Abstract
Background
Intravenous leiomyomatosis (IVL) is a rare endocrine-associated tumor with unique characteristics of intravascular invasion. This study aimed to identify reliable biomarkers to supervise the development or recurrence of IVL in the context of predictive, preventive, and personalized medicine (PPPM/3PM).
Methods
A total of 60 cases were recruited to detect differentially expressed proteins (DEPs) in serum samples from IVL patients. These cases included those with recurrent IVL, non-recurrent IVL, uterine myoma, and healthy individuals without uterine myoma, with 15 cases in each category. Then, weighted gene co-expression network analysis (WGCNA), lasso-penalized Cox regression analysis (Lasso), trend clustering, and a generalized linear regression model (GLM) were utilized to screen the hub proteins involved in IVL progression.
Results
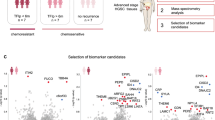
First, 93 differentially expressed proteins (DEPs) were determined from 2582 recognizable proteins, with 54 proteins augmented in the IVL group, and the remaining proteins declined. These proteins were enriched in the modulation of the immune environment, mainly by activating the function of B cells. After the integrated analyses mentioned above, a model based on four proteins (A0A5C2FUE5, A0A5C2GPQ1, A0A5C2GNC7, and A0A5C2GBR3) was developed to efficiently determine the potential of IVL lesions to progress. Among these featured proteins, our results demonstrated that the risk factor A0A5C2FUE5 was associated with IVL progression (OR = 2.64). Conversely, A0A5C2GPQ1, A0A5C2GNC7, and A0A5C2GBR3 might act in a protective manner and prevent disease development (OR = 0.32, 0.60, 0.53, respectively), which was further supported by the multi-class receiver operator characteristic curve analysis.
Conclusion
Four hub proteins were eventually identified based on the integrated bioinformatics analyses. This study potentiates the promising application of these novel biomarkers to predict the prognosis or progression of IVL by a 3PM approach.








Similar content being viewed by others
Data availability
All data generated in this study have been involved in the manuscript or the supplementary files. The raw files of DIA proteomics have been uploaded to the public repository and can be retrieved from the iProX database (https://www.iprox.cn/page/project.html?id=IPX0003343000).
Abbreviations
- IVL :
-
intravenous leiomyomatosis
- CO :
-
control group
- CO-no :
-
control subgroup without uterine myoma
- CO-um :
-
control subgroup with uterine myoma
- IVL-no :
-
IVL subgroup without recurrence
- IVL-re :
-
IVL subgroup with recurrence
- CTA :
-
computerized tomography angiography
- LOH :
-
loss of heterozygosity
- DIA :
-
data-independent acquisition
- FASP :
-
filter-aided sample preparation
- DDA :
-
data dependent acquisition
- AGC :
-
automatic gain control
- LC-MS/MS :
-
liquid chromatography-tandem mass spectrometry
- FDR :
-
false discovery rate
- DEPs :
-
differentially expressed proteins
- FC :
-
fold change
- OPLS-DA :
-
orthogonal partial least-squares discriminant analysis
- GO :
-
gene ontology
- KEGG :
-
Kyoto Encyclopedia of Genes and Genomes
- WGCNA :
-
weighted gene co-expression network analysis
- TOM :
-
topological overlap matrix
- PPI :
-
protein-protein interaction
- FCM :
-
fuzzy c-means
- Lasso :
-
lasso-penalized cox regression
- GLM :
-
generalized linear regression model
- ROC :
-
receiver operator characteristic curve
- AUC :
-
area under the curve
- LTBP2 :
-
latent-transforming growth factor β binding protein 2
- OPN :
-
osteopontin
- NK :
-
natural killer
- Tfh :
-
follicular helper T
- Treg :
-
regulatory T
- GC :
-
germinal center
- Cig :
-
carcinogenic immunoglobulin
- PDAC :
-
pancreatic ductal adenocarcinoma
- LMP1 :
-
latent membrane protein 1
- NF-κB :
-
nuclear factor kappa B
- AP-1 :
-
activating protein-1
References
Ma G, Miao Q, Liu X, Zhang C, Liu J, Zheng Y, et al. Different surgical strategies of patients with intravenous leiomyomatosis. Medicine (Baltimore). 2016;95(37):e4902.
Matsuo K, Fleischman F, Ghattas CS, Gabrielyan AS, Ballard CA, Roman LD, et al. Successful extraction of cardiac-extending intravenous leiomyomatosis through gonadal vein. Fertil Steril. 2012;98(5):1341–5. e1
Yu X, Fu J, Cao T, Huang L, Qie M, Ouyang Y. Clinicopathologic features and clinical outcomes of intravenous leiomyomatosis of the uterus: a case series. Medicine (Baltimore). 2021;100(1):e24228.
