH 10.0–10.2) reaching 40–50 mm diam in 10 d at 25 °C. On MEA (pH 6.5) growth is extremely restricted. Young colonies white; later, black punctuated zones appearing as a result of formation of abundant ascomata, fluffy to woolly. Reverse colourless. Exudate absent. Odour pleasant. Decumbent vegetative hyphae thin-walled, hyaline, 0.5–2.0 μm wide. Mycelium consisting of hyaline, smooth-walled, septate hyphae, 1–3 μm wide, often fasciculate. Chlamydospores 1- and 2-celled, intercalary and terminal, 8–12 μm diam.
Specimen examined. Kenya, soda soil (pH 11) at the edge of Magadi Lake, Jan. 2013, S. Bondarenko (CBS H-21958 – holotype); culture ex-type MAG2 = CBS 137619 = VKM F-4583.
Sodiomyces tronii Bondarenko et al., sp. nov. MycoBank MB 811264 Fig. 28.
Etymology: Name refers to the ‘trona’ salt (carbonate mineral), which is abundant in Magadi Lake in Kenya (Africa), where the fungus was isolated.
Ascomata dark brown, superficial on the substratum, globose, 100–180 (on average 150) μm diam, non-ostiolate, cleistothecial, not changing colour in 3 % KOH and lactic acid. Peridium multi-layered, pseudoparenchymatous, exoperidium up to 40 μm thick, composed of 5–8 layers of angular cells. Strings of thick-walled brown cells can take part in exoperidium formation. Asci thin-walled, without apical apparatus, unitunicate, scattered irregularly in the ascocarp, embedded in a gelatinous matrix. Ascospores not observed.
Asexual morph gliocladium-like. Conidiation abundant mostly plectonematogenous, partially nematogenous and is characterized by considerable variation in the branching patterns of primary and secondary conidiophores. Primary conidiophores in young cultures are predominantly penicillate, symmetrical, with monoverticillate arrangement, 30–60 μm long, phialides somewhat divergent or adpressed, solitary phialides 25–40 μm long are early formed laterally from simple and fasciculate vegetative hyphae. At maturity, secondary conidiophores with narrowly penicillate, biverticillate, branches including the phialides adpressed predominate, secondary conidiophores up to 70–80 μm long, with thick-walled supporting cell and thin-walled branches. Phialides variable, 20–40 μm long, gradually tapering towards the apex from 2 to 4 μm at the base to 1–2 μm at the tip, rather thin-walled. Conidia aseptate, cylindrical, with rounded apices, to oval, allantoid or tapering at times, aggregated in spherical slimy masses, later confluent in common slimy mass, 5–11 × 2–4 μm, smooth as observed by SEM, hyaline.
Culture characteristics — Colonies on alkaline agar (AA, pH 10.0–10.2) rather fast-growing, reaching 40–80 mm diam in 10 d at 25 °C. On MEA (pH 6.0) growth is limited, colonies reaching 10–15 mm diam in 10 d at 25 °C. Young colonies white; after 2–3 month on alkaline agar or after month on Czapek agar (CZ, pH 6.2) darkening punctuated zones appearing as a result of formation of ascomata. Colonies fluffy to woolly. Reverse colourless. Exudate absent. Decumbent vegetative hyphae thin-walled, hyaline, 0.5–2.0 μm wide. Mycelium consisting of hyaline, smooth-walled, septate hyphae, 1–3 μm wide, often fasciculate. Chlamydospores present in old cultures, predominantly in chains, darkening with age, forming bands and microsclerotia.
Specimen examined. Kenya, soda soil (pH 11) at the edge of Magadi Lake, Jan. 2013, S. Bondarenko (CBS H-21957 – holotype); culture ex-type MAG1 = CBS 137618 = VKM F-4582. Kenya, soda soil (pH 11) at the edge of Magadi Lake, Jan. 2013, S. Bondarenko, MAG3 = CBS 137620.
Notes: The comparative morphological chart for the Sodiomyces species is shown in Fig. 29.
Chordomyces Bilanenko et al., gen. nov. MycoBank MB 811265
Etymology: from the English chord and Latin mycetes, referring to the synnemata covering the surface of the fungal colony.
A genus of the Plectosphaerellaceae (Glomerellales, Hypocreomycetidae).
Colonies white, tufted, characterized by bundles of hyaline vegetative hyphae, or synnemata, gradually tapering towards the apex, sometimes repeatedly branched, tufts appearing bristling or fimbriate because of numerous radiating phialides. Contrasting with synnemata in Tilachlidium, the mycelial tufts are not separated from vegetative mycelium by sterile base. Conidiation abundant, mainly synnematogenous, partly plectonematogenous, the tufts bearing simple orthophialides and branched conidiophores. Stroma, setae and hyphopodia absent. Conidiophores branched (in different manner, often verticillate) or consisting of simple phialides. Conidiogenous cells phialidic (schizophialide more rarely), tapering to the apex, hyaline. Conidia catenate or adhering in a slimy mass, ellipsoid to cylindrical, rounded at ends, hyaline, smooth-walled.
Sexual morph: unknown.
Type: Chordomyces antarcticum Bilanenko et al.
Chordomyces antarcticum Bilanenko et al., sp. nov. MycoBank MB 811266 Figs. 30, 31.
Etymology: epithet taken from the Acremonium antarcticum (Speg.) D. Hawksw.
Colonies rather fast-growing on alkaline media, reaching 20–24 mm on AA (pH 10), 16–20 mm on OA in 10 d, slow-growing and restricted on MYA and PDA, white, tomentose to synnematous (especially on AA). Mycelium mainly superficial on AA, partly superficial, partly immersed on OA, webby and mainly immersed on CZ. Odour not present. Conidiation at first with solitary conidiophores, plectonematogenous, later in synnemata, reaching 10 mm in length, hyaline. Synnemata without differential sterile base, sometimes branched, appearing fimbriate due to radiating phialides. Thick parts of synnemata with a sterile tip, thiner ones (branches of synnemata) sporulating all over their length. Conidiophores often verticillate on short broad basal cells or consisting of simple phialides on thickened basal hyphae. Phialides (20)28–30(45) μm long, tapering from 2.5 to 3.0 μm near the base to 0.8–1.0 μm on the tip; tip with a short (length up to 2 μm) collarette, often proliferating sympodially, with branches often longer than the primary phialide tip (schizophialides), usually without a septum at the proliferation. Conidia held in slimy spherical heads (as seen with LM and cryoSEM), ellipsoidal to cylindrical, rounded at ends, with protuberant hilum at detachment (as seen with cryoSEM), hyaline, smooth-walled, 3.8–6.5 × 1.8–2.3 μm, L/W 2.2–2.8, predominantly 1-celled, 2-celled conidia with the middle septa are rare in occurrence. Chlamydospores absent.
