Abstract
Taxol (paclitaxel) is a highly-oxygenated diterpenoid natural product first isolated from the pacific yew tree (Taxus brevifolia). It is one of the most widely used anticancer drugs. Soon after the discovery of its unique mode of action and the resulting high demand, an extensive search was initiated for alternative sources to replace the slow-growing and scarce pacific yew. Thus far, however, Taxol and related compounds have only been found in the genus Taxus, which comprises a small number of slow-growing plants with a broad but generally isolated geographical distribution. In 1993, Stierle and colleagues reported the independent biosynthesis of Taxol in an endophytic fungus isolated from Taxus brevifolia, which resulted in more than 160 subsequent publications and patents addressing the biosynthesis of Taxol and related taxanes by microorganisms. The literature on fungal taxane synthesis contains numerous inconsistencies, prompting us to thoroughly re-examine Taxol biosynthesis in endophytic fungi associated with Taxus spp. Using a combination of phytochemistry, molecular biology and genome sequencing, we were unable to find any evidence for independent taxane biosynthesis in any of the endophytes, including the isolate described in the original publication (Taxomyces andreanae) and several more recent isolates from Taxus trees. Our findings therefore resolve a long-standing mystery concerning the evolution of a complex terpenoid biosynthetic pathway in two distantly-related organisms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Taxol (paclitaxel), a highly-oxygenated diterpenoid natural product first isolated from the pacific yew tree (Taxus brevifolia), is arguably one of the most successful anticancer drugs of all time (Suffness and Wall 1995; Brown 2003). The limited supply of Taxol and related compounds made pharmaceutical development a major challenge (Suffness and Wall 1995). Therefore, soon after its unique mode of action was discovered, an extensive search was launched to find alternative sources because the pacific yew is slow-growing and scarce (Croom 1995; Itokawa 2003). For a long time, Taxol biosynthesis was thought to be restricted to the ancient Taxus genus (Taxaceae, Coniferales), which comprises 11 geographically-isolated species. Fossil records indicate that yew trees have existed for more than 200 million years with little evolutionary change. Taxus grandis from the Quaternary period shared many characteristics with the modern yew, Taxus baccata (Croom 1995). Considering the age and isolation of the genus together with the extreme longevity of individual members (some yew trees live more than 3,000 years), it was believed that the Taxol metabolic pathway was unique to this genus. Members of the closely related genera Pseudotaxus and Austrotaxus do not synthesize Taxol, although simple taxanes lacking the oxetane or D-ring structure have been isolated from Austrotaxus spicata, the only member of the genus Austrotaxus, which is regarded as a primitive ancestor of Taxus (Guéritte-Voegelein et al. 1987). Pseudotaxus spp. do not produce taxanes at all.
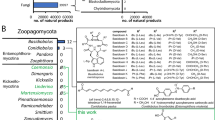
The evolutionary advantage of Taxol biosynthesis in yew trees remains a mystery, particularly in light of the production of the highly cardiotoxic but chemically less complex taxines by several species. More than 360 taxanes have been identified in different Taxus spp. (Baloglu and Kingston 1999; Itokawa 2003) but Taxol (if present at all) represents only a minor fraction of the total taxane complement. The biosynthesis of Taxol and other taxanes is well characterized (Croteau et al. 2006; Kaspera and Croteau 2006; Heinig and Jennewein 2009) and appears to follow an anastamosing pattern that yields several physiologically-active products as well as metabolic dead ends (Fig. 1). Several of the key steps involved in the 20 or more enzymatic reactions required to produce Taxol have been characterized at the biochemical and genetic levels (Croteau et al. 2006; Jennewein et al. 2004b). The biosynthetic pathway, starting with the cyclization of geranylgeranyl diphosphate to form taxa-4(5),11(12)-diene, involves enzymes from several different classes that are located in several different cellular compartments, including the plastid, endoplasmic reticulum and cytosol.
Proposed Taxol/taxoid biosynthesis pathway in Taxus spp. based on the cDNA library sequencing results of taxoid-producing Taxus plant cell cultures and known gene functions. The biosynthesis of Taxol and other taxoids appears to follow an anastamosing pattern, thus representing a pathway with many branches and metabolic dead ends
In 1993, Stierle and colleagues reported the unprecedented isolation of a Taxus spp. endopyhtic fungus (Taxomyces andreanae) that could synthesize Taxol and other taxanes such as baccatin III independently. This was demonstrated using radiolabeled precursors, such as 14C-phenylalanine and 14C-acetate (Stierle et al. 1993). Even more surprisingly, Taxol compromised an unusually high percentage (15–20 %) of the total taxane fraction synthesized by the fungus compared to that synthesized by the yew. The isolated Taxomyces andreanae was subject to a patent application and deposited at the Centraalbureau voor Schimmelcultures (Utrecht, The Netherlands) as number CBS 279.92 (Strobel et al. 1994). Several other groups soon confirmed the findings in this ground-breaking publication and provided additional supporting evidence (Flores-Bustamante et al. 2010). Microbial Taxol and taxane biosynthesis was found in several different genera of fungi, including Alternaria, Aspergillus, Cladosporium, Fusarium, Monochaetia, Pestlotia, Pestalotiopsis, Pithomyces, Penicillium and Xylaria, which were isolated from yew and non-Taxus plants (Flores-Bustamante et al. 2010; Strobel et al. 1996; Soca-Chafre et al. 2011; Zhang et al. 2009; Zhao et al. 2009; Hoffman 2003). Recently, several reports have been published claiming that endophytic fungi contain genes previously identified in Taxus spp. that encode the corresponding pathway enzymes (Zhang et al. 2008; Staniek et al. 2009; Miao et al. 2009; Kumaran et al. 2010). The publication of Stierle and colleagues also resulted in a huge proliferation of studies of endophytes from Taxus species (Rivera-Orduña et al. 2011) and other medicinal plants (Kumar and Hyde 2004; Huang et al. 2009; Lin et al. 2010) as it generally became accepted that horizontal gene transfer was commonplace and that fungal endophytes within these plants could probably also produce the bioactive medicinal compounds produced by the plants (Chandra 2012).
