Abstract
Background
Cancer cell growth, metastasis, and drug resistance are major challenges in treating liver hepatocellular carcinoma (LIHC). However, the lack of comprehensive and reliable models hamper the effectiveness of the predictive, preventive, and personalized medicine (PPPM/3PM) strategy in managing LIHC.
Methods
Leveraging seven distinct patterns of mitochondrial cell death (MCD), we conducted a multi-omic screening of MCD-related genes. A novel machine learning framework was developed, integrating 10 machine learning algorithms with 67 different combinations to establish a consensus mitochondrial cell death index (MCDI). This index underwent rigorous evaluation across training, validation, and in-house clinical cohorts. A comprehensive multi-omics analysis encompassing bulk, single-cell, and spatial transcriptomics was employed to achieve a deeper insight into the constructed signature. The response of risk subgroups to immunotherapy and targeted therapy was evaluated and validated. RT-qPCR, western blotting, and immunohistochemical staining were utilized for findings validation.
Results
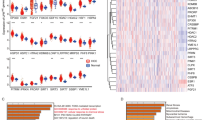
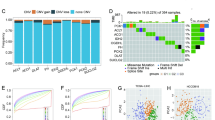
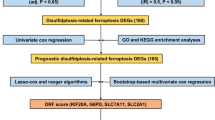
Nine critical differentially expressed MCD-related genes were identified in LIHC. A consensus MCDI was constructed based on a 67-combination machine learning computational framework, demonstrating outstanding performance in predicting prognosis and clinical translation. MCDI correlated with immune infiltration, Tumor Immune Dysfunction and Exclusion (TIDE) score and sorafenib sensitivity. Findings were validated experimentally. Moreover, we identified PAK1IP1 as the most important gene for predicting LIHC prognosis and validated its potential as an indicator of prognosis and sorafenib response in our in-house clinical cohorts.
Conclusion
This study developed a novel predictive model for LIHC, namely MCDI. Incorporating MCDI into the PPPM framework will enhance clinical decision-making processes and optimize individualized treatment strategies for LIHC patients.
Graphical Abstract









Similar content being viewed by others
Data availability
All the data used in this study were collected in this article and supplemental materials.
Availability of data and materials
TCGA datasets enrolled in this study are openly available in the National Cancer Institute GDC Data Portal (https://portal.gdc.cancer.gov/). TCGA data are displayed under the Project IDs “TCGA-LIHC.” GEO datasets are publicly available in the National Center for Biotechnology Information Portal (https://www.ncbi.nlm.nih.gov/geo/), including GSE14520 and GSE156625. ICGC-LIRI dataset is openly available in ICGC (https://dcc.icgc.org/releases/current/Projects). The spatial transcriptomics of LIHC cohorts can be obtained from Mendeley Data (skrx2fz79n). IMvigor210 bladder cancer cohort can be accessed from http://research-pub.gene.com/IMvigor210CoreBiologies.
Code availability
Analyses were conducted using R (version 4.2.1) and GraphPad (GraphPad Prism 8.0). The codes used to support the findings of this study is available from the corresponding author on reasonable request.