Specimen examined. Russia, Altai, Kulunda Steppe, soda soil (pH 9.8) at the edge of Karakul Lake, Nov. 2002, M. Georgieva (CBS H-21956 – holotype); culture ex-type M27 = CBS 120045 = VKM FW-3041. Russia, Altai, Kulunda Steppe, soda soil (pH 9.4) at the edge of Petuchovskoe Lake, Aug. 2002, D. Y. Sorokin, A134 = CBS 137606. Russia, Altai, Kulunda Steppe, soda soil (pH 10.1) at the edge of Berdabay Lake, Aug. 2005, D. Y. Sorokin, A135 = CBS 137607. Russia, Trans-Baikal, soda soil (pH 10.1) at the edge of Nuhe-Nur Lake, Aug. 2003, M. P. Lebedeva, A137 = CBS 137608. Mongolia, North Gobi, Bayan-Zag area, soda soil (pH 9.3), Sep. 2003, I. A. Yamnova, A140 = CBS 137609. Kazakhstan, from Suaeda salsa on the coast of the Aral Lake, Dec. 2003. F. V. Sapozhnikov, A141 = CBS 137610. Kazakhstan, from Suaeda salsa on the coast of the Aral Lake, Dec. 2003, F. V. Sapozhnikov, A142 = CBS 137611. Mongolia, North Gobi, Bayan-Zag area, soda soil (pH 8.9), Aug. 2003, I. A. Yamnova, M10 = CBS 120042 = VKM FW-3039. Russia, Altai, Kulunda Steppe, soda soil (pH 9.8) at the edge of Uzkoe Lake, Aug. 2002, D. Y. Sorokin, M30 = CBS 120046 = VKM FW 3042. Russia, Altai, Kulunda Steppe, soda soil (pH 10) at the edge of Solyonoe Lake, Aug. 2002, D. Y. Sorokin, M31 = CBS 120047 = VKM FW-3906. Russia, Altai, Kulunda Steppe, soda soil (pH 9.9) at the edge of Petuhovskoe Lake, Aug. 2003, D. Y. Sorokin, M41 = CBS 120048 = VKM FW-3034. Russia, Altai, Kulunda Steppe, soda soil (pH 10.1) at the edge of Bezimyannoe Lake, Aug. 2002, D. Y. Sorokin, V213 = CBS 137630.
Notes: The sequences of 12 strains of Chordomyces antarcticum from soda soils differed from the CBS 987.87 isolate identified as Acremonium antarcticum (2 mutations in SSU across 1663 nucleotides, 3 mutations in LSU across 2023 nucleotides, 63 mutations in RPB2 across 1068 nucleotides, 8 mutations in TEF1-α across 903 nucleotides, 4 mutations in ITS across 479 nucleotides), which was isolated from lichen Hypogymnia physodes (the isolate seemed to be mycoparasitized by the hyphomycete Hobsonia christiansenii) in Luxembourg as was reported by Diederich et al. (1991). The species name come from Sporotrichum antarcticum Spegazzini (the type of this taxon is a herbarium specimen LPS 21677 in the herbarium of La Plata, Argentina), growing on Caloplaca cf. regalis (Vain.) Zahlbr. from South Orkney Islands and described by David Hawksworth as a new combination – Acremonium antarcticum Hawksworth (1979). The CBS 987.87 isolate from Luxembourg has no nomenclatural status and there is no ex-type isolate available of the old species name – Sporotrichum antarcticum Speg. It is therefore not possible to say exactly whether or not our isolates and CBS 987.87 have a relation to the type of A. antarcticum, and the matter of linking the name A. antarcticum to any existing isolates is quite problematical. Moreover, the name A. antarcticum is initially associated with lichenicolous species, but the species from Caloplaca might not be strictly lichenicolous but just an opportunist. Both CBS 987.87 and the type of A. antarcticum colonized lichen were parasitized by a more aggressive fungus, and it is possible that the Acremonium in these cases was a mycoparasite on the invader, rather than on the lichen itself. Our 12 isolates from soda soils are not lichenicolous and might be associated with roots of halophyte plants. Despite the differences in sequencing data between our 12 new isolates and CBS 987.87 (especially in RPB2), we established a new genus Chrodomyces with a single species Chordomyces antarcticum, accommodating the name A. antarcticum CBS 987.87 along with 12 our isolates. The clade harbouring 13 isolates received maximal statistical support in our Plectosphaerellaceae tree (LSU, ITS) (see Fig. 5).
Alternaria Section Soda Bilanenko et al., sect. nov. Mycobank MB 811267
Diagnosis: Section Soda contains simple or occasionally branched, short to moderately long, conidiophores with one conidiogenous locus. Apical or lateral short secondary conidiophores with a single conidiogenous locus may occur, conidiogenous tip can be enlarged. Conidia are solitary or in short to long, simple or branched chains, narrowly ellipsoid to long-ovoid or somewhat obclavate, moderate to very large in size, septate, with transverse and longitudinal septa, conspicuously constricted at most of the transverse septa. Microsclerotia or chlamydospores may occur, microsclerotia may germinate by conidial development. Sexual morphs have not been observed.
Notes: Section Soda contains 3 species (A. kulundii, A. petuchovskii, A. shukurtuzii), isolated from the soils at the different soda lakes in Kulunda Steppe in Western Siberia (Russia), pH of soils samples varied from 9.8 to 10.2. Species belonged to this section show strong alkalitolerant to facultative alkaliphilic type of the adaptation. Section Soda is well-defined from sister sections, Infectoriae, Chalastospora, and Embellisia based on partial sequences of SSU, LSU, RPB2, ITS, and GAPDH.
Type species: Alternaria kulundii Bilanenko et al.
