Abstract
The study of winter stress tolerance in perennial legumes needs to consider the complete symbiotic system including both plants and bacteria since these two partners are differentially affected by stress conditions. Here, we compared the regrowth after a freezing stress of four different associations of two alfalfa populations differing in freezing tolerance (A-TF0 and A-TF7) inoculated with two Sinorhizobium (Ensifer) meliloti strains (B399 and NRG34) of contrasted adaptation to cold. To understand the contribution of each partner to a better regrowth performance of an association after freezing, we identified molecular traits having major roles in cold acclimation, freezing tolerance, and those involved in the crosstalk between alfalfa and its symbiotic partner. Regrowth after exposure to a freezing stress was 35% larger in the A-TF7 × NRG34 than in the A-TF0 × B399 association. The metabolomic study of roots, crowns and, more specifically, nodules, revealed profound changes in these organs, switching from a sink to support cold acclimation to a source of reserves enabling regrowth after deacclimation. Marked increases in concentrations of stachyose and raffinose, two sugars of the raffinose-family oligosaccharides (RFO), and in the expression level of a gene of the RFO synthetic pathway were observed in response to cold acclimation supporting the importance of a protective role for RFO in alfalfa. Both cold-adapted partners of the symbiotic association contributed to increases in arginine concentration in nodules in response to cold acclimation and deacclimation underscoring the importance of N storage and remobilization for a successful overwintering in alfalfa.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Alfalfa (Medicago sativa L.) is the most important forage crop species and the fourth largest crop by area in Canada with 3 million hectares of pure stands or alfalfa-based mixtures (Statistics Canada 2022). Alfalfa ability to establish a symbiosis with the nitrogen (N) fixing bacterial partner Sinorhizobium (Ensifer) meliloti reduces the need for N fertilization of this crop as well as for subsequent crops making it a key contributor to the current efforts to reduce agriculture reliance on fossil fuels (Cummings 2005).
In northern latitudes, winter survival, field persistence and yield of alfalfa depend on its ability to tolerate low freezing temperatures (Bélanger et al. 2006; Seppänen et al. 2018). Predicted climate change will increase the risks of winter injury to alfalfa due to higher temperatures in the fall allowing less favorable hardening conditions and to the diminution of insulating snow cover protection during winter due to a higher incidence of freezing rain and thawing events (Bélanger et al. 2002).
Perennial plants such as alfalfa go through the process of cold acclimation that increases their tolerance to freezing temperatures and promotes the accumulation of reserve metabolites to support spring regrowth (Dhont et al. 2002). Cold acclimation elicits several molecular changes in plants (Bertrand et al. 2020a). For instance, low temperature induces starch hydrolysis and modifies the concentration of cryoprotective sugars such as sucrose, and raffinose-family oligosaccharides (RFO) in overwintering roots and crowns of alfalfa (Castonguay et al. 2009). Amino acids like proline, arginine and histidine also accumulate during cold acclimation (Dhont et al. 2006). Proline was shown to stabilize membranes and protect against oxidative stress (Szabados and Savoure 2009). Arginine is known as a precursor of polyamines which are involved in complex signaling networks that regulate stress responses (Alcázar et al. 2011; Menéndez et al. 2019; Pál et al. 2015) and is also an important N reserve to support spring regrowth (Dhont et al. 2006). Cold acclimation elicits major changes in gene expression and several cold regulated (COR) genes have been identified and characterized in alfalfa and other perennial species (Bertrand et al. 2017; Juurakko and Walker 2021). While the process of cold acclimation has been extensively studied in alfalfa, there is a knowledge gap regarding the deacclimation process and associated metabolic changes. Deacclimation occurs rapidly upon the return to warm temperatures in the spring. During that period, plants are more susceptible to damage by freeze–thaw cycles (Kalberer et al. 2006). The unpredictable effects of climate change on spring temperatures urge the need to better understand the deacclimation process since perennials will likely be exposed to more frequent and larger temperature fluctuations in springtime.
Alfalfa has been shown to have a large genetic diversity for freezing tolerance which has been successfully leveraged to improve this complex trait using a recurrent selection approach under controlled conditions (Bertrand et al. 2017). A potential relationship between freezing tolerance and changes in gene expression and metabolite composition in cold-acclimated alfalfa is highlighted by the differential accumulation of COR gene products and of metabolites in alfalfa populations recurrently selected for superior freezing tolerance (Castonguay et al. 2006).
Low temperature stress can not only reduce alfalfa survival and productivity but can also negatively impact its rhizobial partner and hinder the establishment of effective rhizobia-legume interactions (Prévost et al. 2003). The crosstalk between the symbiotic partners is initiated by the secretion of flavonoids by alfalfa (mainly formononetin, medicarpin and coumestrol) which concentrations have been shown to be modified by low temperature causing changes in nod genes activity in the symbiont (Zhang et al. 1996, 2009). Moreover, low temperatures has been shown to decrease Nod factors production by rhizobia while these lipo-chitooligosaccharides compounds are essential for the infection process (Duzan et al. 2006; Zhang et al. 1996). D’Amours et al. (2022) recently showed that choice of the rhizobial strain directly affects the level of nodule damage after a freezing stress. Different proportions of undamaged and necrotic nodules were observed according to the rhizobial strain in symbiosis with alfalfa, after exposure to freezing. This observed in vivo differences in nodule freezing damages was linked with plant yield: alfalfa inoculated with a freezing-tolerant strain had a larger proportion of active nodules with no damage after the freezing stress along with a greater plant vigor and shoot regrowth as compared to symbiosis with freezing-sensitive strains.
Varying responses of legume-rhizobia associations to environmental stresses were reported (Bertrand et al. 2020b; Sanz-Sáez et al. 2012). The underlying molecular bases of differential tolerance between host-symbiont combinations remain to be elucidated. A feedback regulation between host-plants and nodules has been shown to depend on resources availability and environmental conditions (Bertrand et al. 2016; Marquez-Garcia et al. 2015) as well as on the symbiotic rhizobial strain. For instance, when comparing three Bradyrhizobium strains in association with soybean growing under contrasted levels of atmospheric CO2, the strain associated with the highest yield also induced the highest ureides concentration in nodules under elevated CO2 along with the highest nitrogenase activity (Bertrand et al. 2011).This relationship indicated a positive-feedback stimulation: soybean mobilized energy reserves to support more nodules, and in return nodules synthesized more ureides to support plant growth.
Strains of S. meliloti have been shown to directly affect the level of freezing tolerance of alfalfa (Bertrand et al. 2007) and, recently, alfalfa regrowth after freezing was reported to differ depending on the associated S. meliloti strain (D’Amours et al. 2022). To better understand the mechanisms of tolerance of alfalfa-rhizobia associations, this study focused on metabolites and genes that were previously reported to play major roles in cold acclimation and freezing tolerance of alfalfa and nodules. Flavonoids involved in the crosstalk between alfalfa and rhizobia were also quantified as well as the expression of genes of their biosynthetic pathway.
2 Materials and Methods
Inoculum production, plant growth conditions and sampling procedures have been described in D’Amours et al. (2022) and are summarized below.
2.1 Sinorhizobium meliloti strains and plant material
Based on the results of our previous study on the effect of S. meliloti strains on alfalfa yield after a freezing stress (D’Amours et al. 2022), two strains inducing contrasted responses in alfalfa, ‘B399’ and ‘NRG34’, were selected for the present study. Plants inoculated with the commercial strain ‘B399’ (provided by Instituto de Genética “Edwald Alfredo Favret”, INTA, Buenos Aires, Argentina) were compared to plants inoculated with strain ‘NRG34’ isolated from Northwestern Canada (Rice et al. 1995). Both strains were grown in yeast extract mannitol (YEM) broth (Vincent 1970) at 28 °C for 24 to 48 h and a viability count was performed to adjust inoculum at 108 cells mL−1. These strains were used to inoculate two populations of alfalfa contrasted in their levels of freezing tolerance (i.e., A-TF0 and A-TF7). The cultivar ‘Apica’ (A-TF0) was developed at the Quebec Research and Development Centre (QRDC) of Agriculture and Agri-Food Canada (Michaud et al. 1983) and has a freezing lethal temperature for 50% of the plants (LT50) of -20 °C, while population ‘A-TF7’ was obtained after seven cycles of recurrent selection for improved freezing tolerance from the original cultivar Apica and has a LT50 of -26 °C (Bertrand et al. 2020a; D’Amours et al. 2022).
2.2 Plant growth conditions
Sterilized seeds of two alfalfa populations were individually seeded in Ray Leach Cone-tainers TM (SC-10 Super Cell. Stuewe & Sons Inc, Tangent, OR) filled with sterilized Turface® (Profile Products LLC, Buffalo Grove, IL). One week after seeding, plants were inoculated with 1 mL of either strain B399 or NRG34 containing 108 cells. Uninoculated controls for each alfalfa population (A-TF0 and A-TF7) were included to ensure that there was no uncontrolled sources of rhizobia contamination. These control plants failed to grow due to the lack of N input from fixation or nutrient solution and were not included in the statistical analysis. Plants were grown and sampled under the experimental conditions illustrated in Fig. 1 and as described in details in D’Amours et al. (2022). Briefly, plants were grown in a growth chamber under a temperature regime of 21/17 °C day/night and a 16 h-photoperiod and fertilized three times a week with 0.50 N-free Hoagland solution (Hoagland and Arnon 1950). Plants were harvested at the four following sampling events. 1) Non-acclimated (NA) plants were sampled after eight weeks of growth (32 plants: 8 replicates × 2 alfalfa populations × 2 strains). 2) Cold acclimated (CA) plants were sampled after an additional two weeks of growth at 2 °C under a 8 h-photoperiod, followed by two weeks at -2 °C in the dark (Fig. 1). The remaining plants were then exposed to a non-lethal freezing stress in a large programmable freezer in which temperature was gradually reduced from -2 °C to -11 °C (D’Amours et al. 2022), these frozen plants were then thawed for 24 h at 4 °C in darkness and exposed to the initial optimal regrowth conditions by progressively increasing the air temperature from 4 °C to 21/17 °C day/night. 3) Two days after this freezing stress (AFS), 32 plants were sampled to characterize the symbiosis immediately following the freezing stress. 4) The sampling of the remaining 32 plants was made after three weeks of regrowth after freezing (RAF) to study the effect of deacclimation.
Plants growth conditions and sampling events. Plants grown under controlled conditions were harvested at four sampling events: non-acclimated plants (NA) were sampled after being grown for 8 weeks under a 21/17°C, Day/Night (D/N) temperature regime. Plants were then cold acclimated for 4 weeks (CA) and sampled, and then exposed to a freezing stress of -11°C and transferred back to optimal regrowth conditions (21/17°C, D/N). Two days after freezing stress (AFS) and three weeks of regrowth after freezing (RAF) plants were sampled again
2.3 Root exudates sampling and biochemical analyses
2.3.1 Root exudates sampling
In a first step, root exudates were collected at each sampling event (i.e., NA, CA, AFS, and RAF). For this purpose, plants were carefully removed from their cone-tainers and gently shaken to remove the excess of turface and then washed three times in distilled water in order to remove any traces of turface and nutrient solution. Excess water was removed by gently pressing roots in absorbent paper towels. Each plant was then transferred into a 250-mL beaker containing 100 mL of ultrapure water and soaked for 10 min to collect root exudates. Plants were then removed from water and the soaking water containing root exudates was filtered using filter papers with 25 µm particle retention (Cytivia, Whatman™ no.4, Marlborough, MA). The remaining filtrate (75 mL) was separated into three equal volumes of 25 mL and transferred in 50-mL screw cap tubes (Sarstedt Inc Nümbrecht, Germany) in order to proceed to the biochemical analysis of three types of compounds: soluble sugars, amino acids and flavonoids. Tubes were kept on ice during sampling and then covered with perforated parafilm to allow freeze-drying. Root exudates were kept at -40 °C until being freeze-dried during 140 h to obtain dry-powered exudates (Labconco, Model Freezone12, Kansas City, MO).