Abbreviations
- AUC:
-
Area under the curve
- BCLC:
-
Barcelona Clinic Liver Cancer
- CC:
-
Consensus clustering
- CI:
-
Confidence interval
- CNV:
-
Copy number variation
- DCA:
-
Decision curve analysis
- EPO:
-
Erythropoietin
- G6PD:
-
Glucose-6-phosphate dehydrogenase
- GDSC:
-
Genomics of Drug Sensitivity in Cancer
- GEO:
-
Gene Expression Omnibus
- GINS1:
-
GINS complex subunit 1 (Sld5 homolog)
- GO:
-
Gene ontology
- GSVA:
-
Gene set variation analysis
- HR:
-
Hazard ratio
- IHC:
-
Immunohistochemistry
- ICGC:
-
International Cancer Genome Consortium
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- LASSO:
-
Least absolute shrinkage and selection operator
- LOOCV:
-
Leave-one-out cross-validation
- LIHC:
-
Liver hepatocellular carcinoma
- MCD:
-
Mitochondria-associated cell death
- MCD-DEGs:
-
Mitochondria-associated cell death differentially expressed genes
- MCDI:
-
Mitochondrial cell death index
- MCDS:
-
Mitochondria-associated cell death signature genes
- NC:
-
Normal controls
- OS:
-
Overall survival
- PAK1IP1:
-
P21-activated kinase 1 interacting protein 1
- PCA:
-
Principal component analysis
- PPPM:
-
Predictive, preventive, and personalized medicine
- RFS:
-
Recurrence-free survival
- RT-qPCR:
-
Reverse transcription-quantitative polymerase chain reaction
- ROC:
-
Receiver operating characteristic
- RSF:
-
Random survival forest
- TIDE:
-
Tumor Immune Dysfunction and Exclusion
- TIMER:
-
Tumor Immune Estimation Resource
- UMAP:
-
Uniform Manifold Approximation and Projection
- SD:
-
Standard deviation
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F. Global cancer statistics 2020: globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71(3):209–49. https://doi.org/10.3322/caac.21660.
Wang Z, Wang Y, Gao P, Ding J. Immune checkpoint inhibitor resistance in hepatocellular carcinoma. Cancer Lett. 2023;555:216038.
Reiss KA, Yu S, Mamtani R, Mehta R, D’Addeo K, Wileyto EP, Taddei TH, Kaplan DE. Starting dose of sorafenib for the treatment of hepatocellular carcinoma: a retrospective, multi-institutional study. J Clin Oncol. 2017;35(31):3575–81. https://doi.org/10.1200/JCO.2017.73.8245.
Hu D, Wang Y, Shen X, Mao T, Liang X, Wang T, Shen W, Zhuang Y, Ding J. Genetic landscape and clinical significance of cuproptosis-related genes in liver hepatocellular carcinoma. Genes Dis. 2024;11.
Liu X, Nie L, Zhang Y, Yan Y, Wang C, Colic M, Olszewski K, Horbath A, Chen X, Lei G, et al. Actin cytoskeleton vulnerability to disulfide stress mediates disulfidptosis. Nat Cell Biol. 2023;25(3):404–14. https://doi.org/10.1038/s41556-023-01091-2.
Tsvetkov P, Coy S, Petrova B, Dreishpoon M, Verma A, Abdusamad M, Rossen J, Joesch-Cohen L, Humeidi R, Spangler RD, et al. Copper induces cell death by targeting lipoylated tca cycle proteins. Science. 2022;375(6586):1254–61. https://doi.org/10.1126/science.abf0529.
Bock FJ, Tait S. Mitochondria as multifaceted regulators of cell death. Nat Rev Mol Cell Biol. 2020;21(2):85–100. https://doi.org/10.1038/s41580-019-0173-8.
Tang D, Kang R, Berghe TV, Vandenabeele P, Kroemer G. The molecular machinery of regulated cell death. Cell Res. 2019;29(5):347–64. https://doi.org/10.1038/s41422-019-0164-5.
Martinou JC, Youle RJ. Mitochondria in apoptosis: bcl-2 family members and mitochondrial dynamics. Dev Cell. 2011;21(1):92–101. https://doi.org/10.1016/j.devcel.2011.06.017.
Tuzlak S, Kaufmann T, Villunger A. Interrogating the relevance of mitochondrial apoptosis for vertebrate development and postnatal tissue homeostasis. Genes Dev. 2016;30(19):2133–51. https://doi.org/10.1101/gad.289298.116.
Gao W, Wang X, Zhou Y, Wang X, Yu Y. Autophagy, ferroptosis, pyroptosis, and necroptosis in tumor immunotherapy. Signal Transduct Target Ther. 2022;7(1):196. https://doi.org/10.1038/s41392-022-01046-3.
Napoletano F, Baron O, Vandenabeele P, Mollereau B, Fanto M. Intersections between regulated cell death and autophagy. Trends Cell Biol. 2019;29(4):323–38. https://doi.org/10.1016/j.tcb.2018.12.007.