Aydin E, Kose O, Kocaaslan C, Aldag M, Bademci MS. Intravascular leiomyomatosis with extension into the pulmonary artery. J Card Surg. 2018;33(8):453–4.
Li B, Chen X, Chu YD, Li RY, Li WD, Ni YM. Intracardiac leiomyomatosis: a comprehensive analysis of 194 cases. Interact Cardiovasc Thorac Surg. 2013;17(1):132–8.
Burke M, Opeskin K. Death due to intravenous leiomyomatosis extending to the right pulmonary artery. Pathology. 2004;36(2):202–3.
Shi T, Shkrum MJ. A case report of sudden death from intracardiac leiomyomatosis. Am J Forensic Med Pathol. 2018;39(2):119–22.
Kong LY, Chen LL, Xiang W, Liu F. Intravenous leiomyomatosis with paradoxical embolism: unusual presentation of uterine leiomyoma. Circ Cardiovasc Imaging. 2020;13(1):e009930.
Liu J, Liang M, Ma G, Liu X, Cheng N, Cao D, et al. Surgical treatment for intravenous-cardiac leiomyomatosis. Eur J Cardiothorac Surg. 2018;54(3):483–90.
Virzi G, Ragazzi S, Bussichella F, D'Agati P, Caputo S, Scaravilli F, et al. Intravenous leiomyomatosis extending from the inferior caval vein to the pulmonary artery. J Thorac Cardiovasc Surg. 2007;133(3):831–2.
Zhang Y, Clark LH, Sheng X, Zhou C. Successful en bloc venous resection with reconstruction and subsequent radiotherapy for 2 consecutive recurrences of intravenous leiomyoma--a case report. BMC Cancer. 2016;16:6.
Su Q, Zhang X, Zhang H, Liu Y, Dong Z, Li G, et al. Intravenous leiomyomatosis of the uterus: a retrospective single-center study in 14 cases. Biomed Res Int. 2020;2020:9758302.
Guo X, Zhang C, Fang L, Guo L, Zhu W, Fang Q, et al. Echocardiographic characteristics of intravenous leiomyomatosis with intracardiac extension: a single-institution experience. Echocardiography. 2011;28(9):934–40.
Gui T, Qian Q, Cao D, Yang J, Peng P, Shen K. Computerized tomography angiography in preoperative assessment of intravenous leiomyomatosis extending to inferior vena cava and heart. BMC Cancer. 2016;16:73.
Wang H, Nie P, Chen B, Hou F, Dong C, He F, et al. Contrast-enhanced CT findings of intravenous leiomyomatosis. Clin Radiol. 2018;73(5):503 e1–6.
Yu X, Zhang G, Lang J, Liu B, Zhao D. Factors associated with recurrence after surgical resection in women with intravenous leiomyomatosis. Obstet Gynecol. 2016;128(5):1018–24.
Zhang G, Feng F, Wang W, Zhu L. Rapamycin (sirolimus) in treatment of recurrent intravenous leiomyomatosis: a case report. BJOG. 2020;127(6):768–71.
Buza N, Xu F, Wu W, Carr RJ, Li P, Hui P. Recurrent chromosomal aberrations in intravenous leiomyomatosis of the uterus: high-resolution array comparative genomic hybridization study. Hum Pathol. 2014;45(9):1885–92.
Lu B, Liu Q, Tang L, Ma Y, Shi H. Intravenous leiomyomatosis: molecular analysis of 17 cases. Pathology. 2020;52(2):213–7.
Wang W, Wang Y, Chen F, Zhang M, Jia R, Liu X, et al. Intravenous leiomyomatosis is inclined to a solid entity different from uterine leiomyoma based on RNA-seq analysis with RT-qPCR validation. Cancer Med. 2020;9(13):4581–92.
Miro A, Coppola Bottazzi E, Vanella S, Palma T, Noviello A, Apicella I, et al. Intravascular leiomyomatosis with intracardiac extension: a toraco-abdominal approach. J Surg Case Rep. 2021;2021(6):rjab249.
Aslam B, Basit M, Nisar MA, Khurshid M, Rasool MH. Proteomics: technologies and their applications. J Chromatogr Sci. 2017;55(2):182–96.
Chung L, Moore K, Phillips L, Boyle FM, Marsh DJ, Baxter RC. Novel serum protein biomarker panel revealed by mass spectrometry and its prognostic value in breast cancer. Breast Cancer Res. 2014;16(3):R63.