Interestingly, these reports claiming the presence of previously identified Taxus spp. genes in endophytic fungi base their claims on the results of PCR experiments using primers designed according to published sequences from Taxus trees, indicating that fungal genomic DNA yields PCR amplification products virtually identical to the Taxus clones (Staniek et al. 2009; Miao et al. 2009). The presence of these genes would require the extensive horizontal gene transfer (HGT) between the yew trees and multiple endophytic fungi, representing a pathway with more than 20 steps (Croteau et al. 2006). We find it difficult to believe that this entire pathway could have transferred in an arbitrary manner, and therefore we searched for evidence of DNA transfer involving potential taxane-synthesis gene clusters originating from Taxus plants. Whereas biosynthetic gene clusters are a common features in bacterial genomes and have also been described in fungi (Tudzynski and Hölter 1998; Zhang et al. 2004), there have been few reports of clustered metabolic pathways in plants, and those that do exist tend to be spread over larger genomic regions than their microbial counterparts (Field and Osbourn 2008; Field et al. 2011; Chu et al. 2011). The existence of taxane gene clusters in fungi and plants raises intriguing questions about the origin and evolution of these highly-specialized biosynthetic pathways, and the potential for HGT from fungi to Taxus trees. However, HGT between distantly-related organisms is a rare evolutionary event which is also constrained by the amount of genetic information transferred and genetic barriers involving incompatible regulation and codon usage. This contrasts sharply with the widespread observation of Taxol biosynthesis in many different endophytic fungi (Kurland et al. 2003).
Material and methods
Isolation of endophytic fungi from Taxus spp. plant material
Endophytic fungi were isolated as previously described by Guo et al. (2006). Bark segments (0.5 × 0.5 cm) were removed with a sterile scalpel and surface sterilized for 5 min in 70 % ethanol. The inner bark was separated from the outer layer and placed on PDA agar (Carl Roth GmbH, Karlsruhe, Germany) supplemented with 25 mg/L streptomycin. The plates were incubated at room temperature until fungal growth was visible. The mycelium was then transferred to fresh plates using the hyphal tip method.
Cultivation of endophytic fungi
The isolated endophytic fungi were cultivated on solid media, PDA (Carl Roth GmbH) supplemented with streptomycin or on YM-6.3 agar (0.4 % (w/v) glucose, 0.4 % (w/v) yeast extract, 2 % (w/v) malt extract, pH 6.3, 1.5 % (w/v) agar-agar). The fungi were transferred to fresh plates at weekly intervals by cutting out a piece of overgrown agar. In liquid culture, the fungi were grown in 0.6–10 L YM-6.3 medium (120 rpm in the dark) for 3 weeks or until no more glucose could be detected. The fungi were also cultivated in S7 medium as described for taxane-producing endophytes (Stierle et al. 1993).
Taxoid extraction
For taxane analysis, the fungal culture media were extracted twice with an equal volume of chloroform. The organic phase was then dried over magnesium sulfate, evaporated to dryness and the residue was redissolved in 3–5 mL methanol. Plant material (30 g Taxus needles or tobacco leaf tissue) was lyophilized and extracted with 1:1 dichloromethane/methanol in a Soxhlet extractor. The organic solution was evaporated to dryness and redissolved in dichloromethane. After two rounds of extraction with water, the organic layer was dried over magnesium sulfate, evaporated to dryness and the residue was redissolved in methanol (Witherup et al. 1990).
Anti-taxane immunoassay (competitive inhibition enzyme immunoassay, CIEIA)
The anti-taxane immunoassay was carried out according to the manufacturer’s instructions (Cardax Pharmaceuticals, Hawaii). A standard curve for taxane quantitation was made using Taxol concentrations of 111, 37, 12.33, 4.11, 1.37, 0.46 and 1.15 ng/mL (Table S1). The samples were analyzed using three dilutions. Values in the linear range of the standard curve were used to calculate the concentration.
LC/MS/MS analysis
LC/MS/MS was carried out in multiple reaction monitoring scan mode using a QTrap3200 system (Applied Biosystems, Darmstadt, Germany). The three most intensive mass transitions for three standard substances (Taxol, baccatin III and 10-deacetyl-baccatin III; Sigma-Aldrich, Idena) were used for detection (Table S2). Analysis in ESI negative ionization mode was carried out using the following settings: curtain gas 25 psi, CAD gas medium, ionspray voltage −4,500 V, temperature 450 °C, gas1 50 psi, gas2 65 psi. HPLC separation was carried out using a Curosil PFP column (150 × 3 mm, 3 μm; Phenomenex, Aschaffenburg, Germany) under the following conditions: column oven, 25 °C; LC flow rate, 300 μL/min; solvent A, 98 % water and 2 % acetonitrile with 10 mM ammonium acetate; solvent B, 2 % water and 98 % acetonitrile with 10 mM ammonium acetate; gradient, 0 min 70 % A, 0.5 min 70 % A, 15 min 0 % A, 20 min 0 % A, 21 min 70 % A, 23 min 70 % A.