2.3.2 Soluble sugars and amino acids extraction
A first tube of dry-powered root exudates was diluted in 1 mL of ultrapure water to suspend soluble sugars and free amino acids into an aqueous phase. Tubes were immediately heated 20 min at 65 °C to stop enzymatic activity. Tubes were then cooled in an ice water bath before being vortexed and centrifuged 30 s at 2,150 × g and the suspension was transferred into a 1.5 mL-microtube and kept frozen at -80 °C. Samples were centrifuged for 3 min at 11,350 × g prior to chromatographic analyses for soluble sugars and amino acids as described below.
2.3.3 Flavonoids extraction
A second tube of dry-powered root exudates was dissolved in 1 mL of MeOH 80% to extract flavonoids. Tubes were immediately heated 15 min at 65 °C then cooled in an ice water bath before being vortexed and centrifuged 30 s at 2,150 × g. The suspension was transferred to 1.5 mL microtubes and kept frozen at -80 °C. Samples were centrifuged for 3 min at 11,350 × g, prior to UPLC analysis.
2.4 Plant sampling for extraction of metabolites and RNA
Plants were separated into three major sections: roots systems, crowns and shoots that were kept on ice during sampling. For each plant, nodules were detached from roots using tweezers and transferred into 5-mL tubes. Nodules were freeze-dried prior to biochemical analysis and their total dry weight (DW) was recorded. The crowns, considered as the 2-cm subsections between shoots and roots, were sampled and analyzed separately because of their reported key role in the cold acclimation of alfalfa (Castonguay et al. 2009). Crowns were cut into two longitudinal sub-sections, and their fresh weight was recorded. The first crown sub-section (approx. 0.1 g) was flash frozen and manually ground in liquid nitrogen and kept at − 80 °C for RNA extraction. The second sub-section of crowns was freeze-dried (Labconco, Model Freezone12), the DW was recorded and samples were ground using an OMNI Bead Ruptor 24 (PerkinElmer, Inc. Kennesaw, GA) before further biochemical analyses. A 8 cm-long subsample of roots containing a mixture of fine roots and tap roots was weighted (around 0.1 g) and flash frozen in liquid nitrogen for RNA extraction. A second root subsample was weighted (approx. 0.2), freeze-dried, and ground (OMNI Bead Ruptor) prior to biochemical analysis. The remaining part of the root system was dried separately at 55 °C for 72 h for the measurement of total root DW (including the calculated DW of all sub sections). Shoots were dried at 55 °C for 72 h for the measurement of total shoot dry weight (including the calculated DW of crown sub-sections).
Grinded samples of crowns and roots were analyzed for their soluble sugars and amino acids contents after extraction in methanol-chloroform-water as described in Bertrand et al. (2020b). Grinded samples of nodules were analyzed for their content in soluble sugars, amino acids, and flavonoids. For this purpose, the total dry weight of each nodules samples (between 0.02 and 0.09 g) was extracted in 2 mL of MeOH 80%. Tubes were heated 20 min at 65 °C, rapidly cooled on ice and centrifuged (10 min at 1,200 × g at 4 °C). A first subsample of 0.9 mL was evaporated to dryness (Savant Speedvac plus SC210A, Holbrook, NY), solubilized in 0.9 mL of ultrapure water, and kept at -20 °C until the analysis of soluble sugars (HPLC, Waters Inc. Milford, MA) and amino acids (UPLC, Acquity, Waters Inc, Milford, MA). A second subsample of 0.9 mL of the supernatant was transferred into 1.5 mL microtubes and frozen at -20 °C until flavonoid analysis concentration by UPLC. Subsamples of roots were also extracted in MeOH 80% and treated similarly to analyze their flavonoids concentration. Prior to chromatographic separation by HPLC or UPLC, all samples were centrifuged for 3 min at 11,350 × g.
2.5 Metabolites quantification
2.5.1 Quantification of carbohydrates and amino acids
Soluble sugars were separated and quantified on a chromatographic analytical system controlled by the Empower II software (Waters, Milford, MA) as described in Bertrand et al. (2020b). Peak identity and quantity were determined for raffinose, stachyose, sucrose, glucose, fructose and pinitol by comparison to standards (Sigma–Aldrich, Oakville, ON, Canada). Starch was quantified in non-soluble residues following the extraction of soluble sugars. Starch in the residues was hydrolyzed into glucose by adding 3 mL of digestion buffer (200 mM sodium acetate, pH 4.5) containing amyloglucosidase (15 U mL−1; Sigma-Aldrich, Oakville, ON, Canada), and incubation (60 min at 55 °C). After centrifugation, the supernatant was collected for quantification of glucose by HPLC, as described above. Starch concentration was calculated by subtracting the amount of soluble glucose from the total amount of glucose measured following digestion with amyloglucosidase. For crowns, roots and nodules, results of carbohydrates determinations were expressed as concentrations on a dry matter (DM) basis (mg g−1 DM). For root exudates, results were reported on a root DM basis by taking into account the DM of the root systems that were soaked in ultrapure water to extract exudates. Total soluble sugars (SSTot) is the sum of concentrations of individual sugars raffinose, stachyose, sucrose, pinitol, glucose and fructose and non-structural carbohydrates (NSC) is the sum of SSTot and starch.
Twenty-one amino acids (alanine, arginine, asparagine, aspartate, glutamate, glutamine, glycine, γ-aminobutyric acid (GABA), α- aminobutyric acid (AABA), histidine, proline, methionine, lysine, serine, leucine, isoleucine, ornithine, phenylalanine, threonine, tyrosine and valine) were separated and quantities were determined by comparison to a standard mix containing the 21 amino acids. Each individual amino acid were provided by Sigma–Aldrich (Oakville, ON, Canada) and the standard mix was prepared in our laboratory. For crowns, roots and nodules, results were expressed as concentrations on a DM basis (µmol g−1 DM). For root exudates results were reported in µmol on root DM basis by taking into account the DM of the root systems that were soaked in ultrapure water to extract exudates. The total free amino acids (AATot) was the sum of concentrations of the 21 free amino acids.
2.5.2 Quantification of flavonoids
Naringenin, luteolin, echinatin, coumestrol, formononetin and medicarpin and their conjugates were separated and quantified using Waters ACQUITY UPLC analytical system controlled by the Empower II software (WATERS, Milford, MA, USA). Flavonoids were separated using a BEH C8 column (2.1 mm × 100 mm, 1.7 µm, Waters), and detected using a Photodiode Array (PDA) Detector set at 287 nm. The chromatographic conditions were as follows: column temperature, 35 °C flow rate, 0.35 mL min−1, mobile phase A, 0.2% formic acid in water, mobile phase B: methanol 100%. The gradient was of 65% A at 0.2 mL min−1 and a gradual decrease until 2%, before to returning to initial condition of 65% A, for a total run of 8 min. Peak identity and quantity of each flavonoids were determined by comparison to standards. Results for flavonoids determinations in roots were expressed as concentrations on a dry matter (DM) basis (µg g-1DM). For root exudates, the calculation was as described in the above section. Total flavonoids (FlaTot) are the sum of each individual concentration of the seven flavonoids.
2.6 Analysis of gene expression
2.6.1 RNA extraction and cDNA synthesis
Total RNA was extracted from 0.1 g of crowns and roots using CTAB-based protocol (Dubé et al. 2013). Total RNA was quantified using NanoDrop™ One/OneC Microvolume UV–Vis Spectrophotometer (Thermo Fisher Scientific Inc. Waltham, MA). Residual genomic DNA was removed by a treatment with DNaseI (Invitrogen, Burlington, ON, Canada) prior to cDNA synthesis. First strand cDNA was synthesized from 1 μg of total RNA and oligo(dT)18 primers using the Transcriptor First Strand cDNA synthesis Kit (Roche Applied Science, Laval, QC, Canada) following the manufacturer instructions. cDNA synthesis reactions were performed for each sample considered in the subsequent RT-qPCR analyses. Thus, a total of 256 reactions were performed: 8 replicates × 2 alfalfa populations × 2 strains × 4 sampling dates × 2 organs (crowns and roots).
2.6.2 RT- qPCR analysis of COR gene expression
The expression of seven genes of interested (GOI) that are cold-regulated (COR) was measured and compared between the different treatments (Table 1). The five genes related to cold acclimation and freezing stress tolerance were selected from previous studies on alfalfa (Bertrand et al. 2017; Castonguay et al. 2015; Dubé et al. 2013) and their expressions were analyzed in crown samples. Two genes of interest linked to the phenylpropanoid synthetic pathway leading to flavonoid synthesis were selected based on literature (Gifford et al. 2018) and their expressions were measured in alfalfa root samples. Specific qPCR primers were designed using the methodology described in Castonguay et al. (2020), to amplify these last two genes of interests in Medicago sativa samples. Primer design was based on homologous Medicago tranculata genes sequences retrieved from a BLASTn search on NCBI database [Medtr4g088190 (IOMT or 2,7,4′-trihydroxyisoflavanone 4′-O-methyltransferase, EC 2.1.1.212) and Medtr5075450 (C4H or cinnamic acid 4-hydroxylase EC 1.14.13.11)] (Gifford et al. 2018). The sequences of all primers selected for RT-qPCR analysis of GOI as well as of the three reference genes described below are listed in Table 1. The expression of COR genes was measured in crowns as this tissue plays a key role in cold acclimation of alfalfa. The expression of genes involved in flavonoid biosynthesis was measured in roots as this tissue is the synthetic site of flavonoids.
The RT-qPCR was carried out in a Mastercycler® ep realplex system (Eppendorf Canada, Mississauga, ON, Canada) using the QuantiTect® SYBR Green PCR kit (QIAGEN, Toronto, ON, Canada) as described by Dubé et al. (2013). The 10-μl reaction mixture contained 3 μl of first-strand cDNA and 0.5 μM of each of the forward and reverse primers. The thermocycler program was set to: 15 min at 95 °C for denaturation; 40 cycles of 15 s at 95 °C, 15 s annealing at 60 °C; 60 s extension at 72 °C. Reactions with real-time PCR were carried out with control water samples included as checks for potential contamination with genomic DNA. Efficiency of PCR was calculated from the linear regression of a seven fold dilution of PCR products using the following equation: Efficiency % = (10(−1/slope)-1) × 100. The threshold cycle (Cq) values at which the PCR product fluorescence rises over the background fluorescence was determined by the instrument software which was set to default parameters. For the normalization of results, reference genes were selected by screening several candidates identified by Castonguay et al. (2015) for the stability of their expression using the geNormPlus M value program included in the qBasePlus software (Biogazelle, Ghent, Belgium). Results were normalized in each tissues with three reference genes: ubiquitin 5 (crown), eukariotic elongation factor 1-alphas (roots) and H-ATPase (crowns and roots). Relative quantification was calculated with the qBase software using the 2-ΔΔCq or comparative Cq method based on the differences in Cq between the target and the reference genes and corrected for PCR efficiency.