Wu J, Xie X, Liu Y, He J, Benitez R, Buckanovich RJ, et al. Identification and confirmation of differentially expressed fucosylated glycoproteins in the serum of ovarian cancer patients using a lectin array and LC-MS/MS. J Proteome Res. 2012;11(9):4541–52.
da Costa AN, Plymoth A, Santos-Silva D, Ortiz-Cuaran S, Camey S, Guilloreau P, et al. Osteopontin and latent-TGF beta binding-protein 2 as potential diagnostic markers for HBV-related hepatocellular carcinoma. Int J Cancer. 2015;136(1):172–81.
Simon AJ, Parry-Smith WR, Redman CW, Kodampur M, Todd R, Satur C, et al. Intravascular leiomyomatosis: a case report and review of the literature. J Obstet Gynaecol. 2015;35(5):539–40.
Gloria-Bottini F, Pietropolli A, Ammendola M, Saccucci P, Bottini E. PTPN22 and uterine leiomyomas. Eur J Obstet Gynecol Reprod Biol. 2015;185:96–8.
Liu ZQ, Lu MY, Sun RL, Yin ZN, Liu B, Wu YZ. Characteristics of peripheral immune function in reproductive females with uterine leiomyoma. J Oncol. 2019;2019:5935640.
Ueno H, Banchereau J, Vinuesa CG. Pathophysiology of T follicular helper cells in humans and mice. Nat Immunol. 2015;16(2):142–52.
Gitlin AD, Mayer CT, Oliveira TY, Shulman Z, Jones MJ, Koren A, et al. HUMORAL IMMUNITY. T cell help controls the speed of the cell cycle in germinal center B cells. Science. 2015;349(6248):643–6.
Barwick BG, Scharer CD, Martinez RJ, Price MJ, Wein AN, Haines RR, et al. B cell activation and plasma cell differentiation are inhibited by de novo DNA methylation. Nat Commun. 2018;9(1):1900.
Dieu-Nosjean MC, Giraldo NA, Kaplon H, Germain C, Fridman WH, Sautes-Fridman C. Tertiary lymphoid structures, drivers of the anti-tumor responses in human cancers. Immunol Rev. 2016;271(1):260–75.
Fehres CM, van Uden NO, Yeremenko NG, Fernandez L, Franco Salinas G, van Duivenvoorde LM, et al. APRIL induces a novel subset of IgA(+) regulatory B cells that suppress inflammation via expression of IL-10 and PD-L1. Front Immunol. 2019;10:1368.
Ou Z, Wang Y, Liu L, Li L, Yeh S, Qi L, et al. Tumor microenvironment B cells increase bladder cancer metastasis via modulation of the IL-8/androgen receptor (AR)/MMPs signals. Oncotarget. 2015;6(28):26065–78.
Yang C, Lee H, Pal S, Jove V, Deng J, Zhang W, et al. B cells promote tumor progression via STAT3 regulated-angiogenesis. PLoS One. 2013;8(5):e64159.
Gu Y, Liu Y, Fu L, Zhai L, Zhu J, Han Y, et al. Tumor-educated B cells selectively promote breast cancer lymph node metastasis by HSPA4-targeting IgG. Nat Med. 2019;25(2):312–22.
Kang S, Maeng H, Kim BG, Qing GM, Choi YP, Kim HY, et al. In situ identification and localization of IGHA2 in the breast tumor microenvironment by mass spectrometry. J Proteome Res. 2012;11(9):4567–74.
Sheng Z, Liu Y, Qin C, Liu Z, Yuan Y, Hu F, et al. IgG is involved in the migration and invasion of clear cell renal cell carcinoma. J Clin Pathol. 2016;69(6):497–504.
Geng ZH, Ye CX, Huang Y, Jiang HP, Ye YJ, Wang S, et al. Human colorectal cancer cells frequently express IgG and display unique Ig repertoire. World J Gastrointest Oncol. 2019;11(3):195–207.
Cui M, You L, Zheng B, Huang X, Liu Q, Huang J, et al. High expression of cancer-derived glycosylated immunoglobulin G predicts poor prognosis in pancreatic ductal adenocarcinoma. J Cancer. 2020;11(8):2213–21.
Qin C, Sheng Z, Huang X, Tang J, Liu Y, Xu T, et al. Cancer-driven IgG promotes the development of prostate cancer though the SOX2-CIgG pathway. Prostate. 2020;80(13):1134–44.
Tang J, Zhang J, Liu Y, Liao Q, Huang J, Geng Z, et al. Lung squamous cell carcinoma cells express non-canonically glycosylated IgG that activates integrin-FAK signaling. Cancer Lett. 2018;430:148–59.
