Abstract
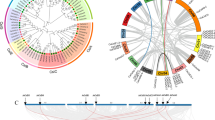
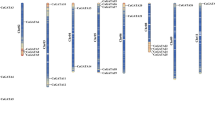
Fructokinase is the main catalytic enzyme for fructose phosphorylation and can also act as a glucose receptor and signal molecule to regulate the metabolism of plants, which plays an important role in plant growth and development. In this study, the CaFRK gene family and their molecular characteristics are systematically identified and analyzed, and the specific expression of CaFRKs under different tissues, abiotic stresses and hormone treatments were explored. Nine FRK genes were authenticated in pepper genome database, which were dispersedly distributed on eight reference chromosomes and predicted to localize in the cytoplasm. Many cis-acting elements that respond to light, different stresses, hormones and tissue-specific expression were found in the promoters of CaFRKs. FRK proteins of four species including Capsicum annuum, Arabidopsis thaliana, Solanum lycopersicum and Oryza sativa were divided into four groups via phylogenetic analysis. The collinearity analysis showed that there were two collinear gene pairs between CaFRKs and AtFRKs. In addition, it was significantly found that CaFRK9 expressed far higher in flower than other tissues, and the relative expression of CaFRK9 was gradually enhanced with the development of flower buds in fertile accessions, 8B, R1 and F1. Nevertheless, CaFRK9 hardly expressed in all stages of cytoplasmic male sterile lines. Based on the quantitative real-time PCR, most of CaFRK genes showed significant up-regulation under low-temperature, NaCl and PEG6000 treatments. On the contrary, the expression levels of most CaFRKs revealed a various trend in response to hormone treatments (IAA, ABA, GA3, SA and MeJA). This study systematically analyzed CaFRK gene family and studied its expression pattern, which lay the foundation of CaFRK genes cloning and functional verification response to abiotic stresses, and provides new insights into exploring the CaFRK genes on the pollen development in pepper.











Similar content being viewed by others
References
Alexandrov NN, Troukhan ME, Brover VV et al (2006) Features of Arabidopsis genes and genome discovered using full-length cDNAs. Plant Mol Biol 60(1):69–85
Ashwell G (1964) Carbohydrate metabolism. Annu Rev Biochem 33(1):101–138
Bagowski CP, Bruins W, Te Velthuis AJW (2010) The nature of protein domain evolution: shaping the interaction network. Curr Genomics 11(5):368–376
Bailey TL, Williams N, Misleh C (2006) MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res 34:W369–W373
Bailey TL, Boden M, Buske FA et al (2009) MEME suite: tools for motif discovery and searching. Nucleic Acids Res 37:W202–W208
Baysdorfer C, Kremer DF, Sicher RC (1989) Partial purification and characterization of fructokinase activity from barley leaves. J Plant Physiol 134(2):156–161
Cao Y, Li S, Han Y et al (2018) A new insight into the evolution and functional divergence of FRK genes in Pyrus bretschneideri. R Soc Open Sci 5(7):171463
Chase CD (2007) Cytoplasmic male sterility: a window to the world of plant mitochondrial-nuclear interactions. Trends Genet 23(2):81–90
Chen C, Chen G, Cao B et al (2015) Transcriptional profiling analysis of genic male sterile–fertile Capsicum annuum reveal candidate genes for pollen development and maturation by RNA-Seq technology. Plant Cell Tissue Organ Cult 122(2):465–476
Chen Y, Zhang Q, Hu W et al (2017) Evolution and expression of the fructokinase gene family in Saccharum. BMC Genomics 18(1):197
Chen C, Chen H, Zhang Y et al (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13(8):1194–1202