Russell RC, Guan KL. The multifaceted role of autophagy in cancer. EMBO J. 2022;41(13):e110031. https://doi.org/10.15252/embj.2021110031.
Stockwell BR, Friedmann AJ, Bayir H, Bush AI, Conrad M, Dixon SJ, Fulda S, Gascon S, Hatzios SK, Kagan VE, et al. Ferroptosis: a regulated cell death nexus linking metabolism, redox biology, and disease. Cell. 2017;171(2):273–85. https://doi.org/10.1016/j.cell.2017.09.021.
Sun X, Ou Z, Chen R, Niu X, Chen D, Kang R, Tang D. Activation of the p62-keap1-nrf2 pathway protects against ferroptosis in hepatocellular carcinoma cells. Hepatology. 2016;63(1):173–84. https://doi.org/10.1002/hep.28251.
Chen L, Zhang C, Xue R, Liu M, Bai J, Bao J, Wang Y, Jiang N, Li Z, Wang W, et al. Deep whole-genome analysis of 494 hepatocellular carcinomas. Nature. 2024;627(8004):586–93. https://doi.org/10.1038/s41586-024-07054-3.
Zhang J, Bajari R, Andric D, Gerthoffert F, Lepsa A, Nahal-Bose H, Stein LD, Ferretti V. The international cancer genome consortium data portal. Nat Biotechnol. 2019;37(4):367–9. https://doi.org/10.1038/s41587-019-0055-9.
Roessler S, Jia HL, Budhu A, Forgues M, Ye QH, Lee JS, Thorgeirsson SS, Sun Z, Tang ZY, Qin LX, et al. A unique metastasis gene signature enables prediction of tumor relapse in early-stage hepatocellular carcinoma patients. Cancer Res. 2010;70(24):10202–12. https://doi.org/10.1158/0008-5472.CAN-10-2607.
Liu J, Sun G, Pan S, Qin M, Ouyang R, Li Z, Huang J. The cancer genome atlas (tcga) based m(6)a methylation-related genes predict prognosis in hepatocellular carcinoma. Bioengineered. 2020;11(1):759–68. https://doi.org/10.1080/21655979.2020.1787764.
Liu Z, Liu L, Weng S, Guo C, Dang Q, Xu H, Wang L, Lu T, Zhang Y, Sun Z, et al. Machine learning-based integration develops an immune-derived lncrna signature for improving outcomes in colorectal cancer. Nat Commun. 2022;13(1):816. https://doi.org/10.1038/s41467-022-28421-6.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cbioportal. Sci Signal. 2013;6(269):l1. https://doi.org/10.1126/scisignal.2004088.
Wilkerson MD, Hayes DN. Consensusclusterplus: a class discovery tool with confidence assessments and item tracking. Bioinformatics. 2010;26(12):1572–3. https://doi.org/10.1093/bioinformatics/btq170.
Hanzelmann S, Castelo R, Guinney J. Gsva: gene set variation analysis for microarray and rna-seq data. BMC Bioinform. 2013;14(7):1–5. https://doi.org/10.1186/1471-2105-14-7.
Balachandran VP, Gonen M, Smith JJ, Dematteo RP. Nomograms in oncology: more than meets the eye. Lancet Oncol. 2015;16(4):e173–80. https://doi.org/10.1016/S1470-2045(14)71116-7.
Blanche P, Dartigues JF, Jacqmin-Gadda H. Estimating and comparing time-dependent areas under receiver operating characteristic curves for censored event times with competing risks. Stat Med. 2013;32(30):5381–97. https://doi.org/10.1002/sim.5958.
Alba AC, Agoritsas T, Walsh M, Hanna S, Iorio A, Devereaux PJ, Mcginn T, Guyatt G. Discrimination and calibration of clinical prediction models: users’’ guides to the medical literature. JAMA. 2017;318(14):1377–84. https://doi.org/10.1001/jama.2017.12126.