Alternaria kulundii Bilanenko et al., sp. nov. Mycobank MB 811268, Fig. 32g–q, 33
Etymology: Name refers to the Kulunda steppe of Western Siberia in Russia, where the fungus was isolated from.
Colonies on CZ, MYA, PDA develop rather fast growing colonies (reaching 40–50 mm in diam after 7 d) with loose aerial brownish-pink-colored mycelium, light beige in colony center, reverse brownish. No sporulation on those media observed. Only microsclerotia and chlamydospores occured. Colonies on AA develop fast growing (colonies reaching 50–60 mm in diam after 7 d) pink-beige-colored mycelium, abundant light felt-fluffy mycelium in the center forming pinkish prostrate mycelium towards the margins of young colonies, reverse is similar in colour. No conidiation, chlamydospores occur. Colonies on SNA with a piece of autoclaved filter paper form poorly growing light beige mycelium. Microsclerotia are less developed in comparison with PDA. Conidia appeared within 7 d of incubation under CoolWhite fluorescent light with an 8 h photoperiod at room temperature. Conidiation on SNA unstable.
Conidiophores solitary simple, moderately long conidiophores with mostly one conidiogenous locus, conidiogenous tip is thick-walled and enlarged. Lateral short secondary conidiophores with a single conidiogenous locus may occur. Conidia solitary and in simple or branched chains, narrowly ellipsoid to long-ovoid or somewhat obclavate, dull pale tan to dark brown, conidial walls smooth to delicately roughened. Mature conidia large in size, mostly 80–110 × 15–25 μm, bulb-like in base, with 8–11, up to 14 transverse and several longitudinal septa, conspicuously constricted at the base transverse septa. Chains of oval chlamydospores form in young culture, microsclerotia are generated as a result of transverse and longitudinal septa formation, bulb-like, dark brown.
Specimen examined. Russia, Altai, the north of Kulunda Steppe, soda soil (pH 10.0), Aug. 2003, M. Georgieva (CBS H-21961 – holotype); culture ex-type M313 = CBS 137525. Russia, Altai, Kulunda Steppe, at the edge of Tanatar Lake, soda soil (pH 10.2) Aug. 2002, M. Georgieva, M310 = CBS 137522. Russia, Altai, Kulunda Steppe, at the edge of Uzkoe Lake, soda soil (pH 9.8) Aug. 2002, M. Georgieva, M309 = CBS 137521.
Alternaria petuchovskii Bilanenko et al., sp. nov. Mycobank MB 811269, Fig. 32a–f
Etymology: Name refers to the Petuchovskoe Lake in Kulunda steppe of Western Siberia (Russia), where the fungus was isolated.
Colonies on CZ, MYA, PDA develop slowly growing (colonies reaching up to 10–20 mm in diam after 7 d) light pink-beige colored mycelium. No sporulation, only microsclerotia or chlamydospores occur on those media. Colonies on AA develop fast growing (colonies reaching 60–70 mm in diam after 7 d) pink-beige-colored fluffy mycelium with prostrate mycelium in the center of colony and funiculose mycelium towards the margins; colony reverse in the similar colors. No conidiation, chlamydospores occur. Colonies on SNA with a piece of autoclaved filter paper form poorly growing light beige mycelium. Microsclerotia are less developed in comparison with PDA. Conidia appeared within 7 d incubation under Cool-White fluorescent light with an 8 h photoperiod at room temperature. Conidiation on SNA unstable.
Conidiophores solitary simple or occasionally branched, short or moderately long conidiophores with mostly one conidiogenous locus, conidiogenous tip is thick-walled and enlarged. Lateral short secondary conidiophores with a single conidiogenous locus may occur. Conidia solitary and in simple or branched chains, narrowly ellipsoid to long-ovoid or somewhat obclavate, with bulb-like base, dull pale tan to dark brown, conidial walls smooth to delicately roughened. Mature conidia large, mostly 90–110 × 20–30 μm, up to 120 μm length, with 9–11, up to 18 or even more transverse and several longitudinal septa, conspicuously constricted at the base transverse septa. Chlamydospores in chains and bulb-like brown-colored microsclerotia present. Microsclerotia are generated as a result of transverse and longitudinal septa formation, bulb-like, dark brown. Microsclerotia may germinate by conidial development.
Specimen examined. Russia, Altai, Kulunda Steppe, soda/chloride soil at the edge of the Petuchovskoe Lake (pH 9.9), Aug. 2003, M. Georgieva (CBS H-21959 – holotype); culture ex-type M304 = CBS 137517.
Alternaria shukurtuzii Bilanenko et al., sp. nov. Mycobank MB 811270, Fig. 32r–t
Etymology: Name refers to the Shukurtuz Lake in Kulunda steppe of Western Siberia (Russia), where the fungus was isolated.
Colonies on CZ, MYA, PDA develop moderately slowly growing (colonies reaching 20–30 mm in diam on MYA and PDA, 40–50 mm on CZ after 7 d) light pink-beige colored mycelium, brownish in center on CZ. No sporulation, only chlamydospores occur on those media. Colonies on AA develop fast growing (colonies reaching 50–60 mm in diam after 7 d) pink-beige-coloured fluffy mycelium, reverse in the similar colours. No sporulation, only chlamydospores occur. Colonies on SNA with a piece of autoclaved filter paper form poorly growing mycelium. Chlamydospores occur. Conidia appeared within 7 d incubation under CoolWhite fluorescent light with an 8 h photoperiod at room temperature. Conidiation on SNA unstable. No microsclerotia observed.
Conidiophores solitary simple or occasionally branched, with mostly one conidiogenous locus, conidiogenous tip is thick-walled and enlarged. Lateral short secondary conidiophores with a single conidiogenous locus may occur. Conidia solitary and in simple or branched chains, narrowly ellipsoid to long-ovoid, dull pale tan to dark brown, conidial walls smooth to delicately roughened. Mature conidia mostly 60–80 × 7–10 μm, with 7–9 transverse and 0–1 longitudinal septa, not conspicuously constricted at the septa. Chlamydospores occur.
Specimen examined. Russia, Altai, Kulunda Steppe, the edge of the Shukurtuz Lake, soda/chloride/sulfate soil (pH 9.9), Aug. 2002, M. Georgieva (CBS H-21960 – holotype); culture ex-type M307 = CBS 137520.