2.7 Statistical analysis
Statistical analysis of biomass were made using a one-way analysis of variance (ANOVA) for a randomized complete block design with the SAS MIXED procedure (SAS® Studio, 2020, Version 3.81, SAS Institute Inc., Cary, NC). The model was used to compare the effects of the associations between alfalfa populations and S. meliloti strains on shoot regrowth and nodules biomass. Concentrations of metabolites and gene expression were analyzed using a two-way analysis of variance (ANOVA) model for a randomized complete block design with the SAS MIXED procedure (SAS® Studio, 2020, Version 3.81, SAS Institute Inc., Cary, NC). The model was used to establish, in a first step, the effects of sampling events. ANOVA were then performed for each sampling events separately to establish the effects of the two alfalfa populations, the two S. meliloti strains, and their interactions on the concentrations of carbohydrates, amino acids, and flavonoids and on genes expression, for each plant organs and root exudates. The Residual normality and variance homogeneity were verified using the UNIVARIATE procedure. The Shapiro–Wilk’s and Kurtosis’s tests were used to verify the normality of the data distribution. Pairwise comparisons of means differences were made using a Fisher’s least significance difference test (LSD) at P ≤ 0.05. The log twofold changes in metabolite concentrations that significantly differed between alfalfa populations and S. meliloti strains were calculated in non-acclimated (NA), cold acclimated (CA), 48 h after freezing stress (AFS) and regrowth after freezing (RAF) plants. The graphical representation of metabolite variations related to either alfalfa populations or S. meliloti strains based on this calculation shows the differential contribution of each alfalfa population (A-TF0 vs A-TF7) and each strain (B399 vs NRG34) to the metabolic changes at each sampling event.
3 Results
3.1 Comparative assessment of shoot and nodule biomass in alfalfa-rhizobia associations under varying environmental conditions
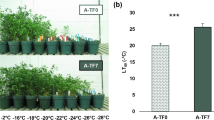
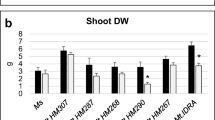
There was no difference in shoot DW between the non-acclimated (NA) four alfalfa-rhizobia associations tested (Table 2). In contrast, shoot regrowth three weeks after exposure to freezing stress (RAF) was significantly different between the populations × strains associations. The shoot DW of the freezing tolerant alfalfa population A-TF7 inoculated with the freezing tolerant strain NRG34 was significantly higher (+ 35%) than the shoot DW observed with the combination of the less freezing tolerant population and freezing sensitive strain A-TF0 × B399 (Fig. 2a). The shoot DW of A-TF0 × NRG34 was also significantly higher (+ 17%) than that of A-TF0 × B399. It is noteworthy that the shoot DW of the A-TF7 × B399 was intermediate between those of A-TF0 × B399 and A-TF0 × NRG34 even though it did not significantly differ with from the shoot DW of these two associations. The four alfalfa-rhizobia associations also differed in nodules DW after a freezing stress (Table 2, Fig. 2b). While no difference in nodules DW was observed in NA and CA plants, the nodules DW of A-TF0 × NRG34 was significantly larger than both associations with strain B399 two days after the freezing stress (AFS). The significant difference in nodules DW between A-TF0 × NRG34 and the other three alfalfa-rhizobia associations was much larger three weeks after the exposure to a sublethal freezing stress (RAF).
Shoot dry weight (DW) (a), and nodules DW (b) of four associations combining two alfalfa populations A-TF0 (in grey) and A-TF7 (in white) inoculated with two S. meliloti strains B399 (lined pattern) and NRG34 (dotted pattern). Alfalfa plants were grown under controlled conditions and sampled at four sampling events: non-acclimated plants (NA) were grown 8 weeks under a 21/17 °C, Day/Night (D/N) temperature regime. Plants were then cold acclimated during 2 weeks at 2 °C followed by two weeks at -2 °C and sampled again (CA). After their exposure to a freezing stress of -11 °C, alfalfa plants were transferred back to optimal regrowth conditions (21/17 °C, D/N) and sampled after two days (AFS), and after three weeks (RAF). Shoots were only sampled for NA and RAF plants since CA and AFS shoots and leaves were killed by the 2 weeks exposure to subfreezing temperature in the dark while nodule dry weight was measured at the four sampling events. Error bars represent the Standard Error of the Mean (SEM), n = 8. No letter means that there is no significant differences between treatments for that event sampling while different letters represent significant differences as determined by the Fisher’s least significant difference (LSD) test at P ≤ 0.05
3.2 Variations in metabolite concentrations between sampling events
3.2.1 Non-structural carbohydrates
Concentrations of starch and soluble sugars in roots, crowns and nodules significantly varied between sampling events (Table 3, Fig. 3). This observation was also true for root exudates except for glucose and fructose concentrations that did not vary between samplings. Starch concentration under NA conditions was much higher in roots (322 mg g−1 DM) and crowns (272 mg g−1 DM) than in nodules (1.6 mg g−1 DM). After four weeks of CA, starch concentrations were reduced by one half in roots and crowns while its concentration doubled in nodules (Fig. 3). Starch concentrations decreased in subsequent AFS and RAF sampling points in nodules, roots and crowns to reach levels that were lower than those measured in NA plants. Concomitantly, a marked increase was observed in total soluble sugars in CA crowns (+ 222%), roots (+ 200%) and nodules (+ 136%) as compared to NA. More specifically, sucrose concentration in nodules, roots and crowns was 2.5 to 4 times higher in CA than in NA plants and then decreased gradually in subsequent AFS and RAF samplings. Concentration of glucose decreased in roots and crowns for CA, AFS and RAF when compared to NA plants while in nodules, a slightly higher glucose concentration was observed in CA as compared to NA. Cold acclimation induced a marked increase of stachyose and raffinose in root exudates, roots, crowns and nodules. While stachyose and raffinose were undetectable in NA nodules, they reached concentrations as high as 9 and 17 mg g−1, respectively, in CA nodules and then decreased progressively to 0 in subsequent sampling events. Generally, the fructose concentration remained stable in nodules, roots, and crowns in response to CA but a significant increase in fructose concentration was observed in nodules of AFS and RAF plants as compared to NA and CA plants. The concentration of pinitol in nodules decreased progressively at each subsequent sampling events in exudates, nodules, roots and crowns as compared to NA.
Average concentration of metabolites measured in exudates, nodules, roots and crowns of the four alfalfa × strain associations at each sampling event (n = 32). Metabolite concentrations of sugars (mg/g DM), amino acids (µmol/g DM) and flavonoids (µg/g DM) were measured in non-acclimated plants (NA), cold-acclimated (CA) plants, plants exposed to a freezing stress and transferred back to optimal regrowth conditions for 48 h (AFS), and after a 3-weeks regrowth (RAF). Colors represent the log twofold change of either a decrease (red) or an increase (blue) of metabolite concentrations compared to NA plants. Color code is shown at the bottom of the last column of the Fig. 3. Ala, alanine; Arg, arginine; Asn, asparagine; Asp, aspartic acid; Formo, formononetin, FlaTot, total flavonoids; Gln, glutamine; Glu, glutamic acid; Gly, glycine; His, histidine; Ile, isoleucine; Leu, leucine; Lys, leucine; Lys, lysine; Met, methionine; NSC, non structural carbohydrates; Phe, phenylalanine; Pro, proline; Ser, serine; Thr, threonine; Tyr, tyrosine; Val, valine; Orn, Ornithine; AATot, Total amino acid; SSTot, total soluble sugars; AABA, α-aminobutyric acid; GABA, γ-aminobutyric acid
3.2.2 Amino acids
Concentrations of total free amino acids in NA samples were much higher in nodules (523.4 μmol g−1 DM) comparatively to roots (143.4 μmol g−1 DM), crowns (131.8 μmol g−1 DM), and root exudates (2.5 µmol g−1 root DM (Fig. 3). Total amino acids concentration decreased with CA in root exudates and in nodules while it remained stable between samplings in roots and crowns (Table 3, Fig. 3). In contrast, individual free amino acids varied in roots, crowns and nodules between samplings except for proline in nodules and crowns, aspartate in nodules, and AABA in roots. Contrarily to the increase that was generally observed in nodules, roots and crowns between NA and the other samplings, individual free amino acids mostly decreased in root exudates at CA and AFS samplings with the exception of methionine and arginine that remained stable. Asparagine was by far the most abundant amino acid, representing 35% of total amino acids in root exudates, 80% in nodules, 58% in roots and 52% in crowns in NA samples. Cold acclimation induced a decline in asparagine concentration in all samples. In crowns, asparagine concentration re-increased after a 3-wks regrowth (RAF) reaching a level higher than in NA samples. Glutamate concentrations slightly increased in CA and AFS roots and crowns compared to NA. An increase in concentrations of glutamine was observed in CA and AFS nodules while a decrease occurred in roots and crowns at those sampling events. Concentrations of ornithine, arginine and histidine increased significantly in CA roots and crowns (avg. of + 82, + 77, and + 42%, respectively) and even more so in nodules (+ 917, + 798, and + 129%, respectively). The concentration of these amino acids remained high in the AFS samples. Concentrations of alanine and glycine increased in CA nodules, remained stable in AFS samples and subsequently decreased to a level lower than that observed in NA nodules in RAF samples. Alanine and glycine concentrations initially decreased in CA roots and crowns and re-increased in AFS samples. Serine concentrations increased with cold acclimation in nodules (+ 219%), roots (+ 70%) and crowns (+ 62%), whereas its concentration decreased in root exudates. An increase in threonine, lysine, isoleucine, leucine, valine, tyrosine and phenylalanine was observed in nodules, roots and crowns 48 h after a freezing stress (AFS) but their levels remained low. The GABA concentration was at a lower level in nodules and roots at RAF as compared to the other samplings. The AABA concentrations decreased in nodules at CA, AFS and RAF compared to NA, while it increased slightly in response to CA and AFS in crowns.
3.2.3 Flavonoids
Total flavonoid concentrations were higher in roots (89 mg g−1 DM) than in nodules (52 mg g−1 nodules DM) and in root exudates (2.7 mg g−1 root DM) of NA plants (Table 3, Fig. 3). In CA, AFS and RAF plants, concentration of total and individual flavonoids differed between samplings. In general, total flavonoids increased at each subsequent sampling in nodules while it decreased in roots and root exudates (Fig. 3). Formononetin was the flavonoid with the highest concentration in all samples, representing slightly more than 75% of all flavonoids in root exudates and nodules, and 95% in roots. Formononetin concentration decreased in response to CA and AFS in nodules and roots while it increased in root exudates in response to CA. Most of individual flavonoids decreased in root exudates with CA and AFS except for formononetin and medicarpin. In roots, concentrations of naringenin and medicarpin were not impacted by the cold acclimation and freezing stress. In nodules, all flavonoids except for fomononetin increased in CA samples and showed an additional increase 48 h after the freezing stress (AFS) with concentrations of naringenin (+ 48%), luteolin (+ 107%), echinatin (+ 367%), coumestrol (+ 253%) showing as compared to NA (Fig. 3). In roots, only luteolin and echinatin increased with CA and freezing stress, while formononetin and coumestrol decreased when compared to NA.