Sharma A, Seow J, Dutertre CA, Pai R, Bleriot C, Mishra A, Wong R, Singh G, Sudhagar S, Khalilnezhad S, et al. Onco-fetal reprogramming of endothelial cells drives immunosuppressive macrophages in hepatocellular carcinoma. Cell. 2020;183(2):377–94. https://doi.org/10.1016/j.cell.2020.08.040.
Zhang J. Clustergvis: one-step to cluster and visualize gene expression matrix. 2022
Patel AP, Tirosh I, Trombetta JJ, Shalek AK, Gillespie SM, Wakimoto H, Cahill DP, Nahed BV, Curry WT, Martuza RL, et al. Single-cell rna-seq highlights intratumoral heterogeneity in primary glioblastoma. Science. 2014;344(6190):1396–401. https://doi.org/10.1126/science.1254257.
Trapnell C, Cacchiarelli D, Grimsby J, Pokharel P, Li S, Morse M, Lennon NJ, Livak KJ, Mikkelsen TS, Rinn JL. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat Biotechnol. 2014;32(4):381–6. https://doi.org/10.1038/nbt.2859.
Gulati GS, Sikandar SS, Wesche DJ, Manjunath A, Bharadwaj A, Berger MJ, Ilagan F, Kuo AH, Hsieh RW, Cai S, et al. Single-cell transcriptional diversity is a hallmark of developmental potential. Science. 2020;367(6476):405–11. https://doi.org/10.1126/science.aax0249.
Zeng D, Ye Z, Shen R, Yu G, Wu J, Xiong Y, Zhou R, Qiu W, Huang N, Sun L, et al. Iobr: multi-omics immuno-oncology biological research to decode tumor microenvironment and signatures. Front Immunol. 2021;12:687975. https://doi.org/10.3389/fimmu.2021.687975.
Topalian SL, Drake CG, Pardoll DM. Immune checkpoint blockade: a common denominator approach to cancer therapy. Cancer Cell. 2015;27(4):450–61. https://doi.org/10.1016/j.ccell.2015.03.001.
Jiang P, Gu S, Pan D, Fu J, Sahu A, Hu X, Li Z, Traugh N, Bu X, Li B, et al. Signatures of t cell dysfunction and exclusion predict cancer immunotherapy response. Nat Med. 2018;24(10):1550–8. https://doi.org/10.1038/s41591-018-0136-1.
Yang W, Soares J, Greninger P, Edelman EJ, Lightfoot H, Forbes S, Bindal N, Beare D, Smith JA, Thompson IR, et al. Genomics of Drug Sensitivity in Cancer (gdsc): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res. 2013;41:D955-61. https://doi.org/10.1093/nar/gks1111.
Liu Y, Xun Z, Ma K, Liang S, Li X, Zhou S, Sun L, Liu Y, Du Y, Guo X, et al. Identification of a tumour immune barrier in the hcc microenvironment that determines the efficacy of immunotherapy. J Hepatol. 2023;78(4):770–82. https://doi.org/10.1016/j.jhep.2023.01.011.
Mariathasan S, Turley SJ, Nickles D, Castiglioni A, Yuen K, Wang Y, Kadel EI, Koeppen H, Astarita JL, Cubas R, et al. Tgfbeta attenuates tumour response to pd-l1 blockade by contributing to exclusion of t cells. Nature. 2018;554(7693):544–8. https://doi.org/10.1038/nature25501.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative pcr and the 2(-delta delta c(t)) method. Methods. 2001;25(4):402–8. https://doi.org/10.1006/meth.2001.1262.
Xiang DM, Sun W, Ning BF, Zhou TF, Li XF, Zhong W, Cheng Z, Xia MY, Wang X, Deng X, et al. The hlf/il-6/stat3 feedforward circuit drives hepatic stellate cell activation to promote liver fibrosis. Gut. 2018;67(9):1704–15. https://doi.org/10.1136/gutjnl-2016-313392.
Menyhart O, Nagy A, Gyorffy B. Determining consistent prognostic biomarkers of overall survival and vascular invasion in hepatocellular carcinoma. R Soc Open Sci. 2018;5(12):181006. https://doi.org/10.1098/rsos.181006.