Chou KC, Shen HB (2010) A new method for predicting the subcellular localization of eukaryotic proteins with both single and multiple sites: Euk-mPLoc 2.0. PLoS ONE 5(4):e9931
Clément C, Audran JC (1995) Anther wall layers control pollen sugar nutrition in Lilium. Protoplasma 187(1):172–181
Clement C, Burrus M (1996) Floral organ growth and carbohydrate content during pollen. Am J Bot 83(4):459
Copeland L, Morell M (1985) Hexose kinases from the plant cytosolic fraction of soybean nodules. Plant Physiol 79(1):114–117
Copeland L, Tanner GJ (1988) Hexose kinases of avocado. Physiol Plant 74(3):531–536
Copeland L, Stone SR, Turner JF (1984) Kinetic studies of fructokinase I of pea seeds. Arch Biochem Biophys 233(2):748–760
Damari-Weissler H, Kandel-Kfir M, Gidoni D et al (2006) Evidence for intracellular spatial separation of hexokinases and fructokinases in tomato plants. Planta 224(6):1495–1502
David-Schwartz R, Weintraub L, Vidavski R et al (2013) The SlFRK4 promoter is active only during late stages of pollen and anther development. Plant Sci 199:61–70
Doehlert DC (1989) Separation and characterization of four hexose kinases from developing maize kernels. Plant Physiol 89(4):1042–1048
Fei Z, Tang X, Alba RM et al (2004) Comprehensive EST analysis of tomato and comparative genomics of fruit ripening. Plant J 10(1):47–59
Feng K, Jb Yu, Cheng Y et al (2016) The SOD gene family in tomato: identification, phylogenetic relationships, and expression patterns. Front Plant Sci 7:1279
Fennington GJ, Hughes TA (1996) The fructokinase from Rhizobium leguminosarum biovar trifolii belongs to group I fructokinase enzymes and is encoded separately from other carbohydrate metabolism enzymes. Microbiology (reading, Engl) 142(2):321–330
Finn RD, Clements J, Eddy SR (2011) HMMER web server: interactive sequence similarity searching. Nucleic Acids Res 39(Suppl 2):W29–W37
Franck C, Nsthan H, Heather AR et al (1995) Partial purification and characterization of fructokinase from developing taproots of sugar beet (Beta vulgaris). Plant Sci 110(2):181–186
Fulda S, Horn R, Stegmann H et al (2011) Physiology and proteomics of drought stress acclimation in sunflower (Helianthus annuus L). Plant Biol 13(4):632–642
Gasteiger E, Hoogland C, Gattiker A et al (2005) Protein identification and analysis tools on the ExPASy server. The proteomics protocols handbook. Humana Press, pp 571–607
German MA, Dai N, Chmelnitsky I et al (2002) LeFRK4, a novel tomato (Lycopersicon esculentum Mill) fructokinase specifically expressed in stamens. Plant Sci 163(3):607–613
German MA, Asher I, Petreikov M et al (2004) Cloning, expression and characterization of LeFRK3, the fourth tomato (Lycopersicon esculentum Mill.) gene encoding fructokinase. Plant Sci 166(2):285–291
Glover J, Grelon M, Craig S et al (1998) Cloning and characterization of MS5 from Arabidopsis: a gene critical in male meiosis. Plant J 15(3):345–356
Goetz M, Godt DE, Guivarc’h A et al (2001) Induction of male sterility in plants by metabolic engineering of the carbohydrate supply. Proc Natl Acad Sci USA 98(11):6522–6527
Gonzali S, Pistelli L, De Bellis L et al (2001) Characterization of two Arabidopsis thaliana fructokinases. Plant Sci 160(6):1107–1114
Guo J, Wang P, Cheng Q et al (2017) Proteomic analysis reveals strong mitochondrial involvement in cytoplasmic male sterility of pepper (Capsicum annuum L.). J Proteomics 168:15–27
Hanson MR, Bentolila S (2004) Interactions of mitochondrial and nuclear genes that affect male gametophyte development. Plant Cell 16(suppl 1):S154-169
He Z, Zhang H, Gao S et al (2016) Evolview v2: an online visualization and management tool for customized and annotated phylogenetic trees. Nucleic Acids Res W1:W236–W241