3.3 Variations in metabolite concentrations related to alfalfa populations in cold acclimated (CA) and deacclimated (RAF) plants
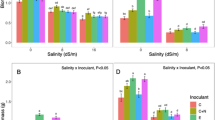
Concentrations of various metabolites differed significantly between alfalfa populations in CA and RAF samples (Figs. 4 and 5 left panels, Supplemental Tables 1–4). In CA plants, the freezing-tolerant population A-TF7 had higher concentrations of total soluble sugars and sucrose in roots and crowns while pinitol and glucose were more abundant only in A-TF7 crowns as compared to the less freezing tolerant A-TF0 (Fig. 4, left panel). Starch was more abundant in roots of A-TF0 than A-TF7. For amino acids, ornithine was the amino acid showing the most striking difference between the two alfalfa populations with higher level in nodules, roots, and crowns of population A-TF7 when compared to A-TF0. Arginine was the second amino acid with highest level in nodules and in roots of population A-TF7 when compared to A-TF0, followed by histidine in nodules and lysine in both nodules and roots. GABA concentration was higher in nodules and roots of population A-TF0 compared to A-TF7. Methionine concentration was also higher in crowns of population A-TF0. We also observed higher concentrations of histidine in root exudates and in nodules of population A-TF7. For flavonoids, higher concentration of medicarpin was observed in root exudates and higher concentration of naringenin and luteolin in nodules associated with A-TF0 than those associated with population A-TF7.
Graphic representation of log twofold changes in metabolites concentration showing the differential contribution of each alfalfa population (A-TF0 vs A-TF7) and each S. meliloti strain (B399 vs NRG34) to the metabolic changes in CA root exudates, nodules, roots and crowns. The left panel compares the two alfalfa populations with significant higher concentration in A-TF7 on the right to zero line (white) and significant higher concentration in A-TF0 on the left (black). The right panel compares the two rhizobial strains with significant higher concentration in response to inoculation with NRG34 on the right to zero line (white) and significant higher concentration in response to B399 on the left (black). Ala, alanine; Arg, arginine; Asn, asparagine; Asp, aspartic acid; Formo, formononetin, FlaTot, total flavonoids, Gln, glutamine; Glu, glutamic acid; Gly, glycine; His, histidine; Ile, isoleucine; Leu, leucine; Lys, leucine; Lys, lysine; Met, methionine; NSC, non structural carbohydrates; Phe, phenylalanine; Pro, proline; Ser, serine; Thr, threonine; Tyr, tyrosine; Val, valine; Orn, ornithine, AATot, Total amino acid; SSTot, total soluble sugars; AABA, α-aminobutyric acid; GABA, γ-aminobutyric acid
Graphic representation of log twofold changes in metabolites concentration showing the differential contribution of each alfalfa population (A-TF0 vs A-TF7) and each S. meliloti strain (B399 vs NRG34) to the metabolic changes in RAF root exudates, nodules, roots and crowns. The left panel compares the two alfalfa populations with significant higher concentration in ATF-7 on the right to zero line (white) and significant higher concentration in ATF-0 on the left (black). The right panel compares the two rhizobial strains with significant higher concentration in response to inoculation with NRG34 on the right to zero line (white) and significant higher concentration in response to B399 on the left (black). Ala, alanine; Arg, arginine; Asn, asparagine; Asp, aspartic acid; Formo, formononetin, FlaTot, total flavonoids, Gln, glutamine; Glu, glutamic acid; Gly, glycine; His, histidine; Ile, isoleucine; Leu, leucine; Lys, leucine; Lys, lysine; Met, methionine; NSC, non structural carbohydrates; Phe, phenylalanine; Pro, proline; Ser, serine; Thr, threonine; Tyr, tyrosine; Val, valine; Orn, ornithine, AATot, Total amino acid; SSTot, total soluble sugars; AABA, α-aminobutyric acid; GABA, γ-aminobutyric acid
Three weeks after the freezing stress (RAF) concentrations of starch in both roots and crowns and raffinose and NSC in crowns were higher in population A-TF7 than in A-TF0 while glucose concentration was higher in A-TF0 roots (Fig. 5 left panel, Supplemental Table 4). Concentrations of arginine and ornithine were higher in both nodules and roots of A-TF7 than in A-TF0. Histidine concentration was also higher in nodules associated with population A-TF7. Concentrations of glutamine in roots and aspartate and glutamate in both roots and crowns were higher in A-TF7 than in A-TF0. GABA concentration was higher in roots and crowns and alanine and serine in crowns of A-TF0. For flavonoids only a difference in naringenin concentration in root exudates was found between the two alfalfa populations with more than twice the concentration in A-TF0 than in A-TF7 (Fig. 5). Variations in metabolite concentrations related to alfalfa populations in NA and AFS plants are presented in supplemental Figs. 1 and 2.
3.4 Variations in metabolite concentrations related to S. meliloti strains in cold acclimated (CA) and deacclimated (RAF) plants
Differences in the metabolite profiles of alfalfa-rhizobia associations established with freezing-sensitive strain B399 and freezing-tolerant strain NRG34 were observed after cold acclimation (CA). More noticeable variations occurred in nodules than in roots and crowns (Fig. 4 right panel, Supplemental Table 2). For non-structural carbohydrates, pinitol concentrations in nodules and crowns as well as glucose and fructose concentrations were higher in nodules of plants inoculated with strain B399 than with NRG34. Differences in cold-induced changes in concentrations of specific free amino acids were also noticeable between strains. Higher concentrations of AABA, aspartate, histidine and threonine in root exudates, AABA and tyrosine in nodules, and GABA and glutamine in roots were detected in the symbiotic association with strain B399. Conversely, higher concentrations of methionine, arginine, and histidine were observed in nodules of alfalfa inoculated with strain NRG34 than in those inoculated with strain B399. As for flavonoid concentrations, luteolin and naringenin concentrations were higher in nodules of plants inoculated with strain NRG34 than in those associated with strain B399.
After three weeks of regrowth after freezing stress (RAF), higher concentrations of pinitol were still detected in crowns and nodules of alfalfa inoculated with strain B399 (Fig. 5 right panel, Supplemental Table 4). For amino acids, concentrations of proline, aspartate, and glycine were higher in nodules of alfalfa inoculated with strain B399 than in nodules of alfalfa inoculated with strain NRG34 (Fig. 5 right panel). Arginine and phenylalanine were alternatively more abundant in nodules of alfalfa inoculated with strain NRG34 than in nodules of alfalfa inoculated with strain B399. Total flavonoid and formononetin concentrations were higher in roots of alfalfa inoculated with strain NRG34 than in those inoculated with strain B399 as well as concentration of luteolin in nodules (Fig. 5). Variations in metabolite concentrations related to S. meliloti strains in NA and AFS plants are presented in supplemental Figs. 1 and 2.
Few interactions between the alfalfa population and the S. meliloti strain on metabolites concentrations were observed in nodules and crowns in response to CA, AFS and RAF. Higher concentrations of fructose, alanine, AABA and medicarpin for the association A-TF0 × B399 were noticeable cases (Supplemental Tables 2–4) as well as variations in the concentration of several free amino acids in root exudates in AFS (Supplemental Table 3).
3.5 Gene expression in crowns and roots in response to sampling event, alfalfa populations and rhizobial strains
The relative expression of all the genes of interest (GOI) measured in crowns was impacted by the sampling events (Table 4). When expression profiles were compared to those in NA plants, the most striking modifications of gene expression were observed in response to CA with the up-regulation of galactinol synthase (GaS) and K3-dehydrin, and the down-regulation of sucrose synthase (SuSy) expressions (Table 4). The expression levels of these three genes returned to a level comparable to NA plants at the subsequent samplings (AFS and RAF). As observed in AFS crowns, freezing stress induced the down-regulation of SPS gene (Table 4). When looking at the effect of alfalfa populations on gene expression, the relative expression of SuSy was higher in A-TF0 than in A-TF7 at the CA and RAF samplings (Figs. 6a and b, Supplemental Table 5). Gene expression of SuSy in crowns did not differ according to strain inoculation (Fig. 6).
Relative expression in alfalfa crowns of sucrose synthase (SuSy) in (a) cold-acclimated (CA) and (b) 48 h after freezing stress in two alfalfa populations contrasted in their freezing tolerance levels (A-TF0 and A-TF7) analyzed by RT-qPCR. Error bars represent the SEM, n = 16. Significant differences between populations at each event, determined by a t-test, are indicated with the following levels of probability: * P ≤ 0.05, ** P < 0.01, ***
In roots, the expression of genes coding for enzymes involved in the biosynthetic pathways of flavonoids and secondary metabolites differed according to sampling events (cinnamic acid 4-hydroxylase (C4H; and 2,7,4′-trihydroxyisoflavanone 4′-O-methyltransferase (IOMT; P = 0.021). CA and AFS roots show that cold acclimation and freezing stress induced slight but significant up-regulation of C4H gene and down-regulation of the IOMT gene (Table 4). The gene expression of both genes in roots were lower in RAF than in NA roots. The relative expression of C4H was higher in A-TF0 than in A-TF7 in the NA and RAF samplings (Figs. 7a and b, Supplemental Table 5) while the expression of IOMT was higher in A-TF0 than in A-TF7 in CA plants (Fig. 7c, Supplemental Table 5). We also observed an interaction between alfalfa population and S. meliloti strains on the relative expression of IOMT in AFS sampling (Fig. 8d and Supplemental Table 5). Relative expression of IOMT was 77% higher for alfalfa population A-TF0 combined with strain B399 when compared to alfalfa population A-TF7 combined with the same strain and both association of alfalfa population inoculated with NRG34 were intermediate and were not different (Fig. 7d).
Relative expression in roots of cinnamic acid 4-hydroxylase (C4H) in non-acclimated NA alfalfa (a) and in RAF alfalfa regrowth (b). Relative expression of isoflavone O-methyltransferase (IOMT) (c) in two CA alfalfa populations contrasted in their freezing tolerance levels (A-TF0 and A-TF7) and of four associations combining two alfalfa populations (grey scale) A-TF0 (in grey) and A-TF7 (in white) inoculated two S. meliloti strains (patterns), B399 (lined) and NRG34 (dotted), sampled 48 h after freezing (AFS) (d) Error bars represent the SEM for n = 16. Significant differences between populations at each event, determined by a t-test, are indicated with the following levels of probability: * P ≤ 0.05, ** P < 0.01, *** P < 0.001. Different letters indicate significant differences in relative expression of IOMT due to pop × strains interactions at P ≤ 0.05 (n = 8)
4 Discussion
The study of winter stress tolerance in legumes needs to consider the complete symbiotic system including both plants and bacteria since these two partners are differentially affected by stress conditions (Hawkins and Oresnik 2022). Here, we compared the regrowth after a freezing stress of four different associations of alfalfa populations and S. meliloti strains and observed up to 35% yield differences between the best (A-TF7 × NRG34) and the worst (A-TF0 × B399) association (Fig. 2a). Both partners were shown to contribute to the recovery after exposure to a freezing stress with a 12% difference that was explained by a population effect (A-TF0 vs A-TF7) and 17% that was attributable to a strain effect (B399 vs NRG34). To understand the contribution of each partner to the better regrowth performance of an association after freezing, a thorough investigation of the mechanisms at play was undertaken by monitoring metabolites and genes having major roles in cold acclimation, freezing tolerance, and those involved in the crosstalk between alfalfa and its symbiotic partner. The metabolomic study of roots, crowns and, more specifically, of nodules, revealed profound changes in these organs, switching from a sink to support cold acclimation to a source of reserves enabling regrowth after deacclimation. Our approach tracing molecular changes separately in alfalfa populations and rhizobial strains provided a novel perspective on their respective contribution to the vigor of spring regrowth of alfalfa. It also allowed the identification of molecular traits potentially conferring superior productivity of alfalfa that was exposed to a sublethal freezing stress.