Discussion
The present study systematically characterizes the alkaliphilic and alkalitolerant filamentous fungi isolated from alkaline (soda) soils. Until now, our understanding of the biodiversity of the alkaliphiles and alkalitolerants has been based on occasional reports that lack both sequencing data and detailed morphological descriptions. The demand for sequencing data is especially high, since most alkaliphiles and alkalitolerants have only simple asexual morphology (polyphyletic acremonium-like development) impairing proper identification of the species. The acremonium-, verticillium-, and gliocladium-type asexual morphological development has also been demonstrated for most of our alkaliphilic and strongly alkalitolerant fungi. To strengthen our descriptions and analyses, we coupled our phylogenetic analyses to the morphology data and pH-dependent growth rate data for essentially every isolate we obtained. The growth experiments at various pH values helped us to assess the potential ecological contribution of the isolated fungi in soda soils and to discriminate between accidental transient species and effective alkaliphiles. Though the growth patterns were often too complex to explicitly interpret, we were able to assign the patterns into one of five ‘growth types’: obligate alkaliphiles and facultative alkaliphiles, strong, moderate and weak alkalitolerants. In several cases this estimation was complicated by the fact that the pH-tolerance is highly affected by the presence of extra Na+ in the growth medium. Based on such adopted classification, we were able to create a combined framework of phylogenetic, morphological and growth data that could serve as a foundation for future studies on physiology and evolution of alkaliphilic filamentous fungi.
Overall, we showed that soda soils are in fact inhabited with a substantial number of filamentous fungal species, which are able not only to withstand high ambient pH, but for some of them high pH is essential for optimal growth. One of the prominent conclusions from our data is that alkaliphilic and alkalitolerant fungi are polyphyletic and spread across several subphyla in Ascomycota. This result suggests independent evolution and several origins of the alkaliphilic trait in filamentous fungi. Still, most of the alkaliphilic species are within the Hypocreomycetidae. This is the taxonomic location where obligate and facultative alkaliphiles are overrepresented. Notably, Emericellopsis (Bionectriaceae) and Plectosphaerellaceae lineages show particular enrichment; nearly half of our isolates belong to either of those two groups. The facultative alkaliphile Emericellopsis alkalina lineage appears to have evolved from marine isolates of Emericellopsis (Grum-Grzhimaylo et al. 2013b). Even though the highest frequency of alkalitolerant fungi was observed within Hypocreomycetidae, we found two obligate alkaliphiles, which belong to the Thielavia of the Chaetomiaceae (Sordariomycetidae) and over ten strong alkalitolerant isolates are nested within the Pleosporaceae (Dothideomycetes).
Interestingly, in the present study, we obtained two Alternaria sect. Phragmosporae isolates (M301 and M312); to our knowledge, this is the only section of Alternaria containing marine-borne isolates, providing extra evidence for the link between marine habitats and soda soils − terrestrial counterparts with similar abiotic conditions. Allied to the Alternaria, six of our isolates are taxonomically spread around Pleospora. Several close reference strains, e.g., Pleospora halimiones CBS 432.77, Chaetodiplodia sp. CBS 453.68 and others, have been isolated from the associated halophytic grasses. The enriched biodiversity of the alkalitolerant fungal strains provides evidence for the ecological overlap between the halophyte-associated isolates and soda lakes in the Pleospora-related species.
The growth experiments using common media coupled with the pH preference test in race tubes allowed us to study the structure of the soda soils fungal community. This approach showed differences in the pH and Na+ preferences of our isolates, which enabled us to reveal true alkaliphiles and distinguish them from the moderate and weak alkalitolerant species. In many cases the preference to grow at high ambient pH relative to neutral pH, coincided with the presence of extra Na+ in the medium, which points towards the fact that high pH and high salts stresses are linked, and probably have co-evolved. Also alkaliphilic bacteria require Na+ cations to tolerate high pH, as the Na+-pumps seem to be one of the major players in pH homeostasis of the cell (Krulwich et al. 2011). Extra evidence for such a tight association has been shown in Fusarium oxysporum (Caracuel et al. 2003). For example, as seen in Fig. 6, strong alkalitolerant Chordomyces isolates prefer alkaline agar (AA) medium (pH 10, which contains 5 g/l NaCl) rather than common neutral media like PDA, MYA without extra NaCl. However, if the NaCl concentration is kept constant, but the pHs are varied, as we set up in race tubes – we see no clear preference to ambient pH, as seen in graphs in Fig. 6. That observation implies that for Chordomyces (Plectosphaerellaceae), Na+ is necessary for optimal growth regardless of the tested ambient pH. Similar observations hold for our Verticillium zaregamsianum, several Alternaria sect. Soda strains, and to some extent, for Acrostalagmus luteoalbus strains. Conversely, we have recovered truly alkaliphilic fungi that prefer high ambient pH regardless of the presence of extra Na+ in the growth medium. For instance, species of Sodiomyces (Plectosphaerellaceae) and sterile Thielavia sp. (Chaetomiaceae) showed clear alkaliphilic phenotype both in the media complemented with NaCl and without it (Figs. 9, 16). As Chordomyces and Sodiomyces belong to taxonomically distinct clades with strong support values, it could be that there is different sensitivity to Na+ ions in these species, which results in utilizing different strategies of dealing with high ambient pH. Strikingly, our newly defined section Soda in Alternaria prominently stands out from the rest, implying specific evolution of this clade within Alternaria of strong alkalitolerance for the members of section Soda.
Our growth experiments also revealed moderate and weak alkalitolerants such as Scopulariopsis brevicaulis, Alternaria alternata (sect. Alternata), Purpureocillium lilacinum, Penicillium sp., Cladosporium sphaerospermum, and Fusarium sp. with the growth optimum at neutral or below neutral pH values. These species partly also appear in the existing reports on the alkalitolerant and halotolerant fungi (Kladwang et al. 2003; Gunde-Cimerman et al. 2009). Darkly pigmented Cladosporium sphaerospermum is a ubiquitous fungus, often isolated from hypersaline environments throughout the world (Zalar et al. 2007; Bensch et al. 2012); however, in our study this species was not observed frequently and did not exhibit alkaliphilic abilities. The inability of the above named weak and moderate alkalitolerants to grow optimally at high pH values as well as their low abundancy, leads us to consider them as transition species in the alkaline soils, as they are also known to inhabit neutral soils worldwide.