Higgins DG, Thompson JD, Gibson TJ (1996) Using CLUSTAL for multiple sequence alignments. Methods Enzymol 266:383–402
Hirsche J, Engelke T, Vller D et al (2009) Interspecies compatibility of the anther specific cell wall invertase promoters from Arabidopsis and tobacco for generating male sterile plants. Theor Appl Genet 118(2):235–245
Hoepfner SW, Botha FC (2004) Purification and characterization of fructokinase from the culm of sugarcane. Plant Sci 167(3):645–654
Hong ST, Chung JE, An G et al (1998) Analysis of 176 expressed sequence tags generated from cDNA clones of hot pepper by single-pass sequencing. J Plant Biol 41(2):116–124
Hu B, Jin J, Guo AY (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31(8):1296–1297
Jiang H, Dian W, Liu F et al (2003) Isolation and characterization of two fructokinase cDNA clones from rice. Phytochemistry 62(1):47–52
Kanayama Y, Dai N, Granot D et al (1997) Divergent fructokinase genes are differentially expressed in tomato. Plant Physiol 113(4):1379–1384
Kaplan CP, Tugal HB, Baker A (1997) Isolation of a cDNA encoding an Arabidopsis galactokinase by functional expression in yeast. Plant Mol Biol 34(3):497–506
Kumar S, Stecher G, Li M et al (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 18(6):1547–1549
Kuo TM, Doehlert DC, Crawford CG (1990) Sugar metabolism in germinating soybean seeds: evidence for the sorbitol pathway in soybean axes. Plant Physiol 93(4):1514–1520
Lescot M, Dehais P, Thijs G (2001) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30(1):325–327
Livak K, Schmittgen T (2000) Analysis of relative gene expression data using real-time quantitative PCR and the 2-△△Ct method. Methods 25(4):402–408
Martinez-Barajas E, Randall DD (1996) Purification and characterization of fructokinase from developing tomato (Lycopersicon esculentum) fruits. Planta 199(3):451–458
McCormick S (2004) Control of male gametophyte development. Plant Cell 16(Suppl 1):S142–S153
Nakamura N, Shimizu M, Suzuki H (1991) Characterization of hexose kinases from camellia and lily pollen grains. Physiol Plant 81(2):215–220
Pino J, Gonzalez M, Ceballos L et al (2006) Characterization of total capsaicinoids, colour and volatile compounds of Habanero chilli pepper (Capsicum chinense Jack) cultivars grown in Yucatan. Food Chem 104(4):1682–1686
Qin QP, Zhang SL, Chen JW et al (2004) Isolation and expression analysis of fructokinase genes from citrus. Acta Botanica Sinica 46(12):1408–1415
Ralph S, Chun HJ, Cooper D et al (2008a) Analysis of 4,664 high-quality sequence-finished poplar full-length cDNA clones and their utility for the discovery of genes responding to insect feeding. BMC Genomics 9(1):57
Ralph S, Chun H, Kolosova N et al (2008b) A conifer genomics resource of 200,000 spruce (Picea spp) ESTs and 6464 high-quality, sequence-finished full-length cDNAs for Sitka spruce (Picea sitchensis). BMC Genomics 9(1):484
Renz A, Stitt M (1993) Substrate specificity and product inhibition of different forms of fructokinases and hexokinases in developing potato tubers. Planta 190(2):166–175
Riggs JW, Cavales PC, Chapiro SM et al (2017) Identification and biochemical characterization of the fructokinase gene family in Arabidopsis thaliana. BMC Plant Biol 17(1):83
Rolland F, Baena-Gonzalez E, Sheen J (2006) Sugar sensing and signaling in plants: conserved and novel mechanisms. Annu Rev Plant Biol 57(1):675–709
Saitou N (1987) The neighbor-joining method. A new method for reconstructing phylogenetic tree. Mol Biol Evol 4(4):406–425
Sanderson MJ (1989) Confidence limits on phylogenies: the bootstrap revisited. Cladistics 5(2):113–129