DNA isolation, construction of genomic phage libraries and hybridization
Fungal and plant genomic DNA was isolated using a modified CTAB method. Plant and fungal samples (1 g) were homogenized with a mortar under liquid nitrogen, supplemented with 10 volumes of CTAB buffer (100 mM Tris pH8, 20 mM EDTA, 1.4 M NaCl, 2 % β-mercaptoethanol, 2 % CTAB) and incubated for 1 h at 65 °C. The cell debris was removed by centrifugation (15 min, 2,000 × g) and the supernatant was extracted twice with an equal volume of 24:1 chloroform:isoamylalcohol. The DNA was then precipitated with isopropanol. Genomic phage libraries were constructed from EF0001, EF0021 and Taxomyces andreanae DNA, and plaque lifting was carried out according to the manufacturer’s guidelines (Lambda Dash® II / Gigapack® III XL, Stratagene). Heat-fixed membranes (Nylon N+, GE Healthcare) were supplemented with 20 mL Roti-Hybri-Quick (Carl Roth GmbH) and 100 μg/mL salmon sperm DNA (Sigma) in hybridization rolls. Pre-hybridization was carried out for 3 h at 55 °C. Probes against taxadiene synthase (TDS) and taxane-13α-hydroxylase (T13H) were prepared by PCR using primers corresponding to specific target genes, i.e. TDS1 (forward 5′-GCA GCG CTG AAG ATG AAT GC-3′, reverse 5′-CGA TTC GAT ACC CCA CGA TCC-3′, bp 22–546), TDS2 (forward 5′-GCC CTC GGC CTC CGA ACC C-3′, reverse 5′-GCC ATG CCG GAT TCT TTC CAC C-3′, bp 1,211–1,710), TDS3 (forward 5′-GGT GGA AGG AAT CCG GCA TGG CAG-3′, reverse 5′-GTC GCC AGC TCA AGG ATA CAA GCT C-3′, bp 1,693–2,263) andT13H (forward 5′-ATG GAT GCC CTT AAG CAA TTG GAA GTT TCC CC-3′, reverse 5′-GCT CCT GCA GGT GCT CC-3′, bp 1–604). The reactions were heated to 94 °C for 2 min followed by 25 cycles of 94 °C for 30 s, 55–60 °C for 30 s, 72 °C for 45 s and finally 72 °C for 5 min. Incorporation of α32P-dATP (Hartmann Analytic, Braunschweig, Germany) was done using the Hexalabel™ DNA Labeling Kit (Fermentas, St. Leon-Rot, Germany). The TDS and T13H probes were purified on G-50 gel filtration columns (GE Healthcare, Karlsruhe, Germany). Probes against taxane-5α-hydroxylase (T5H) were prepared by labeling oligonucleotides with γ-32P-dATP using polynucleotide kinase (oligo1 5′-GGC ATC CCA CAG TAG TAC TCT GCG GCC CTG CGG GAA ACC GGC TTA TTC TGT CCA ACG AGG AGA AGC TGG TGC AGA TGT CG-3′, and oligo2 5′-CCA CCA CTT CGC CAA TGG CTT TGA TTT TCA AGC TCT TGT CTT CCA ATC CAG AAT GCT ATC AAA AAG TAG TTC AAG AGC-3′). Probes were added to the pre-hybridization mix and hybridized against the membranes overnight at 55 °C. The membranes were washed three times for 30 min with 1:2, 1:5 and 1:10 dilutions of hybridization buffer, and then visualized by autoradiography on pre-flashed X-ray films (Hyperfilm MP, GE Healthcare) at −80 °C for 2 days.
Amplification of internal transcribed spacer (ITS) sequences
ITS regions from the isolated Taxus endophytes were amplified by PCR using the universal primers ITS1 (5′-TCC GTA GGT GAA CCT GCG G-3′) and ITS4 (5′-TCC TCC GCT TAT TGA TAT GC-3′) (Sim et al. 2010) in 2× PCR-MasterMix Solution (i-Max II, INtRON Biotechnology) containing 1 μL of each primer (50 μM) and 20 ng genomic DNA, made up to 25 μL with water. Amplification was carried out on the GeneAmp PCR System (Applied Biosystems) at 94 °C for 5 min followed by 35 cycles of 94 °C for 1 min, 55 °C for 1 min and 72 °C for 1.5 min, followed by a final 72 °C step for 7 min. PCR products were purified using NucleoFast 96 PCR plates (Machery-Nagel, Düren, Germany) and sequenced.
Isolation of total RNA and cDNA library construction
Total RNA from endophytes was isolated using the borax method. Mycelia were homogenized under liquid nitrogen using a mortar and pestle, incubated at 42 °C for 1 h in 15 mL borax buffer (0.2 M sodium tetraborate, 30 mM EGTA, 1 % (w/v) SDS, 1 % (w/v) deoxycholate, 1 % (v/v) Nonidet P-40, 2 % (w/v) polyvinylpyrolidone, 10 mM DDT, pH 9.0), supplemented with 1.2 mL 2 M KCl and stored on ice for 1 h. After centrifugation, RNA was selectively precipitated by adding 5 mL 8 M LiCl and storing at −20 °C overnight. The precipitate was washed three times with cold 2 M LiCl and resuspended in 2.8 mL TES buffer (50 mM Tri/HCl pH 5.7, 5 mM EDTA, 50 mM NaCl) supplemented with 1 M CsCl. This suspension was overlaid with 1.2 mL TES buffer supplemented with 5.7 M CsCl and the RNA was purified by density gradient ultracentrifugation at room temperature at 100,000 × g for 16 h. The RNA was dissolved in 500 μL TE buffer (10 mM Tris/HCl pH 8.0, 1 mM EDTA) and mRNA was isolated using the Qiagen Oligotex mRNA Mini Kit (Qiagen, Hilden, Germany). A cDNA-RACE library was constructed using the Clontech Marathon cDNA Amplification Kit (Takara BIO Europe, Saint-Germain-en-Laye, France) according to the manufacturer’s instructions. Primers for the amplification of terpene synthase gene candidates are listed in Table S3.