The morphology studies of our alkalitolerant and alkaliphilic fungi provided evidence for the relation between the morphotype of the strain and its ability to tolerate the pH. The strong alkalitolerants and effective alkaliphiles, e.g., Emericellopsis alkalina, Sodiomyces species, Acrostalagmus luteoalbus, Verticillium zaregamsianum display acremonium-, verticillium-, and gliocladium-type asexual morphology (see examples in Figs. 8, 12 and 28) with conidia being clumped in heads, as better seen with cryogenic scanning electron microscopy. The same type of asexual morphology is observed in moderate alkalitolerants, sister to the Emericellopsis lineage, like Sarocladium sp. and Acremonium sclerotigenum. Ascomata of the isolates Sodiomyces species and Emericellopsis alkalina are of cleistothecial type, see Figs. 27, 28 and 29, and in Grum-Grzhimaylo et al. (2013a, b). Ascospores in those isolates are being formed in asci, walls of which lyse shortly, releasing ascospores in the ascoma matrix. Therefore, by the time the ascoma is fully developed, the ascospores are sunk in the common matrix with no signs of asci left, which in turn would release through the crack in the ascoma. Such type of ascomata, ascospore development and release might be adaptive to harsh alkaline and salty environments. Conidia clumping by slime and common ascospore matrix might create a favorable local environment prolonging the viability of both asexual and sexual spores. Slime can act as a buffering barrier lowering local pH around the spores, and may provide extra protection against osmotic shock. In a similar way, the halophilic basidiomycete Wallemia ichthyophaga, forms sarcina-like cell clumps (Zalar et al. 2005). However, we saw no such gluing slime of conidia in strong alkalitolerants of Alternaria sect. Soda species. They rather often formed abundant dark thick-walled resting mycelium along with the dark microsclerotia, which are known to be stress-related morphological structures (Figs. 32, 33).
The ecology of the obligate alkaliphilic species like members of Sodiomyces is disputable. Our previous growth experiments of Sodiomyces alkalinus ex-type strain CBS 110278 showed the preference towards complex plant material carbon sources, like cotton seeds, alfalfa meal and soybean hulls. That is not surprising, as those substrates are complemented with proteins and vitamins, broadening the diet for the fungus. Most probable substrates for fungi in soda soils include plant biomass from halophytic grasses, chitin carapaces and protein-rich eggs of brine shrimps, and possibly bird feathers. Sodiomyces might be tightly associated with bacteria as we often encountered them along the fungal hypha, which makes it challenging to separate from each other. Another large group of our strains, facultative alkaliphilic Emericellopsis alkalina is associated with marine-borne isolates, probably utilizing similar strategies as coping with marine environments, enabling them to thrive in soda soils as well (Grum-Grzhimaylo et al. 2013b). Acrostalagmus luteoalbus and Verticillium zaregamsianum isolated from soda soils display facultative alkaliphilic and a strong alkalitolerant phenotype. The first one is known to be a widely distributed fungus inhabiting the rizosphere (Domsch et al. 2007), while the latter has only been found in Japan associated with lettuce and ten-week stocks (Inderbitzin et al. 2011b). The facultative alkaliphilic Chordomyces antarcticum fungus is enigmatic. All 12 isolates group unambiguously to the known CBS 987.87 isolate. This isolate was recovered from the Hypogymnia physodes lichen thallus, which was already infected by another fungus. It is therefore unclear whether the CBS 987.87 isolate was mycoparasitizing or was involved in lichen degradation. Acremonium furcatum CBS 122.42 ex-type and Acremonium stromaticum CBS 863.73 ex-type are sister species to Chordomyces antarcticum species group. By our unpublished data, Acremonium furcatum has been isolated frequently from the lichen thalli indeed; however, the type strains of A. furcatum and A. stromaticum were recovered from dune sands and rhizome, respectively. What makes fungi from soda soils and lichen thalli to be possibly associated is a mystery to us.
The present study has provided insight on the morphological and phylogenetic properties of alkalitolerant and alkaliphilic filamentous fungi recovered from soda soils. We attempted to make a systematic overview of the fungal biodiversity in soda soils; however, we expect different fungi to appear depending on the recovery medium. Therefore, this study is a starting point for subsequent recoveries of alkalitolerant and alkaliphilic fungi in the future. Notably, we have shown comprehensive polyphyletic diversity of filamentous fungi at soda lakes, which we hope will initiate more research and lead to subsequent discoveries of this peculiar ecological group of fungi. It is interesting to speculate if future isolations would confirm our observations about the several dominant lineages of strong alkalitolerants and effective alkaliphiles (i. e. families Plectosphaerellaceae and Pleosporaceae; genera Emericellopsis and Thielavia). Another intriguing aspect is the physiology of these fungi. Alkaliphilic fungi are likely to possess industrially valuable metabolites and enzymes, which are active at high pH. Whether alkaliphiles in different taxonomic lineages display convergent evolution in the alkaliphilic traits, or utilize different mechanisms enabling them to survive and grow best at high pH, remains to be discovered.