Qiu X, Zhu X, Zhang L, Mao Y, Zhang J, Hao P, et al. Human epithelial cancers secrete immunoglobulin g with unidentified specificity to promote growth and survival of tumor cells. Cancer Res. 2003;63(19):6488–95.
Kaplan B, Livneh A, Sela BA. Immunoglobulin free light chain dimers in human diseases. Sci World J. 2011;11:726–35.
Kang SY, Suh JT, Lee HJ, Yoon HJ, Lee WI. Clinical usefulness of free light chain concentration as a tumor marker in multiple myeloma. Ann Hematol. 2005;84(9):588–93.
Li M, Feng DY, Ren W, Zheng L, Zheng H, Tang M, et al. Expression of immunoglobulin kappa light chain constant region in abnormal human cervical epithelial cells. Int J Biochem Cell Biol. 2004;36(11):2250–7.
Liu H, Zheng H, Duan Z, Hu D, Li M, Liu S, et al. LMP1-augmented kappa intron enhancer activity contributes to upregulation expression of Ig kappa light chain via NF-kappaB and AP-1 pathways in nasopharyngeal carcinoma cells. Mol Cancer. 2009;8:92.
Ma J, Jiang D, Gong X, Shao W, Zhu Z, Xu W, et al. Free immunoglobulin light chain (FLC) promotes murine colitis and colitis-associated colon carcinogenesis by activating the inflammasome. Sci Rep. 2017;7(1):5165.
Groot Kormelink T, Powe DG, Kuijpers SA, Abudukelimu A, Fens MH, Pieters EH, et al. Immunoglobulin free light chains are biomarkers of poor prognosis in basal-like breast cancer and are potential targets in tumor-associated inflammation. Oncotarget. 2014;5(10):3159–67.
Qiu X, Sun X, He Z, Huang J, Hu F, Chen L, et al. Immunoglobulin gamma heavy chain gene with somatic hypermutation is frequently expressed in acute myeloid leukemia. Leukemia. 2013;27(1):92–9.
Adamovic T, McAllister D, Guryev V, Wang X, Andrae JW, Cuppen E, et al. Microalterations of inherently unstable genomic regions in rat mammary carcinomas as revealed by long oligonucleotide array-based comparative genomic hybridization. Cancer Res. 2009;69(12):5159–67.
Zheng H, Li M, Liu H, Ren W, Hu DS, Shi Y, et al. Immunoglobulin alpha heavy chain derived from human epithelial cancer cells promotes the access of S phase and growth of cancer cells. Cell Biol Int. 2007;31(1):82–7.
Chu J, Li Y, Deng Z, Zhang Z, Xie Q, Zhang H, et al. IGHG1 regulates prostate cancer growth via the MEK/ERK/c-Myc pathway. Biomed Res Int. 2019;2019:7201562.
Acknowledgements
The authors would like to thank AJE (https://www.aje.cn/) for English-language editing.
Code availability
The custom codes performed for R analysis could be retrieved from the Bioconductor website (https://www.bioconductor.org/).
Funding
This work was supported by grants from the National High-Level Hospital Clinical Research Funding (2022-PUMCH-B-064, 2022-PUMCH-C-053, and 2022-PUMCH-B-123) and Natural Science Foundation of Beijing (grant number 7234411).
Author information
Authors and Affiliations
Contributions
Software, data curation, formal analysis, and visualization: ZTG and PHF. Writing-original draft preparation and writing review or editing: ZTG, PHF, and ZJZ. Conceptualization or design, administration, and funding acquisition: JCL, RC, and ZYL.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
The research procedure was carried out in conformity with the principle of the Helsinki Declaration and reviewed by the Ethics Committee of Peking Union Medical College Hospital.
Consent for publication
Written informed consent for publication was obtained from all participants.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information

ESM 1
Supplementary Figure 1. Quality control analysis of the project. A. Quantitative fluctuation evaluation of samples. The abscissa represented different samples, and the ordinate represented the amount of protein expression. Different colors were on behalf of different groups. B. Peak capacity evaluation. The abscissa was the sequence of sample loading. The ordinate was the number of peaks; the green line represented the data of all peptides. A red line was displayed to illustrate iRT internal standard data. C. Protein FDR analysis. Cscore was equivalent to the protein reliability score. The black dotted line represented a 1% Q value (equivalent to 1% FDR) standard line. The higher the Csocre at the standard line, the better. (TIF 1960 kb) (PNG 229 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ge, Z., Feng, P., Zhang, Z. et al. Identification of novel serum protein biomarkers in the context of 3P medicine for intravenous leiomyomatosis: a data-independent acquisition mass spectrometry-based proteomics study. EPMA Journal 14, 613–629 (2023). https://doi.org/10.1007/s13167-023-00338-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13167-023-00338-0