4.1 Effects of cold acclimation, freezing stress and deacclimation on metabolites and gene expression
4.1.1 Sugars
In response to cold acclimation we noted an important increase of soluble sugars concentrations in nodules, roots and crowns of alfalfa. Those observations are concordant with similar results reported of the accumulation of soluble sugars in perennial organs of alfalfa in response to low temperature and photoperiod to provide energy for plant overwintering and regrowth (Fig. 3; Bertrand et al. 2017; Castonguay et al. 2011). The separate biochemical analysis of nodules allowed us to better understand the pivotal role of these belowground organs. For instance, our results highlight the importance of nodules as large carbon sinks during cold acclimation leading not only to the accumulation of sucrose, stachyose, raffinose, and starch in these organs, but also to a significant increase in their biomass after exposure to cold temperatures (CA) (Fig. 2). The allocation of photosynthates into nodules has been linked with the translocation of sugars from other plant organs such as leaves and roots in response to abiotic stress (Bertrand et al. 2016). The large hydrolysis of starch in roots and crowns likely contributed to the new carbohydrates input, including starch, in nodules (Fig. 3). An accumulation of starch grains, amyloplast and oleosomes have been observed in perennial beach pea nodules before winter (Chinnasamy and Bal 2003; Gurusamy et al. 2000) and these storage compounds were proposed to serve as energy source for metabolic activities during winter dormancy of nodules that lasts for prolonged periods under arctic/subarctic conditions. In our study, the concentration of total sugars in nodules decreased to levels below those observed in NA nodules upon the return of alfalfa to optimal regrowth conditions (RAF). This observation makes a compelling case for soluble sugars and starch as key contributors to the nodules survival during overwintering period and as a source of energy to resume N fixation and ultimately support alfalfa regrowth in the spring (D’Amours et al. 2022). An important accumulation of sucrose was observed in nodules, roots and crowns in response to cold acclimation while its concentration decreased in roots exudates with sampling events. Sucrose is a recognized cryoprotectant that can stabilize freeze-dehydrated cells by interacting with cell membrane (Tarkowski and Van den Ende 2015). In alfalfa, a close relationship between sucrose accumulation and superior freezing tolerance has been demonstrated (Castonguay et al. 2011). The decrease in sucrose at regrowth after freezing (RAF) could be partly explained by the quick up-regulation of sucrose synthase (Susy) in nodules after freezing stress to cleave sucrose into fructose and glucose and to provide succinate and malate to the bacteroids, directly feeding into their tricarboxylic acid cycle (TCA) cycle (Geddes and Oresnik 2014; Liu et al. 2018). Interestingly, fructose concentration increased more than three fold in nodules 48 h after freezing stress. Fructose has a high capacity for scavenging superoxide and has been shown to be involved in antioxidative protection in pea under chilling stress (Bogdanović et al. 2008). The marked increase in RFO in root exudates, roots, crowns and nodules in response to CA confirm their importance in the freezing tolerance of alfalfa. Concentrations of RFO are intrinsically linked with the level of freezing tolerance of alfalfa due to their cryoprotective actions (Bertrand et al. 2017). Furthermore, RFO are known to support the growth and survival of symbiotic N-fixing bacteria (e.g. S. meliloti) in the rhizosphere of germinating seeds and alfalfa seedlings (Bringhurst et al. 2001). The larger concentration of RFO in nodules than in other perennial organs (crowns and roots) highlights the importance of the investment of host plant into nodules protection to maximize their survival and allow for a quick return of N fixation in the spring to support alfalfa regrowth.
4.1.2 Amino acids
Amino acids constitute an important source of organic N for spring regrowth and several amino acids possess osmoprotectant and cryoprotectant attributes that help stabilize plant cells under stress (Bertrand et al. 2017, 2020a; Dhont et al. 2006). Cold acclimation has been frequently reported to induce an increase in total amino acids concentration in roots and crowns of alfalfa plants grown under non-limiting nitrogen conditions (Bertrand et al. 2020a; Castonguay et al. 2011; Dhont et al. 2006). In the present study, total amino acids (AATot) did not increase in nodules, roots and crowns in response to cold acclimation which could be due to the fact that the only source of N was provided by the symbiotic N fixation. The conditions of cold acclimation that were used in this experiment (low temperature and short photoperiod) were far from optimal for symbiotic N fixation which is most effective at 25 °C (Alexandre and Oliveira 2013). These conditions likely restricted amino acids synthesis and accumulation, as opposed to what is observed when non-limiting inorganic N source is used with cold acclimated alfalfa plants as in Castonguay et al. (2011). Amino acid concentrations were almost four fold higher in nodules than in roots and crowns, and this was a constant effect at all sampling events, showing the importance of resource investment in nodules by host plants (Fig. 3). Asparagine, which is recognized as the primary N fixed compound as well as the principal amino acid transported in xylem in legume species with indeterminate nodules such as alfalfa, was the most abundant amino acid in all samples for non-acclimated plants and represented up to 80% of the pool of free amino acids (Table 3; Bertrand et al. 2016; Sulieman et al. 2010). Its progressive decrease in concentration at each sampling event (Fig. 3) confirms the slowdown of N fixation under CA as well as the transformation of this major pool of N compounds into other amino acids involved specifically in stress tolerance. For instance, ornithine and arginine as well as their precursor glutamate increased in response to CA and are known to be precursors of polyamines which confer plant resistance to various abiotic stresses (Anwar et al. 2018). Ornithine has also been reported as a signal and regulatory molecule (Majumdar et al. 2016), which could explain why its concentration, while changing significantly in response to CA and freezing, remained low as compared to arginine. Arginine, with its high N to C ratio is considered a storage compound from which N could be readily incorporated into other N-compounds essential for active regrowth, as previously reported in overwintering alfalfa (Castonguay et al. 2011; Dhont et al. 2006). While proline accumulation in responses to low temperatures has previously been observed in alfalfa (Bertrand et al. 2020a; Castonguay et al. 2011; Liu et al. 2019), its direct involvement in the acquisition of freezing tolerance remains unclear. In the present study, proline concentration did not increase in response to decreased temperatures suggesting that proline synthesis is regulated independently of the glutamate-ornithine-arginine pathway (Majumdar et al. 2016). On the other hand, the lack of proline increase could indicate that low temperatures stress increased the degradation of proline to provide a source of carbon and nitrogen to the bacteroid, thereby supporting recent studies suggesting that proline metabolism may play an essential role of energy transfer in the legume-Rhizobium symbiosis under stress (Sabbioni and Forlani 2022). We also noted an increase of alanine, serine and glycine with cold acclimation in nodules and an increase of those three amino acids in roots and crowns 48 h after the freezing stress. Alanine accumulation is recognized as an universal first stress signal in a wide variety of organisms including plants (Table 3, Ben-Izhak Monselise et al. 2003). Moreover, it has been recently proposed that alanine could play an important role in the symbiosis by sustaining bacteroid metabolism under oxygen limitation (Schulte et al. 2021). Serine/glycine metabolism was shown to be an important key player in biochemical adaptation to environmental stress by regulation of intracellular redox, pH regulation and energy levels (Igamberdiev and Kleczkowski 2018). Another interesting observation is a sharp increase of methionine, tyrosine, phenylalanine, threonine, lysine, isoleucine and leucine concentrations 48 h after freezing stress (Fig. 3, AFS). Methionine synthesis, which is provided by the host plant to the nodules, has been reported to be essential for efficient nodulation by various rhizobia (Barra et al. 2006). Tyrosine and phenylalanine are aromatic amino acid produced by the shikimate pathway which require phosphoenolpyruvate as substrate (Dunn 2014). Tyrosine is essential for the nodule formation and can be used by the bacteroid as source of carbon and nitrogen (Saha et al. 2016). Both tyrosine and phenylalanine can act as substrates for the phenylpropanoids biosynthesis pathway and were found in root exudates (Feduraev et al. 2020). Other protein-bounded amino acids have been reported to increase in response to cold and frost stress in A. thaliana and Camellia sinensis and, while the authors suggested a role for these amino acids in freezing stress acquisition, the mechanism involved would need further investigations (Hildebrandt 2018; Samarina et al. 2020).
4.1.3 Flavonoids
Flavonoids are crucial signaling molecules that play an essential role in the Rhizobium-legume symbiosis as chemoattractant and nod gene inducers. Flavonoids are excreted by the plant in the rhizosphere to modulate communications with microorganisms, either to ensure protection against pathogens or to attract beneficial microbes (Falcone Ferreyra et al. 2012). Only few and specific flavonoids excreted by the legume-host will activate the expression of a group of bacterial nod genes, leading to the synthesis of the Nod factor (lipochitooligosaccharides), essential for initiating nodules formation and to maintain symbiotic activity (Reviewed in Liu and Murray 2016). Following a stress, the crosstalk between the host plants and microorganisms is modified to face the new conditions and the measurement of flavonoid concentrations in root exudates could give information on the signals exchanged between the partners. An increase of formononetin and medicarpin was observed in root exudates after CA (Fig. 3). Formononetin is the most abundant flavonoid in alfalfa while its derivative medicarpin is the second most abundant (Gifford et al. 2018). Formononetin has been reported to be involved in nodules organogenesis (Mathesius 2001) and in the activation of nod gene transcription in Rhizobium meliloti (Table 3, Dakora et al. 1993). Thus, the CA-induced increase of formononetin concentration in roots exudates could be linked to the de novo nodules synthesis necessary for the plant to acclimate to cold. On the other hand, formononetin could have been released directly from stressed roots to regulate germination of pathogenic fungi (Tsai and Philipps 1991). Formononetin is also a precursor of medicarpin which was shown to exert an inhibitory effect on incompatible bacterial strains and to repress nod gene transcription in alfalfa roots (Hartwig et al. 1990). The increase in medicarpin concentration in root exudates under CA indicates that protection mechanisms against pathogens are also in place. Those mechanisms play an important role in the host range rhizobia specificity of the symbiotic association since pathogenic bacteria can produce similar signaling molecules to facilitate their invasion of the host plant (Wang et al. 2018). The triggering of protection mechanisms is supported by the observation of large increases in echinatin, coumestrol and medicarpin concomitant with decrease in formononetin in nodules during the experiment. Echinatin is a potent antagonist of Gram + bacteria with antifungal and antibacterial activity (Dong and Song 2020) while coumestrol has been shown to be induced by fungal infection in alfalfa (Table 3, Fields et al. 2018). Increase in medicarpin and decrease in formononetin in CA nodules could have had an inhibitory effect on the nod gene activity and the cellular division of S. meliloti (Zhang et al. 2009). Flavonoids have also been reported to provide stress protection against UV light, drought, salinity, freezing and to act as scavengers of free radicals such as reactive oxygen species (Baskar et al. 2018; Laoué et al. 2022; Schulz et al. 2016; Sharma et al. 2019). Recently, it has been reported that genes involved in the phenylpropanoids pathways are regulated by low temperatures in alfalfa (Liu et al. 2022). Echinatin could potently increase plant tolerance against several biotic and abiotic stresses (Sharma et al. 2019; Tripathi et al. 2016). Luteolin is essential for the nodulation process by controlling the expression of nodABC during the development of the symbiosis between rhizobia and alfalfa and has also been reported as an excellent free-radical scavenger to protect again oxidative stress (Chen et al. 2020), to enhance starch hydrolysis and soluble sugars accumulation (El-Shafey and AbdElgawad 2012), and to be involved in enhanced salt stress tolerance (Song et al. 2022).
