Abstract
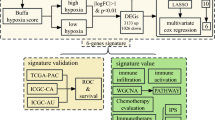
Hepatocellular carcinoma (HCC) is a common cause of cancer-associated death worldwide. The mitochondrial unfolded protein response (UPRmt) not only maintains mitochondrial integrity but also regulates cancer progression and drug resistance. However, no study has used the UPRmt to construct a prognostic signature for HCC. This work aimed to establish a novel signature for predicting patient prognosis, immune cell infiltration, immunotherapy, and chemotherapy response based on UPRmt-related genes (MRGs). Transcriptional profiles and clinical information were obtained from the TCGA and ICGC databases. Cox regression and LASSO regression analyses were applied to select prognostic genes and develop a risk model. The TIMER algorithm was used to investigate immunocytic infiltration in the high- and low-risk subgroups. Here, two distinct clusters were identified with different prognoses, immune cell infiltration statuses, drug sensitivities, and response to immunotherapy. A risk score consisting of seven MRGs (HSPD1, LONP1, SSBP1, MRPS5, YME1L1, HDAC1 and HDAC2) was developed to accurately and independently predict the prognosis of HCC patients. Additionally, the expression of core MRGs was confirmed by immunohistochemistry (IHC) staining, single-cell RNA sequencing, and spatial transcriptome analyses. Notably, the expression of prognostic MRGs was significantly correlated with sorafenib sensitivity in HCC and markedly downregulated in sorafenib-treated HepG2 and Huh7 cells. Furthermore, the knockdown of LONP1 decreased the proliferation, invasion, and migration of HepG2 cells, suggesting that upregulated LONP1 expression contributed to the malignant behaviors of HCC cells. To our knowledge, this is the first study to investigate the consensus clustering algorithm, prognostic potential, immune microenvironment infiltration and drug sensitivity based on the expression of MRGs in HCC. In summary, the UPRmt-related classification and prognostic signature could assist in determining the prognosis and personalized therapy of HCC patients from the perspectives of predictive, preventative and personalized medicine.









Similar content being viewed by others
Data availability
The data used to support the findings of this study are available from the corresponding author upon request.
References
Forner A, Reig M, Bruix J (2018) Hepatocellular carcinoma. Lancet 391(10127):1301–1314
Sung H et al (2021) Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71(3):209–249
Petrick JL et al (2020) International trends in hepatocellular carcinoma incidence, 1978–2012. Int J Cancer 147(2):317–330
Yan T et al (2022) The advanced development of molecular targeted therapy for hepatocellular carcinoma. Cancer Biol Med 19(6):802–817
Ruan L et al (2020) Mitochondria-associated proteostasis. Annu Rev Biophys 49:41–67
Anderson AJ et al (2019) Mitochondria-hubs for regulating cellular biochemistry: emerging concepts and networks. Open Biol 9(8):190126
Wang G et al (2022) Insight into the mitochondrial unfolded protein response and cancer: opportunities and challenges. Cell Biosci 12(1):18
Zhou Z et al (2022) The mitochondrial unfolded protein response: a multitasking giant in the fight against human diseases. Ageing Res Rev 81:101702
Rolland SG et al (2019) Compromised mitochondrial protein import acts as a signal for UPR(mt). Cell Rep 28(7):1659-1669.e5
Köhler F et al (2015) The loss of LRPPRC function induces the mitochondrial unfolded protein response. Aging (Albany NY) 7(9):701–717
Nargund AM et al (2015) Mitochondrial and nuclear accumulation of the transcription factor ATFS-1 promotes OXPHOS recovery during the UPR(mt). Mol Cell 58(1):123–133
Nargund AM et al (2012) Mitochondrial import efficiency of ATFS-1 regulates mitochondrial UPR activation. Science 337(6094):587–590
Inigo JR, Chandra D (2022) The mitochondrial unfolded protein response (UPR(mt)): shielding against toxicity to mitochondria in cancer. J Hematol Oncol 15(1):98
Zhu D, Li X, Tian Y (2022) Mitochondrial-to-nuclear communication in aging: an epigenetic perspective. Trends Biochem Sci 47(8):645–659
Zhu L et al (2021) Mitochondrial unfolded protein response: an emerging pathway in human diseases. Free Radic Biol Med 163:125–134
Muñoz-Carvajal F, Sanhueza M (2020) The mitochondrial unfolded protein response: a hinge between healthy and pathological aging. Front Aging Neurosci 12:581849
Anderson NS, Haynes CM (2020) Folding the mitochondrial UPR into the integrated stress response. Trends Cell Biol 30(6):428–439