Cloning, expression and functional testing of diterpene synthase 0021_TS_1762 and intron-1 splice variants of 0021_TS_1762
Synthetic diterpene synthase 1762 (Genskript, Hong Kong) was amplified using pUC57 as the template, and the product was transferred to the Escherichia coli expression vector pTrc-His2 using the pTrcHis2-TOPO® TA Expression Kit (Invitrogen, Karlsruhe, Germany). The gene was also amplified with primers including Gateway attachment sites allowing the gene to be introduced into the yeast expression vector pYES-Dest52 by homologous recombination. The protein was expressed in E. coli DH5α cells (New England Biolabs, Frankfurt, Germany) and Saccharomyces cerevisiae CEN-PK2-1 cells (EUROSCARF, Frankfurt, German) at 28 °C. Deletion variant 0021_TS_1762_del and intron1 random variants (primers listed in Table S3) were created by whole-plasmid PCR using pTrcHIS2-1762cosyn as the template with Herculase® II Fusion DNA Polymerase (Agilent Technologies, Karlsruhe, Germany) and the following temperature program: 95 °C for 3 min, followed by 30 cycles at 95 °C for 0.5 min, 58 °C for 0.5 min and 72 °C for 4 min, followed by a final step at 72 °C for 7 min. Crude protein extracts were prepared by disrupting the cells with glass beads. One volume of extract was used for in vitro testing with three volumes of assay buffer (100 mM Tris, 10 mM MgCl2, 5 mM β-mercaptoethanol, 50 μM substrate 3H-GGPP, 3H-FPP or 14C-IPP (+DMAPP), total volume 500 μL). Biotransformation reactions were incubated at 30 °C, overnight. After the addition of 500 μL saturated NaCl the reactions were extracted twice with the same volume of ethyl acetate. The extracts were concentrated in a nitrogen stream and analyzed by radio-TLC on silica plates (Merck, Darmstadt, Germany), which were developed with 9:1 cyclohexane:ethyl acetate or 3:1 pentane:diethyl ether. Products were detected using a radio-TLC Scanner RITA Star (Raytest, Straubenhardt, Germany).
Phage insert, ITS and whole genome sequencing
Phage inserts were sequenced using the Sanger method (Functional & Applied Genomics Group, Fraunhofer IME, Aachen, Germany) or shotgun sequencing (Eurofins MWG Operon, Ebersberg, Germany). ITS sequences were determined by Sanger sequencing (Functional & Applied Genomics Group, Fraunhofer IME). The EF0021 genome was sequenced using 454 technology by Seq-It GmbH, Kaiserslautern. The Taxomyces andreanae genome was sequenced by paired-end library sequencing (imagenes GmbH, Berlin, Germany). Each supplier also assembled the sequences they generated.
Sequence analysis
Sequences were analyzed using CLC Combined Workbench v3.6.1, Lasergene 7 Package, NCBI Blast and CloneManager Professional Suite 8. FGENESH was use for ORF and protein prediction (http://linux1.softberry.com/). Phylogenetic analysis was carried out using CLC Combined Workbench v3.6.1 with the protein sequences listed in Supplementary Data S3 and Table S4.
Results and discussion
Inconsistencies in reports describing the distribution of Taxol biosynthesis between distantly-related species prompted us to re-examine the existence of an independent Taxol biosynthesis pathway in endophytic fungi. Previous reports described the isolation of Taxol-producing endophytes from Taxus bark material, so we similarly attempted to isolate endophytic fungi from different Taxus bark materials collected from locations throughout Germany, Poland, the Netherlands and South Korea. Fungal cultures were initiated according to standard protocols and yielded a total of 34 individual cultures (Guo et al. 2006). For further characterization, the genomic DNA from these cultures was isolated and the conserved 18S rDNA internal transcribed spacer (ITS) region was amplified and sequenced (Suppl. Data S1). The isolated endophytic fungi were then transferred into liquid fermentation media for phytochemical analysis. As in previous studies, the isolated fungi were cultivated for up to 21 days or until the glucose source was depleted. The cultures were then extracted with chloroform for phytochemical analysis using a taxane-specific indirect competitive inhibition enzyme immunoassay (CIEIA) featuring a polyclonal antibody (Cardax Pharmaceuticals, Honolulu, Hawaii) (Caruso et al. 2000). We used an organic extract of Taxus baccata needles as a positive control and Nicotiana tabacum leaf material as a negative control. The antibody assay resulted in the identification of two potential taxane-producing fungi, designated EF0001 and EF0016. However, the quantity of taxanes, deduced from the Taxol standard curve, was low in both isolates (less than 10 ng/L of culture medium) compared to the positive control (~170 μg/g plant material; Table 1). Surprisingly, the N. tabacum leaf extract also appeared to contain taxanes, but at approximately five times the level detected in the positive endophytes. This unexpected result probably reflected unanticipated cross reactivity of the polyclonal antibody.
We carried out further characterization of fungal taxane synthesis by LC/MS/MS, using multi-reaction monitoring (MRM) to detect the products Taxol, baccatin III and 10-deacetylbaccatin III as standards with detection limits of 35, 28 and 23 fmol, respectively. We applied this method to organic extracts from all of the isolated fungi and three additional species previously claimed to be capable of independent taxane biosynthesis: Taxomyces andreanae (CBS 279.92; Strobel et al. 1994), UPH-12 (NRRL 30405; Hoffman 2003) and H10BA2 (NRRL 21209; Stierle et al. 2000). Baccatin III and 10-deacetylbaccatin III were detected in the newly-isolated endophytes EF0001 and EF0021, respectively (Fig. 2 and Suppl. Data S2). LC/MS/MS analysis confirmed the initial results obtained with CIEIA for EF0001, but Taxol, baccatin III and 10-deacetylbaccatin III were not detected by CIEIA or LC/MS/MS in any of the other species.