4.1.4 COR genes expression
The expression of the following COR genes of carbohydrate synthetic pathways were affected by the exposure to cold temperature in alfalfa crowns: galactinol synthase (GaS), sucrose synthase (Susy), and sucrose phosphatase synthase (SPS) (Table 4). For instance, the transcript level of GaS, a key enzyme catalyzing the first step of RFO biosynthesis, increased markedly in response to CA, in accordance with the sharp increase in raffinose and stachyose concentrations that we observed in crowns, and in accordance with previous reports (Bertrand et al. 2016; Cunningham et al. 2003; Liu et al. 2019; Xu et al. 2020). Consistently with previous observations in alfalfa (Bertrand et al. 2017), the gene expression of Susy declined in response to CA and, since Susy is responsible for the cleavage of sucrose into glucose and fructose, its down-regulation along with the up regulation of SPS resulted in sucrose accumulation that we observed in storage organs in response to CA. The Susy progressively returned to the non-acclimated level at the next sampling events which is consistent with the progressive decrease in sucrose concentration in crowns, roots and nodules after the freezing stress. Furthermore, it was suggested that Susy activity might be essential for N fixation in root nodules (Gordon et al. 1999). The quick induction of Susy expression that we observed 48 h after the freezing stress concurs with an important role for Susy in the deacclimation process as it would positively activate the reprise of biological N fixation in post-freezing nodules (Gordon et al. 1999). A strong up-regulation of K3-dehydrin, coding for an osmotic-stress protein was also observed in response to cold acclimation. The K3-dehydrin is known to be associated with freezing tolerance in cold-acclimated alfalfa trough membrane-stabilizing effect (Bertrand et al. 2016; Dubé et al. 2013; Xu et al. 2020). The transcript levels of K3-dehydrin did not vary between alfalfa-rhizobia associations and, as such, the up-regulation of K3-dehydrin seems to be part of a general protective process of cold acclimation in alfalfa. In general, our results confirmed previous studies showing that COR genes involved in carbohydrate metabolic pathways are crucial for cold acclimation and freezing stress tolerance and that they are also key actors in the deacclimation process. It would be interesting to investigate the level of expression of those genes in nodules and in response to cold/freezing/deacclimation processes to better understand their role in the recovery of the symbiotic association after freezing.
4.2 Contribution of each partner to the increased freezing tolerance of the association
4.2.1 Sugars and amino acids
We monitored the independent effects of populations and strains on metabolic changes that occurred during cold acclimation (Fig. 4) and deacclimation (Fig. 5) to better understand their respective contribution to the recovery of the symbiotic association after a freezing stress. In general, metabolites concentration in crowns significantly differed mainly in response to alfalfa populations while strains affected mainly nodules metabolites. For instance, RFOs and sucrose increased in alfalfa crowns in response to CA (Fig. 3) but only sucrose was more abundant in freezing-tolerant population A-TF7 as compared to A-TF0, indicating that under conditions of the current study the accumulation of sucrose was more determinant for regrowth after freezing than RFO. Our observations concur with previous reports on cold-induced accumulation of cryoprotective sucrose and RFO in alfalfa populations recurrently selected for superior freezing tolerance (Castonguay et al. 2011). Interestingly, the accumulation of glucose and fructose in CA nodules in response to the inoculation with freeze-sensitive strain B399 seems to indicate that the strain has an effect on the hydrolysis of sucrose by the plant. In response to CA, both symbiotic partners of the most freezing tolerant association (A-TF7 × NRG34) induced an increase in arginine and histidine, confirming the importance of a strategy for N storage during cold acclimation of alfalfa. The response to deacclimation (RAF) also shows a greater remobilisation of N through arginine, ornithine, and histidine in A-TF7 as compared to A-TF0 as well as through arginine for NRG34. During cold acclimation and deacclimation, both symbiotic partners of the most freezing sensitive association (A-TF0 × B399) showed higher accumulation of osmoprotectants and scavenging reacting oxygen species amino acids like GABA, serine, alanine and proline in crowns and proline in nodules (Figs. 4 and 5). These reactions could indicate that these plants suffered more damage by freezing that triggered reparation mechanisms such as ROS scavenging. On the other hand, higher contents of cryoprotective substances after freezing stress could be beneficial to reduce the risks of damages caused by abrupt freeze–thaw episode in early autumn or spring (Xu et al. 2020).
4.2.2 Flavonoids and regulation of C4H and IOMT genes
Production of flavonoids by the phenylpropanoid pathway was influenced by both symbiotic partners at different sampling events. With cold acclimation we found a higher concentration of the nodulation repressor medicarpin in nodules of the less freezing tolerant association (A-TF0 × B399) when compared to the most freezing tolerant association (A-TF7 × NRG34) (Supplemental Table 2). The inhibitory and antimicrobial effect of medicarpin could be linked with the higher incidence of freezing damage and necrosis of nodules of those plants as observed in D’Amours et al. (2022). The up-regulation of the cinnamic acid 4-hydroxylase (C4H) gene was observed in roots in response to CA, which is coherent with the observed concentration increases of the flavonoid precursor tyrosine as well as of total flavonoids concentrations in roots and nodules observed at this sampling point. Low temperature has been reported as the main factor responsible for the accumulation of flavonoids with antioxidant properties, either through induced expression of the encoding gene or increase enzymatic activity of C4H (He et al. 2022). However, as a major rate-limiting enzyme in the phenylpropanoid biosynthesis that separates the pinocembrin pathway from the lignin/monolignol synthesis pathway, the up-regulation of C4H could also be linked to the accumulation of other secondary metabolites including lignin (Gifford et al. 2018). In alfalfa, isoflavone O-methyltransferase (IOMT) catalyses the reaction leading to 7-O-methyl daidzein, the precursor of formononetin and, further down the pathway to medicarpin (He et al. 1998). The IOMT was down-regulated with CA and its expression was lower in freezing-tolerant population A-TF7 than in A-TF0. This is consistent with the decrease of formononetin observed in both roots and nodules at CA sampling and with the higher concentration of medicarpin in root exudates of A-TF0. A lesser repression of expression of the gene involved in synthesis of formononetin and medicarpin by cold and freezing stress could be associated with less freezing tolerance. Upon return to conditions allowing regrowth, at RAF, the expression of IOMT further increased. A larger formononetin concentration in roots of plants inoculated with NRG34 at RAF suggests that strain-induced signals are more active during regrowth after freezing in the specific A-TF7 × NRG34 association. However further investigations are necessary to better understand the role of flavonoids in plant stress tolerance (He et al. 2022).
5 Conclusion
This study highlights the importance to consider the rhizobial symbiosis in strategies aimed at improving stress tolerance in legumes. Rhizobia and plants play complementary roles to ensure enhanced regrowth after freezing. It was shown that root nodules, while accumulating a large pool of free amino acids and carbohydrates during cold acclimation, turned into a source of reserves that enable regrowth in spring after deacclimation. The study also identified metabolic and genetic traits that confer superior yields to alfalfa populations exposed to a sublethal freezing stress such as the accumulation and remobilization of N storage amino acids during cold acclimation and deacclimation, respectively. Choosing the right partners when applying rhizobial inoculants may contribute in improving the stress tolerance of alfalfa.
Data availability
The datasets generated during and analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- Ala:
-
Alanine
- Arg:
-
Arginine
- Asn:
-
Asparagine
- Asp:
-
Aspartic acid
- DM:
-
Dry matter
- DW:
-
Dry weight
- Formo:
-
Formononetin
- FlaTot:
-
Total flavonoids
- Gln:
-
Glutamine
- Glu:
-
Glutamic acid
- Gly:
-
Glycine
- His:
-
Histidine
- Ile:
-
Isoleucine
- Leu:
-
Leucine
- Lys:
-
Lysine
- Met:
-
Methionine
- N:
-
Nitrogen
- NSC:
-
Non structural carbohydrates
- Phe:
-
Phenylalanine
- Pro:
-
Proline
- RFO:
-
Raffinose-family oligosaccharides
- Ser:
-
Serine
- Thr:
-
Threonine
- Tyr:
-
Tyrosine
- Val:
-
Valine
- Orn:
-
Ornithine
- AATot:
-
Total amino acids
- SSTot:
-
Total soluble sugars
- AABA:
-
α-Aminobutyric acid
- GABA:
-
γ-Aminobutyric acid
References
Abd Elbar OH, Elkelish A, Niedbała G, Farag R, Wojciechowski T, Mukherjee S, Abou-Hadid AF, El-Hennawy HM, Abou El-Yazied A, Abd El-Gawad HG, Azab E, Gobouri AA, El-Sawy AM, Bondok A, Ibrahim MFM (2021) Protective effect of γ-aminobutyric acid against chilling stress during reproductive stage in tomato plants through modulation of sugar metabolism, chloroplast integrity, and antioxidative defense systems. Front Plant Sci 12:663750. https://doi.org/10.3389/fpls.2021.663750
Alcázar R, Cuevas JC, Planas J, Zarza X, Bortolotti C, Carrasco P, Salinas J, Tiburcio AF, Altabella T (2011) Integration of polyamines in the cold acclimation response. Plant Sci 180:31–38. https://doi.org/10.1016/j.plantsci.2010.07.022
Alexandre A, Oliveira S (2013) Response to temperature stress in rhizobia. Crit Rev Microbiol 39(3):219–228. https://doi.org/10.3109/1040841X.2012.702097
Anwar A, She M, Wang K, Riaz B, Ye X (2018) Biological roles of ornithine aminotransferase (OAT) in plant stress tolerance: present progress and future perspectives. Int J Mol Sci 19(11):3681. https://doi.org/10.3390/ijms1911368