Wu R, Guo W, Qiu X, Wang S, Sui C, Lian Q, Wu J, Shan Y, Yang Z, Yang S, et al. Comprehensive analysis of spatial architecture in primary liver cancer. Sci Adv. 2021;7(51):eabg3750. https://doi.org/10.1126/sciadv.abg3750.
Hu D, Zhang T, Yan Z, Wang L, Wang Y, Meng N, Tu B, Teng Y, Li Z, Lou X, et al. Multimolecular characteristics of cell-death related hub genes in human cancers: a comprehensive pan-cancer analysis. Cell Cycle. 2022;21(22):2444–54. https://doi.org/10.1080/15384101.2022.2101337.
Zou Y, Xie J, Zheng S, Liu W, Tang Y, Tian W, Deng X, Wu L, Zhang Y, Wong CW, et al. Leveraging diverse cell-death patterns to predict the prognosis and drug sensitivity of triple-negative breast cancer patients after surgery. Int J Surg. 2022;107:106936. https://doi.org/10.1016/j.ijsu.2022.106936.
Pan H, Pan J, Li P, Gao J. Characterization of panoptosis patterns predicts survival and immunotherapy response in gastric cancer. Clin Immunol. 2022;238:109019. https://doi.org/10.1016/j.clim.2022.109019.
Yang J, Huang Y, Song M, Pan Q, Zhao J, He J, Ouyang D, Yang C, Han Y, Tang Y, et al. Spc25 promotes proliferation and stemness of hepatocellular carcinoma cells via the dna-pk/akt/notch1 signaling pathway. Int J Biol Sci. 2022;18(14):5241–59. https://doi.org/10.7150/ijbs.71694.
Yang H, Liu X, Zhu X, Zhang M, Wang Y, Ma M, Lv K. Gins1 promotes the proliferation and migration of glioma cells through usp15-mediated deubiquitination of top2a. iScience. 2022;25(9):104952. https://doi.org/10.1016/j.isci.2022.104952.
Nagahama Y, Ueno M, Miyamoto S, Morii E, Minami T, Mochizuki N, Saya H, Takakura N. Psf1, a dna replication factor expressed widely in stem and progenitor cells, drives tumorigenic and metastatic properties. Cancer Res. 2010;70(3):1215–24. https://doi.org/10.1158/0008-5472.CAN-09-3662.
Zhou YF, Song SS, Tian MX, Tang Z, Wang H, Fang Y, Qu WF, Jiang XF, Tao CY, Huang R, et al. Cystathionine beta-synthase mediated prrx2/il-6/stat3 inactivation suppresses tregs infiltration and induces apoptosis to inhibit hcc carcinogenesis. J Immunother Cancer. 2021;9(8).https://doi.org/10.1136/jitc-2021-003031.
Yang HC, Wu YH, Yen WC, Liu HY, Hwang TL, Stern A, Chiu DT. The redox role of g6pd in cell growth, cell death, and cancer. Cells. 2019;8(9):1055. https://doi.org/10.3390/cells8091055.
Arcasoy MO. Erythropoiesis-stimulating agent use in cancer: preclinical and clinical perspectives. Clin Cancer Res. 2008;14(15):4685–90. https://doi.org/10.1158/1078-0432.CCR-08-0264.
Yang L, Zhang Z, Sun Y, Pang S, Yao Q, Lin P, Cheng J, Li J, Ding G, Hui L, et al. Integrative analysis reveals novel driver genes and molecular subclasses of hepatocellular carcinoma. Aging (Albany NY). 2020;12(23):23849–71. https://doi.org/10.18632/aging.104047.
Hui Y, Zeng H, Feng Y, Qin W, Chen P, Huang L, Zhong W, Lin L, Lv H, Qin X. Regulatory role of sfn gene in hepatocellular carcinoma and its mechanism. Biotechnol Bioprocess Eng. 2021;26(3):375–83. https://doi.org/10.1007/s12257-020-0292-2.
Wong LL, Lam IP, Wong TY, Lai WL, Liu HF, Yeung LL, Ching YP. Ipa-3 inhibits the growth of liver cancer cells by suppressing pak1 and nf-kappab activation. PLoS ONE. 2013;8(7):e68843. https://doi.org/10.1371/journal.pone.0068843.