Sara EG, Jaina M, Alex B et al (2019) The Pfam protein families database in 2019. Nucleic Acids Res 47(D1):D427–D432
Schaffer AA, Petreikov M (1997) Sucrose-to-starch metabolism in tomato fruit undergoing transient starch accumulation. Plant Physiol 113(3):739–746
Schnarrenberger C (1990) Characterization and compartmentation, in green leaves, of hexokinases with different specificities for glucose, fructose, and mannose and for nucleoside triphosphates. Planta 181(2):249–255
Shi L, Cao S, Shao J et al (2014) Relationship between sucrose metabolism and anthocyanin biosynthesis during ripening in Chinese bayberry fruit. J Agric Food Chem 62(43):10522–10528
Swamy BN, Hedau NK, Chaudhari GV et al (2017) CMS system and its stimulation in hybrid seed production of Capsicum annuum L. Sci Hortic 222:175–179
Taylor MA, Ross HA, Gardner A et al (1995) Characterisation of a cDNA encoding fructokinase from potato (Solanum tuberosum L.). J Plant Physiol 145(3):253–256
Truernit E, Stadler R, Baier K et al (1999) A male gametophyte-specific monosaccharide transporter in Arabidopsis. Plant J 17(2):191–201
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93(1):77–78
Wei BQ, Wang LL, Bosland PW et al (2019) Comparative transcriptional analysis of Capsicum flower buds between a sterile flower pool and a restorer flower pool provides insight into the regulation of fertility restoration. BMC Genomics 20(1):837
Wei BQ, Wang LL, Bosland PW et al (2020) A joint segregation analysis of the inheritance of fertility restoration for cytoplasmic male sterility in pepper. J Am Soc Hortic Sci 145(1):3–11
Yang WQ, Lai Y, Li MN et al (2008) A novel C2-domain phospholipid-binding protein, OsPBP.1 is required for pollen fertility in rice. Mol Plant 1(5):770–785
Yang X, Ye J, Zhang L et al (2020) Blocked synthesis of sporopollenin and jasmonic acid leads to pollen wall defects and anther indehiscence in genic male sterile wheat line 4110S at high temperatures. Funct Integr Genomics 20(3):383–396
Yao Y, Geng MT, Wu XH et al (2017) Identification, expression, and functional analysis of the fructokinase gene family in Cassava. Int J Mol Sci 18(11):2398
Ye XY, Zhou WB (2021) Research advances in plant fructokinases. Chin Sci Bull 66(22):2820–2831 (in Chinese)
Zhang S, Nichols SE, Dong JG (2003) Cloning and characterization of two fructokinases from maize. Plant Sci 165(5):1051–1058
Zhang X, Chen B, Zhang L et al (2015) Identification of proteins associated with cytoplasmic male sterility in pepper (Capsicum annuum L.). S Afr J Bot 100:1–6
Zoerb C, Schmitt S, Muehling KH (2010) Proteomic changes in maize roots after short-term adjustment to saline growth conditions. Proteomics 10(24):4441–4444
Funding
The work was supported by the National Natural Sciences Foundation of China (31760572) and the scientific research start-up funds for openly recruited doctors (GAU-KYQD-2018-17) of Science and Technology Innovation Funds of Gansu Agricultural University, China.
Author information
Authors and Affiliations
Contributions
SZ and BW conceived and designed the experiments. SZ wrote the manuscript. BG and YW performed the qRT-PCR experiments. NY and PD and analyzed gene expression. MW and GZ shared their expertise in editing and revising the content of the manuscript. BW critically revised the manuscript. The manuscript was read and approved by all the authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhao, S., Gou, B., Wang, Y. et al. Identification and relative expression analysis of CaFRK gene family in pepper. 3 Biotech 12, 137 (2022). https://doi.org/10.1007/s13205-022-03196-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-022-03196-1