LC/MS/MS-multi-reaction monitoring (MRM) analysis of an organic extract from the Taxus endophyte EF0021. a LC/MS/MS-MRM chromatogram of 10-deacetylbaccatin III (10-DABIII, authentic standard (Idena, Milano, Italy), dissolved in methanol at a concentration of 1 mg/mL, injection volume 10 μL) eluting from the HPLC column at 4.72 min. The insert shows the three monitored ion transitions (m/z = 76.2, 120.8 and 391.2) of the 10-DABIII parent ion (m/z = 543.2) (M-H). b LC/MS/MS-MRM chromatogram with the observed mass pattern (shown in insert) at 4.72 min obtained with the organic extract of Taxus endophyte EF0021
Without delay (assuming potential genetic instability in the fungi), we extracted genomic DNA from EF0001 and EF0021. To avoid potential contamination leading to PCR artifacts, we established genomic phage libraries for both species and used conventional hybridization as the screening method. We used three probes specific for Taxol biosynthesis: taxadiene synthase (Wildung and Croteau 1996), taxane-5α-hydroxylase (Jennewein et al. 2004a), and taxane-13α-hydroxylase (Jennewein et al. 2001). For EF0001, we screened a total of 300,000 phage plaques (average insert size, 23 kb) corresponding to ~6,900 Mb of endophyte genomic sequence. Assuming an average fungal genome size of 50 Mb, this strategy achieved >130-fold genome coverage. For EF0021, we screened a total of 40,000 phage plaques, corresponding to 920 Mb of genomic sequence and 18-fold genome coverage. Several potential positive inserts were sequenced, but none of them corresponded to known Taxus spp. genes involved in taxane biosynthesis. Given that we were unable to identify taxane-related genomic sequences in EF0001 and ER0021, we constructed a T. andreanae genomic phage library and screened 162,000 phage plaques (average insert size 20.3 kb, corresponding to 3,300 Mb of genomic sequence and 66-fold genome coverage) using the same probes as above and did not identify any positive clones.
Our failure to identify fungal genomic sequence related to known taxane-specific sequences from yew trees led us to conclude that taxane biosynthesis in endophytes may have evolved independently, as is the case for gibberellins, whose biosynthesis pathway differs between microbes and plants (Tudzynski and Hölter 1998; Bömke and Tudzynski 2009). To further examine the potential for independent taxane biosynthesis by endophytes, we sequenced the EF0021 genome using a shotgun sequencing approach, yielding 2,234,101 sequence reads with an average length of 390 bp. Sequence alignment of the raw data achieved 98.55 % aligned reads and 2,623 contigs covering 44.45 Mb of genomic DNA, corresponding to an estimated genome size of 45.9 Mb. Analysis of the resulting contigs revealed the absence of any genes with significant homology to Taxus spp. genes involved in taxane biosynthesis, confirming the negative results of the library screening experiment. Further analysis of the EF0021 genome sequence resulted in the identification of six putative terpene synthases, two of which were closely related to Aspergillus nidulans lanosterol synthase (and were therefore likely to be involved in sterol biosynthesis). The four others have potential roles in secondary metabolism, including one related to a previously-isolated fungal diterpene synthase (fusicoccadiene synthase) from the plant–pathogen Phomopsis amygdali (Toyomasu et al. 2007) (Suppl. Data S3). Fusicoccadiene synthase is a unique terpene synthase because it possesses both terpene synthase and prenyltransferase activity. The three other identified terpene synthases showed significant homology to fungal sesquiterpene synthases.
Functional analysis was carried out by constructing an EF0021 cDNA library, but it proved impossible to isolate cDNAs corresponding to the above genomic clones using gene-specific primers, indicating that the genes may not have been expressed under the cultivation conditions we used. The genomic sequence was therefore used to design a synthetic open reading frame for the putative diterpene synthase that was codon-optimized for expression in E. coli. Several variants were constructed due to an obscure intron/exon border at one position reflecting variability in the original sequence. Crude extracts from recombinant E. coli cells were examined for diterpene synthase activity using 3H-geranylgeranyl diphosphate (GGPP), and for prenyltransferase activity using 14C-isopentenyl diphosphate and dimethylallyl diphosphate. The synthetic genes were also expressed in Saccharomyces cerevisiae. None of the heterologous expression assays in either host showed any evidence for diterpene synthase enzymatic activity.
In addition to the functional characterization of the potential prenyltransferase/diterpene synthase from endophyte EF0021, we also compared the gene and enzyme architecture with the known taxadiene synthase from Taxus spp., revealing several major differences. The intron/exon structure differed significantly with regard to the number and size of coding and non-coding regions (Fig. 3a, b) and the predicted proteins were also fundamentally distinct (Fig. 3c). Whereas Taxus spp. taxadiene synthase is a typical plant-derived terpene synthase based on the location of the catalytic DDXXD motif and characteristic domains such as the conifer diterpene internal sequence domain and the plastid leader sequence, the terpene synthase component of the EF0021 enzyme comprises only 300 amino acids containing the features relevant for synthase activity (Trapp and Croteau 2001). Furthermore, the chimeric nature of the endophyte protein appeared typical of fungal synthases and this has not been observed previously in plants, where condensation towards the diterpene prenyl precursor and the cyclization reaction are always catalyzed by two individual enzymes (Toyomasu et al. 2007). The active site of terpene synthase is sensitive to modifications, and even minor changes result in different product structures or complete inactivity. The significant differences in the geometry of the active site in plants and fungi therefore raise doubts about the ability of these enzymes to catalyze the synthesis of a complex product such as taxadiene (Seemann et al. 2002; Fellicetti and Cane 2004). Having been unable to identify a Taxus-related sequence in the EF0021 genome or to isolate a functional and active diterpene synthase, we concluded that EF0021 is incapable of independent Taxol biosynthesis.