References
Antony CP, Murrell JC, Shouche YS (2012) Molecular diversity of methanogens and identification of Methanolobus sp. as active methylotrophic Archaea in Lonar crater lake sediments. FEMS Microbiol Ecol 81:43–51. doi:10.1111/j.1574-6941.2011.01274.x
Antony CP, Kumaresan D, Hunger S, Drake HL, Murrell JC, Shouche YS (2013) Microbiology of Lonar Lake and other soda lakes. ISME J 7:468–476. doi:10.1038/ismej.2012.137
Aono R (1990) Taxonomic distribution of alkali-tolerant yeasts. Syst Appl Microbiol 13:394–397. doi:10.1016/S0723-2020(11)80239-0
Bensch K, Braun U, Groenewald JZ, Crous PW (2012) The genus Cladosporium. Stud Mycol 72:1–401. doi:10.3114/sim0003
Berbee ML, Pirseyedi M, Hubbard S (1999) Cochliobolus phylogenetics and the origin of known, highly virulent pathogens, inferred from ITS and glyceraldehyde-3-phosphate dehydrogenase gene sequences. Mycologia 91:964–977. doi:10.2307/3761627
Cannon PF, Buddie AG, Bridge PD, de Neergaard E, Lübeck M, Askar MM (2012) Lectera, a new genus of the Plectosphaerellaceae for the legume pathogen Volutella colletotrichoides. Mycokeys 3:23–36. doi:10.3897/mycokeys.3.3065
Caracuel Z, Casanova C, Roncero MIG, Di Pietro A, Ramos J (2003) pH response transcription factor PacC controls salt stress tolerance and expression of the P-Type Na+-ATPase ena1 in Fusarium oxysporum. Eukaryot Cell 2:1246–1252. doi:10.1128/EC.2.6.1246-1252.2003
Crous PW, Gams W, Stalpers JA, Robert V, Stegehuis G (2004) MycoBank: an online initiative to launch mycology into the 21st century. Stud Mycol 50:19–22
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9:772. doi:10.1038/nmeth.2109
De Hoog GS, Ahmed SA, Najafzadeh MJ, Sutton DA, Keisari MS, Fahal AH, Eberhardt U, Verkleij GJ, Xin L, Stielow B, van de Sande WW (2013) Phylogenetic findings suggest possible new habitat and routes of infection of human eumycetoma. PLoS Negl Trop Dis 7:e2229. doi:10.1371/journal.pntd.0002229
Di Menna ME (1959) Some physiological characters of yeasts from soils and allied habitats. J Gen Microbiol 20:13–23. doi:10.1099/00221287-20-1-13
Diederich P, Sérusiaux E, van den Boom P (1991) Lichens et champignons lichénicoles nouveaux ou intéressants pour la flore de la Belgique et des régions voisines. V. Lejeunia, nouvelle série 136:1–47
Domsch KH, Gams W, Anderson T-H (2007) Compendium of soil fungi. IHW-Verl, Eching
Dong H, Zhang G, Jiang H, Yu B, Chapman LR, Lucas CR, Fields MW (2006) Microbial diversity in sediments of saline Qinghai Lake, China: linking geochemical controls to microbial ecology. Microb Ecol 51:65–82. doi:10.1007/s00248-005-0228-6
Duckworth AW, Grant WD, Jones BE, Steenbergen RV (1996) Phylogenetic diversity of soda lake alkaliphiles. FEMS Microbiol Ecol 19:181–191. doi:10.1111/j.1574-6941.1996.tb00211.x
Goto S, Aono R, Sugiyama J, Horikoshi K (1981) Exophiala alcalophila, a new black, yeast-like hyphomycete with an accompanying Phaeococcomyces alcalophilus morph, and its physiological characteristics. Trans Mycol Soc Jpn 22:429–439
Grum-Grzhimaylo AA, Debets AJM, van Diepeningen AD, Georgieva ML, Bilanenko EN (2013a) Sodiomyces alkalinus, a new holomorphic alkaliphilic ascomycete within the Plectosphaerellaceae. Persoonia 31:147–158. doi:10.3767/003158513X673080
Grum-Grzhimaylo AA, Georgieva ML, Debets AJM, Bilanenko EN (2013b) Are alkalitolerant fungi of the Emericellopsis lineage (Bionectriaceae) of marine origin? IMA Fungus 4:213–228. doi:10.5598/imafungus.2013.04.02.07
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704. doi:10.1080/10635150390235520
Gunde-Cimerman N, Ramos J, Plemenitaš A (2009) Halotolerant and halophilic fungi. Mycol Res 113:1231–1241. doi:10.1016/j.mycres.2009.09.002
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hammer UT (1986) Saline Lake ecosystems of the world. Monographiae biologicae 59. Dr W Junk Publishers, Dordrecht
Hawksworth DL (1979) The lichenicolous hyphomycetes. Bulletin of the British Museum (Natural history). Bot Ser 6:183–300
Horikoshi K (1999) Alkaliphiles: some applications of their products for biotechnology. Microbiol Mol Biol Rev 63:735–750
Horikoshi K, Akiba T (1982) Alkalophilic microorganisms: a new microbial world. Japan Scientific Societies Press, Tokyo and Springer-Verlag, New York
Huelsenbeck JP, Ronquist F (2001) MrBayes: bayesian inference of phylogenetic trees. Bioinformatics 17:754–755
Hyde KD, Jones EBG, Liu JK, Ariyawansa H, Boehm E, Boonmee S, Braun U, Chomnunti P, Crous PW, Dai DQ, Diederich P, Dissanayake A, Doilom M, Doveri F, Hongsanan S, Jayawardena R, Lawrey JD, Li YM, Liu YX, Lücking R, Monkai J, Muggia L, Nelsen MP, Pang KL, Phookamsak R, Senanayake IC, Shearer CA, Suetrong S, Tanaka K, Thambugala KM, Wijayawardene NN, Wikee S, Wu HX, Zhang Y, Aguirre-Hudson B, Alias SA, Aptroot A, Bahkali AH, Bezerra JL, Bhat DJ, Camporesi E, Chukeatirote E, Gueidan C, Hawksworth DL, Hirayama K, Hoog SD, Kang JC, Knudsen K, Li WJ, Li XH, Liu ZY, Mapook A, McKenzie EHC, Miller AN, Mortimer PE, Phillips AJL, Raja HA, Scheuer C, Schumm F, Taylor JE, Tian Q, Tibpromma S, Wanasinghe DN, Wang Y, Xu JC, Yacharoen S, Yan JY, Zhang M (2013) Families of Dothideomycetes. Fungal Divers 63:1–313. doi:10.1007/s13225-013-0263-4