Barra L, Fontenelle C, Ermel G, Trautwetter A, Walker GC, Blanco C (2006) Interrelations between glycine betaine catabolism and methionine biosynthesis in Sinorhizobium meliloti strain 102F34. J Bacteriol 188(20):7195–7204. https://doi.org/10.1128/JB.00208-06
Baskar V, Rajendran V, Ramalingam S (2018) Flavonoids (antioxidants systems) in higher plants and their response to stresses. In: Gupta D, Palma J, Corpas F (eds) Antioxidants and antioxidant enzymes in higher plants. Springer, Cham, pp 253–268. https://doi.org/10.1007/978-3-319-75088-0_12
Bélanger G, Rochette P, Castonguay Y, Bootsma A, Mongrain D, Ryan DAJ (2002) Climate change and winter survival of perennial forage crops in eastern Canada. Agron J 94:1120–1130. https://doi.org/10.2134/agronj2002.1120
Bélanger G, Castonguay Y, Bertrand A, Dhont C, Rochette P, Couture L, Drapeau R, Mongrains D, Chalifour FP, Michaud R (2006) Winter damage to perennial forage crops in eastern Canada: causes, mitigation, and prediction. Can J Plant Sci 86:33–47. https://doi.org/10.4141/P04-171
Ben-Izhak Monselise E, Parola AH, Kost D (2003) Low-frequency electromagnetic fields induce a stress effect upon higher plants, as evident by the universal stress signal, alanine. Biochem Biophys Res Commun 302(2):427–434. https://doi.org/10.1016/S0006-291X(03)00194-3
Bertrand A, Prévost D, Bigras FJ, Castonguay Y (2007) Elevated atmospheric CO2 and strain of rhizobium alter freezing tolerance and cold-induced molecular changes in alfalfa (Medicago sativa L.). Ann Bot 99:275–284. https://doi.org/10.1093/aob/mcl254
Bertrand A, Prévost D, Juge C, Chalifour FP (2011) Impact of elevated CO2 on carbohydrate and ureide concentrations in soybean inoculated with different strains of Bradyrhizobium japonicum. Botany 89(7):481–490. https://doi.org/10.1139/b11-034
Bertrand A, Bipfubusa M, Dhont C, Chalifour FP, Drouin P, Beauchamp CJ (2016) Rhizobial strains exert a major effect on the amino acid composition of alfalfa nodules under NaCl stress. Plant Physiol Biochem 108:344–352. https://doi.org/10.1016/j.plaphy.2016.08.002
Bertrand A, Bipfubusa M, Claessens A, Rocher S, Castonguay Y (2017) Effect of photoperiod prior to cold acclimation on freezing tolerance and carbohydrate metabolism in alfalfa (Medicago sativa L.). Plant Sci 264:122–128. https://doi.org/10.1016/j.plantsci.2017.09.003
Bertrand A, Gatzke C, Bipfubusa M, Lévesque V, Chalifour FP, Claessens A, Rocher S, Tremblay GF, Beauchamp CJ (2020b) Physiological and biochemical responses to salt stress of alfalfa populations selected for salinity tolerance and grown in symbiosis with salt-tolerant rhizobium. Agronomy 10(4):569. https://doi.org/10.3390/agronomy10040569
Bertrand A, Claessens A, Bourassa J, Rocher S, Baron V (2020a) A whole-plant screening test to select freezing-tolerant and low-dormant genotypes. In: Hincha DK, Zuther E (Eds), Plant cold acclimation, methods and protocols. Methods in molecular biology 2156, 2nd edition, Humana Press, Springer Science+Business Media, LLC, part of Springer Nature 2020a, New York pp 53–60
Bogdanović J, Mojović M, Milosavić N, Mitrović A, Vucinić Z, Spasojević I (2008) Role of fructose in the adaptation of plants to cold-induced oxidative stress. Eur Biophys J 37(7):1241–1246. https://doi.org/10.1007/s00249-008-0260-9
Bringhurst RM, Cardon ZG, Gage DJ (2001) Galactosides in the rhizosphere: utilization by Sinorhizobium meliloti and development of a biosensor. Proc Natl Acad Sci USA 98(8):4540–4545. https://doi.org/10.1073/pnas.071375898
Castonguay Y, Laberge S, Brummer EC, Volenec JJ (2006) Alfalfa winter hardiness: a research retrospective and integrated perspective. Adv Agron 90:203–265. https://doi.org/10.1016/S0065-2113(06)90006-6
Castonguay Y, Michaud R, Nadeau P, Bertrand A (2009) An indoor screening method for improvement of freezing tolerance in alfalfa. Crop Sci 49:809–818. https://doi.org/10.2135/cropsci2008.09.0539
Castonguay Y, Bertrand A, Michaud R, Laberge S (2011) Cold-induced biochemical and molecular changes in alfalfa populations selectively improved for freezing tolerance. Crop Sci 51:2132–2144. https://doi.org/10.2135/cropsci2011.02.0060
Castonguay Y, Michaud J, Dubé M (2015) Reference genes for RT-qPCR analysis of environmentally and developmentally regulated gene expression in alfalfa. Am J Plant Sci 06:132–143. https://doi.org/10.4236/ajps.2015.61015
Castonguay Y, Rocher S, Bertrand A, Michaud J (2020) Identification of transcripts associated with the acquisition of superior freezing tolerance in recurrently-selected populations of alfalfa. Euphytica 216:27. https://doi.org/10.1007/s10681-020-2559-2
Chen HI, Hu WS, Hung MY, Ou HC, Huang SH, Hsu PT, Day CH, Lin KH, Viswanadha VP, Kuo WW, Huang CY (2020) Protective effects of luteolin against oxidative stress and mitochondrial dysfunction in endothelial cells. Nutr Metab Cardiovasc Dis 30(6):1032–1043. https://doi.org/10.1016/j.numecd.2020.02.014
Chinnasamy G, Bal AK (2003) Seasonal changes in carbohydrates of perennial root nodules of beach pea. J Plant Physiol 160(10):1185–1192. https://doi.org/10.1078/0176-1617-01090
Cummings SP (2005) The role and future potential of nitrogen fixing bacteria to boost productivity in organic and low-input sustainable farming systems. Environ Biotechnol 1(1):1–10
Cunningham SM, Nadeau P, Castonguay Y, Laberge S, Volenec JJ (2003) Raffinose and stachyose accumulation, galactinol synthase expression, and winter injury of contrasting alfalfa germplasms. Crop Sci 43:562–570. https://doi.org/10.2135/cropsci2003.5620
D’Amours E, Bertrand A, Cloutier J, Classens A, Rocher S, Seguin P (2022) Impact of Sinorhizobium meliloti strains and plant population on regrowth and nodule regeneration of alfalfa after a freezing event. Plant Soil. https://doi.org/10.1007/s11104-022-05662-4
Dakora FD, Joseph CM, Phillips DA (1993) Alfalfa (Medicago sativa L.) Root exudates contain isoflavonoids in the presence of Rhizobium meliloti. Plant Physiol 101(3):819–824. https://doi.org/10.1104/pp.101.3.819
Dhont C, Castonguay Y, Nadeau P, Bélanger G, Chalifour FP (2002) Alfalfa root carbohydrates and regrowth potential in response to fall harvests. Crop Sci 42(3):754–765. https://doi.org/10.2135/cropsci2002.7540
Dhont C, Castonguay Y, Nadeau P, Bélanger G, Drapeau R, Laberge S, Avice JC, Chalifour FP (2006) Nitrogen reserves, spring regrowth and winter survival of field-grown alfalfa (Medicago sativa) defoliated in the autumn. Ann Bot 97(1):109–120. https://doi.org/10.1093/aob/mcj006
Dong W, Song Y (2020) The significance of flavonoids in the process of biological nitrogen fixation. Int J Mol Sci 21(16):5926. https://doi.org/10.3390/ijms21165926
Dubé MP, Castonguay Y, Cloutier J, Michaud J, Bertrand A (2013) Characterization of two novel cold-inducible K3 dehydrin genes from alfalfa (Medicago sativa spp. sativa L.). Theor Appl Genet 126:823–835. https://doi.org/10.1007/s00122-012-2020-6
Dunn MF (2014) Key roles of microsymbiont amino acid metabolism in rhizobia-legume interactions. Crit Rev Microbiol 41(4):411–451. https://doi.org/10.3109/1040841X.2013.856854
Duzan HM, Mabood F, Souleimanov A, Smith DL (2006) Nod Bj-V (C18:1, MeFuc) production by Bradyrhizobium japonicum (USDA110, 532C) at suboptimal growth temperatures. J Plant Physiol 163(1):107–111. https://doi.org/10.1016/j.jplph.2005.04.029
El-Shafey NM, AbdElgawad H (2012) Luteolin, a bioactive flavone compound extracted from Cichorium endivia L. subsp. divaricatum alleviates the harmful effect of salinity on maize. Acta Physiol Plant 34:2165–2177. https://doi.org/10.1007/s11738-012-1017-8
Falcone Ferreyra ML, Rius SP, Casati P (2012) Flavonoids: biosynthesis, biological functions, and biotechnological applications. Front Plant Sci 3:222. https://doi.org/10.3389/fpls.2012.00222
Feduraev P, Skrypnik L, Riabova A, Pungin A, Tokupova E, Maslennikov P, Chupakhina G (2020) Phenylalanine and tyrosine as exogenous precursors of wheat (Triticum aestivum L.) secondary metabolism through PAL-associated pathways. Plants 9(4):476. https://doi.org/10.3390/plants9040476
Fields RL, Barrell GK, Gash A, Zhao J, Moot DJ (2018) Alfalfa coumestrol content in response to development stage, fungi, aphids, and cultivar. Agron J 110:910–921. https://doi.org/10.2134/agronj2017.09.0535
Galili G (2011) The aspartate-family pathway of plants: linking production of essential amino acids with energy and stress regulation. Plant Signal Behav 6(2):192–195. https://doi.org/10.4161/psb.6.2.14425
Geddes BA, Oresnik IJ (2014) Physiology, genetics, and biochemistry of carbon metabolism in the alphaproteobacterium Sinorhizobium meliloti. Can J Microbiol 60(8):491–507. https://doi.org/10.1139/cjm-2014-0306
Gifford I, Battenberg K, Vaniya A, Wilson A, Tian L, Fiehn O, Am B (2018) Distinctive patterns of flavonoid biosynthesis in roots and nodules of Datisca glomerata and medicago spp. Revealed by metabolomic and gene expression profiles. Front Plant Sci 9:1463. https://doi.org/10.3389/fpls.2018.01463
Gordon AJ, Minchin FR, James CL, Komina O (1999) Sucrose synthase in legume nodules is essential for nitrogen fixation. Plant Phys 120(3):867–878. https://doi.org/10.1104/pp.120.3.867
Gurusamy C, Davis PJ, Bal AK (2000) Seasonal changes in perennial nodules of beach pea (Lathyrus maritimus [L.] Bigel.) with special reference to oleosomes. Int J Plant Sci 161(4):631–638. https://doi.org/10.1086/314291
Hartwig UA, Maxwell CA, Joseph CM, Phillips DA (1990) Effects of alfalfa nod gene-inducing flavonoids on nodABC transcription in Rhizobium meliloti strains containing different nodD genes. J Bacteriol 172(5):2769–2773. https://doi.org/10.1128/jb.172.5.2769-2773.1990
Hawkins JP, Oresnik IJ (2022) The rhizobium-legume symbiosis: co-opting successful stress management. Front Plant Sci 12:796045. https://doi.org/10.3389/fpls.2021.796045
He XZ, Reddy JT, Dixon RA (1998) Stress responses in alfalfa (Medicago sativa L.) XXII. cDNA cloning and characterization of an elicitor-inducible isoflavone 7-O-methyltransferase. Plant Mol Biol 36(1):43–54. https://doi.org/10.1023/a:1005938121453
He J, Yao L, Pecoraro L, Liu C, Wang J, Huang L, Gao W (2023) Cold stress regulates accumulation of flavonoids and terpenoids in plants by phytohormone, transcription process, functional enzyme, and epigenetics. Crit Rev Biotechnol 43(5):680–697. https://doi.org/10.1080/07388551.2022.2053056
Hildebrandt TM (2018) Synthesis versus degradation: directions of amino acid metabolism during Arabidopsis abiotic stress response. Plant Mol Biol 98:121–135. https://doi.org/10.1007/s11103-018-0767-0