Tran HC, Van Aken O (2020) Mitochondrial unfolded protein-related responses across kingdoms: similar problems, different regulators. Mitochondrion 53:166–177
Wodrich APK et al (2022) The unfolded protein responses in health, aging, and neurodegeneration: recent advances and future considerations. Front Mol Neurosci 15:831116
Urbina-Varela R et al (2020) Impact of mitophagy and mitochondrial unfolded protein response as new adaptive mechanisms underlying old pathologies: sarcopenia and non-alcoholic fatty liver disease. Int J Mol Sci 21(20):7704
Richards BJ et al (2023) Mitochondrial protein import and UPR(mt) in skeletal muscle remodeling and adaptation. Semin Cell Dev Biol 143:28–36
Kenny TC, Gomez ML, Germain D (2019) Mitohormesis, UPR(mt), and the complexity of mitochondrial DNA landscapes in cancer. Cancer Res 79(24):6057–6066
Keerthiga R, Pei DS, Fu A (2021) Mitochondrial dysfunction, UPR(mt) signaling, and targeted therapy in metastasis tumor. Cell Biosci 11(1):186
Chattopadhyay M et al (2022) The portrait of liver cancer is shaped by mitochondrial genetics. Cell Rep 38(3):110254
Wu Y et al (2022) Spatiotemporal immune landscape of colorectal cancer liver metastasis at single-cell level. Cancer Discov 12(1):134–153
Liu M et al (2021) Ferroptosis inducer erastin sensitizes NSCLC cells to celastrol through activation of the ROS-mitochondrial fission-mitophagy axis. Mol Oncol 15(8):2084–2105
Li D et al (2021) Nuciferine protects against folic acid-induced acute kidney injury by inhibiting ferroptosis. Br J Pharmacol 178(5):1182–1199
Tan K et al (2015) Mitochondrial SSBP1 protects cells from proteotoxic stresses by potentiating stress-induced HSF1 transcriptional activity. Nat Commun 6:6580
Naresh NU, Haynes CM (2019) Signaling and regulation of the mitochondrial unfolded protein response. Cold Spring Harb Perspect Biol 11(6):a033944
Vyas S, Zaganjor E, Haigis MC (2016) Mitochondria and cancer. Cell 166(3):555–566
Lee HY et al (2021) Mitochondrial metabolic signatures in hepatocellular carcinoma. Cells 10(8):1901
Chen H et al (2021) SIRT3-mediated mitochondrial unfolded protein response weakens breast cancer sensitivity to cisplatin. Genes Genomics 43(12):1433–1444
Llovet JM et al (2008) Sorafenib in advanced hepatocellular carcinoma. N Engl J Med 359(4):378–390
Tang W et al (2020) The mechanisms of sorafenib resistance in hepatocellular carcinoma: theoretical basis and therapeutic aspects. Signal Transduct Target Ther 5(1):87
Rikimaru T et al (2007) Clinical significance of histone deacetylase 1 expression in patients with hepatocellular carcinoma. Oncology 72(1–2):69–74
Teng L et al (2022) Development and validation of a microenvironment-related prognostic model for hepatocellular carcinoma patients based on histone deacetylase family. Transl Oncol 26:101547
Sun L et al (2022) The prognostic value of lysine acetylation regulators in hepatocellular carcinoma. Front Mol Biosci 9:840412
Zhang J, Qiao W, Luo Y (2023) Mitochondrial quality control proteases and their modulation for cancer therapy. Med Res Rev 43(2):399–436
Li S et al (2022) Impaired oxygen-sensitive regulation of mitochondrial biogenesis within the von Hippel–Lindau syndrome. Nat Metab 4(6):739–758
Acknowledgements
The authors would like to thank the researchers and staff of the above software and databases.
Funding
This study was supported by the Guiding Funds of the Central Government for Supporting the Development of the Local Science and Technology (Grant No. 236Z3003G), the Hebei Province Higher Education Scientific Research Special Task Project (Grant No. JZX2024020), the Hebei Administration of Traditional Chinese Medicine (Grant No. 2023072) and the Innovation and Entrepreneurship Training Program for Undergraduates (Grant No. 202310094004).
Author information
Authors and Affiliations
Contributions
Study concept and design: KT, SZ, HG, HW and XL. Acquisition of data: KT, SZ, HG and HW. Analysis and interpretation of the data: KT, SZ, HG, HW, XL and KT. RT-PCR analysis: SZ and MW. Statistical analysis: KT, HG, SZ and HW. Drafting of the article: KT. Critical revision and final approval of the article: HG, XL, YF and KT. Obtained funding: KT and HW. Study supervision: KT. All the authors contributed to the article and approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no known competing financial interests.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, S., Guo, H., Wang, H. et al. A novel mitochondrial unfolded protein response-related risk signature to predict prognosis, immunotherapy and sorafenib sensitivity in hepatocellular carcinoma. Apoptosis (2024). https://doi.org/10.1007/s10495-024-01945-6
Accepted:
Published:
DOI: https://doi.org/10.1007/s10495-024-01945-6