Structure of diterpene synthase 0021_TS_1762_del from EF0021 compared to taxadiene synthase (TDS), including the intron/exon structures of TDS (a) and 0021_TS_1762_del (b). Schematic protein domain structures are also shown for both enzymes (c), including the catalytic DDXXD/E motifs and the annotation of domains according to Trapp and Croteau (2001) for TDS and from a comparison with Phomopsis amygdali fusicoccadiene synthase (Toyomasu et al. 2007)
We repeated the above strategy for T. andreanae, which was previously reported to produce taxanes independently (CBS 279.92; US Patent 5322779(A)). Shotgun sequencing of the T. andreanae paired-end library yielded 235 million sequence reads with an average length of 100 bp. Assembly of the raw sequence data generated 2,274 contigs with an average size of 18 kbp, covering 93.5 % of the sequence reads. Contig alignment covered a cumulative sequence of 45.08 Mb, corresponding to an approximate genome size of 45 Mb. As was the case for EF0021, the T. andreanae genome did not contain any sequences with significant homology to taxane biosynthesis genes from Taxus spp., but in contrast to EF0021, further analysis of the T. andreanae genome revealed the presence of several additional terpene synthase genes (Suppl. Data S3). All of these sequences were homologous to other known fungal sesquiterpene synthases, although none of them were closely related to known diterpene synthases. As was the case for Taxus endophyte EF0021, we were therefore unable to identify any potential genes related or non-related to taxane biosynthesis in yew that could confer upon T. andreanae the ability to synthesize Taxol independently.
We next used phylogenetic analysis to compare the predicted terpene synthases from endophyte EF0021 and Taxomyces andreanae (Supplementary Fig. 2). All the predicted terpene synthases were aligned with the protein sequences initially used for targeted screening (Table S4). A phylogenetic tree was constructed based on the aligned dataset using UPGMA (unweighted pair group method with arithmetic means) with bootstrapping (100 replicates, bootstrap values shown at the nodes, Suppl. Fig. 2).
The terpene synthase sequences clustered into three major clades (A–C). Clade A consisted of proteins annotated as sesquiterpene synthases with the greatest similarity to Cop6 from Coprinopsis cinereus, including two proteins from EF0021 and eight from Taxomyces andreanae, whereas all other sequences formerly annotated as sesquiterpene synthases clustered in clade C along with Cop1–Cop5 from Coprinopsis cinereus and protoilludene synthase from Armillaria gallica (Agger et al. 2009; Engels et al. 2011). Because Cop1–5 differ from Cop6 mechanistically, using all-trans-farnesyl diphosphate (FPP) or cis-FPP as a substrate to form trichodiene-like or germancrene-like cyclization products, the new terpene synthases clustering in clades A and C are probably grouped on the basis of conserved functionally-relevant motifs as well as their fungal origin. Only two sequences, one each from EF0021 and Taxomyces andreanae, were similar to proteins in clade B, which contained all plant and fungal sequences related to diterpene biosynthesis. Clade B comprised three sub-clades, based either on origin (fungi vs. plants) or specific function (e.g. their role in gibberellin biosynthesis). The abovementioned diterpene synthase from EF0021 and prenyltransferase from T. andreanae clustered with the fungal prenyltransferases and fusicoccadiene synthases. However, since these special chimeric synthases contain a prenyltransferase domain, clustering probably reflected the stronger conservation of this domain which sets these proteins aside from the other terpene synthases. The presence of this domain also confers greater similarity e.g. to plant geranylgeranyl diphosphate and copalyl diphosphate synthases than other fungal sesquiterpene synthases in clades A and C. Our data clearly showed no evidence for homology to plant terpene synthases, and thus for trans-kingdom gene transfer, as initially proposed as a possible explanation for the evolution of Taxol biosynthesis in plants and fungi. Furthermore, we found no evidence for similarities between the terpene synthases in the two endophytes we investigated. Terpene synthase 0021_TS_1762 remains the only candidate for an enzyme that might be involved in diterpenoid metabolism, although the absence of a Taxomyces andreanae ortholog argues against the hypothesis that this enzyme is a fungal taxadiene synthase. Even if the pathway evolved independently in fungi and plants, as is thought to be the case for gibberellin biosynthesis (Bömke and Tudzynski 2009), enzymes that catalyze the complex synthesis of taxadiene should have a common evolutionary origin and should therefore show evidence of significant sequence similarity.
Excluding any evolutionary scenario discussed above, the detection of minute amounts of taxanes in our fungal isolates is best explained by residual taxanes synthesized by the host yew tree. Taxol and related taxanes are highly lipophilic compounds that accumulate in endophyte cell wall structures. The interaction between Taxol and membranes is dependent on the chemical composition of the membranes, and Taxol-lipid complexes can be stable for months (Sharma and Straubinger 1994; Crosasso et al. 2000). Furthermore, low Taxol concentrations, comparable to the levels we detected in endophyte extracts, did not affect the physiological properties of the membrane (Balasubramanian and Straubinger 1994; Sharma and Straubinger 1994; Bernsdorf et al. 1999; Crosasso et al. 2000; Zhao and Feng 2004). Although these experiments involved artificial membranes, there is also evidence that fungi can take up non-polar compounds by passive transport and store them in vesicles. For example, Fusarium solani can absorb polyaromatic compounds from the cell culture medium and store them within intracellular compartments with no impact on growth (Verdin et al. 2005). In the endophytes we studied, the accumulation of non-polar taxoid molecules in lipophilic cell structures combined with the high sensitivity of our analytical methods, immunological detection and LC/MS/MS-based multi-reaction monitoring (MRM) ensured that these carry-overs could be detected. After the first and second passages of the fungal cultures, no taxanes could be detected by LC/MS/MS. The fungi were no longer associated with the Taxol source and hence the trace amounts of taxanes detected initially were diluted below the detection limit.