Hoagland DR, Arnon DI (1950) The water-culture method for growing plants without soil. Calif Agric Exp Stn Circ 347
Igamberdiev AU, Kleczkowski LA (2018) The glycerate and phosphorylated pathways of serine synthesis in plants: the branches of plant glycolysis linking carbon and nitrogen metabolism. Front Plant Sci 9:318. https://doi.org/10.1007/s11103-018-0767-0
Juurakko CL, Walker VK (2021) Cold acclimation and prospects for cold-resilient crops. Plant Stress 2:100028. https://doi.org/10.1016/j.stress.2021.100028
Kalberer SR, Wisniewski M, Arora R (2006) Deacclimation and reacclimation of cold-hardy plants: current understanding and emerging concepts. Plant Sci 171:3–16. https://doi.org/10.1016/j.plantsci.2006.02.013
Klein W, Weber MH, Marahiel MA (1999) Cold shock response of Bacillus subtilis: isoleucine-dependent switch in the fatty acid branching pattern for membrane adaptation to low temperatures. J Bacteriol 181(17):5341–5349. https://doi.org/10.1128/JB.181.17.5341-5349.1999
Laoué J, Fernandez C, Ormeño E (2022) Plant flavonoids in mediterranean species: a focus on flavonols as protective metabolites under climate stress. Plants 11(2):172. https://doi.org/10.3390/plants11020172
Liu A, Contador CA, Fan K, Lam HM (2018) Interaction and regulation of carbon, nitrogen, and phosphorus metabolisms in root nodules of legumes. Front Plant Sci 9:1860. https://doi.org/10.3389/fpls.2018.01860
Liu CW, Murray JD (2016) The role of flavonoids in nodulation host-range specificity: an update. Plants (Basel) 5(3):33. https://doi.org/10.3390/plants5030033
Liu YS, Geng JC, Sha XY, Zhao YX, Hu TM, Yang PZ (2019) Effect of Rhizobium symbiosis on low-temperature tolerance and antioxidant response in alfalfa (Medicago sativa L.). Front Plant Sci 10:538. https://doi.org/10.3389/fpls.2019.00538
Liu J, Wang T, Weng Y, Liu B, Gao Q, Ji W, Wang Z, Wang Y, Ma X (2022) Identification and characterization of regulatory pathways controlling dormancy under lower temperature in alfalfa (Medicago sativa L.). Front Plant Sci 13:872839. https://doi.org/10.3389/fpls.2022.872839
Majumdar R, Barchi B, Turlapati SA, Gagne M, Minocha R, Long S, Minocha SC (2016) Glutamate, ornithine, arginine, proline, and polyamine metabolic interactions: the pathway is regulated at the post-transcriptional level. Front Plant Sci 7:78. https://doi.org/10.3389/fpls.2016.00078
Marquez-Garcia B, Shaw D, Cooper JW, Karpinska B, Quain MD, Makgopa EM, Kunert K, Foyer CH (2015) Redox markers for drought-induced nodule senescence, a process occurring after drought-induced senescence of the lowest leaves in soybean (Glycine max). Ann Bot 116(4):497–510. https://doi.org/10.1093/aob/mcv030
Mathesius U (2001) Flavonoids induced in cells undergoing nodule organogenesis in white clover are regulators of auxin breakdown by peroxidase. J Exp Bot 52(1):419–426. https://doi.org/10.1093/jxb/52.suppl_1.419
Menéndez AB, Calzadilla PI, Sansberro PA, Espasandin FD, Gazquez A, Bordenave CD, Maiale SJ, Rodríguez AA, Maguire VG, Campestre MP, Garriz A, Rossi FR, Romero FM, Solmi L, Salloum MS, Monteoliva MI, Debat JH, Ruiz OA (2019) Polyamines and legumes: joint stories of stress, nitrogen fixation and environment. Front Plant Sci 10:1415. https://doi.org/10.3389/fpls.2019.01415
Michaud R, Richard C, Willhmot C, Gasser HR (1983) Apica Alfalfa. Can J Plant Sci 63(2):547–549. https://doi.org/10.4141/cjps83-066
Pál M, Szalai G, Janda T (2015) Speculation: polyamines are important in abiotic stress signaling. Plant Sci 237:16–23. https://doi.org/10.1016/j.plantsci.2015.05.003
Peters NK, Frost JW, Long SR (1986) A plant flavone, luteolin, induces expression of Rhizobium meliloti nodulation genes. Science 233(4767):977–980. https://doi.org/10.1126/science.3738520
Prévost D, Drouin P, Laberge S, Bertrand A, Cloutier J, Levesque G (2003) Cold-adapted rhizobia for nitrogen fixation in temperate regions. Can J Bot 81:1153–1161. https://doi.org/10.1139/b03-113
Rice W, Olsen P, Collins M (1995) Symbiotic effectiveness of Rhizobium meliloti at low root temperature. Plant Soil 170(2):351–358. https://doi.org/10.1007/BF00010489
Sabbioni G, Forlani G (2022) The emerging role of proline in the establishment and functioning of legume-rhizobium symbiosis. Front Plant Sci 13:888769. https://doi.org/10.3389/fpls.2022.888769
Saha S, Paul A, Herring L, Dutta A, Bhattacharya A, Samaddar S, Goshe MB, DasGupta M (2016) Gatekeeper tyrosine phosphorylation of SYMRK is essential for synchronizing the epidermal and cortical responses in root nodule symbiosis. Plant Phys 171(1):71–81. https://doi.org/10.1104/pp.15.01962
Samarina LS, Malyukova LS, Efremov AM, Simonyan TA, Matskiv AO, Koninskaya NG, Rakhmangulov RS, Gvasaliya MV, Malyarovskaya VI, Ryndin AV, Orlov YL, Tong W, Hanke MV (2020) Physiological, biochemical and genetic responses of Caucasian tea (Camellia sinensis (L.) Kuntze) genotypes under cold and frost stress. PeerJ 8:e9787. https://doi.org/10.7717/peerj.9787
Sanz-Sáez Á, Erice G, Aguirreolea J, Irigoyen JJ, Sánchez-Díaz M (2012) Alfalfa yield under elevated CO2 and temperature depends on the Sinorhizobium strain and growth season. Environ Exp Bot 77:267–273. https://doi.org/10.1016/j.envexpbot.2011.11.017
Schulte CCM, Borah K, Wheatley RM, Terpolilli JJ, Saalbach G, Crang N, de Groot DH, Ratcliffe RG, Kruger NJ, Papachristodoulou A, Poole PS (2021) Metabolic control of nitrogen fixation in rhizobium-legume symbioses. Sci Adv 7(31):eabh2433. https://doi.org/10.1126/sciadv.abh2433
Schulz E, Tohge T, Zuther E, Fernie AR, Hincha DK (2016) Flavonoids are determinants of freezing tolerance and cold acclimation in Arabidopsis thaliana. Sci Rep 6:34027. https://doi.org/10.1038/srep34027
Seppänen MM, Alitalo V, Bäckström HK, Mäkiniemi K, Jokela V, Falghera-Winseman L, Khazaei H (2018) Growth, freezing tolerance, and yield performance of alfalfa (Medicago sativa L.) cultivars grown under controlled and field conditions in northern latitudes. Can J Plant Sci 98(5):1109–1118. https://doi.org/10.1139/cjps-2017-0305
Sharma A, Shahzad B, Rehman A, Bhardwaj R, Landi M, Zheng B (2019) Response of phenylpropanoid pathway and the role of polyphenols in plants under abiotic stress. Molecules 24(13):2452. https://doi.org/10.3390/molecules24132452
Song Z, Yang Q, Dong B, Li N, Wang M, Du T, Liu N, Niu L, Jin H, Meng D, Fu Y (2022) Melatonin enhances stress tolerance in pigeon pea by promoting flavonoid enrichment, particularly luteolin in response to salt stress. J Exp Bot 73(17):5992–6008. https://doi.org/10.1093/jxb/erac276
Statistics Canada (2022) Alfalfa and alfalfa mixture. In: 32–10–0309–01 Field crops and hay, Census of Agriculture, 2021. Available via. https://www150.statcan.gc.ca/t1/tbl1/en/tv.action. Accessed 12 September 2022
Sulieman S, Fischinger SA, Gresshoff PM, Schulze J (2010) Asparagine as a major factor in the N-feedback regulation of N2 fixation in Medicago truncatula. Physiol Plant 140:21–31. https://doi.org/10.1111/j.1399-3054.2010.01380.x
Szabados L, Savoure A (2009) Proline: a multifunctional amino acid. Trends Plant Sci 15:89–97. https://doi.org/10.1016/j.tplants.2009.11.009
Tarkowski ŁP, Van den Ende W (2015) Cold tolerance triggered by soluble sugars: a multifaceted countermeasure. Front Plant Sci 6:203. https://doi.org/10.3389/fpls.2015.00203
Tripathi P, Rabara RC, Reese RN, Miller MA, Rohila JS, Subramanian S, Shen QJ, Morandi D, Bücking H, Shulaev V, Rushton PJ (2016) A toolbox of genes, proteins, metabolites and promoters for improving drought tolerance in soybean includes the metabolite coumestrol and stomatal development genes. BMC Genomics 17:102. https://doi.org/10.1186/s12864-016-2420-0
Tsai SM, Philips DA (1991) Flavonoids released naturally from alfalfa promote development of symbiotic glomus spores in vitro. App Environ Microbiol 57:1485–1488. https://doi.org/10.1128/aem.57.5.1485-1488.1991
Vincent JM (1970) A manual for the practical study of the root-nodule bacteria. Blackwell Sci Pub, Oxford
Wang Q, Liu J, Zhu H (2018) Genetic and molecular mechanisms underlying symbiotic specificity in legume-rhizobium interactions. Front Plant Sci 9:313. https://doi.org/10.3389/fpls.2018.00313
Xu HY, Tong ZY, He F, Li XL (2020) Response of alfalfa (Medicago sativa L.) to abrupt chilling as reflected by changes in freezing tolerance and soluble sugars. Agronomy 10:255–270. https://doi.org/10.3390/agronomy10020255
Zhang F, Charles TC, Pan B, Smith DL (1996) Inhibition of the expression of Bradyrhizobium japonicum nod genes at low temperatures. Soil Biol Biochem 28:1579–1583. https://doi.org/10.1016/S0038-0717(96)00261-1
Zhang J, Subramanian S, Stacey G, Yu O (2009) Flavones and flavonols play distinct critical roles during nodulation of Medicago truncatula by Sinorhizobium meliloti. Plant J 57(1):171–183. https://doi.org/10.1111/j.1365-313X.2008.03676.x
Zuanazzi JAS, Clergeot PH, Quirion JC, Husson HP, Kondorosi A, Ratet P (1998) Production of Sinorhizobium meliloti nod gene activator and repressor flavonoids from Medicago sativa roots. MPMI 11(8):784–794. https://doi.org/10.1094/MPMI.1998.11.8.784
Acknowledgements
This work was supported by a grant program of Agriculture and Agri-Food Canada. The first author was financially supported by a scholarship from the Fonds de Recherche Québécois Nature et Technologies. The authors sincerely thank Josée Bourassa, Josée Michaud and Sandra Delaney for their technical assistance and Dr Yves Castonguay for his thorough revision of the manuscript. We also thank, Dr Nicolas Ayub from Instituto de Genética “Edwald Alfredo Favret”, INTA who kindly provided us the commercial S. meliloti strain, B399.
Funding
Open Access funding provided by Agriculture & Agri-Food Canada. This work was supported by a grant program of Agriculture and Agri-Food Canada. The first author was financially supported by scholarship from the Fonds de Recherche Québécois Nature et Technologies.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by Emmanuelle D’Amours, Annick Bertrand and Jean Cloutier. The first draft of the manuscript was written by Emmanuelle D’Amours and all authors revised the previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
D’Amours, E., Bertrand, A., Cloutier, J. et al. Metabolic and genetic responses to simulated overwintering conditions of alfalfa-rhizobia associations contrasted in their freezing tolerance. Symbiosis 90, 321–343 (2023). https://doi.org/10.1007/s13199-023-00939-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13199-023-00939-3