Abstract
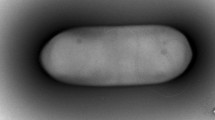
Gram-negative, rod-shaped bacterial strain ptl-2T, was isolated from soil of the Himalayan valley, India. The bacterial strain ptl-2T has been characterized using a polyphasic taxonomic approach including morphological characterization, fatty acid analysis, biochemical tests, 16S rRNA and nifH gene sequence analysis. 16S rRNA gene sequence analysis showed that the strain ptl-2T belonged to the genus Pseudacidovorax and is closely related to Pseudacidovorax intermedius (99.3 % similarity). It showed <97 % similarity to species of the genera Acidovorax, Alicycliphilus, Xylophilus, Giesbergeria and Simplicispira. The generic assignment has been confirmed on the basis of chemotaxonomic data, which revealed the fatty acid profile, characteristic of the genus Pseudacidovorax, consisting of C16:0 (21.78) and C18:1w7c (19.78) as major fatty acids. Phylogenetic, chemotaxonomic and phenotypic analysis based on signature sequences, DNA-DNA hybridization and physiological characterizations, confirms that strain ptl-2T represents a different species of the genus Pseudacidovorax for which the name Pseudacidovorax austerolens sp. nov. is proposed. The type strain ptl-2T (= CCUG 58759T, = DSM 24877T) has been submitted to two culture collection centres. GenBank accession numbers for the 16S rRNA and nifH sequence of strain Pseudacidovorax austerolens ptl-2T are FJ581042 and GQ249664, respectively.



Similar content being viewed by others
References
Alam S, Brailsford SR, Whiley RA, Beighton D (1999) PCR-based methods for genotyping viridans group streptococci. J Clin Microbiol 37:2772–2776
Alexander SK, Strete D, Niles MJ (2003) Laboratory exercises in organismal molecular microbiology. The McGraw-Hill, pp 77–83
Arden-Jones MP, McCarthy AJ, Cross T (1979) Taxonomic and Serological studies on Micropolyspora faeni and Micropolyspora strains from soil bearing the specific epithet rectivirgula. J Gen Microbiol 115:343–354
Burris RH (1972) Nitrogen fixation assay – methods and techniques. Methods Enzymol 24:415–431
Cashion P, Hodler-Franklin MA, McCully J, Franklin M (1977) A rapid method for base ratio determination of bacterial DNA. Anal Biochem 81:461–466
Chun J, Lee J-H, Jung Y, Kim S, Kim BK, Lim YW (2007) EzTaxon: a web based tool for the identification of prokaryotes based on 16 S ribosomal RNA gene sequences. Int J Syst Evol Microbiol 57:2259–2261
Gonzalez JM, Saiz-Jimenez C (2002) A fluorimetric method for the estimation of G + C mol% content in microorganisms by thermal denaturation temperature. Environ Microbiol 4:770–773
Kämpfer P, Thummes K, Chu H-I, Tan C-C, Arun AB, Chen W-M, Lai W-A, Shen F-T, Rekha PD, Young C-C (2008) Pseudacidovorax intermedius gen. nov., sp. nov., a novel nitrogen-fixing betaproteobacterium isolated from soil sequences. Int J Syst Evol Microbiol 58:491–495
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Öhlinger R (1996) Methods in soil physics; methods in soil chemistry. In: Schinner F (ed) Methods in soil biology. Springer, NY, pp 383–396
Paisley R (1996) MIS whole cell fatty acid analysis by gas chromatography training manual. MIDI, Newark
Pal R, Bala S, Dadhwal M (2005) Hexachlorocyclohexane-degrading bacterial strains Sphingomonas paucimobilis B90A, UT26 and Sp+, having similar lin genes, represent three distinct species, Sphingobium indicum sp. nov., Sphingobium japonicum sp. nov. and Sphingobium francense sp. nov., and reclassification of [Sphingomonas] chungbukensis as Sphingobium chungbukense comb. nov. Int J Syst Evol Microbiol 55:1965–1972
Pearson WR, Lipman DJ (1988) Improved tools for biological sequence comparison. Proc Natl Acad Sci 85:2444–2448
Poly F, Monrozier LJ, Bally R (2001) Improvement in the RFLP procedure for studying the diversity of nifH genes in communities of nitrogen fixers in soil. Res Microbiol 152:95–103
Prakash O, Verma M, Sharma P, Kumar M, Kumari K, Singh A, Kumari H, Jit S, Gupta K, Khanna M, Lal R (2007) Polyphasic approach of bacterial classification—an overview of recent advances. Indian J Microbiol 47:98–108
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Stackebrandt E, Goebel BM (1994) Taxonomic note: a place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int J Syst Evol Microbiol 44:846–849
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE, Stackebrandt E, Starr MP, Truper HG (1987) International committee on systematic bacteriology. Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
Acknowledgements
Financial assistance from the Department of Biotechnology (BT/PR-12522/BCE/08/05/2009) is highly acknowledged. Shivani is thankful to the Council of Scientific and Industrial Research, New Delhi, for providing fellowship as JRF/SRF to her during this work. We would like to thank SIAF-DST (Sophisticated Analytical Instrumentation Facility - Department of Science and Technology, Department of Anatomy, AIIMS) for providing the electron microscopy facility. We are highly thankful Mr. Jean P. Euzeby (Ecole Nationale Veterinaire, Toulelouse, France) for etymological advice. We are also thankful to Dr. Sangeeta Pal, Division of Microbiology, IARI, New Delhi for her kind help during experimentation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tyagi, S., Singh, D.K. Pseudacidovorax austerolens sp. nov., a nifH bacterium isolated from Himalayan valley soil, India. Ann Microbiol 65, 217–223 (2015). https://doi.org/10.1007/s13213-014-0852-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13213-014-0852-9