Our results and conclusions therefore offer a satisfactory explanation for the contradictory results in earlier publications, some providing evidence for independent taxane biosynthesis in different endophytic fungi and others lacking this evidence.
References
Agger S, Lopez-Gallego F, Schmidt-Dannert C (2009) Diversity of sesquiterpene synthases in the basidiomycete Coprinus cinereus. Mol Microbiol 72(5):1181–1195
Balasubramanian SV, Straubinger RM (1994) Taxol–lipid interactions: taxol-dependent effects on the physical properties of model membranes. Biochemistry 33:8941–8947
Baloglu E, Kingston DGI (1999) Taxane diterpenoids. J Nat Prod 62:1448–1472
Bernsdorf C, Reszka R, Winter R (1999) Interaction of the anticancer agent Taxol (paclitaxel) with phospholipid bilayers. J Biomed Mater Res 46:141–149
Bömke C, Tudzynski B (2009) Diversity, regulation, and evolution of the gibberellin biosynthetic pathway in fungi compared to plants and bacteria. Phytochemistry 70:1876–1893
Brown DT (2003) Preclinical and clinical studies of the taxanes. In: Itokawa H, Lee K-H (eds) The genus Taxus. Taylor & Francis, London, pp 387–435
Caruso M, Colombo AL, Fedeli L, Pavesi A, Quaroni S, Saracchi M, Ventrella G (2000) Isolation of endophytic fungi actinomycetes taxane producers. Ann Microbiol 50:3–13
Chandra S (2012) Endophytic fungi: novel sources for anticancer lead molecules. Appl Microbiol Biotechnol 95:47–59
Chu HY, Wegel E, Osbourn A (2011) From hormones to secondary metabolism: the emergence of metabolic gene clusters in plants. Plant J 66:66–79
Croom EM Jr (1995) Taxus for taxol and taxoids. In: Suffness M (ed) Taxol® science and applications. CRC Press, Boca Raton, pp 37–70
Crosasso P, Ceruti M, Brusa P, Arpicco S, Dosio F, Cattel L (2000) Preparation, characterization and properties of sterically stabilized paclitaxel-containing liposomes. J Control Release 63:19–30
Croteau R, Ketchum R, Long R, Kaspera R, Wildung M (2006) Taxol biosynthesis and molecular genetics. Phytochem Rev 5:75–97
Engels B, Heinig U, Grothe T, Stadler M, Jennewein S (2011) Cloning and characterization of an Armillaria gallica cDNA encoding protoilludene synthase, which catalyzes the first committed step in the synthesis of antimicrobila melleolides. J Biol Chem 286:6871–6878
Fellicetti B, Cane DE (2004) Aristolochene synthase: mechanistic analysis of active site residues by site-directed mutagenesis. J Am Chem Soc 126(23):7212–7221
Field B, Osbourn AE (2008) Metabolic diversification—independent assembly of operon-like gene clusters in different plants. Science 320:543–547
Field B, Fiston-Lavier AS, Kemen A, Geisler K, Quesneville H, Osbourn AE (2011) Formation of plant metabolic gene clusters within dynamic chromosomal regions. Proc Natl Acad Sci U S A 108:16116–16121
Flores-Bustamante FZ, Rivera-Orduna FN, Martinez-Cádenas A, Flores-Cotera LB (2010) Microbial paclitaxel: advances and perspectives. J Antibiot 63:460–467
Guéritte-Voegelein F, Guénard D, Potier P (1987) Taxol and derivatives: a biogenetic hypothesis. J Nat Prod 50:9–18
Guo BH, Wang YC, Zhou XW, Hu K, Tan F, Miao ZQ, Tang KX (2006) An endophytic Taxol-producing fungus BT2 isolated from Taxus chinensis var. mairei. Afr J Biotechnol 5:875–877
Heinig U, Jennewein S (2009) Taxol: a complex diterpenoid natural product with an evolutionarily obscure origin. Afr J Biotechnol 8:1370–1385
Hoffman A (2003) Methods for obtaining taxanes. US patent 6638742 (B1)
Huang WY, Cai YZ, Surveswaran S, Hyde KD, Corke H, Sun M (2009) Molecular phylogenetic identification of endophytic fungi isolated from three Artemisia species. Fungal Divers 36:69–88
Itokawa H (2003) Taxoids occurring in the genus Taxus. In: Itokawa H, Lee K-H (eds) The genus Taxus. Taylor & Francis, London, pp 35–78
Jennewein S, Rithner CD, Williams RM, Croteau R (2001) Taxol biosynthesis: taxane 13α-hydroxylase is a cytochrome P450-dependent monooxygenase. Proc Natl Acad Sci U S A 98:13595–13600
Jennewein S, Long RM, Williams RM, Croteau R (2004a) Cytochrome P450 taxadiene 5α-hydroxylase, a mechanistically unusual monooxygenase catalyzing the first oxygenation step of taxol biosynthesis. Chem Biol 11:379–387
Jennewein S, Wildung MR, Chau M, Walker K, Croteau R (2004b) Random sequencing of an induced Taxus cell cDNA library for identification of clones involved in Taxol biosynthesis. Proc Natl Acad Sci U S A 101:9149–9154
Kaspera R, Croteau R (2006) Cytochrome P450 oxygenases of Taxol biosynthesis. Phytochem Rev 5:433–444
Kumar DDS, Hyde KD (2004) Biodiversity and tissue-recurrence of endophytic fungi in Tripterygium wilfordii. Fungal Divers 17:69–90
Kumaran RS, Kim HJ, Hur B-K (2010) Taxol promising fungal endophyte, Pestalotiopsis species isolated from Taxus cuspidata. J Biosci Bioeng 110:541–546