Pan JA, Sun Y, Jiang YP, Bott AJ, Jaber N, Dou Z, Yang B, Chen JS, Catanzaro JM, Du C, et al. Trim21 ubiquitylates sqstm1/p62 and suppresses protein sequestration to regulate redox homeostasis. Mol Cell. 2016;61(5):720–33. https://doi.org/10.1016/j.molcel.2016.02.007.
Ying Z, Tian H, Li Y, Lian R, Li W, Wu S, Zhang HZ, Wu J, Liu L, Song J, et al. Cct6a suppresses smad2 and promotes prometastatic tgf-beta signaling. J Clin Invest. 2017;127(5):1725–40. https://doi.org/10.1172/JCI90439.
Pinter M, Scheiner B, Peck-Radosavljevic M. Immunotherapy for advanced hepatocellular carcinoma: a focus on special subgroups. Gut. 2021;70(1):204–14. https://doi.org/10.1136/gutjnl-2020-321702.
Liu J, Shi Y, Zhang Y. Multi-omics identification of an immunogenic cell death-related signature for clear cell renal cell carcinoma in the context of 3p medicine and based on a 101-combination machine learning computational framework. EPMA J. 2023;14(2):275–305. https://doi.org/10.1007/s13167-023-00327-3.
Ge Z, Feng P, Zhang Z, Liang Z, Chen R, Li J. Identification of novel serum protein biomarkers in the context of 3p medicine for intravenous leiomyomatosis: a data-independent acquisition mass spectrometry-based proteomics study. EPMA J. 2023;14(4):613–29. https://doi.org/10.1007/s13167-023-00338-0.
Kurysheva NI, Rodionova OY, Pomerantsev AL, Sharova GA, Golubnitschaja O. Machine learning-couched treatment algorithms tailored to individualized profile of patients with primary anterior chamber angle closure predisposed to the glaucomatous optic neuropathy. EPMA J. 2023;14(3):527–38. https://doi.org/10.1007/s13167-023-00337-1.
Lu M, Zhan H, Liu B, Li D, Li W, Chen X, Zhou X. N6-methyladenosine-related non-coding rnas are potential prognostic and immunotherapeutic responsiveness biomarkers for bladder cancer. EPMA J. 2021;12(4):589–604. https://doi.org/10.1007/s13167-021-00259-w.
Goldstein E, Yeghiazaryan K, Ahmad A, Giordano FA, Frohlich H, Golubnitschaja O. Optimal multiparametric set-up modelled for best survival outcomes in palliative treatment of liver malignancies: unsupervised machine learning and 3 pm recommendations. EPMA J. 2020;11(3):505–15. https://doi.org/10.1007/s13167-020-00221-2.
Funding
This work was supported by a grant from the National Natural Science Foundation of China (82002588, 32300731), the Natural Science Foundation project of Shanghai (23ZR1477200), the State Key Laboratory of the Cancer Biology Project (CBSKL2022ZDKF05) and the Shanghai Key Laboratory of Cell Engineering (14DZ2272300).
Author information
Authors and Affiliations
Contributions
All authors searched the literature, designed the study, interpreted the findings, and revised the manuscript. DTH, XS, and PG carried out data management and statistical analysis and drafted the manuscript. DTH, XS, PG, TTM, YC, WFS, YGZ, and JD helped with cohort identification and data management. YGZ and JD contributed to the critical revision of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
All participants were enrolled from the Eastern Hepatobiliary Surgery Hospital (EHBH), and the sample collection procedure was approved by the ethics committee of EHBH.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Dingtao Hu, Xu Shen, and Peng Gao share co-first authorship.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hu, D., Shen, X., Gao, P. et al. Multi-omic profiling reveals potential biomarkers of hepatocellular carcinoma prognosis and therapy response among mitochondria-associated cell death genes in the context of 3P medicine. EPMA Journal (2024). https://doi.org/10.1007/s13167-024-00362-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13167-024-00362-8