Hyde KD, Nilsson RH, Alias SA, Ariyawansa HA, Blair JE, Cai L, de Cock AWAM, Dissanayake AJ, Glockling SL, Goonasekara ID, Gorczak M, Hahn M, Jayawardena RS, van Kan JAL, Laurence MH, Lévesque CA, Li X, Liu JK, Maharachchikumbura SSN, Manamgoda DS, Martin FN, McKenzie EHC, McTaggart AR, Mortimer PE, Nair PVR, Pawłowska J, Rintoul TL, Shivas RG, Spies CFJ, Summerell BA, Taylor PWJ, Terhem RB, Udayanga D, Vaghefi N, Walther G, Wilk M, Wrzosek M, Xu JC, Yan JY, Zhou N (2014) One stop shop: backbones trees for important phytopathogenic genera: I. Fungal Divers 67:21–125. doi:10.1007/s13225-014-0298-1
Inderbitzin P, Bostock RM, Davis RM, Usami T, Platt HW, Subbarao KV (2011a) Phylogenetics and taxonomy of the fungal vascular wilt pathogen Verticillium, with the descriptions of five new species. PLoS One 6:e28341. doi:10.1371/journal.pone.0028341
Inderbitzin P, Davis RM, Bostock RM, Subbarao KV (2011b) The ascomycete Verticillium longisporum is a hybrid and a plant pathogen with an expanded host range. PLoS One 6:e18260. doi:10.1371/journal.pone.0018260
Jones BE, Grant WD, Duckworth AW, Owenson GG (1998) Microbial diversity of soda lakes. Extremophiles 2:191–200. doi:10.1007/s007920050060
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. doi:10.1093/molbev/mst010
Keresztes ZG, Felföldi T, Somogyi B, Székely G, Dragoş N, Márialigeti K, Bartha C, Vörös L (2012) First record of picophytoplankton diversity in Central European hypersaline lakes. Extremophiles 16:759–769. doi:10.1007/s00792-012-0472-x
Kis-Papo T, Weig AR, Riley R, Peršoh D, Salamov A, Sun H, Lipzen A, Wasser SP, Rambold G, Grigoriev IV, Nevo E (2014) Genomic adaptations of the halophilic Dead Sea filamentous fungus Eurotium rubrum. Nat Commun 5. doi: 10.1038/ncomms4745
Kladwang W, Bhumirattana A, Hywel-Jones N (2003) Alkaline-tolerant fungi from Thailand. Fungal Divers 13:69–83
Kompantseva EI, Komova AV, Novikov AA, Kostrikina NA (2012) Rhodovulum tesquicola sp. nov., a haloalkaliphilic purple non-sulfur bacterium from brackish steppe soda lakes. Int J Syst Evol Microbiol 62:2962–2966. doi:10.1099/ijs. 0.035857-0
Krulwich TA, Sachs G, Padan E (2011) Molecular aspects of bacterial pH sensing and homeostasis. Nat Rev Microbiol 9:330–343. doi:10.1038/nrmicro2549
Kruys Å, Huhndorf SM, Miller AN (2014) Coprophilous contributions to the phylogeny of Lasiosphaeriaceae and allied taxa within Sordariales (Ascomycota, Fungi). Fungal Divers 1–13. doi: 10.1007/s13225-014-0296-3
Lanzén A, Simachew A, Gessesse A, Chmolowska D, Jonassen I, Øvreås L (2013) Surprising prokaryotic and eukaryotic diversity, community structure and biogeography of Ethiopian soda lakes. PLoS One 8:e72577. doi:10.1371/journal.pone.0072577
Lisichkina GA, Bab’eva IP, Sorokin DY (2003) Alkalitolerant yeasts from natural biotopes. Microbiology 72:618–620. doi:10.1023/A:1026011821021
Liu TT, Hub DM, Liu F, Cai L (2013) Polyphasic characterization of Plectosphaerella oligotrophica, a new oligotrophic species from China. Mycoscience 54:387–393. doi:10.1016/j.myc.2013.01.003
Maddison WP, Maddison DR (2011) Mesquite: a modular system for evolutionary analysis. Version 2.75. http://mesquiteproject.org
Mueller GM, Bills GF, Foster MS (2004) Biodiversity of fungi: inventory and monitoring methods. Elsevier Academic Press, Amsterdam
Nagai K, Sakai T, Rantiatmodjo RM, Suzuki K, Gams W, Okada G (1995) Studies on the distribution of alkalophilic and alkali-tolerant soil fungi I. Mycoscience 36:247–256. doi:10.1007/BF02268598
Nagai K, Suzuki K, Okada G (1998) Studies on the distribution of alkaliphilic and alkali-tolerant soil fungi II: fungal flora in two limestone caves in Japan. Mycoscience 39:293–298. doi:10.1007/BF02464011
Nagy L, Peterfi LS, Momeu L (2006) Structure, seasonal dynamics and vertical distribution of diatoms in lake Durgau, near Turda (Cluj County, Romania). Contribuţii Botanice XLI:63–69
Nirenberg HI (1976) Untersuchungen über die morphologische und biologische Differenzierung in der Fusarium-Sektion Liseola. Mitteilungen aus der Biologischen Bundesanstalt für Land- und Forstwirtschaft Berlin-Dahlem 169:1–117
Okada G, Niimura Y, Sakata T, Uchimura T, Ohara N, Suzuki H, Kozaki M (1993) Acremonium alcalophilum, a new alkalophilic cellulolytic hyphomycete. Trans Mycol Soc Jpn 34:171–185
Perkins DD, Pollard VC (1986) Linear growth rates of strains representing ten Neurospora species. Fungal Genet Newsl 33:41–43
Rambaut A, Drummond AJ (2007) Tracer v. 1.5. Computer program and documentation distributed by the authors: http://beast.bio.ed.ac.uk/Tracer
Rees HC, Grant WD, Jones BE, Heaphy S (2004) Diversity of Kenyan soda lake alkaliphiles assessed by molecular methods. Extremophiles 8:63–71. doi:10.1007/s00792-003-0361-4
Sandoval-Denis M, Sutton DA, Fothergill AW, Cano-Lira J, Gené J, Decock CA, de Hoog GS, Guarro J (2013) Scopulariopsis, a poorly known opportunistic fungus: spectrum of species in clinical samples and in vitro responses to antifungal drugs. J Clin Microbiol 51:3937–3943. doi:10.1128/JCM. 01927-13
Schneider T, Herzig A, Koinig KA, Sommaruga R (2012) Copepods in turbid shallow soda lakes accumulate unexpected high levels of carotenoids. PLoS One 7:e43063. doi:10.1371/journal.pone.0043063
Sorokin DY, Muyzer G (2010) Desulfurispira natronophila gen. nov. sp. nov.: an obligately anaerobic dissimilatory sulfur-reducing bacterium from soda lakes. Extremophiles 14:349–355. doi:10.1007/s00792-010-0314-7
Sorokin DY, Panteleeva AN, Tourova TP, Kaparullina EN, Muyzer G (2011) Natronoflexus pectinivorans gen. nov. sp. nov., an obligately anaerobic and alkaliphilic fermentative member of Bacteroidetes from soda lakes. Extremophiles 15:691–696. doi:10.1007/s00792-011-0399-7