Kurland CG, Canback B, Berg OG (2003) Horizontal gene transfer. A critical review. Proc Natl Acad Sci U S A 100:9658–9662
Lin X, Huang YJ, Zheng ZH, Su WJ, Qian XM, Shen YM (2010) Endophytes from the pharmaceutical plant, Annona squamosa: isolation, bioactivity, identification and diversity of its polyketide synthase gene. Fungal Divers 41:41–51
Miao Z, Wang Y, Yu X, Guo B, Tang K (2009) A new endophytic taxane producing fungus from Taxus chinensis. Appl Biochem Micobiol 45:81–86
Rivera-Orduña FN, Suarez-Sanchez RA, Flores-Bustamante ZR, Gracida-Rodriguez JN, Flores-Cotera LB (2011) Diversity of endophytic fungi of Taxus globosa (Mexican yew). Fungal Divers 47:65–74
Seemann M, Zhai G, De Kraker JW, Paschall CM, Christianson DW, Cane DE (2002) Pentalenene synthase. Analysis of active site residues by site-directed mutagenesis. J Am Chem Soc 124(26):7681–7689
Sharma A, Straubinger RM (1994) Novel taxol formulations: preparation and characterization of taxol-containing liposomes. Pharm Res 11:889–896
Sim JH, Khoo CH, Lee LH, Cheah YK (2010) Molecular diversity of fungal endophytes isolated from Garcinia mangostana and Garcinia parvifolia. J Microbiol Biotechnol 20:651–658
Soca-Chafre G, Rivera-Orduna FN, Hidalgo-Lara ME, Hernandez-Rodriguez C, Marsch R, Flores-Cotera LB (2011) Molecular phlogeny and paclitaxel screening of fungal endophytes from Taxus globosa. Fungal Biol 115:143–156
Staniek A, Woerdenbag HJ, Kayser O (2009) Taxomyces andreanae: a presumed paclitaxel producer demystified? Plant Med 75:1–6
Stierle A, Strobel G, Stierle D (1993) Taxol and taxane production by Taxomyces andreanae, an endophytic fungus of pacific yew. Science 260:214–216
Stierle A, Stierle D, Strobel G (2000) Taxol production by a microbe. US patent 6013493(A)
Strobel G, Stierle AA, Stierle DB (1994) Taxol production by Taxomyces andreanae. US patent 5322779 (A)
Strobel GA, Yang XS, Sears J, Kramer R, Sidhu RS, Hess WM (1996) Taxol from Pestalotiopsis microspora, an endophytic fungus of Taxus wallichiana. Microbiology 142:435–440
Suffness M, Wall ME (1995) Discovery and development of Taxol. In: Suffness M (ed) Taxol® science and applications. CRC Press, Boca Raton, pp 3–25
Toyomasu T, Tsukahara M, Kaneko A, Niida R, Mitsuhashi W, Dairi T, Kato N, Sassa T (2007) Fusicoccins are biosynthesized by an unusual chimera diterpene synthase in fungi. Proc Natl Acad Sci U S A 104:3084–3088
Trapp S, Croteau R (2001) Genomic organization of plant terpene synthases and molecular evolutionary implications. Genetics 158(2):811–832
Tudzynski B, Hölter K (1998) Gibberellin biosynthetic pathway in Gibberella fujikuroi: evidence for a gene cluster. Fungal Genet Biol 25:157–170
Verdin A, Loundes-Hadj Sahraoui A, Newsam R, Robinson G, Durand R (2005) Polycyclic aromatic hydrocarbons storage by Fusarium solani in intracellular lipid vesicles. Environ Pollut 133:283–291
Wildung MR, Croteau R (1996) cDNA clone for taxadiene synthase, the diterpene cyclase that catalyzes the committed step of taxol biosynthesis. J Biol Chem 271:9201–9204
Witherup KM, Look SA, Stasko MW, Ghiorzi TJ, Muschik GM, Cragg GM (1990) Taxus spp. needles contain amounts of taxol comparable to the bark of Taxus brevifolia: analysis and isolation. J Nat Prod 53:1249–1255
Zhang S, Monahan B, Tkacz JS, Scott B (2004) Indol-diterpene gene cluster from Aspergillus flavus. Appl Environ Microbiol 70:6875–6883
Zhang P, Zhou P-P, Jiang C, Yu H, Yu L-J (2008) Screening of Taxol-producing fungi based on PCR amplification from Taxus. Biotechnol Lett 30:2119–2123
Zhang P, Zhou P-P, Yu L-J (2009) An endophytic Taxol-producing fungus from Taxus media, Cladosporium cladosporioides MD2. Curr Microbiol 59:227–232
Zhao L, Feng SS (2004) Effects of lipid chain length on molecular interactions between paclitaxel and phospholipid within model biomembranes. J Colloid Interface Sci 274:55–68
Zhao K, Ping W, Li Q, Hao S, Zhao L, Gao T, Zhou D (2009) Aspergillus niger var. taxi, a new species variant of Taxol-producing fungus isolated from Taxus cuspidata in China. J Appl Microbiol 4:1202–1207
Acknowledgments
U.H. was supported by a pre-doctoral fellowship from the Volkswagen Foundation, Hannover, Germany (AZ.: I/82 754).
Author information
Authors and Affiliations
Corresponding author
Additional information
Data deposition
The sequences reported in this paper have been deposited in GenBank under accession nos. PRJNA77805 and PRJNA77807.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 666 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Heinig, U., Scholz, S. & Jennewein, S. Getting to the bottom of Taxol biosynthesis by fungi. Fungal Diversity 60, 161–170 (2013). https://doi.org/10.1007/s13225-013-0228-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13225-013-0228-7