Spatafora JW, Sung GH, Johnson D, Hesse C, O’Rourke B, Serdani M, Spotts R, Lutzoni F, Hofstetter V, Miadlikowska J, Reeb V, Gueidan C, Fraker E, Lumbsch T, Lücking R, Schmitt I, Hosaka K, Aptroot A, Roux C, Miller AN, Geiser DM, Hafellner J, Hestmark G, Arnold AE, Büdel B, Rauhut A, Hewitt D, Untereiner WA, Cole MS, Scheidegger C, Schultz M, Sipman H, Schoch CL (2006) A five-gene phylogeny of Pezizomycotina. Mycologia 98:1018–1028. doi:10.3852/mycologia.98.6.1018
Steiman R, Ford L, Ducros V, Lafond JL, Guiraud P (2004) First survey of fungi in hypersaline soil and water of Mono Lake area (California). Anton Leeuw Int J G 85:69–83. doi:10.1023/B:ANTO.0000020150.91058.4d
Stöver BC, Müller KF (2010) TreeGraph 2: Combining and visualizing evidence from different phylogenetic analyses. BMC Bioinformatics 11:7. doi:10.1186/1471-2105-11-7
Sukumaran J, Holder MT (2010) DendroPy: A Python library for phylogenetic computing. Bioinformatics 26:1569–1571. doi:10.1093/bioinformatics/btq228
Summerbell RC, Gueidan C, Schroers HJ, De Hoog GS, Starink M, Rosete YA, Guarro J, Scott JA (2011) Acremonium phylogenetic overview and revision of Gliomastix, Sarocladium, and Trichothecium. Stud Mycol 68:139–162. doi:10.3114/sim.2011.68.06
Wang XW, Wang XL, Liu FJ, Zhao XM, Li J, Cai L (2014) Phylogenetic assessment of Chaetomium indicum and allied species, with the introduction of three new species and epitypification of C. funicola and C. indicum. Mycol Progress 1–14. doi: 10.1007/s11557-013-0955-x
Wani AA, Surakasi VP, Siddharth J, Raghavan RG, Patole MS, Ranade D, Shouche YS (2006) Molecular analyses of microbial diversity associated with the Lonar soda lake in India: an impact crater in a basalt area. Res Microbiol 157:928–937. doi:10.1016/j.resmic.2006.08.005
Woudenberg JHC, Groenewald JZ, Binder M, Crous PW (2013) Alternaria redefined. Stud Mycol 75:171–212. doi:10.3114/sim0015
Xiong J, Liu Y, Lin X, Zhang H, Zeng J, Hou J, Yang Y, Yao T, Knight R, Chu H (2012) Geographic distance and pH drive bacterial distribution in alkaline lake sediments across Tibetan Plateau. Environ Microbiol 14:2457–2466. doi:10.1111/j.1462-2920.2012.02799.x
Zalar P, de Hoog GS, Schroers HJ, Frank JM, Gunde-Cimerman N (2005) Taxonomy and phylogeny of the xerophilic genus Wallemia (Wallemiomycetes and Wallemiales, cl. et ord. nov.). Anton Leeuw Int J G 87:311–328. doi:10.1007/s10482-004-6783-x
Zalar P, de Hoog GS, Schroers HJ, Crous PW, Groenewald JZ, Gunde-Cimerman N (2007) Phylogeny and ecology of the ubiquitous saprobe Cladosporium sphaerospermum, with descriptions of seven new species from hypersaline environments. Stud Mycol 58:157–183. doi:10.3114/sim.2007.58.06
Zare R, Gams W, Schroers HJ (2004) The type species of Verticillium is not congeneric with the plant-pathogenic species placed in Verticillium and it is not the anamorph of ‘Nectria’ inventa. Mycol Res 108:576–582. doi:10.1017/S0953756204009839
Zare R, Gams W, Starink-Willemse M, Summerbell RC (2007) Gibellulopsis, a suitable genus for Verticillium nigrescens, and Musicillium, a new genus for V. theobromae. Nova Hedwigia 85:463–489. doi:10.1127/0029-5035/2007/0085-0463
Zavarzin GA (1993) Epicontinental soda lakes as probable relict biotopes of terrestrial biota formation. Microbiology 62:473–479
Zwickl DJ (2006) Genetic algorithm approaches for the phylogenetic analysis of large biological sequence datasets under the maximum likelihood criterion. University of Texas at Austin, Dissertation
Acknowledgments
We thank Tiny Franssen-Verheijen for her invaluable help with the cryogenic SEM imaging. Marijke Slakhorst and Bertha Koopmanschap are thanked for their technical assistance. We are much obliged for the discussions on the Acremonium antarcticum species given by David Hawksworth. We acknowledge Joyce Woudenberg for her suggestions regarding the Alternaria isolates and help with the imaging. Trix Merkx helped with the strains deposition for which we are grateful. Denis Landin is thanked for the assistance with the image processing. Finally, we are indebted to Dmitry Sorokin for providing the soil samples and initiation of the study in the field of the biodiversity of microorganisms in soda soils. The work was supported by the Laboratory of Genetics at Wageningen University (The Netherlands), RSCF grant № 14-50-00029, RFBR grants № 15-04-06975, № 15-04-06260 (Russia).
Conflict of interest
The authors declare no conflict of interest in the present manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
Ascomycota tree GenBank Accessions (PDF 436 kb)
ESM 2
Scopulariopsis tree GenBank Accessions (PDF 126 kb)
ESM 3
Plectosphaerellaceae tree GenBank Accessions (PDF 119 kb)
ESM 4
Verticillium sensu stricto tree GenBank Accessions (PDF 113 kb)
ESM 5
Sodiomyces tree GenBank Accessions (PDF 106 kb)
ESM 6
Pleosporaceae tree GenBank Accessions (PDF 129 kb)
ESM 7
Lasiosphaeriaceae & Chaetomiaceae GenBank Accessions (PDF 120 kb)
ESM 8
Alternaria section Soda GenBank Accessions (PDF 112 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Grum-Grzhimaylo, A.A., Georgieva, M.L., Bondarenko, S.A. et al. On the diversity of fungi from soda soils. Fungal Diversity 76, 27–74 (2016). https://doi.org/10.1007/s13225-015-0320-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13225-015-0320-2